Abstract
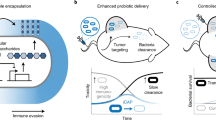
Engineered bacteria for therapeutic applications would benefit from control mechanisms that confine the growth of the bacteria within specific tissues or regions in the body. Here we show that the tropism of engineered bacteria can be enhanced by coupling bacterial growth with genetic circuits that sense oxygen, pH or lactate through the control of the expression of essential genes. Bacteria that were engineered with pH or oxygen sensors showed preferential growth in physiologically relevant acidic or oxygen conditions, and reduced growth outside the permissive environments when orally delivered to mice. In syngeneic mice bearing subcutaneous tumours, bacteria engineered with both hypoxia and lactate biosensors coupled through an AND gate showed increased tumour specificity. The multiplexing of genetic circuits may be more broadly applicable for enhancing the localization of bacteria to specified niches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The raw fluorescence and OD600 data in Fig. 2b, and the raw c.f.u. counts and measured weights in Supplementary Fig. 12 are provided as Supplementary Information. Additional data are available from the corresponding author on request.
Code availability
The MATLAB code used in this study is available from the corresponding author on request.
References
Riglar, D. T. & Silver, P. A. Engineering bacteria for diagnostic and therapeutic applications. Nat. Rev. Microbiol. 16, 214–225 (2018).
Ruder, W. C., Lu, T. & Collins, J. J. Synthetic biology moving into the clinic. Science 333, 1248–1252 (2011).
Khalil, A. S. & Collins, J. J. Synthetic biology: applications come of age. Nat. Rev. Genet. 11, 367–379 (2010).
Kitada, T., DiAndreth, B., Teague, B. & Weiss, R. Programming gene and engineered-cell therapies with synthetic biology. Science 359, eaad1067 (2018).
Garbeva, P., van Veen, J. A. & van Elsas, J. D. Microbial diversity in soil: selection microbial populations by plant and soil type and implications for disease suppressiveness. Annu. Rev. Phytopathol. 42, 243–270 (2004).
Martinez-Hidalgo, P., Maymon, M., Pule-Meulenberg, F. & Hirsch, A. M. Engineering root microbiomes for healthier crops and soils using beneficial, environmentally safe bacteria. Can. J. Microbiol. 65, 91–104 (2019).
Vijay, K., Murmu, M. & Deo, S. V. Bacteria based self healing concrete—a review. Constr. Build. Mater. 152, 1008–1014 (2017).
Lawson, C. E. et al. Common principles and best practices for engineering microbiomes. Nat. Rev. Microbiol. 17, 725–741 (2019).
Cao, Y. et al. Programmable assembly of pressure sensors using pattern-forming bacteria. Nat. Biotechnol. 35, 1087–1093 (2017).
Riglar, D. T. et al. Engineered bacteria can function in the mammalian gut long-term as live diagnostics of inflammation. Nat. Biotechnol. 35, 653–658 (2017).
Daeffler, K. N. et al. Engineering bacterial thiosulfate and tetrathionate sensors for detecting gut inflammation. Mol. Syst. Biol. 13, 923 (2017).
Hwang, I. Y. et al. Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models. Nat. Commun. 8, 15028 (2017).
Mao, N., Cubillos-Ruiz, A., Cameron, D. E. & Collins, J. J. Probiotic strains detect and suppress cholera in mice. Sci. Transl. Med. 10, eaao2586 (2018).
Din, M. O. et al. Synchronized cycles of bacterial lysis for in vivo delivery. Nature 536, 81–85 (2016).
Danino, T. et al. Programmable probiotics for detection of cancer in urine. Sci. Transl. Med. 7, 289ra284 (2015).
Anderson, J. C., Clarke, E. J., Arkin, A. P. & Voigt, C. A. Environmentally controlled invasion of cancer cells by engineered bacteria. J. Mol. Biol. 355, 619–627 (2006).
Low, K. B. et al. Lipid A mutant Salmonella with suppressed virulence and TNFα induction retain tumor-targeting in vivo. Nat. Biotechnol. 17, 37–41 (1999).
