Abstract
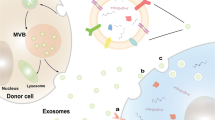
Cancer and other cells residing in the same niche engage various modes of interactions to synchronize and buffer the negative effects of environmental changes. Extracellular microRNAs (miRNAs) have recently been implicated in the intercellular crosstalk. Here we show a mechanistic model involving breast-cancer-secreted, extracellular-vesicle-encapsulated miR-105, which is induced by the oncoprotein MYC in cancer cells and, in turn, activates MYC signalling in cancer-associated fibroblasts (CAFs) to induce a metabolic program. This results in the capacity of CAFs to display different metabolic features in response to changes in the metabolic environment. When nutrients are sufficient, miR-105-reprogrammed CAFs enhance glucose and glutamine metabolism to fuel adjacent cancer cells. When nutrient levels are low and metabolic by-products accumulate, these CAFs detoxify metabolic wastes, including lactic acid and ammonium, by converting them into energy-rich metabolites. Thus, the miR-105-mediated metabolic reprogramming of stromal cells contributes to sustained tumour growth by conditioning the shared metabolic environment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Barcellos-Hoff, M. H., Lyden, D. & Wang, T. C. The evolution of the cancer niche during multistage carcinogenesis. Nat. Rev. Cancer 13, 511–518 (2013).
Chin, A. R. & Wang, S. E. Cancer tills the premetastatic field: mechanistic basis and clinical implications. Clin. Cancer Res. 22, 3725–3733 (2016).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell. Biol. 9, 654–659 (2007).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell. Biol. 10, 1470–1476 (2008).
Tkach, M. & Thery, C. Communication by extracellular vesicles: where we are and where we need to go. Cell 164, 1226–1232 (2016).
S, E. L. A., Mager, I., Breakefield, X. O. & Wood, M. J. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug. Discov. 12, 347–357 (2013).
Becker, A. et al. Extracellular vesicles in cancer: cell-to-cell mediators of metastasis. Cancer Cell. 30, 836–848 (2016).
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 18, 883–891 (2012).
Webber, J., Steadman, R., Mason, M. D., Tabi, Z. & Clayton, A. Cancer exosomes trigger fibroblast to myofibroblast differentiation. Cancer Res. 70, 9621–9630 (2010).
Fong, M. Y. et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat. Cell. Biol. 17, 183–194 (2015).
Zhou, W. et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell. 25, 501–515 (2014).
Chow, A. et al. Macrophage immunomodulation by breast cancer-derived exosomes requires Toll-like receptor 2-mediated activation of NF-kappaB. Sci. Rep. 4, 5750 (2014).
Redzic, J. S., Balaj, L., van der Vos, K. E. & Breakefield, X. O. Extracellular RNA mediates and marks cancer progression. Semin. Cancer Biol. 28, 14–23 (2014).
Chin, A. R. & Wang, S. E. Cancer-derived extracellular vesicles: the ‘soil conditioner’ in breast cancer metastasis?. Cancer Metast. Rev. 35, 669–676 (2016).
Costa-Silva, B. et al Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat. Cell. Biol. 17, 816–826 (2015).
Le, M. T. et al. miR-200-containing extracellular vesicles promote breast cancer cell metastasis. J. Clin. Invest. 124, 5109–5128 (2014).
Zhuang, G. et al. Tumour-secreted miR-9 promotes endothelial cell migration and angiogenesis by activating the JAK-STAT pathway. EMBO J. 31, 3513–3523 (2012).
Tominaga, N. et al. Brain metastatic cancer cells release microRNA-181c-containing extracellular vesicles capable of destructing blood-brain barrier. Nat. Commun. 6, 6716 (2015).
Wu, X. et al. De novo sequencing of circulating miRNAs identifies novel markers predicting clinical outcome of locally advanced breast cancer. J. Transl. Med. 10, 42 (2012).
Kalluri, R. The biology and function of fibroblasts in cancer. Nat. Rev. Cancer 16, 582–598 (2016).
Wise, D. R. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl Acad. Sci. USA 105, 18782–18787 (2008).
DeBerardinis, R. J., Lum, J. J., Hatzivassiliou, G. & Thompson, C. B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell. Metab. 7, 11–20 (2008).
