Abstract
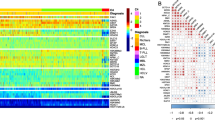
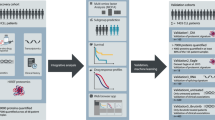
Acute myelogenous leukaemia (AML) is associated with risk factors that are largely unknown and with a heterogeneous response to treatment. Here, we provide a comprehensive quantitative understanding of AML proteomic heterogeneities and hallmarks by using the AML Proteome Atlas, a proteomics database that we have newly derived from MetaGalaxy analyses, for the proteomic profiling of 205 patients with AML and 111 leukaemia cell lines. The analysis of the dataset revealed 154 functional patterns based on common molecular pathways, 11 constellations of correlated functional patterns and 13 signatures that stratify the outcomes of patients. We find limited overlap between proteomics data and both cytogenetics and genetic mutations. Moreover, leukaemia cell lines show limited proteomic similarities with cells from patients with AML, suggesting that a deeper focus on patient-derived samples is needed to gain disease-relevant insights. The AML Proteome Atlas provides a knowledge base for proteomic patterns in AML, a guide to leukaemia cell line selection, and a broadly applicable computational approach for quantifying the heterogeneities of protein expression and proteomic hallmarks in AML.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated and analysed in the study are available on the AML Proteome Atlas: LeukemiaAtlas.org (direct dataset-download at https://www.leukemiaatlas.org/adultaml). Source data for the figures in this study are also provided in .xlsx and .csv formats in figshare at https://figshare.com/s/5ec75fb971747383c0be53 and are accessible directly from the AML Proteome Atlas (https://www.leukemiaatlas.org/adultaml).
Code availability
Access to the entire code used in the MetaGalaxy analysis is listed in Supplementary Table 7. In particular, the Progeny Clustering code is available on the R repository at https://cran.r-project.org/web/packages/progenyClust/index.html, and the MetaGalaxy pipeline is available in a R Shiny portal at https://meta-galaxy-analysis.shinyapps.io/mpa_r/, with a corresponding tutorial and demo files (main.csv and pathway.csv) available in the supplement and online at the AML Proteome Atlas (https://www.leukemiaatlas.org/code).
References
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Cancer Facts and Figures 2018 (American Cancer Society, 2018).
Byrd, J. C. et al. Pretreatment cytogenetic abnormalities are predictive of induction success, cumulative incidence of relapse, and overall survival in adult patients with de novo acute myeloid leukemia: results from Cancer and Leukemia Group B (CALGB 8461). Blood 100, 4325–4336 (2002).
Grimwade, D. et al. The importance of diagnostic cytogenetics on outcome in AML: analysis of 1,612 patients entered into the MRC AML 10 trial. The Medical Research Council Adult and Children’s Leukaemia Working Parties. Blood 92, 2322–2333 (1998).
Slovak, M. L. et al. Karyotypic analysis predicts outcome of preremission and postremission therapy in adult acute myeloid leukemia: a Southwest Oncology Group/Eastern Cooperative Oncology Group Study. Blood 96, 4075–4083 (2000).
Mardis, E. R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 361, 1058–1066 (2009).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Valk, P. J. et al. Prognostically useful gene-expression profiles in acute myeloid leukemia. N. Engl. J. Med. 350, 1617–1628 (2004).
Paweletz, C. P. et al. Reverse phase protein microarrays which capture disease progression show activation of pro-survival pathways at the cancer invasion front. Oncogene 20, 1981–1989 (2001).
Tibes, R. et al. Reverse phase protein array: validation of a novel proteomic technology and utility for analysis of primary leukemia specimens and hematopoietic stem cells. Mol. Cancer Ther. 5, 2512–2521 (2006).
Masuda, M. & Yamada, T. Signaling pathway profiling by reverse-phase protein array for personalized cancer medicine. Biochim. Biophys. Acta 1854, 651–657 (2015).
Byers, L. A. et al. Proteomic profiling identifies dysregulated pathways in small cell lung cancer and novel therapeutic targets including PARP1. Cancer Discov. 2, 798–811 (2012).
Carey, M. S. et al. Functional proteomic analysis of advanced serous ovarian cancer using reverse phase protein array: TGF-beta pathway signaling indicates response to primary chemotherapy. Clin. Cancer Res. 16, 2852–2860 (2010).
Grubb, R. L. et al. Signal pathway profiling of prostate cancer using reverse phase protein arrays. Proteomics 3, 2142–2146 (2003).
