Abstract
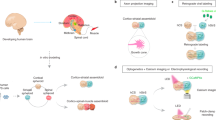
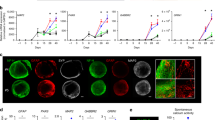
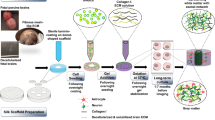
Understanding neurological diseases requires tractable genetic systems, and engineered three-dimensional (3D) neural tissues are an attractive choice. Yet how the cellular transcriptomic profiles in these tissues are affected by the encapsulating materials and are related to the human brain transcriptome is not well understood. Here, we report the characterization of the effects of different culturing conditions on the transcriptomic profiles of induced neuronal cells and developed a method for the rapid generation of 3D co-cultures of neuronal and astrocytic cells from the same pool of human embryonic stem cells. By comparing the gene-expression profiles of neuronal cells in culture conditions relevant to the developing human brain, we found that modifying the degree of crosslinking of composite hydrogels can tune expression patterns so that they correlate with those of specific brain regions and developmental stages. Moreover, single-cell-sequencing results showed that our engineered tissues recapitulate transcriptional patterns of cell types in the human brain. Analyses of culturing conditions will inform the development of 3D neural tissues for use as tractable models of brain diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
25 June 2018
In the version of this Article originally published, the acronym ‘CHs’ was not defined; the acronym should have been included in the sentence beginning “The effects of the addition of hyaluronic acid (HA) and the formation of composite hydrogels (CHs) of Matrigel and alginate...” This has now been corrected.
References
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Quadrato, G., Brown, J. & Arlotta, P. The promises and challenges of human brain organoids as models of neuropsychiatric disease. Nat. Med. 22, 1220–1228 (2016).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
McCarro, S. A., Feng, G. P. & Hyman, S. E. Genome-scale neurogenetics: methodology and meaning. Nat. Neurosci. 17, 756–763 (2014).
Gandhi, S. & Wood, N. W. Genome-wide association studies: the key to unlocking neurodegeneration?. Nat. Neurosci. 13, 789–794 (2010).
Choi, S. H. et al. A three-dimensional human neural cell culture model of Alzheimer’s disease. Nature 515, 274–278 (2014).
Tang-Schomer, M. D. et al. Bioengineered functional brain-like cortical tissue. Proc. Natl Acad. Sci. USA 111, 13811–13816 (2014).
Schwartz, M. P. et al. Human pluripotent stem cell-derived neural constructs for predicting neural toxicity. Proc. Natl Acad. Sci. USA 112, 12516–12521 (2015).
Kim, S. H. et al. Anisotropically organized three-dimensional culture platform for reconstruction of a hippocampal neural network. Nat. Commun. 8, 14346 (2017).
Grealish, S. et al. Human ESC-derived dopamine neurons show similar preclinical efficacy and potency to fetal neurons when grafted in a rat model of Parkinson’s disease. Cell Stem Cell 15, 653–665 (2014).
Dimos, J. T. et al. Induced pluripotent stem cells generated from patients with ALS can be differentiated into motor neurons. Science 321, 1218–1221 (2008).
Pasca, A. M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Qian, X. Y. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Kraehenbuehl, T. P., Langer, R. & Ferreira, L. S. Three-dimensional biomaterials for the study of human pluripotent stem cells. Nat. Methods 8, 731–736 (2011).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53 (2017).
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Busskamp, V. et al. Rapid neurogenesis through transcriptional activation in human stem cells. Mol. Syst. Biol. https://doi.org/10.15252/msb.20145508 (2014).
Zhang, Y. S. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Pang, Z. P. P. et al. Induction of human neuronal cells by defined transcription factors. Nature 476, 220–223 (2011).
Chanda, S. et al. Generation of induced neuronal cells by the single reprogramming factor ASCL1. Stem Cell Reports 3, 282–296 (2014).
Lam, R. S., Töpfer, F. M., Wood, P. G., Busskamp, V. & Bamberg, E. Functional maturation of human stem cell-derived neurons in long-term cultures. PLoS ONE 12, e0169506 (2017).
Yi, F. et al. Autism-associated SHANK3 haploinsufficiency causes I h channelopathy in human neurons. Science 352, aaf2669 (2016).
Huang, Y.-W. A., Zhou, B., Wernig, M. & Südhof, T. C. ApoE2, ApoE3, and ApoE4 differentially stimulate APP transcription and Aβ secretion. Cell 168, 427–441.e421 (2017).
Carlson, A. L. et al. Generation and transplantation of reprogrammed human neurons in the brain using 3D microtopographic scaffolds. Nat. Commun. 7, 10862 (2016).
