Abstract
In this work, we present a quantum circuit model for inferring gene regulatory networks (GRNs) from single-cell transcriptomic data. The model employs qubit entanglement to simulate interactions between genes, resulting in competitive performance and promising potential for further exploration. We applied our quantum GRN modeling approach to single-cell transcriptomic data from human lymphoblastoid cells, focusing on a small set of genes involved in innate immunity regulation. Our quantum circuit model successfully predicted the presence and absence of regulatory interactions between genes, while also estimating the strength of these interactions. We argue that the application of quantum computing in biology has the potential to provide a better understanding of single-cell GRNs by more effectively approaching the relationship between fully interconnected genes compared to conventional statistical methods such as correlation and regression. Our results encourage further investigation into the creation of quantum algorithms that utilize single-cell data, paving the way for future research into the intersection of quantum computing and biology.
Similar content being viewed by others
Introduction
A gene regulatory network (GRN) defines the ensemble of regulatory relationships between genes in a biological system. Inferring GRNs is a powerful approach for studying transcriptional regulation and the molecular basis of the regulatory mechanism, to understand the function of genes in processes of cellular activities1,2. A GRN is often represented as a graph—which can be signed, directed, and weighted—to depict relationships between transcription factors or regulators and their targets whose expression level is regulated. However, because the regulatory activity inside a cell is difficult to observe, measurements of static, intracellular gene expression are often used as a proxy, and the statistical dependencies are used to infer real regulatory relationships between genes.
Single-cell technologies, which have recently been developed and improved, open up opportunities for studying biology at remarkable resolution and scale. Single-cell RNA sequencing (scRNA-seq), for example, allows us to measure the expression of thousands of genes in each of thousands of cells3. Computational methods for constructing GRNs can adopt scRNA-seq data and leverage the information from the sheer number of cells to improve the inference power4,5,6. Thus, the utilization of single-cell data can lead to the development of more detailed and precise network models, which will help us gain a better understanding of the molecular mechanisms involved in cellular activities.
Numerous computational methods have been developed for constructing GRNs. These methods use statistical approaches to detect dependencies between expression profiles of genes and establish potential regulatory relationships between genes. The typical strategies that have been employed broadly fall into several categories such as correlation, regression, information theory, Gaussian graphical model, and Bayesian and Boolean networks4,5,6,7,8,9,10,11,12. For a broader perspective on the topic, readers are referred to several review articles13,14,15,16. It is important to note that each method has its own set of assumptions and limitations that are not always explicitly stated17,18,19. More importantly, none of these conventional methods fully exploits simultaneous, inter-regulatory connections between all genes. There is still a need for a general and principled approach to model GRNs.
Quantum computing has become an emerging technology and an intense field of research constantly seeking applications20. Researchers have developed quantum algorithms with applications in areas such as finance, cryptography, machine learning, drug discovery, chemistry, and material science21,22,23,24,25. A theoretical speedup is expected in certain types of computation using quantum algorithms versus classical algorithms because a quantum computer takes advantage of superposition and entanglement phenomena during the computation26,27. Given the potential of quantum computing, conventional strategies for inferring GRNs might be expanded by taking advantage of the quantum computing framework.
In this work, we introduce a quantum single-cell GRN (qscGRN) modeling method, which is based on a parameterized quantum circuit and uses the quantum computing framework to infer biological GRNs from scRNA-seq data. In our qscGRN model, each gene is represented using a qubit, and the circuit structure is divided into two types of layers: the encoder layer that translates the scRNA-seq data into a superposition state, and the regulation layers that entangle qubits to model gene-gene interactions in the quantum framework. Our qscGRN model maps binarized gene expression values onto a large vector space, known as Hilbert space, making full use of the information in the individual cells. Thus, the signal from thousands of cells is leveraged to improve the mapping of regulatory relationships between genes. The parameterization of our qscGRN model allows gene-to-gene regulatory relationships to be inferred all at once by fitting the superposition state probabilities onto the distribution observed in the scRNA-seq data. We include a quantum-classical framework for optimizing the parameters of our qscGRN model for a given scRNA-seq data set. The classical component of our framework uses the Laplace smoothing28 and the gradient descent algorithm29 to perform optimization by minimizing a loss function based on Kullback-Leibler (KL) divergence30. We apply the quantum-classical framework to real scRNA-seq data sets31,32 to show that gene regulatory relationships can be modeled using quantum computing, and the network recovered from the parameter-optimized quantum circuit is largely consistent with a previously published GRN33,34.
