Abstract
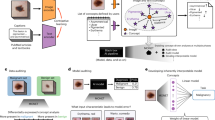
The inferences of most machine-learning models powering medical artificial intelligence are difficult to interpret. Here we report a general framework for model auditing that combines insights from medical experts with a highly expressive form of explainable artificial intelligence. Specifically, we leveraged the expertise of dermatologists for the clinical task of differentiating melanomas from melanoma ‘lookalikes’ on the basis of dermoscopic and clinical images of the skin, and the power of generative models to render ‘counterfactual’ images to understand the ‘reasoning’ processes of five medical-image classifiers. By altering image attributes to produce analogous images that elicit a different prediction by the classifiers, and by asking physicians to identify medically meaningful features in the images, the counterfactual images revealed that the classifiers rely both on features used by human dermatologists, such as lesional pigmentation patterns, and on undesirable features, such as background skin texture and colour balance. The framework can be applied to any specialized medical domain to make the powerful inference processes of machine-learning models medically understandable.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The images used in this study were obtained from publicly available repositories. ISIC images are available at https://challenge.isic-archive.com/data. Fitzpatrick17k images are available at https://github.com/mattgroh/fitzpatrick17k. The DDI images are available at https://stanfordaimi.azurewebsites.net/datasets/35866158-8196-48d8-87bf-50dca81df965. Model weights for the DeepDerm classifier are available at https://zenodo.org/record/6784279#.ZFrDc9LMK-Z. The weights and model specification for the ModelDerm classifier are available at https://figshare.com/articles/Caffemodel_files_and_Python_Examples/5406223. Model weights for our retrained variant of the SIIM-ISIC competition classifier are available at https://zenodo.org/doi/10.5281/zenodo.10049216. Scanoma and Smart Skin Cancer Detection are third-party software for which we cannot redistribute model weights. At the time of writing, both are apps that are available for download with no fee from the Google Play store and from third-party APK-package download sites.
Code availability
Custom codes, including a PyTorch implementation of explanation by progressive exaggeration and of classes for loading datasets and classifiers, are available at https://github.com/suinleelab/derm_audit. The weights for the trained generative models and the re-trained SIIM-ISIC classifier are available at https://zenodo.org/doi/10.5281/zenodo.10049216.
References
Wu, E. et al. How medical AI devices are evaluated: limitations and recommendations from an analysis of FDA approvals. Nat. Med. 27, 582–584 (2021).
Reddy, S. Explainability and artificial intelligence in medicine. Lancet Digit. Health 4, E214–E215 (2022).
Young, A. T. et al. Stress testing reveals gaps in clinic readiness of image-based diagnostic artificial intelligence models. npj Digit. Med. 4, 10 (2021).
DeGrave, A. J., Janizek, J. D. & Lee, S.-I. AI for radiographic COVID-19 detection selects shortcuts over signal. Nat. Mach. Intell. 3, 610–619 (2021).
Singh, N. et al. Agreement between saliency maps and human-labeled regions of interest: applications to skin disease classification. In 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), 3172–3181 (IEEE, 2020).
Bissoto, A., Fornaciali, M., Valle, E. & Avila, S. (De) constructing bias on skin lesion datasets. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), 2766–2774 (IEEE, 2019).
Winkler, J. K. et al. Association between surgical skin markings in dermoscopic images and diagnostic performance of a deep learning convolutional neural network for melanoma recognition. JAMA Dermatol. 155, 1135–1141 (2019).
Singla, S., Pollack, B., Chen, J. & Batmanghelich, K. Explanation by progressive exaggeration. In International Conference on Learning Representations (ICLR, 2020).
Mertes, S., Huber, T., Weitz, K., Heimerl, A., & Andr, E. GANterfactual—counterfactual explanations for medical non-experts using generative adversarial learning. Front. Artif. Intell. 5, 825565 (2022).
Ghoshal, B. & Tucker, A. Estimating uncertainty and interpretability in deep learning for coronavirus (COVID-19) detection. Preprint at arXiv:2003.10769 (2020).
Ozturk, T. et al. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 121, 103792 (2020).
Brunese, L., Mercaldo, F., Reginelli, A. & Santone, A. Explainable deep learning for pulmonary disease and coronavirus COVID-19 detection from X-rays. Comput. Methods Programs Biomed. 196, 105608 (2020).
Karim, M. et al. DeepCOVIDExplainer: explainable COVID-19 diagnosis from chest X-ray images. In 2020 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 1034–1037 (IEEE, 2020).