Lau, L. W., Cua, R., Keough, M. B., Haylock-Jacobs, S. & Yong, V. W. Pathophysiology of the brain extracellular matrix: a new target for remyelination. Nat. Rev. Neurosci. 14, 722–729 (2013).
Tang, X. et al. Astroglial cells regulate the developmental timeline of human neurons differentiated from induced pluripotent stem cells. Stem Cell Res. 11, 743–757 (2013).
Margolis, R. U., Margolis, R. K., Chang, L. B. & Preti, C. Glycosaminoglycans of brain during development. Biochemistry 14, 85–88 (1975).
Bozza, A. et al. Neural differentiation of pluripotent cells in 3D alginate-based cultures. Biomaterials 35, 4636–4645 (2014).
Brannvall, K. et al. Enhanced neuronal differentiation in a three-dimensional collagen–hyaluronan matrix. J. Neurosci. Res. 85, 2138–2146 (2007).
Seidlits, S. K. et al. The effects of hyaluronic acid hydrogels with tunable mechanical properties on neural progenitor cell differentiation. Biomaterials 31, 3930–3940 (2010).
Khetan, S. et al. Degradation-mediated cellular traction directs stem cell fate in covalently crosslinked three-dimensional hydrogels. Nat. Mater. 12, 458–465 (2013).
Chaudhuri, O. et al. Extracellular matrix stiffness and composition jointly regulate the induction of malignant phenotypes in mammary epithelium. Nat. Mater. 13, 970–978 (2014).
Chaudhuri, O. et al. Hydrogels with tunable stress relaxation regulate stem cell fate and activity. Nat. Mater. 15, 326–334 (2016).
Huebsch, N. et al. Harnessing traction-mediated manipulation of the cell/matrix interface to control stem-cell fate. Nat. Mater. 9, 518–526 (2010).
Shaltouki, A., Peng, J., Liu, Q. Y., Rao, M. S. & Zeng, X. M. Efficient generation of astrocytes from human pluripotent stem cells in defined conditions. Stem Cells 31, 941–952 (2013).
Chojnacki, A. & Weiss, S. Production of neurons, astrocytes and oligodendrocytes from mammalian CNS stem cells. Nat. Protoc. 3, 935–940 (2008).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Zetsche, B. et al. Multiplex gene editing by CRISPR-Cpf1 using a single crRNA array. Nat. Biotechnol. 35, 31–34 (2017).
Zetsche, B. et al. Cpf1 Is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Renton, A. E., Chio, A. & Traynor, B. J. State of play in amyotrophic lateral sclerosis genetics. Nat. Neurosci. 17, 17–23 (2014).
Katz, J. S., Katzberg, H. D., Woolley, S. C., Marklund, S. L. & Andersen, P. M. Combined fulminant frontotemporal dementia and amyotrophic lateral sclerosis associated with an I113T SOD1 mutation. Amyotroph. Lateral Scler. 13, 567–569 (2012).
Mackenzie, I. R. A., Rademakers, R. & Neumann, M. TDP-43 and FUS in amyotrophic lateral sclerosis and frontotemporal dementia. Lancet Neurol. 9, 995–1007 (2010).
Freischmidt, A. et al. Haploinsufficiency of TBK1 causes familial ALS and fronto-temporal dementia. Nat. Neurosci. 18, 631–636 (2015).
Burdick, J. A. & Prestwich, G. D. Hyaluronic acid hydrogels for biomedical applications. Adv. Mater. 23, H41–H56 (2011).
Brigham, M. D. et al. Mechanically robust and bioadhesive collagen and photocrosslinkable hyaluronic acid semi-interpenetrating networks. Tissue Eng. Part A 15, 1645–1653 (2009).
Back, S. A. et al. Hyaluronan accumulates in demyelinated lesions and inhibits oligodendrocyte progenitor maturation. Nat. Med. 11, 966–972 (2005).
Zaman, M. H. et al. Migration of tumor cells in 3D matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proc. Natl Acad. Sci. USA 103, 10889–10894 (2006).
Catanzano, O. et al. Alginate–hyaluronan composite hydrogels accelerate wound healing process. Carbohydr. Polym. 131, 407–414 (2015).
Gao, L. Y. et al. Engineered Cpf1 variants with altered PAM specificities. Nat. Biotechnol. 35, 789–792 (2017).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J. P. GSEA-P: a desktop application for gene set enrichment analysis. Bioinformatics 23, 3251–3253 (2007).
Abel, O., Powell, J. F., Andersen, P. M. & Al-Chalabi, A. ALSoD: a user-friendly online bioinformatics tool for amyotrophic lateral sclerosis genetics. Hum. Mutat. 33, 1345–1351 (2012).
