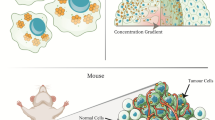
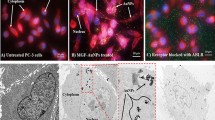
Abstract
Phosphatase and tensin homologue deleted on chromosome 10 (PTEN) is a well-characterized tumour-suppressor gene that is lost or mutated in about half of metastatic castration-resistant prostate cancers and in many other human cancers. The restoration of functional PTEN as a treatment for prostate cancer has, however, proven difficult. Here, we show that PTEN messenger RNA (mRNA) can be reintroduced into PTEN-null prostate cancer cells in vitro and in vivo via its encapsulation in polymer–lipid hybrid nanoparticles coated with a polyethylene glycol shell. The nanoparticles are stable in serum, elicit low toxicity and enable high PTEN mRNA transfection in prostate cancer cells. Moreover, significant inhibition of tumour growth is achieved when delivered systemically in multiple mouse models of prostate cancer. We also show that the restoration of PTEN function in PTEN-null prostate cancer cells inhibits the phosphatidylinositol 3-kinase (PI3K)–AKT pathway and enhances apoptosis. Our findings provide proof-of-principle evidence of the restoration of mRNA-based tumour suppression in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the paper and its Supplementary.
Change history
23 November 2018
The authors wish to add the following sentence into the ‘Competing interests’ section of this Article: “P.W.K. has investment interest in Context Therapeutics LLC, DRGT, Placon, Seer Biosciences and Tarveda Therapeutics, is a company board member for Context Therapeutics LLC, is a consultant and scientific advisory board member for BIND Biosciences, Inc., BN Immunotherapeutics, DRGT, GE Healthcare, Janssen, Metamark, New England Research Institutes, Inc., OncoCellMDX, Progenity, Sanofi, Seer Biosciences, Tarveda Therapeutics and Thermo Fisher, and serves on data safety monitoring boards for Genentech/Roche and Merck.” This has now been included.
References
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nat. Rev. Mol. Cell Biol. 13, 283–296 (2012).
McCall, P., Witton, C. J., Grimsley, S., Nielsen, K. V. & Edwards, J. Is PTEN loss associated with clinical outcome measures in human prostate cancer? Br. J. Cancer 99, 1296–1301 (2008).
Yoshimoto, M. et al. Interphase FISH analysis of PTEN in histologic sections shows genomic deletions in 68% of primary prostate cancer and 23% of high-grade prostatic intra-epithelial neoplasias. Cancer Genet. Cytogenet. 169, 128–137 (2006).
Han, B. et al. Fluorescence in situ hybridization study shows association of PTEN deletion with ERG rearrangement during prostate cancer progression. Mod. Pathol. 22, 1083–1093 (2009).
Verhagen, P. C. et al. The PTEN gene in locally progressive prostate cancer is preferentially inactivated by bi-allelic gene deletion. J. Pathol. 208, 699–707 (2006).
Yoshimoto, M. et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br. J. Cancer. 97, 678–685 (2007).
Sircar, K. et al. PTEN genomic deletion is associated with p-Akt and AR signalling in poorer outcome, hormone refractory prostate cancer. J. Pathol. 218, 505–513 (2009).
Schmitz, M. et al. Complete loss of PTEN expression as a possible early prognostic marker for prostate cancer metastasis. Int. J. Cancer 120, 1284–1292 (2007).
Lotan, T. L. et al. PTEN protein loss by immunostaining: analytic validation and prognostic indicator for a high risk surgical cohort of prostate cancer patients. Clin. Cancer Res. 17, 6563–6573 (2011).
Stambolic, V. et al. Negative regulation of PKB/Akt-dependent cell survival by the tumour suppressor PTEN. Cell 95, 29–39 (1998).
Furnari, F. B., Lin, H., Huang, H. S. & Cavenee, W. K. Growth suppression of glioma cells by PTEN requires a functional phosphatase catalytic domain. Proc. Natl Acad. Sci. USA 94, 12479–12484 (1997).
Sun, H. et al. PTEN modulates cell cycle progression and cell survival by regulating phosphatidylinositol 3,4,5,-trisphosphate and Akt/protein kinase B signaling pathway. Proc. Natl Acad. Sci. USA 96, 6199–6204 (1999).
Suzuki, A. et al. High cancer susceptibility and embryonic lethality associated with mutation of the PTEN tumour suppressor gene in mice. Curr. Biol. 8, 1169–1178 (1998).
Maehama, T. & Dixon, J. E. The tumour suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J. Biol. Chem. 273, 13375–13378 (1998).