Stine, Z. E., Walton, Z. E., Altman, B. J., Hsieh, A. L. & Dang, C. V. MYC, metabolism, and cancer. Cancer Discov. 5, 1024–1039 (2015).
Pavlides, S. et al. The reverse Warburg effect: aerobic glycolysis in cancer associated fibroblasts and the tumor stroma. Cell. Cycle 8, 3984–4001 (2009).
Zhang, D. et al. Metabolic reprogramming of cancer-associated fibroblasts by IDH3alpha downregulation. Cell. Rep. 10, 1335–1348 (2015).
Lisanti, M. P., Martinez-Outschoorn, U. E. & Sotgia, F. Oncogenes induce the cancer-associated fibroblast phenotype: metabolic symbiosis and “fibroblast addiction” are new therapeutic targets for drug discovery. Cell. Cycle 12, 2723–2732 (2013).
Martinez-Outschoorn, U. E., Lisanti, M. P. & Sotgia, F. Catabolic cancer-associated fibroblasts transfer energy and biomass to anabolic cancer cells, fueling tumor growth. Semin. Cancer Biol. 25, 47–60 (2014).
Martins, D. et al. Loss of caveolin-1 and gain of MCT4 expression in the tumor stroma: key events in the progression from an in situ to an invasive breast carcinoma. Cell. Cycle 12, 2684–2690 (2013).
Tsuyada, A. et al. CCL2 mediates cross-talk between cancer cells and stromal fibroblasts that regulates breast cancer stem cells. Cancer Res. 72, 2768–2779 (2012).
Zervos, A. S., Gyuris, J. & Brent, R. Mxi1, a protein that specifically interacts with Max to bind Myc–Max recognition sites. Cell 72, 223–232 (1993).
Schreiber-Agus, N. et al. An amino-terminal domain of Mxi1 mediates anti-Myc oncogenic activity and interacts with a homolog of the yeast transcriptional repressor SIN3. Cell 80, 777–786 (1995).
Ayer, D. E., Lawrence, Q. A. & Eisenman, R. N. Mad-Max transcriptional repression is mediated by ternary complex formation with mammalian homologs of yeast repressor Sin3. Cell 80, 767–776 (1995).
Lee, T. C. & Ziff, E. B. Mxi1 is a repressor of the c-Myc promoter and reverses activation by USF. J. Biol. Chem. 274, 595–606 (1999).
Conacci-Sorrell, M., McFerrin, L. & Eisenman, R. N. An overview of MYC and its interactome. Cold Spring Harb. Perspect. Med. 4, a014357 (2014).
Yang, L. et al. Targeting stromal glutamine synthetase in tumors disrupts tumor microenvironment-regulated cancer cell growth. Cell. Metab. 24, 685–700 (2016).
Kung, H. N., Marks, J. R. & Chi, J. T. Glutamine synthetase is a genetic determinant of cell type-specific glutamine independence in breast epithelia. PLoS Genet. 7, e1002229 (2011).
Bott, A. J. et al. Oncogenic Myc induces expression of glutamine synthetase through promoter demethylation. Cell. Metab. 22, 1068–1077 (2015).
Faubert, B. Lactate metabolism in human lung tumors. Cell 171, 358–371 (2017).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Spinelli, J.B. et al. Metabolic recycling of ammonia via glutamate dehydrogenase supports breast cancer biomass. Science 358, 941–946 (2017).
Zhao, H. et al. Tumor microenvironment derived exosomes pleiotropically modulate cancer cell metabolism. eLife 5, e10250 (2016).
Valencia, T. et al. Metabolic reprogramming of stromal fibroblasts through p62-mTORC1 signaling promotes inflammation and tumorigenesis. Cancer Cell. 26, 121–135 (2014).
Booth, A. M. et al. Exosomes and HIV Gag bud from endosome-like domains of the T cell plasma membrane. J. Cell. Biol. 172, 923–935 (2006).
Ramakrishnaiah, V. et al. Exosome-mediated transmission of hepatitis C virus between human hepatoma Huh7.5 cells. Proc. Natl Acad. Sci. USA 110, 13109–13113 (2013).
Kahlert, C. et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J. Biol. Chem. 289, 3869–3875 (2014).
