Abstract
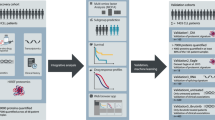
The chronic lymphocytic leukemia (CLL) armamentarium has evolved significantly, with novel therapies that inhibit Bruton Tyrosine Kinase, PI3K delta and/or the BCL2 protein improving outcomes. Still, the clinical course of CLL patients is highly variable and most previously recognized prognostic features lack the capacity to predict response to modern treatments indicating the need for new prognostic markers. In this study, we identified four epigenetically distinct proteomic signatures of a large cohort of CLL and related diseases derived samples (n = 871) using reverse phase protein array technology. These signatures are associated with clinical features including age, cytogenetic abnormalities [trisomy 12, del(13q) and del(17p)], immunoglobulin heavy-chain locus (IGHV) mutational load, ZAP-70 status, Binet and Rai staging as well as with the outcome measures of time to treatment and overall survival. Protein signature membership was identified as predictive marker for overall survival regardless of other clinical features. Among the analyzed epigenetic proteins, EZH2, HDAC6, and loss of H3K27me3 levels were the most independently associated with poor survival. These findings demonstrate that proteomic based epigenetic biomarkers can be used to better classify CLL patients and provide therapeutic guidance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rozman C, Montserrat E. Chronic lymphocytic leukemia. N. Engl J Med. 1995;333:1052–7.
Jain N, O’Brien S. Initial treatment of CLL: integrating biology and functional status. Blood. 2015;126:463–70.
Wierda WG, O’Brien S, Wang X, Faderl S, Ferrajoli A, Do KA, et al. Multivariable model for time to first treatment in patients with chronic lymphocytic leukemia. J Clin Oncol: Off J Am Soc Clin Oncol. 2011;29:4088–95.
Condoluci A, Terzi di Bergamo L, Langerbeins P, Hoechstetter MA, Herling CD, De Paoli L, et al. International prognostic score for asymptomatic early-stage chronic lymphocytic leukemia. Blood. 2020;135:1859–69.
Kanduri M, Cahill N, Göransson H, Enström C, Ryan F, Isaksson A, et al. Differential genome-wide array-based methylation profiles in prognostic subsets of chronic lymphocytic leukemia. Blood. 2010;115:296–305.
Pei L, Choi JH, Liu J, Lee EJ, McCarthy B, Wilson JM, et al. Genome-wide DNA methylation analysis reveals novel epigenetic changes in chronic lymphocytic leukemia. Epigenetics. 2012;7:567–78.
Landau DA, Clement K, Ziller MJ, Boyle P, Fan J, Gu H, et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer cell. 2014;26:813–25.
Cahill N, Kanduri M, Mansouri L, Sundstrom C, Rosenquist R, Ryan F, et al. 450K-array analysis of chronic lymphocytic leukemia cells reveals global DNA methylation to be relatively stable over time and similar in resting and proliferative compartments. Leukemia. 2013;27:150–8.
Kulis M, Heath S, Bibikova M, Queirós AC, Navarro A, Clot G, et al. Epigenomic analysis detects widespread gene-body DNA hypomethylation in chronic lymphocytic leukemia. Nat Genet. 2012;44:1236–42.
Queiros AC, Villamor N, Clot G, Martinez-Trillos A, Kulis M, Navarro A, et al. A B-cell epigenetic signature defines three biologic subgroups of chronic lymphocytic leukemia with clinical impact. Leukemia. 2015;29:598–605.
Smith EN, Ghia EM, DeBoever CM, Rassenti LZ, Jepsen K, Yoon K, et al. Genetic and epigenetic profiling of CLL disease progression reveals limited somatic evolution and suggests a relationship to memory-cell development. Blood Cancer J. 2015;5:e303.
Papakonstantinou N, Ntoufa S, Chartomatsidou E, Kotta K, Agathangelidis A, Giassafaki L, et al. The histone methyltransferase EZH2 as a novel prosurvival factor in clinically aggressive chronic lymphocytic leukemia. Oncotarget. 2016;7:35946–59.
Rabello DdA, Lucena-Araujo AR, Alves-Silva JCR, Souza dEVBA, de Vasconcellos MC, de Oliveira FM, et al. Overexpression of EZH2 associates with a poor prognosis in chronic lymphocytic leukemia. Blood Cells Mol Dis. 2015;54:97–102.
Mansouri L, Wierzbinska JA, Plass C, Rosenquist R. Epigenetic deregulation in chronic lymphocytic leukemia: Clinical and biological impact. Semin Cancer Biol. 2018;51:1–11.
van Dijk AD, Hu CW, de Bont, ESJM, Qiu Y, Hoff FW, Yoo SY, et al. Histone modification patterns using RPPA-based profiling predict outcome in acute myeloid leukemia patients. Proteomics. 2018;18:e1700379.
van Dijk AD, Hoff FW, Qiu Y, de Bont ES, Bruggeman SWM, Horton TM, et al. Loss of H3K27 methylation identifies poor outcome in adult-onset acute myeloid leukemia. Blood. 2020;136:24.
Tibes R, Qiu Y, Lu Y, Hennessy B, Andreeff M, Mills GB, et al. Reverse phase protein array: validation of a novel proteomic technology and utility for analysis of primary leukemia specimens and hematopoietic stem cells. Mol cancer therapeutics. 2006;5:2512–21.
Kornblau SM, Coombes KR. Use of reverse phase protein microarrays to study protein expression in leukemia: technical and methodological lessons learned. Methods Mol Biol. 2011;785:141–55.