Cao, Z., Cheng, S., Wang, X., Pang, Y. & Liu, J. Camouflaging bacteria by wrapping with cell membranes. Nat. Commun. 10, 3452 (2019).
Mimee, M., Tucker, A. C., Voigt, C. A. & Lu, T. K. Programming a human commensal bacterium, bacteroides thetaiotaomicron, to sense and respond to stimuli in the murine gut microbiota. Cell Syst. 1, 62–71 (2015).
Nakatsuji, T. et al. A commensal strain of Staphylococcus epidermidis protects against skin neoplasia. Sci. Adv. 4, eaao4502 (2018).
Dang, L. H., Bettegowda, C., Huso, D. L., Kinzler, K. W. & Vogelstein, B. Combination bacteriolytic therapy for the treatment of experimental tumors. Proc. Natl Acad. Sci. USA 98, 15155–15160 (2001).
Lee, J. W., Chan, C. T. Y., Slomovic, S. & Collins, J. J. Next-generation biocontainment systems for engineered organisms. Nat. Chem. Biol. 14, 530–537 (2018).
Mandell, D. J. et al. Biocontainment of genetically modified organisms by synthetic protein design. Nature 518, 55–60 (2015).
Stirling, F. et al. Rational design of evolutionarily stable microbial kill switches. Mol. Cell 68, 686–697 (2017).
Chan, C. T., Lee, J. W., Cameron, D. E., Bashor, C. J. & Collins, J. J. ‘Deadman’ and ‘Passcode’ microbial kill switches for bacterial containment. Nat. Chem. Biol. 12, 82–86 (2016).
Zhao, M. et al. Tumor-targeting bacterial therapy with amino acid auxotrophs of GFP-expressing Salmonella Typhimurium. Proc. Natl Acad. Sci. USA 102, 755–760 (2005).
Piraner, D. I., Abedi, M. H., Moser, B. A., Lee-Gosselin, A. & Shapiro, M. G. Tunable thermal bioswitches for in vivo control of microbial therapeutics. Nat. Chem. Biol. 13, 75–80 (2017).
Yu, B. et al. Explicit hypoxia targeting with tumor suppression by creating an ‘obligate’ anaerobic Salmonella Typhimurium strain. Sci. Rep. 2, 436 (2012).
Gallagher, R. R., Patel, J. R., Interiano, A. L., Rovner, A. J. & Isaacs, F. J. Multilayered genetic safeguards limit growth of microorganisms to defined environments. Nucleic Acids Res. 43, 1945–1954 (2015).
Huang, S. et al. Coupling spatial segregation with synthetic circuits to control bacterial survival. Mol. Syst. Biol. 12, 859 (2016).
Wang, B., Barahona, M. & Buck, M. A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals. Biosens. Bioelectron. 40, 368–376 (2013).
De Santis, V. & Singer, M. Tissue oxygen tension monitoring of organ perfusion: rationale, methodologies, and literature review. Br. J. Anaesth. 115, 357–365 (2015).
Carreau, A., El Hafny-Rahbi, B., Matejuk, A., Grillon, C. & Kieda, C. Why is the partial oxygen pressure of human tissues a crucial parameter? Small molecules and hypoxia. J. Cell. Mol. Med. 15, 1239–1253 (2011).
Andersen, L. W. et al. Etiology and therapeutic approach to elevated lactate levels. Mayo Clin. Proc. 88, 1127–1140 (2013).
Bakker, J., Nijsten, M. W. & Jansen, T. C. Clinical use of lactate monitoring in critically ill patients. Ann. Intensive Care 3, 12 (2013).
Aoi, W. & Marunaka, Y. Importance of pH homeostasis in metabolic health and diseases: crucial role of membrane proton transport. BioMed. Res. Int. 2014, 598986 (2014).
Crack, J. C. et al. Signal perception by FNR: the role of the iron-sulfur cluster. Biochem. Soc. Trans. 36, 1144–1148 (2008).