Geirhos, R. et al. Shortcut learning in deep neural networks. Nat. Mach. Intell. 2, 665–673 (2020).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Liu, Y. et al. A deep learning system for differential diagnosis of skin diseases. Nat. Med. 26, 900–908 (2020).
Han, S. S. et al. Augmented intellignece dermatology: deep neural networks empower medical professionals in diagnosing skin cancer and predicting treatment options for 134 skin disorders. J. Invest. Dermatol. 140, 1753–1761 (2020).
Sun, M. D. et al. Accuracy of commercially available smartphone applications for the detection of melanoma. Br. J. Dermatol. 186, 744–746 (2022).
Freeman, K. et al. Algorithm based smartphone apps to assess risk of skin cancer in adults: systematic review of diagnostic accuracy studies. Br. Med. J. 368, m127 (2020).
Beltrami, E. J. et al. Artificial intelligence in the detection of skin cancer. J. Am. Acad. Dermatol. 87, 1336–1342 (2022).
Daneshjou, R. et al. Disparities in dermatology AI performance on a diverse, curated clinical image set. Sci. Adv. 8, eabq6147 (2022).
Han, S. S. et al. Classification of the clinical images for benign and malignant cutaneous tumors using a deep learning algorithm. J. Invest. Dermatol. 138, 1529–1538 (2018).
Ha, Q., Liu, B. & Liu, F. Identifying melanoma images using EfficientNet ensemble: winning solution to the SIIM-ISIC melanoma classification challenge. Preprint at arXiv:2010.05351 (2020).
Rotemberg, V. et al. A patient-centric dataset of images and metadata for identifying melanomas using clinical context. Sci. Data 8, 34 (2021).
Tschandl, P., Rosendahl, C. & Kittler, H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 5, 180161 (2018).
Combalia, M. et al. BCN20000: dermoscopic lesions in the wild. Preprint at arXiv:1908.02288 (2019).
Groh, M. et al. Evaluating deep neural networks trained on clinical images in dermatology with the Fitzpatrick 17k dataset. In Proceedings of the Computer Vision and Pattern Recognition (CVPR) Sixth ISIC Skin Image Analysis Workshop (IEEE, 2021).
Karras, T. et al. Analyzing and improving the image quality of StyleGAN. In 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 8107–8116 (IEEE, 2020).
Shi, K. et al. A retrospective cohort study of the diagnostic value of different subtypes of atypical pigment network on dermoscopy. J. Am. Acad. Dermatol. 83, 1028–1034 (2020).
Yélamos, O. et al. Usefulness of dermoscopy to improve the clinical and histopathologic diagnosis of skin cancers. J. Am. Acad. Dermatol. 80, 365–377 (2019).
Halpern, A. C., Marghoob, A. A. & Reiter, O. Melanoma Warning Signs: What You Need to Know About Early Signs of Skin Cancer (Skin Cancer Foundation, 2021); https://www.skincancer.org/skin-cancer-information/melanoma/melanoma-warningsigns-and-images/. Accessed April 2023.
Massi, D., De Giorgi, V., Carli, P. & Santucci, M. Diagnostic significance of the blue hue in dermoscopy of melanocytic lesions: a dermoscopic-pathologic study. Am. J. Dermatopathol. 23, 463–469 (2001).
Marghoob, N. G., Liopyris, K. & Jaimes, N. Dermoscopy: a review of the structures that facilitate melanoma detection. J. Osteopath. Med. 119, 380–390 (2019).
Oliveria, S. A., Saraiya, M., Geller, A. C., Heneghan, M. K. & Jorgensen, C. Sun exposure and risk of melanoma. Arch. Dis. Child. 91, 131–138 (2006).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV) 2223–2232 (IEEE, 2017).
Illumination, I. C. on. ISO/CIE 11664-5:2016(e) Colorimetry—part 5: CIE 1976 L*u*v* colour space and u’, v’ uniform chromaticity scale diagram (2016).
Deng, Z., Gijsenij, A. & Zhang, J. Source camera identification using auto-white balance approximation. In 2011 IEEE International Conference on Computer Vision 57–64 (IEEE, 2011).
Rader, R. K. et al. The pink rim sign: location of pink as an indicator of melanoma in dermoscopic images. J. Skin Cancer 2014, 719740 (2014).
Tschandl, P. et al. Human–computer collaboration for skin cancer recognition. Nat. Med. 26, 1229–1234 (2020).