Lill, C. M., Abel, O., Bertram, L. & Al-Chalabi, A. Keeping up with genetic discoveries in amyotrophic lateral sclerosis: the ALSoD and ALSGene databases. Amyotroph. Lateral Scler. 12, 238–249 (2011).
Van Cauwenberghe, C., Van Broeckhoven, C. & Sleegers, K. The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet. Med. 18, 421–430 (2016).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323.e30 (2016).
Acknowledgements
We thank R. Macrae for critical reading of the manuscript and R. Belliveau for technical support. We thank R. Langer for input during the preparation of the manuscript and the entire Zhang Laboratory for assistance in the laboratory and helpful discussions. F.Z. is a New York Stem Cell Foundation–Robertson Investigator. F.Z. is supported by the following grants and institutes: NIH grant nos. 1R01-HG009761, 1R01-MH110049 and 1DP1-HL141201; the Howard Hughes Medical Institute; the New York Stem Cell, Simons, Paul G. Allen Family, and Vallee Foundations; and J. and P. Poitras, R. Metcalfe, and D. Cheng. J.Z.L and J.Q.P. are supported by The Stanley Center for Psychiatric Research at the Broad Institute. We thank the Klarman Cell Observatory for supporting experiments using the 10× Chromium Instrument. Reagents are available through Addgene and codes can be accessed via GitHub.
Author information
Authors and Affiliations
Contributions
H.T. and F.Z. conceived the study. H.T. designed the experiments, developed 3D and 2D cultures and 3D co-cultures of iN cells and the method to derive astrocytic cells, analysed the RNA-seq data, performed qPCR, immunostaining and imaging. S.S. filtered mouse reads from bulk RNA-seq data of iN cells co-cultured with mouse astrocytes. B.C. and H.T. performed and interpreted the comparisons between the transcriptomes of 3D cultures and co-cultures of iN cells and the human brain developmental transcriptome. X.A., C.C.H. and J.Z.L. constructed bulk RNA-seq libraries and performed sequencing. D.D. constructed scRNA-seq libraries and C.C.H. performed sequencing. J.Z.L aligned the scRNA-seq data to reference genomes. S.S. and H.T. analysed and interpreted the scRNA-seq data. L.G., S.R.C. and M.H. cloned the DNA constructs. L.G. and H.T. tested single guide RNAs in HEK cells. S.R.C. produced the AAVs. H.T. developed 3D tissues of iN cells and astrocytic cells, performed AAV infection of iN cells in 3D tissues and isolated targeted iN cells by FACS. L.G. performed next-generation sequencing and indel analyses. A.G. and J.Q.P. performed electrophysiology experiments. V.Y. and H.T. performed the mechanical characterization of the hydrogels. N.E.S. and X.S. developed hESCs with inducible expression of NGN1 and NGN2. C.L. isolated and expanded mouse glia. H.T. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
H.T. and F.Z. are co-inventors in a patent application relating to work in this manuscript. The remaining authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary text, methods, figures, tables captions, video captions and references.
Supplementary Tables
Differentially expressed genes, gene rankings and qPCR probes.
Supplementary Video 1
Z-section sequences of a 3D neural tissue of iN cells cultured in 4.6 mg ml–1 Matrigel for 40 days stained for MAP2 (green, MAP2) (using 10× objective).
Supplementary Video 2
Z-section sequences of a 3D neural tissue of iN cells cultured in 4.6 mg ml–1 Matrigel for 40 days stained for MAP2 (green, MAP2) (using 20× objective).
Supplementary Video 3
Z-section sequences of a 3D neural tissue of iN cells cultured in 4.6 mg ml–1 Matrigel for 40 days stained for MAP2 (green, MAP2) (using 63× objective).
Rights and permissions
About this article
Cite this article
Tekin, H., Simmons, S., Cummings, B. et al. Effects of 3D culturing conditions on the transcriptomic profile of stem-cell-derived neurons. Nat Biomed Eng 2, 540–554 (2018). https://doi.org/10.1038/s41551-018-0219-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-018-0219-9
This article is cited by
-
A custom-made AAV1 variant (AAV1-T593K) enables efficient transduction of Japanese quail neurons in vitro and in vivo
Communications Biology (2023)
-
In-silico study of asymmetric remodeling of tumors in response to external biochemical stimuli
Scientific Reports (2023)
-
Fabrication of Hydrogel Microchannels Using Aqueous Two-Phase Printing for 3D Blood Brain Barrier
BioChip Journal (2023)
-
Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Nature Communications (2022)
-
Bioengineered models of Parkinson’s disease using patient-derived dopaminergic neurons exhibit distinct biological profiles in a 3D microenvironment
Cellular and Molecular Life Sciences (2022)