Abstract
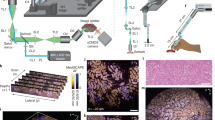
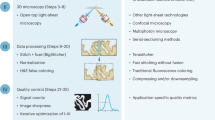
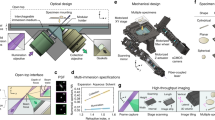
For the 1.7 million patients per year in the US who receive a new cancer diagnosis, treatment decisions are largely based on histopathological specimen examinations. Unfortunately, the gold standard of slide-based microscopic pathology suffers from high inter-observer variability and limited prognostic value due to sampling limitations and the inability to visualize tissue structures and molecular targets in their native 3D context. Here, we show that an open-top light-sheet microscope optimized for non-destructive slide-free pathology of clinical specimens enables the rapid imaging of intact tissues at high resolution over large 2D and 3D fields of view, with the same level of detail as traditional pathology. We demonstrate the utility of this technology for various applications: wide-area surface microscopy to triage surgical specimens (with ~200 μm surface irregularities), rapid intraoperative assessment of tumour-margin surfaces (12.5 s cm−2), and volumetric assessment of optically cleared core-needle biopsies (1 mm in diameter, 2 cm in length). Light-sheet microscopy can be a versatile tool for both rapid surface microscopy and deep volumetric microscopy of human specimens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Surveillance Research Program, N.C.I. Fast Stats: An Interactive Tool for Access to SEER Cancer Statistics (2016); https://seer.cancer.gov/faststats/
Barakat, F. H., Sulaiman, I. & Sughayer, M. A. Reliability of frozen section in breast sentinel lymph node examination. Breast Cancer 21, 576– 582 (2014).
McKenney, J. K. et al. The potential impact of reproducibility of Gleason grading in men with early stage prostate cancer managed by active surveillance: a multi-institutional study. J. Urol. 186, 465–469 (2011).
Shah, R. B. et al. Diagnosis of Gleason pattern 5 prostate adenocarcinoma on core needle biopsy: an interobserver reproducibility study among urologic pathologists. Am. J. Surg. Pathol. 39, 1242–1249 (2015).
Meyer, J. S. et al. Breast carcinoma malignancy grading by Bloom-Richardson system vs proliferation index: reproducibility of grade and advantages of proliferation index. Mod. Pathol. 18, 1067–1078 (2005).
Tozbikian, G . et al. Atypical ductal hyperplasia bordering on ductal carcinoma in situ: interobserver variability and outcomes in 105 cases. Int. J. Surg. Pathol. 25, 100–107 (2016).
Bedossa, P., Dargere, D. & Paradis, V. Sampling variability of liver fibrosis in chronic hepatitis C. Hepatology 38, 1449–1457 (2003).
Roberts, N. et al. Toward routine use of 3D histopathology as a research tool. Am. J. Pathol. 180, 1835–1842 (2012).
Carlson, R. O., Amirahmadi, F. & Hernandez, J. S. A primer on the cost of quality for improvement of laboratory and pathology specimen processes. Am. J. Clin. Pathol. 138, 347–354 (2012).
Gareau, D. S. et al. Confocal mosaicing microscopy in Mohs skin excisions: feasibility of rapid surgical pathology. J. Biomed. Opt. 13, 054001 (2008).
Van Royen, M. E. et al. Three-dimensional microscopic analysis of clinical prostate specimens. Histopathology 69, 985–992 (2016).
Fereidouni, F . et al. Microscopy with UV Surface Excitation (MUSE) for slide-free histology and pathology imaging. Proc. SPIE 9318, 93180F (2015).
Wang, M. et al. High-resolution rapid diagnostic imaging of whole prostate biopsies using video-rate fluorescence structured illumination microscopy. Cancer Res. 75, 4032–4041 (2015).
Wang, M. et al. Gigapixel surface imaging of radical prostatectomy specimens for comprehensive detection of cancer-positive surgical margins using structured illumination microscopy. Sci. Rep. 6, 27419 (2016 ).
Tao, Y. K. et al. Assessment of breast pathologies using nonlinear microscopy. Proc. Natl Acad. Sci. USA 111, 15304–15309 (2014).
Orringer, D. A. et al. Rapid intraoperative histology of unprocessed surgical specimens via fibre-laser-based stimulated Raman scattering microscopy. Nat. Biomed. Eng. 1, 0027 (2017).
Tu, H. et al. Stain-free histopathology by programmable supercontinuum pulses. Nat. Photon. 10, 534–540 (2016).
Olson, E., Levene, M. J. & Torres, R. Multiphoton microscopy with clearing for three dimensional histology of kidney biopsies. Biomed. Opt. Express 7, 3089–3096 (2016).
Jonkman, J. & Brown, C. M. Any way you slice it—a comparison of confocal microscopy techniques. J. Biomol. Tech. 26, 54–65 (2015).
Mertz, J. Optical sectioning microscopy with planar or structured illumination. Nat. Methods 8, 811–819 (2011).