Tschandl, P. et al. Comparison of the accuracy of human readers versus machine-learning algorithms for pigmented skin lesion classification: an open, web-based international, diagnostic study. Lancet Oncol. 20, 938–947 (2019).
Weber, P., Sinz, C., Rinner, C., Kittler, H. & Tschandl, P. Perilesional sun damage as a diagnostic clue for pigmented actinic keratosis and Bowen’s disease. J. Eur. Acad. Dermatol. Venereol. 35, 2022–2026 (2021).
Fitzpatrick, J. E., High, W. A. & Kyle, W. L. Urgent Care Dermatology: Symptom-Based Diagnosis. 477–488 (Elsevier, 2018).
Wu, E. et al. Toward Stronger FDA Approval Standards for AI Medical Devices (Stanford University Human-centered Artificial Intelligence (2022).
Bansal, G. et al. Does the whole exceed its parts? The effect of AI explanations on complementary team performance. In Proceedings of the 2021 CHI Conference on Human Factors in Computing Systems (ACM, 2021).
Rok, R. & Weld, D. S. In search of verifiability: explanations rarely enable complementary performance in AI-advised decision making. Preprint at arXiv:2305.07722v3 (2023).
Roth, L. Looking at Shirley, the ultimate norm: colour balance, image technologies, and cognitive equity. Can. J. Commun. 34, 111–136 (2009).
Lester, J. C., Clark, L., Linos, E. & Daneshjou, R. Clinical photography in skin of colour: tips and best practices. Br. J. Dermatol. 184, 1177–1179 (2021).
Poplin, R. et al. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nat. Biomed. Eng. 2, 158–164 (2018).
Yamashita, T. et al. Factors in color fundus photographs that can be used by humans to determine sex of individuals. Transl Vis. Sci. Technol. 9, 4 (2020).
Codella, N. C. F. et al. Skin lesion analysis toward melanoma detection: a challenge at the 2017 International Symposium on Biomedical Imaging (ISBI), hosted by the International Skin Imaging Collaboration (ISIC). In 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI), 168–172 (IEEE, 2018).
Tan, M. et al. MnasNet: platform-aware neural architecture search for mobile. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 2820–2828 (IEEE, 2019).
Jacob, B. et al. Quantization and training of neural networks for efficient integer-arithmetic-only inference. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2704–2713 (IEEE, 2018)
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Tan, M. & Le, Q. EfficientNet: rethinking model scaling for convolutional neural networks. In Proceedings of the 36th International Conference on Machine Learning (ICML 2019) 6105–6114 (PMLR, 2019).
Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 7132–7141 (IEEE, 2018).
Zhang, H. et al. ResNeSt: split-attention networks. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW) 2735–2745 (IEEE, 2022).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2818–2826 (IEEE, 2016).
Giotis, I. et al. MED-NODE: a computer-assisted melanoma diagnosis system using non-dermoscopic images. Expert Syst. Appl. 42, 6578–6585 (2015).
Acknowledgements
A.J.D., J.D.J., and S.-I.L. were supported by the National Science Foundation (CAREER DBI-1552309 and DBI-1759487) and the National Institutes of Health (R35 GM 128638 and R01 AG061132). R.D. was supported by the National Institutes of Health (5T32 AR007422-38) and the Stanford Catalyst Program.
Author information
Authors and Affiliations
Contributions
A.J.D., J.D.J., R.D. and S.-I.L. conceived the initial study. A.J.D. prepared data and developed software for the reproduction of dermatology AI classifiers, for their counterfactual analysis and for confirmatory experiments. A.J.D. and J.D.J. developed software for the generation of saliency maps. Z.R.C. and R.D. analysed counterfactual images and examined saliency maps. A.J.D., Z.R.C., J.D.J., R.D. and S.-I.L. analysed data and designed additional experiments. Z.R.C. and R.D. provided dermatological insights and clinical context. A.J.D., Z.R.C., J.D.J., R.D., and S.-I.L. wrote the manuscript. S.-I.L. secured funding, and R.D. and S.-I.L. jointly supervised the study.
Corresponding authors
Ethics declarations
Competing interests
R.D. reports fees from L’Oreal, Frazier Healthcare Partners, Pfizer, DWA and VisualDx for consulting; stock options from MDAcne and Revea for advisory board; and research funding from UCB. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of insights from counterfactuals and saliency maps.