San Lucas, F. A. et al. Minimally invasive genomic and transcriptomic profiling of visceral cancers by next-generation sequencing of circulating exosomes. Ann. Oncol. 27, 635–641 (2016).
Thakur, B. K. et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell. Res. 24, 766–769 (2014).
Zhang, H. et al. Exosome-delivered EGFR regulates liver microenvironment to promote gastric cancer liver metastasis. Nat. Commun. 8, 15016 (2017).
Balaj, L. et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat. Commun. 2, 180 (2011).
Slaughter, D. P., Southwick, H. W. & Smejkal, W. Field cancerization in oral stratified squamous epithelium; clinical implications of multicentric origin. Cancer 6, 963–968 (1953).
Debnath, J. et al. The role of apoptosis in creating and maintaining luminal space within normal and oncogene-expressing mammary acini. Cell 111, 29–40 (2002).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Tauro, B. J. et al. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 56, 293–304 (2012).
Kowal, J. et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl Acad. Sci. USA 113, 968–977 (2016).
Li, X. et al. Na+–H+ exchanger 3 (NHE3) is present in lipid rafts in the rabbit ileal brush border: a role for rafts in trafficking and rapid stimulation of NHE3. J. Physiol. 537, 537–552 (2001).
Karamanos, Y. Purification and characterisation of lactate dehydrogenase: an undergraduate biochemistry laboratory experiment. Adv. Biochem. 2, 14–23 (2014).
Beckonert, O. et al. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2, 2692–2703 (2007).
Liu, X., Romero, I. L., Litchfield, L. M., Lengyel, E. & Locasale, J. W. Metformin targets central carbon metabolism and reveals mitochondrial requirements in human cancers. Cell. Metab. 24, 728–739 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Kramer, A., Green, J., Pollard, J. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, 90–97 (2016).
Chen, E. Y.et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, (2013).
Haug, K. et al. MetaboLights-an open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res 41, 781–786 (2013).
Acknowledgements
This work was supported by the National Institutes of Health (NIH)/National Cancer Institute (NCI) grants R01CA218140 (S.E.W.), R01CA166020 (S.E.W.), R01CA206911 (S.E.W.) and R01CA163586 (S.E.W.), California Breast Cancer Research Program grant 20IB-0118 (S.E.W.), Breast Cancer Research Foundation-AACR grant 12-60-26-WANG (S.E.W.) and the National Key Technology R&D Program of China Grant 2015BAI12B12 (X.R.). Research reported in this publication included work performed in Core facilities supported by the NIH/NCI under grant number P30CA23100 (UCSD Cancer Center) and P30CA33572 (City of Hope Cancer Center). We acknowledge the ENCODE Consortium and the ENCODE production laboratories generating the data sets used herein for the analysis.
Author information
Authors and Affiliations
Contributions
S.E.W. and W.Y. conceived ideas, and J.W.L., Y.C. and X.R. contributed to project planning. W.Y. and S.E.W. designed and performed most of the experiments. X.W. performed all bioinformatics analyses. W.Z. performed some experiments with the MCF10A-derived lines. M.Y.F., M.C., J.W. and L.L. assisted with cell line construction and mouse experiments. Xuxiang L. and A.R.C. assisted with the nanoparticle tracking analysis and EV gradient separation. J.L., Xiaojing L. and J.W.L. performed LC/HRMS and data analysis. C.-H.C. and Y.C. assisted with NMR analysis. M.Y.F., D.P.P. and O.F. performed IHC and pathological evaluation. S.E.W. and W.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
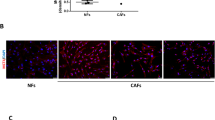
Supplementary Figure 1 Characterization of primary CAFs.
(a-b) CAFs in culture were examined by immunofluorescence (a) and flow cytometry (b; gating performed according to bead-based compensation controls) for the positivity of PDGFβ and vimentin and negativity of EpCAM and CD31. Bar = 100 μm. The experiment was repeated independently for three times with similar results.
Supplementary Figure 2 Characterization of EVs.