Results
The qscGRN model and its optimization framework
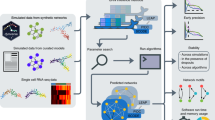
Our qscGRN model is a quantum circuit consisting of n qubits and models a biological GRN for n genes in the framework of quantum computation giving a qubit-gene equivalence (Fig. 1). A complete quantum-classical framework, which employs the qscGRN model to infer the corresponding biological GRN, is also introduced (Fig. 2). The methods section provides a detailed explanation of the model and its optimization framework.
The input matrix X is binarized into matrix Xb and n genes are selected to be modelled. a Observed distribution pobs and activation ratios \({{\rm{act}}}_{k}\) are computed using the labels and binarized values, respectively. The ratios \({{\rm{act}}}_{k}\) are used in the initial setup of the parameter \({\mathbf{\theta}}_{0}\). b The qscGRN is trained to fit the output distribution pout into pobs by minimizing a loss function based on KL-divergence. c The adjacency and network representation of the biological GRN using the optimal parameter \({\mathbf{\theta}}\).
Applying qscGRN model to scRNA-seq data of lymphoblastoid cells
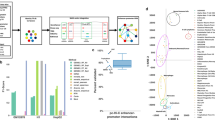
This section outlines the practical application of our qscGRN model in constructing a 6-gene GRN from real scRNA-seq data sets. The process began by feeding an input expression matrix, containing the expression values of 6 genes in over 28,000 lymphoblastoid cells, into the framewok. The 6 genes, IRF4, REL, PAX5, RELA, PRDM1, and AICDA, are members of the NF-κB signaling pathway. The pobs distribution was used to show the frequencies of the 26 = 64 possible cell states mapped into a vector space. The pobs is represented in blue in Fig. 3a, in which only the states with a probability greater than 0.01 are shown. The qscGRN model schema for the data set was a 6-qubit system and consisted of an encoder layer and six regulation layers. We measured the output register of the qscGRN model to recover the output distribution pout from the quantum framework. Then, we optimized the parameter \({\mathbf{\theta}}\) in the qscGRN model using 1087 iterations to minimizing the loss function \(L\left({\mathbf{\theta}}\right)\). The distribution \({\hat{p}}^{{\rm{out}}}\) was fitted into \({\hat{p}}^{{\rm{obs}}}\) during the optimization—smoothed distributions for pout and pobs show the similarity of the two distributions after optimization. The pout after optimization is also represented in pink in Fig. 3a. The similarity is quantified using the loss function and error metrics that reached values of 4.25e − 3 and 3.21e − 4, respectively (Fig. 3b). We validated the optimized parameter \({\mathbf{\theta}}\) running a quantum simulator that uses the Aer Simulator backend (colored in yellow in Fig. 3a).
a The observed, output, and simulated frequency distributions (pobs, pout and pqiskit) of cell activation states, colored in blue, pink and yellow respectively. b Loss function changes during training until optimization. c The adjacency matrix of the biological GRN. The heatmap shows the strength of gene-gene interactions. The diagonal elements are colored in black due to these parameters are not trained. d A weighted representation of the biological GRN recovered from the quantum circuit, where the thickness is proportional to the corresponding adjacency matrix. e Evolution of parameters in the qscGRN model recovered from the quantum framework during the optimization.
The value of the parameter \({\mathbf{\theta}}\) after optimization retrieved an adjacency matrix (Fig. 3c), which was used to construct the biological GRN. Then, we constructed a weighted network from the quantum framework using the non-diagonal elements of \({\mathbf{\theta}}\), as shown in Fig. 3d. We compared the sign of the element of each pair of genes with the corresponding regulatory effect in the previously published network, i.e., the baseline GRN33,34. Figure 3e shows the evolution of parameters for 10 regulator-target gene pairs in the qscGRN model during the optimization. These gene pairs are among the relationships recovered from the quantum framework, or present in the baseline NF-κB network33,34. Gene pairs IRF4-PRDM1, REL-AICDA, PAX5-PRDM1, REL-PRDM1 and PAX5-AICDA are correctly recovered, IRF4-AICDA incorrectly recovered, while IRF4-REL, REL-PAX5, PAX5-RELA and PRDM1-AICDA are predicted in our workflow. These recovered relationships are supported by previous studies, for example, PAX5 plays a role in the B-lineage-specific control of AICDA transcription as suggested by a previous study35. PRDM1 is a master regulator that represses PAX5 expression in B cells36. IRF4-PRDM1’s regulatory relationship might be through a third-party modulator. Indeed, IRF4 is known to inhibit BCL6 expression, and because BCL6 can repress PRDM1refs. 37,38, it has been formally speculated that the effects of IRF4 on PRDM1 expression might have been mediated through inhibition of BCL6 expression39. Although, several relationships are correctly recovered, IRF4 is known to induce AICDA expression through an indirect mechanism in the NF-κB signaling cascade40, suggesting the inference power is still limited.