Kornblau SM, Tibes R, Qiu YH, Chen W, Kantarjian HM, Andreeff M, et al. Functional proteomic profiling of AML predicts response and survival. Blood. 2009;113:154–64.
Hu CW, Kornblau SM, Slater JH, Qutub AA. Progeny clustering: a method to Identify Biological Phenotypes. Sci Rep. 2015. https://doi.org/10.1038/srep12894.
Hartigan JA, Wong MAA. K-Means clustering algorithm. J R Stat Soc: Ser C (Appl Stat). 1979;28:100–8.
Therneau PM. Modeling survival data: extending the Cox model. Springer; 2000.
Swerdlow SH, WHO Classification of tumors of haematopoietic and lymphoid tissues.4th ed. IARC; 2008.
Scarfò L, Ferreri AJM, Ghia P. Chronic lymphocytic leukaemia. Crit Rev Oncol / Hematol. 2016;104:169–82.
Kipps TJ, Fraser G, Coutre SE, Brown JR, Barrientos JC, Barr PM, et al. Long-Term Studies Assessing Outcomes of Ibrutinib Therapy in Patients With Del(11q) Chronic Lymphocytic Leukemia. Clin Lymphoma Myeloma Leuk. 2019;19:715–22.e6.
Burger JA, Barr PM, Robak T, Owen C, Ghia P, Tedeschi A, et al. Long-term efficacy and safety of first-line ibrutinib treatment for patients with CLL/SLL: 5 years of follow-up from the phase 3 RESONATE-2 study. Leukemia 2020;34:787–98.
van Dijk AD, Hoff FW, Qiu YH, Chandra J, Jabbour E, de Bont, ESJM, et al. Loss of H3K27 methylation identifies poor outcomes in adult-onset acute leukemia. Clin Epigenet. 2021. https://doi.org/10.1186/s13148-021-01011-x.
Li Y, Schulz VP, Deng C, Li G, Shen Y, Tusi BK, et al. Setd1a and NURF mediate chromatin dynamics and gene regulation during erythroid lineage commitment and differentiation. Nucleic Acids Res. 2016;44:7173–88.
Li BE, Gan T, Ernst P, Meyerson M, Rabbitts TH. Distinct pathways regulated by menin and by MLL1 in hematopoietic stem cells and developing B cells. Blood. 2013;122:2039–46.
Tusi BK, Deng C, Salz T, Zeumer L, Li Y, So CWE, et al. Setd1a regulates progenitor B-cell-to-precursor B-cell development through histone H3 lysine 4 trimethylation and Ig heavy-chain rearrangement. FASEB J. 2015;29:1505–15.
Yang W, Ernst P. Distinct functions of histone H3, lysine 4 methyltransferases in normal and malignant hematopoiesis. Curr Opin Hematol. 2017;24:322–8.
Su IH, Basavaraj A, Krutchinsky AN, Hobert O, Ullrich A, Chait BT, et al. Ezh2 controls B cell development through histone H3 methylation and Igh rearrangement. Nat Immunol. 2003;4:124–31.
Zhou M. Small-Molecule Modulators of Methyl-Lysine Binding for the CBX7 Chromodomain. Chem Biol. 2015;22:161–8.
Maharaj K, Powers JJ, Achille A, Deng S, Fonseca R, Pabon-Saldana M, et al. Silencing of HDAC6 as a therapeutic target in chronic lymphocytic leukemia. Blood Adv. 2018;2:3012–24.
Fiskus W, Buckley K, Rao R, Mandawat A, Yang Y, Joshi R, et al. Panobinostat treatment depletes EZH2 and DNMT1 levels and enhances decitabine mediated de-repression of JunB and loss of survival of human acute leukemia cells. Cancer Biol Ther. 2009;8:939–50.
Fiskus W, Pranpat M, Balasis M, Herger B, Rao R, Chinnaiyan A, et al. Histone deacetylase inhibitors deplete enhancer of zeste 2 and associated polycomb repressive complex 2 proteins in human acute leukemia cells. Mol cancer therapeutics. 2006;5:3096–104.
Lee DH, Kim GW, Kwon SH. The HDAC6-selective inhibitor is effective against non-Hodgkin lymphoma and synergizes with ibrutinib in follicular lymphoma. Mol Carcinog. 2019;58:944–56.
Cahill N, Rosenquist R. Uncovering the DNA methylome in chronic lymphocytic leukemia. Epigenetics. 2013;8:138–48.
Author information
Authors and Affiliations
Contributions
ADvD, TLG, JWL, ESJMdB, and SMK designed and supervised the study. SMK, JAB, and WW collected data, ADvD, TLG, FWH, ET, KR, PPR, and YHQ performed research. ADvD and SMK analyzed data and wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
van Dijk, A.D., Griffen, T.L., Qiu, Y.H. et al. RPPA-based proteomics recognizes distinct epigenetic signatures in chronic lymphocytic leukemia with clinical consequences. Leukemia 36, 712–722 (2022). https://doi.org/10.1038/s41375-021-01438-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-021-01438-4
This article is cited by
-
Proteomics for optimizing therapy in acute myeloid leukemia: venetoclax plus hypomethylating agents versus conventional chemotherapy
Leukemia (2024)
-
Proteomic profiling based classification of CLL provides prognostication for modern therapy and identifies novel therapeutic targets
Blood Cancer Journal (2022)
-
Proteogenomics refines the molecular classification of chronic lymphocytic leukemia
Nature Communications (2022)