(a) EVs pelleted at 110,000 ×g were analysed by nanoparticle tracking analysis (n = 3 independent experiments). The black line indicates mean, whereas the red shaded area indicates SEM. (b) Numbers of secreted EVs determined by nanoparticle tracking analysis (n = 3 independent experiments). (c) A schematic representation of the gradient separation procedure to further characterize the 110,000 ×g pellets. (d) Western blot, density measurement, and RT-qPCR of fractions collected from gradient separation indicating miR-105 enrichment in fractions containing EV markers (n = 3 independent experiments). (e) RT-qPCR-determined levels of miR-105 and miR-155 in equal amounts of EVs from indicated cells (normalized to cel-miR-39-3p spike-in control) and in cell extracts (normalized to U6) (one-way ANOVA, n = 3 independent experiments). For the entire figure, data are shown as mean ± SD; **P < 0.01, ***P < 0.001. †undetectable. Unprocessed original scans of blots are shown in Supplementary Figure 9. Source data are shown in Supplementary Table 5.
Supplementary Figure 3 Overexpression of miR-105 and MYC induce overlapping gene profiles.
(a) GSEA demonstrating the enrichment of MYC target gene sets in MCF10A cells overexpressing miR-105 vs. control cells overexpressing empty vector or GFP (n = 1 biologically independent miR-105-overexpressing cell line and n = 2 biologically independent control cell lines, i.e., empty-vector- and GFP-expressing cell lines). Genes were ranked by signed P value score from edgeR and subjected to GSEA interrogation, which generated the indicated P value, q value and normalized enrichment score (NES) for each gene set based on 1,000 random permutations. (b) A Venn diagram showing the numbers of genes whose levels are altered by ≥2-fold by overexpression of miR-105 or MYC in MCF10A cells.
Supplementary Figure 4 Schematic representatives of the constructs.
(a) Sequence of the putative miR-105 binding sites in MXI1 3’UTR and schematic representatives of the reporter constructs. (b) Sequence of the putative E-box in GABRA3/hsa-mir-105/mir-767 gene promoter and schematic representatives of the reporter constructs. (c) Schematic representatives of the genetic deletion of hsa-mir-105-1 and hsa-mir-105-2 genes and the confirmed genomic DNA sequence in MDA-MB-231ΔmiR-105 cells (the deleted sequences are indicated by strikethrough).
Supplementary Figure 5 Cancer-secreted miR-105 enhances glycolysis and glutaminolysis in NIH3T3 cells.
(a) ECAR and OCR assays in NIH3T3 cells expressing anti-miR-105 or control and treated with indicated EVs or PBS for 48 h (n = 3 independent experiments). **ECAR P < 0.01, ***ECAR P < 0.001, †OCR P < 0.001 (when individually compared to both PBS and MCF10A/vec EV control groups). (b) Changes of metabolite levels in the medium over 72 h by NIH3T3 cells that have been transduced with anti-miR-105 or control and treated with indicated EVs or PBS (n = 3 independent experiments). (c) Levels of metabolites in NIH3T3 cells pretreated as indicated and cultured in regular medium containing 3 g/L glucose and 4 mM glutamine were measured by 1D NMR. Data was normalized to the PBS group (n = 3 independent experiments). (d) Relative RNA levels in EV-treated NIH3T3 cells were detected by RT-qPCR and compared to PBS-treated cells (n = 3 independent experiments). (e) Western blots showing indicated protein levels in EV-treated NIH3T3 cells. For the entire figure, data are shown as mean ± SD; statistical significance was assessed using one-way ANOVA. *P < 0.05, **P < 0.01, ***P < 0.001 (when individually compared to both PBS and MCF10A/vec EV control groups unless indicated). Unprocessed original scans of blots are shown in Supplementary Figure 9. Source data are shown in Supplementary Table 5.
Supplementary Figure 6 miR-105-reprogrammed NIH3T3 cells have an enhanced ability to consume extracellular LA.
