Abstract
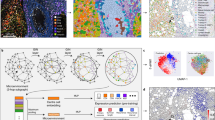
Methods of computational pathology applied to the analysis of whole-slide images (WSIs) do not typically consider histopathological features from the tumour microenvironment. Here, we show that a graph deep neural network that considers such contextual features in gigapixel-sized WSIs in a semi-supervised manner can provide interpretable prognostic biomarkers. We designed a neural-network model that leverages attention techniques to learn features of the heterogeneous tumour microenvironment from memory-efficient representations of aggregates of highly correlated image patches. We trained the model with WSIs of kidney, breast, lung and uterine cancers and validated it by predicting the prognosis of 3,950 patients with these four different types of cancer. We also show that the model provides interpretable contextual features of clear cell renal cell carcinoma that allowed for the risk-based retrospective stratification of 1,333 patients. Deep graph neural networks that derive contextual histopathological features from WSIs may aid diagnostic and prognostic tasks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The TCGA WSI datasets, which were generated by the TCGA Research Network (http://cancergenome.nih.gov), are publicly available through the Genomic Data Commons portal (https://gdc-portal.nci.nih.gov). The NLST WSI datasets are available through the Cancer Imaging Archive (TCIA, https://wiki.cancerimagingarchive.net). The SNUH WSI datasets are not publicly available, in accordance with institutional requirements governing human-subject privacy protection. Source data are provided with this paper.
Code availability
The custom codes are available at https://doi.org/10.5281/zenodo.6799191 and at https://github.com/taliq/TEA-graph68.
References
Yuan, Y. et al. Quantitative image analysis of cellular heterogeneity in breast tumors complements genomic profiling. Sci. Transl. Med. 4, 157ra143 (2012).
Beck, A. H. et al. Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci. Transl. Med. 3, 108ra113 (2011).
Lu, M. Y. et al. AI-based pathology predicts origins for cancers of unknown primary. Nature 594, 106–110 (2021).
Coudray, N. et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
AbdulJabbar, K. et al. Geospatial immune variability illuminates differential evolution of lung adenocarcinoma. Nat. Med. 26, 1054–1062 (2020).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Mobadersany, P. et al. Predicting cancer outcomes from histology and genomics using convolutional networks. Proc. Natl Acad. Sci. USA 115, 201717139 (2018).
Courtiol, P. et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat. Med. 25, 1519–1525 (2019).
Medeiros, L. J., Gelb, A. B. & Weiss, L. M. Renal cell carcinoma. Prognostic significance of morphologic parameters in 121 cases. Cancer 61, 1639–1651 (1988).
Moch, H., Cubilla, A. L., Humphrey, P. A., Reuter, V. E. & Ulbright, T. M. The 2016 WHO classification of tumours of the urinary system and male genital organs—part A: renal, penile, and testicular tumours. Eur. Urol. 70, 93–105 (2016).
Sobottka, B., Lorch, A., Silina, K., van den Broek, M. & Moch, H. Renal cell carcinoma pathology in 2021: ‘new need for renal cancer immune profiling’. Curr. Opin. Urol. 31, 228–235 (2021).
Lu, W., Graham, S., Bilal, M., Rajpoot, N. & Minhas, F. Capturing cellular topology in multi-gigapixel pathology images. CVPRW, 1049–1058 (2020).
Pati, P. et al. HACT-Net: a hierarchical cell-to-tissue graph neural network for histopathological image classification. Uncertainty for Safe Utilization of Machine Learning in Medical Imaging, and Graphs in Biomedical Image Analysis, 208–219 (2020).
Chen, R. J. et al. Pathomic fusion: an integrated framework for fusing histopathology and genomic features for cancer diagnosis and prognosis. IEEE Trans. Med. Imaging (2019).
Zhao, Y. et al. Predicting lymph node metastasis using histopathological images based on multiple instance learning with deep graph convolution. CVPR, 4836–4845 (2020).
Ozen, Y., Aksoy, S., Kösemehmetoğlu, K., Önder, S. & Üner, A. Self-supervised learning with graph neural networks for region of interest retrieval in histopathology. ICPR, 6329–6334 (2021).