Nishizuka, S. et al. Proteomic profiling of the NCI-60 cancer cell lines using new high-density reverse-phase lysate microarrays. Proc. Natl Acad. Sci. USA 100, 14229–14234 (2003).
Gonzalez-Angulo, A. M. et al. Open-label randomized clinical trial of standard neoadjuvant chemotherapy with paclitaxel followed by FEC versus the combination of paclitaxel and everolimus followed by FEC in women with triple receptor-negative breast cancer. Ann. Oncol. 25, 1122–1127 (2014).
Pierobon, M. et al. Pilot phase I/II personalized therapy trial for metastatic colorectal cancer: evaluating the feasibility of protein pathway activation mapping for stratifying patients to therapy with imatinib and panitumumab. J. Proteome Res. 13, 2846–2855 (2014).
Posadas, E. M. et al. A phase II and pharmacodynamic study of gefitinib in patients with refractory or recurrent epithelial ovarian cancer. Cancer 109, 1323–1330 (2007).
Kornblau, S. M. et al. Highly phosphorylated FOXO3A is an adverse prognostic factor in acute myeloid leukemia. Clin. Cancer Res. 16, 1865–1874 (2010).
Kornblau, S. M. et al. Functional proteomic profiling of AML predicts response and survival. Blood 113, 154–164 (2009).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Benito, J. et al. Targeting hypoxia in the leukemia microenvironment. Int. J. Hematol. Oncol. 2, 279–288 (2013).
Hu, C. W., Kornblau, S. M., Slater, J. H. & Qutub, A. A. Progeny clustering: a method to identify biological phenotypes. Sci. Rep. 5, 12894 (2015).
Harris, K. F. et al. Ubiquitin-mediated degradation of active Src tyrosine kinase. Proc. Natl Acad. Sci. USA 96, 13738–13743 (1999).
Kim, N. G. & Gumbiner, B. M. Adhesion to fibronectin regulates Hippo signaling via the FAK–Src–PI3K pathway. J. Cell Biol. 210, 503–515 (2015).
Gjertsen, B. T. et al. Analysis of acute myelogenous leukemia: preparation of samples for genomic and proteomic analyses. J. Hematother. Stem Cell Res. 11, 469–481 (2002).
Rai, A. J. et al. HUPO Plasma Proteome Project specimen collection and handling: towards the standardization of parameters for plasma proteome samples. Proteomics 5, 3262–3277 (2005).
Dvinge, H. et al. Sample processing obscures cancer-specific alterations in leukemic transcriptomes. Proc. Natl Acad. Sci USA 111, 16802–16807 (2014).
Aasebo, E. et al. Freezing effects on the acute myeloid leukemia cell proteome and phosphoproteome revealed using optimal quantitative workflows. J. Proteomics 145, 214–225 (2016).
Lanza, F. et al. Assessment of distribution of CD34 epitope classes in fresh and cryopreserved peripheral blood progenitor cells and acute myeloid leukemic blasts. Haematologica 84, 969–977 (1999).
Xia, Z., Baer, M. R., Block, A. W., Baumann, H. & Wetzler, M. Expression of signal transducers and activators of transcription proteins in acute myeloid leukemia blasts. Cancer Res. 58, 3173–3180 (1998).
Hunyady, B., Krempels, K., Harta, G. & Mezey, E. Immunohistochemical signal amplification by catalyzed reporter deposition and its application in double immunostaining. J. Histochem. Cytochem. 44, 1353–1362 (1996).
Eyre, T. A. et al. The HUGO gene nomenclature database, 2006 updates. Nucleic Acids Res. 34, D319–D321 (2006).
Hermjakob, H. et al. The HUPO PSI’s molecular interaction format–a community standard for the representation of protein interaction data. Nat. Biotechnol. 22, 177–183 (2004).
Jayapandian, M. et al. Michigan Molecular Interactions (MiMI): putting the jigsaw puzzle together. Nucleic Acids Res. 35, D566–D571 (2007).
Akbani, R. et al. A pan-cancer proteomic perspective on The Cancer Genome Atlas. Nat. Commun. 5, 3887 (2014).
Neeley, E. S., Baggerly, K. A. & Kornblau, S. M. Surface adjustment of reverse phase protein arrays using positive control spots. Cancer Inform. 11, 77–86 (2012).
Neeley, E. S., Kornblau, S. M., Coombes, K. R. & Baggerly, K. A. Variable slope normalization of reverse phase protein arrays. Bioinformatics 25, 1384–1389 (2009).