Aguilera, L. et al. Dual role of LldR in regulation of the lldPRD operon, involved in l-lactate metabolism in Escherichia coli. J. Bacteriol. 190, 2997–3005 (2008).
Goers, L. et al. Whole-cell Escherichia coli lactate biosensor for monitoring mammalian cell cultures during biopharmaceutical production. Biotechnol. Bioeng. 114, 1290–1300 (2017).
Weghoff, M. C., Bertsch, J. & Muller, V. A novel mode of lactate metabolism in strictly anaerobic bacteria. Environ. Microbiol. 17, 670–677 (2015).
Xu, Y. et al. An acid-tolerance response system protecting exponentially growing Escherichia coli. Nat. Commun. 11, 1496 (2020).
Stirling, F. et al. Synthetic cassettes for pH-mediated sensing, counting, and containment. Cell Rep. 30, 3139–3148 (2020).
Schlundt, A. et al. Structure-function analysis of the DNA-binding domain of a transmembrane transcriptional activator. Sci. Rep. 7, 1051 (2017).
Viala, J. P. et al. Sensing and adaptation to low pH mediated by inducible amino acid decarboxylases in Salmonella. PLoS ONE 6, e22397 (2011).
Lee, Y. H., Kim, J. H., Bang, I. S. & Park, Y. K. The membrane-bound transcriptional regulator CadC is activated by proteolytic cleavage in response to acid stress. J. Bacteriol. 190, 5120–5126 (2008).
Ryan, R. M. et al. Bacterial delivery of a novel cytolysin to hypoxic areas of solid tumors. Gene Ther. 16, 329–339 (2009).
Mengesha, A. et al. Development of a flexible and potent hypoxia-inducible promoter for tumor-targeted gene expression in attenuated Salmonella. Cancer Biol. Ther. 5, 1120–1128 (2006).
Petrova, V., Annicchiarico-Petruzzelli, M., Melino, G. & Amelio, I. The hypoxic tumour microenvironment. Oncogenesis 7, 10 (2018).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Kim, K. et al. A novel balanced-lethal host-vector system based on glmS. PLoS ONE 8, e60511 (2013).
Winslow, M. M. et al. Suppression of lung adenocarcinoma progression by Nkx2-1. Nature 473, 101–104 (2011).
Sonnenborn, U. Escherichia coli strain Nissle 1917—from bench to bedside and back: history of a special Escherichia coli strain with probiotic properties. FEMS Microbiol. Lett. 363, fnw212 (2016).
Wassenaar, T. M. Insights from 100 years of research with probiotic E. coli. Eur. J. Microbiol Immunol. 6, 147–161 (2016).
Sheth, R. U. et al. Spatial metagenomic characterization of microbial biogeography in the gut. Nat. Biotechnol. 37, 877–883 (2019).
Yasuda, K. et al. Biogeography of the intestinal mucosal and lumenal microbiome in the rhesus macaque. Cell Host Microbe 17, 385–391 (2015).
Mark Welch, J. L., Hasegawa, Y., McNulty, N. P., Gordon, J. I. & Borisy, G. G. Spatial organization of a model 15-member human gut microbiota established in gnotobiotic mice. Proc. Natl Acad. Sci. USA 114, E9105–E9114 (2017).
Hohmann, E. L., Oletta, C. A. & Miller, S. I. Evaluation of a phoP/phoQ-deleted, aroA-deleted live oral Salmonella Typhi vaccine strain in human volunteers. Vaccine 14, 19–24 (1996).
Zheng, J. H. et al. Two-step enhanced cancer immunotherapy with engineered Salmonella Typhimurium secreting heterologous flagellin. Sci. Transl. Med. 9, eaak9537 (2017).
Zhao, M. et al. Monotherapy with a tumor-targeting mutant of Salmonella Typhimurium cures orthotopic metastatic mouse models of human prostate cancer. Proc. Natl Acad. Sci. USA 104, 10170–10174 (2007).
Hoffman, R. M. Tumor-targeting Salmonella Typhimurium A1-R: an overview. Methods Mol. Biol. 1409, 1–8 (2016).