Zhou, Y. et al. CGC-Net: cell graph convolutional network for grading of colorectal cancer histology images. CVPRW, 388–398 (2019).
Adnan, M., Kalra, S. & Tizhoosh, H. Representation learning of histopathology images using graph neural networks. CVPRW, 4254–4261 (2020).
Chen, R. J. et al. Whole slide images are 2D point clouds: context-aware survival prediction using patch-based graph convolutional networks. MICCAI, 339–349 (2021).
Jaume, G. et al. Towards explainable graph representations in digital pathology. ICML 2020 Workshop on Computational Biology, 1–5 (2020).
Ying, R., Bourgeois, D., You, J., Zitnik, M. & Leskovec, J. GNNExplainer: generating explanations for graph neural networks. NeurIPS 32, 9240–9251 (2019).
Li, K., Wu, Z., Peng, K., Ernst, J. & Fu, Y. Tell me where to look: guided attention inference network. CVPR, 9215–9223 (2018).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vis. 128, 336–359 (2019).
Yuan, H., Tang, J., Hu, X. & Ji, S. XGNN: Towards model-level explanations of graph neural networks. Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining 430–438 (2020).
Fortunato, S. Community detection in graphs. Phys. Rep. 486, 75–174 (2010).
Stanley, N., Kwitt, R., Niethammer, M. & Mucha, P. J. Compressing networks with super nodes. Sci. Rep. 8, 10892 (2018).
Veličković, P. et al. Graph attention networks. ICLR (2018).
Kokhlikyan, N. et al. Captum: a unified and generic model interpretability library for PyTorch. arXiv https://doi.org/10.48550/arXiv.2009.07896 (2020).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. PMLR (2017).
B, W. Alexander et al. Evaluating attribution for graph neural networks. NeurIPS 33, 5898–5910 (2020).
Fuhrman, S. A., Lasky, L. C. & Limas, C. Prognostic significance of morphologic parameters in renal cell carcinoma. Am. J. Surg. Pathol. 6, 655–664 (1982).
Delahunt, B. et al. The International Society of Urological Pathology (ISUP) grading system for renal cell carcinoma and other prognostic parameters. Am. J. Surg. Pathol. 37, 1490–1504 (2013).
Cai, Q. et al. Ontological analyses reveal clinically-significant clear cell renal cell carcinoma subtypes with convergent evolutionary trajectories into an aggressive type. EBioMedicine 51, 102526 (2020).
Trpkov, K. et al. New developments in existing WHO entities and evolving molecular concepts: The Genitourinary Pathology Society (GUPS) update on renal neoplasia. Mod. Pathol. 34, 1392–1424 (2021).
Zaheer, M. et al. Deep sets. NeurIPS 30, 3394–3404 (2017).
Li, R., Yao, J., Zhu, X., Li, Y. & Huang, J. Graph CNN for survival analysis on whole slide pathological images. MICCAI 174–182 (2018).
Ilse, M., Tomczak, J. M. & Welling, M. Attention-based deep multiple instance learning. PMLR 80, 2127–2136 (2018).
Zhang, S. et al. Immune infiltration in renal cell carcinoma. Cancer Sci. 110, 1564–1572 (2019).
Braun, D. A. et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med. 26, 909–918 (2020).
Qi, Y. et al. Tumor-infiltrating CD39+CD8+ T cells determine poor prognosis and immune evasion in clear cell renal cell carcinoma patients. Cancer Immunol. Immunother. 69, 1565–1576 (2020).
Luo, X. et al. Comprehensive computational pathological image analysis predicts lung cancer prognosis. J. Thorac. Oncol. 12, 501–509 (2017).
Travis, W. D. et al. The 2015 World Health Organization classification of lung tumors impact of genetic, clinical and radiologic advances since the 2004 classification. J. Thorac. Oncol. 10, 1243–1260 (2015).
Hakimi, A. A. et al. An integrated metabolic atlas of clear cell renal cell carcinoma. Cancer Cell 29, 104–116 (2016).
Tjota, M. et al. Eosinophilic renal cell tumors with a TSC and MTOR gene mutations are morphologically and immunohistochemically heterogenous. Am. J. Surg. Pathol. 44, 943–954 (2020).