(a) LDH activity assays to measure the interconversion between pyruvate and lactate in EV-treated NIH3T3 cells under high-glucose (3 g/L glucose; no LA) or high-LA (1 g/L glucose; 25 mM LA) conditions (n = 3 independent experiments). (b) Extracellular (in the non-buffered conditioned medium) and intracellular pH of treated NIH3T3 cells following 24-h incubation in medium containing indicated levels of glucose and LA (n = 3 independent experiments). (c) NIH3T3 cells pretreated with EVs were cultured in medium containing indicated levels of glucose and LA. Cell numbers were determined after 24 h and compared to those of PBS-treated cells (n = 3 independent experiments). (d) MDA-MB-231 cells were cultured in high-LA medium (1 g/L glucose; 25 mM LA) that had been conditioned by NIH3T3 cells treated as indicated. Survival of MDA-MB-231 cancer cells in the CM was determined after 24 h by comparing to cells grown in high-glucose medium (3 g/L glucose; no LA) (n = 3 independent experiments). For the entire figure, data are shown as mean ± SD; statistical significance was assessed using one-way ANOVA. *P < 0.05, **P < 0.01, ***P < 0.001. Source data are shown in Supplementary Table 5.
Supplementary Figure 7 ROS levels in CAFs, migration of CM-treated cancer cells, and GLUD1 expression.
(a) Fluorescence intensity of the ROS indicator CM-H2DCFDA in CAFs treated as indicated in glutamine/glutamate-free medium containing 5 mM NH4Cl (n = 3 independent experiments). H2O2 was used as a positive control to induce ROS. (b) MDA-MB-231 cells were cultured in NH4+-containing medium (3 g/L glucose; glutamine/glutamate-free; 5 mM NH4Cl) that had been conditioned by CAFs treated as indicated. After 24 h, MDA-MB-231 cells were subjected to transwell migration assay and cells that had migrated within 16 h were quantified (n = 3 independent experiments). (c) Left: Relative RNA levels of GLUD1 in EV-treated CAFs grown in regular medium (3 g/L glucose; 4 mM glutamine) (n = 3 independent experiments). Right: Western blots showing protein levels of GLUD1 in EV-treated CAFs grown in regular medium. For the entire figure, data are shown as mean ± SD; statistical significance was assessed using one-way ANOVA. **P < 0.01, ***P < 0.001. Unprocessed original scans of blots are shown in Supplementary Figure 9. Source data are shown in Supplementary Table 5.
Supplementary Figure 8 miR-105-reprogrammed NIH3T3 cells promote tumour growth.
(a) Female NSG mice received mammary fat pad injection of 1 × 106 MDA-MB-231 cells mixed with 1 × 106 NIH3T3 cells stably expressing anti-miR-105 or control. Tumour volume was measured over the indicated time course (two-way ANOVA, n = 7 mice per condition). (b) Representative IHC images showing Ki67 staining and the overall percentage of tumour cells positive for human Ki67 (two-sided t-test, n = 7 mice per condition). Bar = 100 μm. For the entire figure, data are shown as mean ± SD; ***P < 0.001. Source data are shown in Supplementary Table 5.
Supplementary Figure 9 Unprocessed original scans of blots.
Unprocessed images of all Western blots as indicated. Molecular size markers in kDa.
Supplementary information
Supplementary Information
Supplementary Figures 1–9 and Supplementary Table legends.
Supplementary Table 1
Predicted lists of miRNA targeting MXI1.
Supplementary Table 2
IPA and ENCODE analyses of genes associated with miR-105 overexpression.
Supplementary Table 3
Lists of primers, antibodies and other key reagents.
Supplementary Table 4
LC/HRMS metabolomics data summary.
Supplementary Table 5
Source data.
Rights and permissions
About this article
Cite this article
Yan, W., Wu, X., Zhou, W. et al. Cancer-cell-secreted exosomal miR-105 promotes tumour growth through the MYC-dependent metabolic reprogramming of stromal cells. Nat Cell Biol 20, 597–609 (2018). https://doi.org/10.1038/s41556-018-0083-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0083-6
This article is cited by
-
Metabolic heterogeneity in cancer
Nature Metabolism (2024)
-
Extracellular vesicles as tools and targets in therapy for diseases
Signal Transduction and Targeted Therapy (2024)
-
The clinical significance and oncogenic function of LRRFIP1 in pancreatic cancer
Discover Oncology (2024)
-
The biological function of tumor-derived extracellular vesicles on metabolism
Cell Communication and Signaling (2023)
-
Transcription factors in fibroblast plasticity and CAF heterogeneity
Journal of Experimental & Clinical Cancer Research (2023)