Engelman, J. A., Luo, J. & Cantley, L. C. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat. Rev. Genet. 7, 606–619 (2006).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Grasso, C. S. et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature 487, 239–243 (2012).
Backman, S. A. et al. Deletion of Pten in mouse brain causes seizures, ataxia and defects in soma size resembling Lhermitte-Duclos disease. Nat. Genet. 29, 396–403 (2001).
Liliental, J. et al. Genetic deletion of the Pten tumour suppressor gene promotes cell motility by activation of Rac1 and Cdc42 GTPases. Curr. Biol. 10, 401–404 (2000).
Tamura, M. et al. Inhibition of cell migration, spreading, and focal adhesions by tumour suppressor PTEN. Science 280, 1614–1617 (1998).
Hamada, K. et al. The PTEN/PI3K pathway governs normal vascular development and tumour angiogenesis. Genes Dev. 19, 2054–2065 (2005).
Jiang, B. H. & Liu, L. Z. PI3K/PTEN signaling in angiogenesis and tumourigenesis. Adv. Cancer Res. 102, 19–65 (2009).
Yin, H. et al. Non-viral vectors for gene-based therapy. Nat. Rev. Genet. 15, 541–555 (2014).
Quabius, E. S. & Krupp, G. Synthetic mRNAs for manipulating cellular phenotypes: an overview. Nat. Biotechnol. 32, 229–235 (2015).
Lee, J., Boczkowski, D. & Nair, S. Programming human dendritic cells with mRNA. Methods Mol. Biol. 969, 111–125 (2013).
Yamamoto, A., Kormann, M., Rosenecker, J. & Rudolph, C. Current prospects for mRNA gene delivery. Eur. J. Pharm. Biopharm. 71, 484–489 (2009).
Leonhardt, C. et al. Single-cell mRNA transfection studies: delivery, kinetics and statistics by numbers. Nanomedicine 10, 679–688 (2014).
Ligon, T. S., Leonhardt, C. & Radler, J. O. Multi-level kinetic model of mRNA delivery via transfection of lipoplexes. PLoS ONE 9, e107148 (2014).
Islam, M. A. et al. Biomaterials for mRNA delivery. Biomater. Sci. 3, 1519–1533 (2015).
Davis, M. E. et al. Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature 464, 1067–1070 (2010).
Zuckerman, J. E. et al. Correlating animal and human phase Ia/Ib clinical data with CALAA-01, a targeted, polymer-based nanoparticle containing siRNA. Proc. Natl Acad. Sci. USA 111, 11449–11454 (2014).
Tabernero, J. et al. First-in-humans trial of an RNA interference therapeutic targeting VEGF and KSP in cancer patients with liver involvement. Cancer Discov. 3, 406–417 (2013).
Strumberg, D. et al. Phase I clinical development of Atu027, a siRNA formulation targeting PKN3 in patients with advanced solid tumours. Int. J. Clin. Pharmacol. Ther. 50, 76 (2012).
Schultheis, B. et al. First-in-human phase I study of the liposomal RNA interference therapeutic Atu027 in patients with advanced solid tumours. J. Clin. Oncol. 32, 4141–4148 (2014).
Tolcher, A. W. et al. A phase 1 study of the BCL2-targeted deoxyribonucleic acid inhibitor (DNAi) PNT2258 in patients with advanced solid tumours. Cancer Chemother. Pharmacol. 73, 363–371 (2014).
Akinc, A. et al. A combinatorial library of lipid-like materials for delivery of RNAi therapeutics. Nat. Biotechnol. 26, 561–569 (2008).
Whitehead, K. A. et al. Degradable lipid nanoparticles with predictable in vivo siRNA delivery activity. Nat. Commun. 5, 4277 (2014).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nat. Rev. Drug Discov. 8, 129–138 (2009).
Zuckerman, J. E. & Davis, M. E. Clinical experiences with systemically administered siRNA-based therapeutics in cancer. Nat. Rev. Drug Discov. 14, 843–856 (2015).
Conde, J., Oliva, N., Atilano, M., Song, H. S. & Artzi, N. Self-assembled RNA-triple-helix hydrogel scaffold for microRNA modulation in the tumour microenvironment. Nat. Mater. 15, 353–363 (2016).
Kauffman, K. J. et al. Optimization of lipid nanoparticle formulations for mRNA delivery in vivo with fractional factorial and definitive screening designs. Nano Lett. 15, 7300–7306 (2015).
Kormann, M. S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 29, 154–157 (2011).
Li, B. et al. An orthogonal array optimization of lipid-like nanoparticles for mRNA delivery in vivo. Nano Lett. 15, 8099–8107 (2015).