Rathmell, W. K., Rathmell, J. C. & Linehan, W. M. Metabolic pathways in kidney cancer: current therapies and future directions. J. Clin. Oncol. 36, 3540–3546 (2018).
You, J., Ying, R. & Leskovec, J. Position-aware graph neural networks. PMLR, 7134–7143 (2019).
Xu, H., Jiang, C., Liang, X. & Li, Z. Spatial-aware graph relation network for large-scale object detection. CVPR, 9290–9299 (2019).
Rong, Y., Huang, W., Xu, T. & Huang, J. DropEdge: towards deep graph convolutional networks on node classification. ICLR (2020).
You, Y. et al. Graph contrastive learning with augmentations. NeurIPS 33, 5812–5823 (2020).
Lu, W., Toss, M., Rakha, E., Rajpoot, N. & Minhas, F. SlideGraph+: whole slide image level graphs to predict HER2 status in breast cancer. Med. Image Anal. 80, 102486 (2022).
Jaume, G. et al. Quantifying explainers of graph neural networks in computational pathology. CVPR, 8102–8112 (2021).
Ying, C. et al. Do transformers really perform bad for graph representation? NeurIPS 34, 28887–28888 (2021).
Dwivedi, V. P. & Bresson, X. A generalization of transformer networks to graphs. AAAI Workshop on Deep Learning on Graphs: Methods and Applications (2021).
Esteva, A. et al. Deep learning-enabled medical computer vision. NPJ Digit. Med. 4, 1–9 (2021).
Skrede, O.-J. et al. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet 395, 350–360 (2020).
Yamashita, R. et al. Deep learning model for the prediction of microsatellite instability in colorectal cancer: a diagnostic study. Lancet Oncol. 22, 132–141 (2021).
Bulten, W. et al. Automated deep-learning system for Gleason grading of prostate cancer using biopsies: a diagnostic study. Lancet Oncol. 21, 233–241 (2020).
Gary, B. The OpenCV Library. Dr. Dobb’s J. Softw. Tools 25, 120–123 (2000).
Adam, G., Benjamin, G., Jan, H., Drazen, J. & Mahadev, S. OpenSlide: a vendor-neutral software foundation for digital pathology. J. Pathol. Inform. 4, 1–8 (2013).
Dwivedi, V. P., Joshi, C. K., Laurent, T., Bengio, Y. & Bresson, X. Benchmarking graph neural networks. arXiv https://doi.org/10.48550/arXiv.2003.00982 (2020).
You, J., Ying, Z. & Leskovec, J. Design space for graph. Neural Netw. NeurIPS 33, 17009–17021 (2020).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. PMLR 37, 448–456 (2015).
Fey, M. & Lenssen, J. E. Fast graph representation learning with PyTorch Geometric. ICLR Workshop on Representation Learning on Graphs and Manifolds (2019).
Davidson-Pilon, C. lifelines: survival analysis in Python. J. Open Source Softw. 4, 1317 (2019).
Aric, H., Pieter, S. & Daniel, S. C. Exploring network structure, dynamics, and function using networkx. Proceedings of the 7th Python in Science Conference (2008).
Amin, M. B. et al. AJCC Cancer Staging Manual 8th edn (Springer Cham, 2017).
Lee, Y. et al. Derivation of prognostic contextual histopathological features from whole-slide images of tumours via graph deep learning. Zenedo https://doi.org/10.5281/zenodo.6799191 (2022).
Acknowledgements
We thank A. Choi and N. Kim for many helpful discussions and suggestions. S.K. received funding support for the publication of this study and for the research described in this study from the Ministry of Science and ICT (MSIT) of the Republic of Korea and the National Research Foundation of Korea (NRF-2020R1A3B3079653) and from the BK21 FOUR programme of the Education and Research Program for Future ICT Pioneers, Seoul National University in 2022. J.H.P. received funding support for the research described in this study from the Seoul Metropolitan Government Seoul National University (SMG-SNU) Boramae Medical Center (03-2020-18).