We calculated feature attributions using three popular techniques, Expected Gradients, Kernel SHAP, and GradCAM (see Supplementary Methods) and then produced our best-effort visualizations of the resulting saliency maps. We failed to gather insights from the saliency maps, except that the AI classifier may focus on the lesion (but perhaps not always, depending on the saliency technique). In contrast, the counterfactuals provided more granular and medically interpretable insights: for instance, based on the malignant counterfactuals we inferred that multiple colors of pigment (top + bottom), erythema (middle + bottom), darker pigmentation (all), and blue-white veil (bottom) tend to elicit more malignant predictions. In this figure, all saliency maps and counterfactuals were generated in reference to our AI classifier ‘SIIM-ISIC’. Figure adapted with permission from ref. 25, ViDIR Group, Department of Dermatology, Medical University of Vienna.
Extended Data Fig. 2 Attributes identified by the joint expert–XAI auditing procedure as key influences on the output of individual dermatology AI classifiers, when evaluated on the ISIC dataset.
In contrast to main text Fig. 2, attributes are ordered by the proportion of counterfactual pairs from the specified AI classifier in which experts noted that attribute differs, enabling examination of attributes relevant to a particular AI classifier but not necessarily to most AI classifiers (for example, prominence of skin grooves or dermatoglyphs, which influences Scanoma and ModelDerm).
Extended Data Fig. 3 Attributes identified by the join expert–XAI auditing procedure as key influences on the output of individual dermatology AI classifiers, when evaluated on the Fitzpatrick17k dataset.
In contrast to main text Fig. 2, attributes are ordered by the proportion of counterfactual pairs from the specified AI classifier in which experts noted that attribute differs, enabling examination of attributes relevant to a particular AI classifier but not necessarily to other AI classifiers.
Extended Data Fig. 4 Analysis of inter-reader variability, displaying the two readers’ individual conclusions side-by-side for each attribute.
For each reader, we separately determine whether that attribute was 'predominant' in benign or malignant counterfactuals, that is, present to a greater extent in benign (malignant) counterfactuals in at least twice as many images as malignant (benign) counterfactuals. The size of each rectangle (the 'fraction of counterfactual pairs') is then determined as the proportion of counterfactual pairs with a difference noted in the predominant direction, for that reader alone. While readers typically do not attain quantitative agreement on the fraction of counterfactual pairs for a given attribute, the presence and direction of an attribute’s effect typically remains consistent. For conciseness, attribute names are shortened as described in Supplementary Table 1.
Extended Data Fig. 5 Effect of the programmatic modification of image brightness on the predictions of the AI classifier.
We separately applied three methods of image brightness modification (see Supplementary Methods), then calculated the mean change in AI classifier output relative to the original, unaltered images. For modifications in linear RGB or Jzazbz space, we modified brightness by applying a multiplicative factor B = 2n; we display AI classifier responses as a function of n. For modifications in CIELUV space, we add a constant ΔL* to the perceptual lightness L*, where the maximum value of L* is 100. To facilitate visualization, the vertical axis is normalized to the maximum absolute change in AI classifier output observed for a given method; the normalization factors are displayed at bottom right. Images indicate the effect of each given brightness modification.
Extended Data Fig. 6 Effect of the programmatic modification of image chromaticity on the predictions of the AI classifier.
We separately applied three methods of image chromaticity modification (see Supplementary Methods), then calculated the mean change in AI classifier output relative to the original, unaltered images. Each method of chromaticity modification reflects the chromatic adaptation transform (white balancing method) provided by the corresponding color appearance model (CIE 1976 L* u* v*, CIE 1976 L* a* b*, or CAM16). To facilitate visualization, the vertical axis is normalized to the maximum absolute change in AI classifier output observed for a given method; the normalization factors are displayed at bottom right. Images indicate the effect of each given chromaticity modification. Color bars indicate the hue to which a neutral color (white) is shifted by the chromaticity modification; colorfulness in the color bar (but not example images) is exaggerated for ease of viewing.
Supplementary Information
Supplementary Information
Supplementary methods, Tables 1–4, Figs. 1–9 and references.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
DeGrave, A.J., Cai, Z.R., Janizek, J.D. et al. Auditing the inference processes of medical-image classifiers by leveraging generative AI and the expertise of physicians. Nat. Biomed. Eng (2023). https://doi.org/10.1038/s41551-023-01160-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41551-023-01160-9
This article is cited by
-
Hail the AI journal editor
Nature Biomedical Engineering (2024)
-
Transparent medical image AI via an image–text foundation model grounded in medical literature
Nature Medicine (2024)