Zhu, X. et al. Long-circulating siRNA nanoparticles for validating Prohibitin1-targeted non-small cell lung cancer treatment. Proc. Natl Acad. Sci. USA 112, 7779–7784 (2015).
Islam, M. A. et al. The role of osmotic polysorbitol-based transporter in RNAi silencing via caveolae-mediated endocytosis and COX-2 expression. Biomaterials 33, 8868–8880 (2012).
Islam, M. A. et al. Accelerated gene transfer through a polysorbitol-based transporter mechanism. Biomaterials 32, 9908–9924 (2011).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Wang, Y. et al. Systemic delivery of modified mRNA encoding herpes simplex virus 1 thymidine kinase for targeted cancer gene therapy. Mol. Ther. 21, 358–367 (2013).
Luo, X. et al. Dual-functional lipid-like nanoparticles for delivery of mRNA and MRI contrast agents. Nanoscale 9, 1575–1579 (2017).
Huang, H. et al. PTEN induces chemosensitivity in PTEN-mutated prostate cancer cells by suppression of Bcl-2 expression. J. Biol. Chem. 276, 38830–38836 (2001).
Sturge, J., Caley, M. P. & Waxman, J. Bone metastasis in prostate cancer: emerging therapeutic strategies. Nat. Rev. Clin. Oncol. 8, 357–368 (2011).
Smukste, I. & Stockwell, B. R. Restoring functions of tumour suppressors with small molecules. Cancer Cell 4, 419–420 (2003).
Guo, X. E., Ngo, B., Modrek, A. S. & Lee, W. H. Targeting tumour suppressor networks for cancer therapeutics. Curr. Drug Targets 15, 2–16 (2014).
Bettinger, T., Carlisle, R. C., Read, M. L., Ogris, M. & Seymour, L. W. Peptide-mediated RNA delivery: a novel approach for enhanced transfection of primary and post-mitotic cells. Nucleic Acids Res. 29, 3882–3891 (2001).
Rejman, J., Tavernier, G., Bavarsad, N., Demeester, J. & De Smedt, S. C. mRNA transfection of cervical carcinoma and mesenchymal stem cells mediated by cationic carriers. J. Control. Release 147, 385–391 (2010).
Zou, S., Scarfo, K., Nantz, M. H. & Hecker, J. G. Lipid-mediated delivery of RNA is more efficient than delivery of DNA in non-dividing cells. Int. J. Pharm. 389, 232–243 (2010).
Read, M. L. et al. A versatile reducible polycation-based system for efficient delivery of a broad range of nucleic acids. Nucleic Acids Res. 33, e86 (2005).
Kong, G., Braun, R. D. & Dewhirst, M. W. Hyperthermia enables tumour-specific nanoparticle delivery: effect of particle size. Cancer Res. 60, 4440–4445 (2000).
Alexis, F., Pridgen, E., Molnar, L. K. & Farokhzad, O. C. Factors affecting the clearance and biodistribution of polymeric nanoparticles. Mol. Pharm. 5, 505–515 (2008).
Shi, J., Kantoff, P. W., Wooster, R. & Farokhzad, O. C. Cancer nanomedicine: progress, challenges and opportunities. Nat. Rev. Cancer 17, 20–37 (2017).
Bertrand, N., Wu, J., Xu, X., Kamaly, N. & Farokhzad, O. C. Cancer nanotechnology: the impact of passive and active targeting in the era of modern cancer biology. Adv. Drug Deliv. Rev. 66, 2–25 (2014).
Knop, K., Hoogenboom, R., Fischer, D. & Schubert, U. S. Poly(ethylene glycol) in drug delivery: pros and cons as well as potential alternatives. Angew. Chem. Int. Ed. Engl. 49, 6288–6308 (2010).
Guo, X. & Huang, L. Recent advances in nonviral vectors for gene delivery. Acc. Chem. Res. 45, 971–979 (2012).
Liu, H. et al. Structure-based programming of lymph-node targeting in molecular vaccines. Nature 507, 519–522 (2014).
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1947 (1997).
Steck, P. A. et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat. Genet. 15, 356–362 (1997).
Di Cristofano, A., De Acetis, M., Koff, A., Cordon-Cardo, C. & Pandolfi, P. P. Pten and p27KIP1 cooperate in prostate cancer tumour suppression in the mouse. Nat. Genet. 27, 222–224 (2001).
Hopkins, B. D. et al. A secreted PTEN phosphatase that enters cells to alter signaling and survival. Science 341, 399–402 (2013).
Masson, G. R., Perisic, O., Burke, J. E. & Williams, R. L. The intrinsically disordered tails of PTEN and PTEN-L have distinct roles in regulating substrate specificity and membrane activity. Biochem. J. 473, 135–144 (2016).