Hu, J. et al. Non-parametric quantification of protein lysate arrays. Bioinformatics 23, 1986–1994 (2007).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Hartigan, J. A. & Wong, M. A. Algorithm AS 136: a k-means clustering algorithm. J. R. Stat. Soc. Ser. C Appl. Stat. 28, 100–108 (1979).
Franceschini, A. et al. STRINGv9. 1: protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2012).
Friedman, J., Hastie, T. & Tibshirani, R. Sparse inverse covariance estimation with the graphical lasso. Biostatistics 9, 432–441 (2008).
Liu, H., Roeer, K. & Wasserman, L. Stability approach to regularization selection (StARS) for high dimensional graphical models. In Proc. 23rd International Conference on Neural Information Processing Systems—Volume 2 (eds Lafferty, J. D. et al) 1432–1440 (Curran Associates, 2010).
Zuo, Y., Cui, Y., Yu, G., Li, R. & Ressom, H. W. Incorporating prior biological knowledge for network-based differential gene expression analysis using differentially weighted graphical LASSO. BMC Bioinformatics 18, 99 (2017).
Sulaimanov, N. & Koeppl, H. Graph reconstruction using covariance-based methods. EURASIP J. Bioinform. Syst. Biol. 2016, 19 (2016).
Hill, S. M. et al. Inferring causal molecular networks: empirical assessment through a community-based effort. Nat. Methods 13, 310–318 (2016).
York, H., Kornblau, S. M. & Qutub, A. A. Network analysis of reverse phase protein expression data: characterizing protein signatures in acute myeloid leukemia cytogenetic categories t(8;21) and inv(16). Proteomics 12, 2084–2093 (2012).
Kornblau, S. M. et al. Proteomic profiling identifies distinct protein patterns in acute myelogenous leukemia CD34+CD38– stem-like cells. PLoS ONE 8, e78453 (2013).
Govaert, G. & Nadif, M. Clustering with block mixture models. Pattern Recog. 36, 463–473 (2003).
Therneau, T., Atkinson, B. & Ripley, B. rpart: Recursive Partitioning and Regression Trees. R version 4.1–10 (2015).
Hu, C. W. et al. Dataset for “A quantitative analysis of heterogeneities and hallmarks in acute myelogenous leukaemia.” f igshare https://figshare.com/s/5ec75fb971747383c0be (2019).
Acknowledgements
This research was funded in part by a translational research grant from the Leukemia and Lymphoma Society to S.M.K., NSF CAREER 1150645, NSF NCS 1533708 and NIH R01 GM106027 grants to A.A.Q., and a HHMI Med-into-Grad fellowship to C.W.H.
Author information
Authors and Affiliations
Contributions
A.A.Q., S.M.K. and C.W.H. conceived and designed the study. Y.Q. performed the experiments. C.W.H., S.Y.Y., A.L., A.Y.R. and K.R.C. performed the computational and statistical analyses. C.W.H., S.M.K. and A.A.Q wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures, tables and software tutorial.
Supplementary Dataset 1
Antibody nomenclature table.
Supplementary Dataset 2
Correlation between post-translational modifications and total protein expression levels.
Supplementary Dataset 3
Functional group memberships.
Supplementary Dataset 4
Constellation memberships for cell lines.
Supplementary Dataset 5
Functional pattern memberships for cell lines.
Supplementary Dataset 6
Demographic information of patients overall and in each signature (S1–S13).
Supplementary Dataset 7
Method implementation and parameter specification.
Supplementary Dataset 8
Stability scores from Progeny Clustering for co-clustering.
Supplementary Dataset 9
Demo. 1
Supplementary Dataset 10
Demo. 2
Rights and permissions
About this article
Cite this article
Hu, C.W., Qiu, Y., Ligeralde, A. et al. A quantitative analysis of heterogeneities and hallmarks in acute myelogenous leukaemia. Nat Biomed Eng 3, 889–901 (2019). https://doi.org/10.1038/s41551-019-0387-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-019-0387-2
This article is cited by
-
Proteomics for optimizing therapy in acute myeloid leukemia: venetoclax plus hypomethylating agents versus conventional chemotherapy
Leukemia (2024)
-
Proteomic profiling based classification of CLL provides prognostication for modern therapy and identifies novel therapeutic targets
Blood Cancer Journal (2022)
-
Loss of H3K27 methylation identifies poor outcomes in adult-onset acute leukemia
Clinical Epigenetics (2021)
-
A clinical transcriptome approach to patient stratification and therapy selection in acute myeloid leukemia
Nature Communications (2021)
-
Early stratification of radiotherapy response by activatable inflammation magnetic resonance imaging
Nature Communications (2020)