Vakoc, B. J. et al. Three-dimensional microscopy of the tumor microenvironment in vivo using optical frequency domain imaging. Nat. Med. 15, 1219–1223 (2009).
Assayag, O. et al. Large field, high resolution full-field optical coherence tomography: a pre-clinical study of human breast tissue and cancer assessment. Technol. Cancer Res. Treat. 13, 455–468 (2014).
Zysk, A. M. et al. Optical coherence tomography: a review of clinical development from bench to bedside. J. Biomed. Opt. 12, 051403 (2007).
Siedentopf, H. & Zsigmondy, R. Uber Sichtbarmachung und Größenbestimmung ultramikoskopischer Teilchen, mit besonderer Anwendung auf Goldrubingläser. Annal. Phys. 315, 1–39 (1902).
Dodt, H. U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Keller, P. J. et al. Reconstruction of zebrafish early embryonic development by scanned light sheet microscopy. Science 322, 1065–1069 (2008).
Cella Zanacchi, F. et al. Live-cell 3D super-resolution imaging in thick biological samples. Nat. Methods 8, 1047–1049 (2011).
Huisken, J. et al. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Glaser, A. K., Wang, Y. & Liu, J. T. Assessing the imaging performance of light sheet microscopies in highly scattering tissues. Biomed. Opt. Express 7, 454–466 (2016).
Pitrone, P. G. et al. OpenSPIM: an open-access light-sheet microscopy platform. Nat. Methods 10, 598–599 (2013).
Reynaud, E. G. et al. Guide to light-sheet microscopy for adventurous biologists. Nat. Methods 12, 30–34 (2015).
Kumar, A. et al. Dual-view plane illumination microscopy for rapid and spatially isotropic imaging. Nat. Protoc. 9, 2555–2573 (2014).
Strnad, P. et al. Inverted light-sheet microscope for imaging mouse pre-implantation development. Nat. Methods 13, 139–142 (2016).
Yang, Z. et al. An inverted light sheet microscope optimized for studies in neuroscience. Conf. CLEO Atu3O.5 (2016).
Wu, Y. et al. Inverted selective plane illumination microscopy (iSPIM) enables coupled cell identity lineaging and neurodevelopmental imaging in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 108, 17708–17713 (2011).
Power, R. M. & Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat. Methods 14, 360–373 (2017).
McGorty, R. et al. Open-top selective plane illumination microscope for conventionally mounted specimens. Opt. Express 23, 16142–16153 (2015).
Kino, G. S. Applications and theory of the solid immersion lens. Proc. SPIE 3609, 56 (1999).
Liu, J. T. et al. Efficient rejection of scattered light enables deep optical sectioning in turbid media with low-numerical-aperture optics in a dual-axis confocal architecture. J. Biomed. Opt. 13, 034020 (2008).
Hall, G. S., Kramer, C. E. & Epstein, J. I. Evaluation of radical prostatectomy specimens. A comparative analysis of sampling methods. Am. J. Surg. Pathol. 16, 315–324 (1992).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Elfer, K. N. et al. DRAQ5 and eosin ('D&E') as an analog to hematoxylin and eosin for rapid fluorescence histology of fresh tissues. PLoS ONE 11, e0165530 (2016).
Moffitt, J. R. et al. High-performance multiplexed fluorescence in situ hybridization in culture and tissue with matrix imprinting and clearing. Proc. Natl Acad. Sci. USA 113, 14456–14461 (2016).
Chen, F. et al. Nanoscale imaging of RNA with expansion microscopy. Nat. Methods 13, 679–684 (2016).
Schmid, H. P. & McNeal, J. E. An abbreviated standard procedure for accurate tumor volume estimation in prostate cancer. Am. J. Surg. Pathol. 16, 184–191 (1992).
Sehdev, A. E., Pan, C. C. & Epstein, J. I. Comparative analysis of sampling methods for grossing radical prostatectomy specimens performed for nonpalpable (stage T1c) prostatic adenocarcinoma. Hum. Pathol. 32, 494–499 (2001).
Jacobs, L. Positive margins: the challenge continues for breast surgeons. Ann. Surg. Oncol. 15, 1271–1272 (2008).
Moran, M. S. et al. Society of Surgical Oncology-American Society for Radiation Oncology consensus guideline on margins for breast-conserving surgery with whole-breast irradiation in stages I and II invasive breast cancer. J. Clin. Oncol. 32, 1507–1515 (2014).
Adams, B. J. et al. The role of margin status and reexcision in local recurrence following breast conservation surgery. Ann. Surg. Oncol. 20, 2250–2255 (2013).
Singletary, S. E. et al. Revision of the American Joint Committee on Cancer staging system for breast cancer. J. Clin. Oncol. 20, 3628–3636 (2002).
Zhou, M. et al. Diagnosis of "poorly formed glands" Gleason pattern 4 prostatic adenocarcinoma on needle biopsy: an interobserver reproducibility study among urologic pathologists with recommendations. Am. J. Surg. Pathol. 39, 1331–1339 (2015).
Vettenburg, T. et al. Light-sheet microscopy using an Airy beam. Nat. Methods 11, 541–544 (2014).