The qscGRN model predicted four regulatory relationships between genes that were not present in the published baseline GRN. These included the gene pair PRDM1 and AICDA, which may indeed interact as shown that PRDM1 can silence AICDA expression a dose-dependent manner41. These results indicate that our qscGRN method has the potential to uncover regulatory relationships that were previously missed in the baseline model.
Discussion
Finding ways to apply quantum computing in biological research is an active research area42,43,44,45,46. Many questions in biology can benefit from quantum computing by exploring many possible parallel computational paths, but identifying such questions remains challenging. Especially, understanding how to exploit quantum computers for progress in solving important biological questions is crucial. The latest development of scRNA-seq technology has made it possible to gather transcriptome information from tens of thousands of individual cells per assay in a high-throughput manner. These complex data sets with higher detail are driving the development of new computational and statistical tools that are revolutionizing our understanding of cellular processes. However, quantum computation has not yet received enough attention in the face of this single-cell big data revolution.
Here, we present our qscGRN method for modeling interactions between genes to derive the quantum computing framework for constructing GRNs. In the GRN inference, the interaction between two genes determines the level of production of the target gene based on the expression of a control gene, whether this interaction is promotion or repression. Similarly, the parameter in a c-Ry gate indicates the degree of rotation of a target qubit based on the state of a control qubit. We took inspiration from the analogy between these two phenomena to design the quantum circuit in the quantum algorithm and used probability distribution to constrain the parameter of the circuit. Below we discuss three aspects of application issues.
Conventional correlation- or regression-based methods for GRN construction can handle a large number of genes because, for these methods, the gene-gene interaction is calculated as a single summary statistic from the expression profile of genes in measured cells. In contrast, our quantum approach for GRN inference can only model a small number of genes due to the vector space size—which is equal to the number of basis states—increases exponentially with the number of genes. In other words, cells in binarized scRNA-seq matrix may only be mapped to a moderate number of basis states such that each basis state is occupied by at least one cell. For example, a 15-qubit qscGRN model offers 215 = 32,768 basis states, while a scRNA-seq data set with 20,000 cells can take at most 61% of activations states in the best case. Thus, our qscGRN model may retrieve an observed distribution with no biological information mapped to many basis states. Insufficient mapping may happen even though the latest scRNA-seq technology has the capacity to allow the transcriptome of millions of cells to be measured. To obtain enough cells, we can merge multiple scRNA-seq data sets as long as they are from the same cell types or similar biological sources and the batch effect can be corrected47. On the other hand, we can select most biologically informative genes such as highly variable genes48 to be included in the analysis, reducing the burden of a large number of genes in the model while maintaining the biological relevance.
To simulate the regulatory relationship between two genes, we use a c-Ry gate to create a link between each pair of qubits in the regulation layers. The rotation angle of the c-Ry gate indicates the strength of interaction between the control gene and the target gene. The rotation angles of c-Ry gates are parameterized and mapped to the adjacency matrix after optimization to form the GRN. Throughout the paper, we assume that the rotation angle reflects the interaction strength—this is, the greater the angle, the stronger the interaction. However, we discovered that this is not always the case. We provide a simple example in Fig. 4 to illustrate the problem. Figure 4a shows the basic unit circuit, initialized in |00〉 state, that consists of a control qubit (1st qubit, rotated using an Ry gate with an angle \({\phi }_{1}\)), a target qubit (2nd qubit, rotated using an Ry gate with an angle \({\phi }_{2}\)) and a c-Ry gate with rotation angle \(\theta\). Figure 4b–f show the effect of rotation \(\theta\) in the c-Ry gate on the amplitude of |1〉 of the 2nd qubit, µ, under different settings with various combinations of \({\phi }_{1}\) and \({\phi }_{2}\). When considering µ as a function of \(\theta\), we can see in most cases, µ increases with increasing \(\theta\) or vice versa which is consistent with our assumption in gene regulation simulation. However, in some cases with specific combinations of \({\phi }_{1}\) and \({\phi }_{2}\) (as indicated with red triangles in Fig. 4e, f, the pattern is opposite—µ increases with decreasing \(\theta\) or vice versa. The opposite pattern becomes evident when \({\phi }_{1}\) approaches \({\rm{\pi }}\). We regard this phenomenon “boundary effect”, which may influence the interpretation of our modelling results. However, it should not have a great impact on our analysis. This is because: First, the boundary effect happens when the absolute value of rotation angle \(\theta\) of c-Ry gate approaches \({\rm{\pi }}/2\). In our real-data study, as shown in Fig. 3e, we found the values of \(\theta\) for all genes are in the range between \(-{\rm{\pi }}/4\) and \({\rm{\pi }}/4\), in which the boundary effect is neglectable. Second, the boundary effect only happens in limited areas in the regions with specific combinations of states of control and target qubits. The phenomenon becomes pronounced when the rotation angle of Ry gate for the control qubit, i.e. \({\phi }_{1}\), is greater than \(0.75{\rm{\pi }}\), which means the gene is being activated in more than \(85 \%\) of cells. In the case of the rotation angle is \({\rm{\pi }}\), the corresponding gene is always activated in all cells. In single-cell biology, a fully activated gene is most likely to happen for so called “house-keeping” genes, which are consistently expressed in a high level. These genes are essential for cell survival but are less likely to play any important regulatory role. Our previous study48 provides evidence that highly variable genes such as those expressed in 50% of cells and inactivated in the other 50% of cells are most functionally important for any given cell type. Taken together, we acknowledge the potential impact of the boundary effect in our model but argue that, as the impact is likely to be limited, the interpretability of the c-Ry rotation angle as a measure of interacting strength remains largely intact.
a The circuit configuration consists of a control quit (the 1st qubit) and a target qubit (the 2nd qubit), each rotated by an Ry gate with angles \({\phi }_{1}\) and \({\phi }_{2}\), respectively. b–f The heatmap panels display various combinatios of \({\phi }_{1}\) (0, 0.25π, 0.5π, 0.75π and 1.0π) and \({\phi }_{2}\) (0–1.0π) settings. The heatmap colors indicate the amplitude of |1〉 state of the target qubit with respect to \(\theta\). In general, the amplitude of |1〉 state of the target qubit increases or decreases monotonically with respect of \(\theta\). In regions marked by red trangles in (e and f), the pattern is reversed.
Correlation- and regression-based are the most widely used methods for GRN inference, owing to in part their computational efficiency. These methods typically compute correlation or regression coefficients for gene pairs using the total number of cells in the data. The issue with these methods is that they deal with gene pairs across cells, not fully exploiting complex expression patterns by incorporating another degree of information. The relationship between any two genes is measured using a single value of summary statistics such as correlation or regression coefficient. Once computed, the coefficient becomes independent of the total number of cells. Increasing the number of cells would have little influence on correlation or regression coefficient. The other issue is that the coefficient is computed only between the two genes, regardless of the expression values of other genes in the same biological system. Not considering other genes in the computation may result in a biased coefficient, which does not represent the true behavior of underlying interactions. There are methods such as partial correlation7, principal component regression5, and LASSO49 that may correct this. But, the correcting effect is limited given that all-to-all interactions cannot be easily modeled.
Methods
The implementation of our package QuantumGRN is achieved using NumPy, Pandas, Matplotlib, iGraph and Qiskit—an open-source library for working with quantum computer simulators. Our package uses the Aer Simulator backend for a noisy circuit simulator. More details about code implementation and dataset can be found in data and code availability sections.
Quantum computation theory
In this section, we introduce broad-audience background of quantum computation. In classical computation, a bit is the unit of information being |0〉 or |1〉 in Dirac notation, defined as (1 0) T and (0 1) T respectively50,51,52. In quantum computation, a qubit is the unit of information being |ψ〉 = c0 | 0〉 + c1 | 1〉 in superposition, where |ψ〉 is the quantum state, c0 and c1 are complex numbers, and |c0 | 2 + |c1 | 2 = 1. The measurement of |ψ〉 results in 0 with a probability to be observed of |c0 | 2 and 1 of |c1 | 2.
The Hadamard gate H is a single-qubit gate frequently used in quantum algorithms and is defined as \(\tfrac{1}{\sqrt{2}}\left(\begin{array}{cc}1 & 1\\ 1 & -1\end{array}\right)\), creating superpositions of the basis states (i.e., \(H\left|0\right\rangle =\tfrac{1}{\sqrt{2}}\left(\begin{array}{cc}1 & 1\\ 1 & -1\end{array}\right)\left(\begin{array}{c}1\\ 0\end{array}\right)=\tfrac{\left|0\right\rangle +{\rm{|}}1{{\rangle }}}{\sqrt{2}}\)). Furthermore, the rotation gate Ry is also a single-qubit gate that uses a rotation parameter \(\theta\) and is defined as \({R}_{{\rm{y}}}\left(\uptheta \right)=\left(\begin{array}{cc}{\rm{cos }}\uptheta /2 & -{\rm{sin }}\uptheta /2\\ {\rm{sin }}\uptheta /2 & {\rm{cos }}\uptheta /2\end{array}\right)\). In addition, a controlled gate is a 2-qubit gate that performs an operation on a target qubit when the control qubit is in state |1〉, where the operation is typically a single-qubit gate. For example, Table 1 shows the mapping of basis states when using a controlled-Ry gate that has the first qubit as control and the second qubit as target. The Ry operation is performed in basis states |10〉 and |11〉 because the control qubit is 1, no operation is performed otherwise.