Author information
Authors and Affiliations
Contributions
Y.L., J.H.P., S.O., K.S., K.C.M. and S.K. designed the experiments. Y.L. and K.S. wrote the code, performed the experiments and analysed the results. Y.L. designed and performed the deep-learning model. K.S. analysed the graphical features according to the context of the histopathological features. J.H.P. collected the ccRCC data at the SNUH and TCGA. H.K. and J.-H.C. reviewed the NLST cases and selected the risk-related pathological features. J.H.P. and K.C.M. reviewed the ccRCC cases. M.J. updated the patients’ metadata. C.L. provided the metastasis-related metadata. S.O. and J.S. analysed and reviewed the statistical model used in the study. S.K. and K.C.M. conceived the project. All authors contributed to the preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.L., J.H.P., S.O., K.S., K.C.M. and S.K. are listed as inventors on patents (1020220029619) related to the work applied by the Seoul National University covering the technology. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Joachim Buhmann, Faisal Mahmood and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
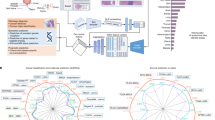
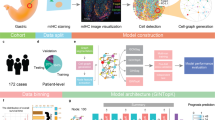
Extended Data Fig. 1 Workflow of TEA-graph and WSI interpretation.
a, Pathologist’s workflow to decide the prognostic region on WSI. b, Workflow of TEA-graph to learn and interpret the context feature on WSI. c, Risk-visualized WSI and example of risk region detection using a connected graph.
Extended Data Fig. 2 Correlation measurement between the risk and IG value.
a, Numbers of patches in the low, mid, and high IG groups for each risk group. IQR of box plot is between Q1 and Q3 and center line indicates median value. Maxima is Q3 + 1.5*IQR and minima is Q1 – 1.5*IQR (n = 259 (Low risk, Mid risk), n = 223 (High risk)). b, Scatter plot between the risk and IG values. c, Merged scatter plot between the risk values and numbers of patches in each IG group. d-f, Scatter plots between the risk values and numbers of patches for each IG group. g, Kaplan-Meier plot analysis according to IG values quantized by 10% of entire IG values. P-values were calculated through two-sided log-rank test (n = 831).
Extended Data Fig. 3 Validation of the TEA-graph on the external NLST dataset.
a, Kaplan-Meier survival analysis using the TEA-graph predicted-risk value (right) and the original stage (left). P-values were calculated through two-sided log-rank test (n = 445). b, Number of patches belong to low, mid, and high IG group for each risk group. IQR of box plot is between Q1 and Q3 and center line indicates median value. Maxima is Q3 + 1.5*IQR and minima is Q1 – 1.5*IQR (n = 378 for each risk group). c, Merged scatter plot between the risk value and the number of patches belonging to each IG group. d, Predicted risk heat map of NLST patients. Scale bar, 4 mm e, Risk-related contextual features predicted by the TEA-graph. Scale bar, 400 μm.
Extended Data Fig. 4 Heterogeneous tumoral architecture features extracted by TEA-graph and effect of attention mechanism to extract the context features.
a, The node IG value is represented by the color of each node, and the edge attention score is represented by the color of each edge. Hemorrhagic cyst (top), patchy stromal hemorrhage (bottom). Scale bar, 100 μm (left), 400 μm (right). Patch size of last column is 80 μm. b, Portion of pairs that had low or high correlation within pairs that had high or low attention. c, Median feature correlation between two nodes connected with low or high attention edge within low, mid, and high IG group. (b-c), The p-value is calculated by two sample t-test (n = 944).
Extended Data Fig. 5 Histopathological prognosis feature comparison between the survival event and metastasis event.
a, Predicted risk heat map of two different events in the same patient who experienced metastasis of cancer to the lung. Scale bar, 4 mm. b, Pathological features of the connected graph that had a high IG value and appeared in both survival and metastasis events. Scale bar, 400 μm. c, Pathological features of the connected graph that had a high IG value and appeared predominantly in survival events. Scale bar, 400 μm.