Chen, L. Q. et al. Evaluations of tumor acidosis within in vivo tumor models using parametric maps generated with acido CEST MRI. Mol. Imaging Biol. 17, 488–496 (2015).
Romero-Garcia, S., Moreno-Altamirano, M. M., Prado-Garcia, H. & Sanchez-Garcia, F. J. Lactate contribution to the tumor microenvironment: mechanisms, effects on immune cells and therapeutic relevance. Front. Immunol. 7, 52 (2016).
Sonveaux, P. et al. Targeting lactate-fueled respiration selectively kills hypoxic tumor cells in mice. J. Clin. Invest. 118, 3930–3942 (2008).
Harimoto, T. et al. Rapid screening of engineered microbial therapies in a 3D multicellular model. Proc. Natl Acad. Sci. USA 116, 9002–9007 (2019).
Walenta, S., Doetsch, J., Mueller-Klieser, W. & Kunz-Schughart, L. A. Metabolic imaging in multicellular spheroids of oncogene-transfected fibroblasts. J. Histochem. Cytochem. 48, 509–522 (2000).
Riffle, S., Pandey, R. N., Albert, M. & Hegde, R. S. Linking hypoxia, DNA damage and proliferation in multicellular tumor spheroids. BMC Cancer 17, 338 (2017).
Wan, X. et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals. Nat. Chem. Biol. 15, 540–548 (2019).
Chen, Y. et al. Tuning the dynamic range of bacterial promoters regulated by ligand-inducible transcription factors. Nat. Commun. 9, 64 (2018).
Hicks, M., Bachmann, T. T. & Wang, B. Synthetic biology enables programmable cell-based biosensors. ChemPhysChem 21, 132–144 (2020).
Runa, F. et al. Tumor microenvironment heterogeneity: challenges and opportunities. Curr. Mol. Biol. Rep. 3, 218–229 (2017).
Stanta, G. & Bonin, S. Overview on clinical relevance of intra-tumor heterogeneity. Front. Med. 5, 85 (2018).
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Haldimann, A. & Wanner, B. L. Conditional-replication, integration, excision, and retrieval plasmid-host systems for gene structure-function studies of bacteria. J. Bacteriol. 183, 6384–6393 (2001).
Craggs, T. D. Green fluorescent protein: structure, folding and chromophore maturation. Chem. Soc. Rev. 38, 2865–2875 (2009).
Coralli, C., Cemazar, M., Kanthou, C., Tozer, G. M. & Dachs, G. U. Limitations of the reporter green fluorescent protein under simulated tumor conditions. Cancer Res. 61, 4784–4790 (2001).
Misra, T. et al. A genetically encoded biosensor for visualising hypoxia responses in vivo. Biol. Open 6, 296–304 (2017).
Lee, J. Y. et al. A novel chimeric promoter that is highly responsive to hypoxia and metals. Gene Ther. 13, 857–868 (2006).
Leventhal, D. S. et al. Immunotherapy with engineered bacteria by targeting the STING pathway for anti-tumor immunity. Nat. Commun. 11, 2739 (2020).
Prewitt, J. M. & Mendelsohn, M. L. The analysis of cell images. Ann. N. Y. Acad. Sci. 128, 1035–1053 (1966).
Yen, J.-C., Chang, F.-J. & Chang, S. A new criterion for automatic multilevel thresholding. IEEE Trans. Image Process. 4, 370–378 (1995).
Carvajal-Aldaz, D. G. et al. Simultaneous delivery of antibiotics neomycin and ampicillin in drinking water inhibits fermentation of resistant starch in rats. Mol. Nutr. Food Res. 61, 1600609 (2017).
Ho, C. L. et al. Engineered commensal microbes for diet-mediated colorectal-cancer chemoprevention. Nat. Biomed. Eng. 2, 27–37 (2018).
Alon, U. An Introduction to Systems Biology: Design Principles of Biological Circuits 3–19 (Chapman & Hall/CRC, 2007).
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008).
Cookson, N. A. et al. Queueing up for enzymatic processing: correlated signaling through coupled degradation. Mol. Syst. Biol. 7, 561 (2011).