Juric, D. et al. Convergent loss of PTEN leads to clinical resistance to a PI(3)Kalpha inhibitor. Nature 518, 240–244 (2015).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov 6, 202–216 (2016).
Campeau, E. et al. A versatile viral system for expression and depletion of proteins in mammalian cells. PLoS ONE 4, e6529 (2009).
Ramaswamy, S. et al. Regulation of G1 progression by the PTEN tumour suppressor protein is linked to inhibition of the phosphatidylinositol 3-kinase/Akt pathway. Proc. Natl Acad. Sci. USA 96, 2110–2115 (1999).
Xu, X. et al. Enhancing tumour cell response to chemotherapy through nanoparticle-mediated codelivery of siRNA and cisplatin prodrug. Proc. Natl Acad. Sci. USA 110, 18638–18643 (2013).
Cox, T. R. et al. The hypoxic cancer secretome induces pre-metastatic bone lesions through lysyl oxidase. Nature 522, 106–110 (2015).
Krzywinski, M. & Altman, N. Points of significance: nonparametric tests. Nat. Methods 11, 467–468 (2014).
Tammela, T. et al. A Wnt-producing niche drives proliferative potential and progression in lung adenocarcinoma. Nature 545, 355–359 (2017).
Acknowledgements
This work was in part supported by the Prostate Cancer Foundation (PCF) Young Investigator Award (J.S.); the David Koch-PCF Award in Nanotherapeutics (O.C.F., R.L. and P.W.K.); the US National Institutes of Health (NIH) grants CA200900 (J.S. and B.R.Z.), HL127464 (O.C.F.) and R00CA160350 (J.S.); the National Research Foundation of Korea (K1A1A2048701) (O.C.F.); and the DoD Prostate Cancer Research Program Postdoctoral Training Award (W81XWH-14-1-0268) (Y.X.). The authors would also like to thank E. Reesor, S. Guillemette, J. Rice, Y. Li, M. Zaffagni, D. Bielenberg and J. Wang for their assistance. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
M.A.I., Y.X., O.C.F., B.R.Z. and J.S. conceived the idea, designed the study and directed the project. M.A.I., Y.X. and W.T. performed all the experiments and analysed data. W.T., J.M.U., K.Z. and G.Y.L. assisted with the metastatic and orthotopic PCa experiments in vivo. M.L., D.A., J.T.O. and W.C. helped in nanoparticle preparation and experimental assays. R.L. provided reagents and conceptual advice. W.T., H.Z., M.Y., M.D., M.M., P.W.K. provided technical support and corrections of manuscript. M.A.I., Y.X. and W.T. wrote the manuscript and revised according to the comments of R.L., P.W.K., O.C.F., B.R.Z. and J.S.
Corresponding authors
Ethics declarations
Competing interests
O.C.F. and R.L. declare financial interests in Selecta Biosciences, Tarveda Therapeutics, Placon Therapeutics and Seer. R.L. declares financial interests in Moderna Therapeutics. P.W.K. has investment interest in Context Therapeutics LLC, DRGT, Placon, Seer Biosciences and Tarveda Therapeutics, is a company board member for Context Therapeutics LLC, is a consultant and scientific advisory board member for BIND Biosciences, Inc., BN Immunotherapeutics, DRGT, GE Healthcare, Janssen, Metamark, New England Research Institutes, Inc., OncoCellMDX, Progenity, Sanofi, Seer Biosciences, Tarveda Therapeutics and Thermo Fisher, and serves on data safety monitoring boards for Genentech/Roche and Merck.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures.
Rights and permissions
About this article
Cite this article
Islam, M.A., Xu, Y., Tao, W. et al. Restoration of tumour-growth suppression in vivo via systemic nanoparticle-mediated delivery of PTEN mRNA. Nat Biomed Eng 2, 850–864 (2018). https://doi.org/10.1038/s41551-018-0284-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-018-0284-0
This article is cited by
-
Synergy and Coordination Between Biomimetic Nanoparticles and Biological Cells/Tissues/Organs/Systems: Applications in Nanomedicine and Prospect
Biomedical Materials & Devices (2024)
-
mRNA nanodelivery systems: targeting strategies and administration routes
Biomaterials Research (2023)
-
CREBZF mRNA nanoparticles suppress breast cancer progression through a positive feedback loop boosted by circPAPD4
Journal of Experimental & Clinical Cancer Research (2023)
-
mRNA-based cancer therapeutics
Nature Reviews Cancer (2023)
-
Controlling the biodistribution and clearance of nanomedicines
Nature Reviews Bioengineering (2023)