Extended Data Fig. 6 Contextual pathological characteristics of high IG group.
a, Additional pathological images that represents patch-level cluster characteristics of the high IG group. Patch size is 80 μm. b, The edge distribution (connectivity) heat map of all subgraph cluster of high IG group. c, Graph-level Kaplan-Meier analysis of selected subgraph in graph cluster six in high IG group. P-values were calculated through two-sided log-rank test (n = 831). d, Additional example of pathological features of subgraphs that had high similarity with selected subgraph in graph cluster six in high IG group. Scale bar, 400 μm.
Extended Data Fig. 7 Contextual pathological characteristics of low IG group.
a, Additional pathological images that represents patch-level cluster characteristics of the low IG group. Patch size is 80 μm. b, The edge distribution (connectivity) heat map of all subgraph cluster of low IG group. c, Graph-level Kaplan-Meier analysis of selected subgraph in graph cluster three in low IG group. P-values were calculated through two-sided log-rank test (n = 831). d, Additional example of pathological features of subgraphs that had high similarity with selected subgraph in graph cluster three in low IG group. Scale bar, 400 μm.
Extended Data Fig. 8 Unfavorable risk-related environmental markers on the lung adenocarcinoma predicted by the TEA-graph.
a, Patch-level cluster characteristics of the high IG group. b, Plot of the difference between the areas under the curve of the low- and high-count Kaplan–Meier plots, which reflects the risk of the clusters (left), Kaplan-Meier plot of subgraph cluster three (right) (n = 514). c, Example of connected patch cluster in subgraph cluster three (left) and edge distribution (connectivity) of subgraph cluster three (right). The connectivity shows which patch clusters interact with each other more frequently. d, t-SNE plot of high IG subgraph clustered by k-means clustering method using graph features. e, Examples of the subgraph-level pathological features of subgraph cluster three. Scale bar, 400 μm. f, The edge distribution (connectivity) heat map of the other subgraph cluster of unfavorable (high IG) group. (a,c), Patch size is 80 μm.
Extended Data Fig. 9 Angiogenesis-related environmental markers predicted by the TEA-graph.
a, Kaplan-Meier plot of subgraph cluster zero of high IG group (n = 831). b, Graph-level Kaplan-Meier analysis of selected subgraph related to the angiogenesis in subgraph cluster zero of high IG group (n = 831). c, Edge distribution (connectivity) of subgraph cluster zero and angiogenesis-related connectivity. d, Example of the subgraph-level angiogenesis-related pathological features (left) and connected patch cluster (right). Patch size is 80 μm. e, Additional example of pathological features of subgraphs that had high similarity with selected angiogenesis-related subgraph in graph cluster zero in high IG group. Scale bar, 400 μm. (a-b), P-values were calculated through two-sided log-rank test.
Extended Data Fig. 10 Pathological features of each IG group misclassified by TEA-graph.
a, False positive clusters of the low IG group, which have a small area under the curve value for the Kaplan–Meier plot. b-c, Pathological features of low IG group cluster five. d, False positive cluster of the high IG group, which has a negative area under the curve value for the Kaplan–Meier plot. e, Pathological features of high IG group cluster two. (b, c, e), Scale bar, 400 μm.
Supplementary information
Supplementary Information
Supplementary figures and tables.
Supplementary dataset
Source data for supplementary figures.
Source data
Source data for Figs. 2, 3 and 6 and for Extended Data Figs. 2–4 and 6–9
Source data and statistics for Figs. 2, 3 and 6, and for Extended Data Figs. 2–4 and 6–9.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, Y., Park, J.H., Oh, S. et al. Derivation of prognostic contextual histopathological features from whole-slide images of tumours via graph deep learning. Nat. Biomed. Eng (2022). https://doi.org/10.1038/s41551-022-00923-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41551-022-00923-0
This article is cited by
-
Mapping cancer biology in space: applications and perspectives on spatial omics for oncology
Molecular Cancer (2024)
-
Deep learning on tertiary lymphoid structures in hematoxylin-eosin predicts cancer prognosis and immunotherapy response
npj Precision Oncology (2024)
-
AI-based histopathology image analysis reveals a distinct subset of endometrial cancers
Nature Communications (2024)
-
A transformer-based weakly supervised computational pathology method for clinical-grade diagnosis and molecular marker discovery of gliomas
Nature Machine Intelligence (2024)
-
Holotomography
Nature Reviews Methods Primers (2024)