Chen, Y., Kim, J. K., Hirning, A. J., Josic, K. & Bennett, M. R. Synthetic biology. Emergent genetic oscillations in a synthetic microbial consortium. Science 349, 986–989 (2015).
Bernstein, J. A., Khodursky, A. B., Lin, P. H., Lin-Chao, S. & Cohen, S. N. Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays. Proc. Natl Acad. Sci. USA 99, 9697–9702 (2002).
Acknowledgements
We thank J. Zhang and W. Mather for technical assistance and the members of the Danino laboratory for reading the manuscript. This work was supported by the DoD Idea Development Award (LC160314), DoD Era of Hope Scholar Award (BC160541), Honjo International Foundation Scholarship (to T.H.) and NIH F99CA253756 (to T.H.).
Author information
Authors and Affiliations
Contributions
T.C., T.H. and T.D. conceived and designed the study. T.C., T.H., B.K. and T.A. performed in vitro characterization of biosensor. T.C., T.H., K.G., C.C., K.P. and A.N. performed in vivo experiments for this study. T.C., T.H., N.H. and T.D. developed the computational model for this study. T.C., T.H., M.P. and T.D. analysed experimental data. T.C., T.H. and T.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
T.C., T.H. and T.D. have filed a provisional patent application (number 62/930,665) with the US Patent and Trademark Office related to this work.
Additional information
Peer review information Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Computational modelling of biosensors.
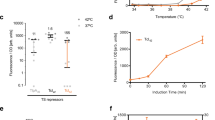
Biosensors are modelled under regulation of transcription activators (FNR, CadC) or repressors (LldR) with varying environmental conditions. (Left) biosensor circuit schematics as shown in Fig. 1 with (middle) modeling in silico predictions compared to (right) in vitro experimental results (n = 3, ±S.E.M). See supplementary materials for detailed equations and parameters used in this study. Lactate and pH biosensor in vitro GFP fluorescent result is normalized by OD. Hypoxia biosensor in vitro GFP fluorescence was measured in an anaerobic chamber and are normalized by data from constitutive promoter pTac.
Extended Data Fig. 2 Computational simulation of three 2-input AND-gate containment circuits.
a, Computational modelling of lactate hypoxia AND gate containment strain growth over time in permissive (high lactate and hypoxia) and non-permissive (none, high lactate or hypoxia only) conditions. b, Growth simulation of pH hypoxia AND gate containment strain in permissive (low pH and hypoxia) and non-permissive (none, low pH or hypoxia only) conditions. c, Growth simulation of lactate pH AND gate containment strain in permissive (high lactate and low pH) and non-permissive (none, high lactate or low pH only) conditions. Lines not shown are covered by overlapping lines.
Supplementary information
Supplementary Information
Supplementary figures, tables and references.
Supplementary Data 1
Relevant gene sequences used in this study.
Supplementary Data 2
Raw fluorescence and OD600 data for Fig. 2b.
Supplementary Data 3
Raw c.f.u. counts and measured weight (in g) for Supplementary Fig. 12.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chien, T., Harimoto, T., Kepecs, B. et al. Enhancing the tropism of bacteria via genetically programmed biosensors. Nat Biomed Eng 6, 94–104 (2022). https://doi.org/10.1038/s41551-021-00772-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-021-00772-3
This article is cited by
-
Promising dawn in tumor microenvironment therapy: engineering oral bacteria
International Journal of Oral Science (2024)
-
Enhancing tumor-specific recognition of programmable synthetic bacterial consortium for precision therapy of colorectal cancer
npj Biofilms and Microbiomes (2024)
-
Reprogramming a Doxycycline-Inducible Gene Switch System for Bacteria-Mediated Cancer Therapy
Molecular Imaging and Biology (2024)
-
Engineered live bacteria as disease detection and diagnosis tools
Journal of Biological Engineering (2023)
-
Bacterial therapies at the interface of synthetic biology and nanomedicine
Nature Reviews Bioengineering (2023)