Fahrbach, F. O. & Rohrbach, A. Propagation stability of self-reconstructing Bessel beams enables contrast-enhanced imaging in thick media. Nat. Commun. 3, 632 (2012).
Fu, Q. et al. Imaging multicellular specimens with real-time optimized tiling light-sheet selective plane illumination microscopy. Nat. Commun. 7, 11088 (2016).
De Medeiros, G. et al. Confocal multiview light-sheet microscopy. Nat. Commun. 6, 8881 (2015).
Tomer, R. et al. Quantitative high-speed imaging of entire developing embryos with simultaneous multiview light-sheet microscopy. Nat. Methods 9, 755–763 (2012).
Dean, K. M. et al. Imaging subcellular dynamics with fast and light-efficient volumetrically parallelized microscopy. Optica 4, 263–271 (2017).
Munch, B. et al. Stripe and ring artifact removal with combined wavelet—Fourier filtering. Opt. Express 17, 8567–8591 (2009).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Aguet, F., Van De Ville, D. & Unser, M. Model-based 2.5-d deconvolution for extended depth of field in brightfield microscopy. IEEE Trans. Image Process. 17, 1144–1153 (2008).
Giacomelli, M. G. et al. Virtual hematoxylin and eosin transillumination microscopy using epi-fluorescence imaging. PLoS ONE 11, e0159337 (2016).
Acknowledgements
The authors thank N. Sanai of the Barrow Neurological Institute, St. Joseph’s Hospital and Medical Center (Phoenix, Arizona) for providing the human glioma sample, and B. Najafian for providing the human kidney tissue sample. Human breast specimens were provided by the NorthWest BioTrust, which is supported in part by the NCI of the National Institutes of Health (P30CA015704). Human prostate specimens were provided by the GU Specimen Biorepository, University of Washington, which is supported by resources of the Department of Defense Prostate Cancer Research Program (W81XWH-14-2-0183), the Pacific Northwest Prostate Cancer SPORE (P50CA97186), a PO1 NIH grant (PO1 CA163227), and the Institute for Prostate Cancer Research of the University of Washington. This work was also supported by resources from the NIH/NCI (R01 CA175391 and F32 CA213615), the NIH/NIDCR (R01 DE023497), the University of Washington Royalty Research Fund, and a University of Washington CoMotion Innovation Award.
Author information
Authors and Affiliations
Contributions
A.K.G., N.P.R. and J.T.C.L. designed the studies. A.K.G. and J.T.C.L. designed the open-top light-sheet microscope. A.K.G., Y.C., C.Y. and L.W. fabricated the microscope. A.K.G., N.P.R. and Y.C. imaged the tissue samples. E.F.M. prepared the optically cleared tissue samples. N.P.R. and L.D.T. performed the blinded evaluation of prostate samples. N.P.R. and L.D.T. provided pathological diagnosis of all samples. A.K.G., N.P.R., Y.C., E.F.M., C.Y., L.W., Y.W., L.D.T. and J.T.C.L. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary methods, figures, tables, video captions and references (PDF 3357 kb)
Supplementary Video 1
Ease-of-use and scanning trajectory of the open-top light-sheet microscope. The sample is a fresh prostate tissue slice, stained with 1 mM acridine orange for 20 s, and rinsed for 10 s in 1× PBS. (MP4 6702 kb)
Supplementary Video 2
Open-top light-sheet microscopy imaging dataset from a piece of fresh human prostate tissue. High-magnification regions of benign glands and stroma are shown. (MP4 18966 kb)
Supplementary Video 3
Open-top light-sheet microscopy imaging dataset from a piece of fresh human breast tissue. High-magnification regions of benign lobules and invasive ductal carcinoma are shown (MP4 17755 kb)
Supplementary Video 4
Volumetric visualization of a human prostate core-needle biopsy with two representative zoomed-in (high-magnification) depth-varying image stacks (sagittal) revealing 3D glandular morphology. (MP4 7631 kb)
Supplementary Video 5
Zoomed-in region of a human prostate core-needle biopsy, in which a stack of depth-varying ‘sections’ is shown (at ~5-μm increments) to mimic a stack of conventional H&E–stained tissue sections on glass slides. (MP4 8209 kb)
Rights and permissions
About this article
Cite this article
Glaser, A., Reder, N., Chen, Y. et al. Light-sheet microscopy for slide-free non-destructive pathology of large clinical specimens. Nat Biomed Eng 1, 0084 (2017). https://doi.org/10.1038/s41551-017-0084
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41551-017-0084
This article is cited by
-
An end-to-end workflow for nondestructive 3D pathology
Nature Protocols (2024)
-
DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
Nature Communications (2024)
-
A novel micro-CT analysis for evaluating the regenerative potential of bone augmentation xenografts in rabbit calvarias
Scientific Reports (2024)
-
Content-aware frame interpolation (CAFI): deep learning-based temporal super-resolution for fast bioimaging
Nature Methods (2024)
-
Benchtop mesoSPIM: a next-generation open-source light-sheet microscope for cleared samples
Nature Communications (2024)