In classical computation, a circuit is a model composed of a sequence of gates (NOT, AND, OR operations) having input bits that flow though such a sequence eventually computing the output bits for a given task53. Similarly, a quantum circuit is a model consisting of a sequence of quantum gates that perform operations on input qubits54. In a quantum algorithm, the input qubits are usually initialized to |0〉n, meaning a string of n bits of zeros. Then, the register flows through the sequence of gates, computing an output register that is measured and decoded to interpret the result.
The qscGRN model: a parameterized quantum circuit
Here, we introduce the quantum single-cell gene regulatory network (qscGRN) model that is a quantum circuit consisting of n qubits and models a biological GRN for n genes in the framework of quantum computation giving a qubit-gene equivalence (Fig. 1, Algorithm 1). The sequence of gates is grouped into 2 types of layers: The encoder layer Lenc consists of a Ry gate in each qubit and translates biological information (i.e., the frequency of gene actively expressed among cells) onto a superposition state. The regulation layer Lk consists of a sequence of c-Ry gates that have the kth qubit as control and a corresponding target such that the kth qubit is fully connected to other qubits. In the Lk layer, a c-Ry gate—that has the kth qubit as control and the pth qubit as the target—models the regulation interaction in the corresponding gene-gene pair. In particular, the parameter of the c-Ry gate quantifies the strength of the gene-gene interaction.
In this work, we used the notation \({\theta }_{k,{k}}\) for the parameter of the Ry gate on the kth qubit in the Lenc layer, and \({\theta }_{k,{p}}\) for the c-Ry,n gate with the kth qubit as control and the pth qubit as target, in the layer Lk of a n-qubit system. Thus, two layers were defined respectively as
where the ⊗ operator is the tensor product, and
The computation of Lenc and Lk is noncommutative due to the needed operations are matrix multiplication and tensor product.
The qscGRN model was initialized to |0〉n and put into a superposition state using the Lenc layer. Then, the gene-gene interactions were modeled using regulation layers L0, L1, ⋯, Ln-1. Thus, the qscGRN model is a quantum circuit that has n2 quantum gate parameters given by a matrix representation \({\mathbf{\theta}}\) for the set of parameters \({\uptheta }_{k,{p}}\) in the quantum gates:
where the diagonal elements belong to the Ry gates in the Lenc layer, and the non-diagonal elements to the c-Ry,n gates in the regulation layers L0, L1, ⋯, Ln-1. We recognized the matrix \({\mathbf{\theta}}\) as the adjacency matrix of the biological GRN.
Therefore, the output register \({{|}}{{\psi}}_{{\rm{out}}}{{\rangle}}\) of the qscGRN model encodes the gene-gene interactions in superposition as a function of the matrix \({\mathbf{\theta}}\) and was defined as
Algorithm 1
Construction of qscGRN model
Require: Number of qubits n, Parameter \({\mathbf{\theta}}\)
1: Create n-qubit quantum circuit qscGRN
2: for \(k=0,\,1,\,\cdots ,{n}-1\) do
3: Create and append \({R}_{{\rm{y}}}\left({\theta }_{k,k}\right)\) gate in qubit k
4: end for
5: for \(k=0,\,1,\,\cdots ,{n}-1\) do
6: for \(p=0,\,1,\,\cdots ,{n}-1\) do
7: if \(k\,\ne\, p\) then
8: Create and append \({{\rm{c}}\hbox{-}R}_{{\rm{y}},n}\left({\theta }_{k,p}\right)\) gate having k as control and p as target
9: end if
10: end for
11: end for
12: return qscGRN
Quantum-classical framework for optimization of the qscGRN model
In this section, we introduce the complete quantum-classical framework using the qscGRN model to infer the corresponding biological GRN (Fig. 2).
Gene selection
The input data of the workflow was a scRNA-seq expression data matrix X that contains expression values of N genes in m cells. The data matrix X was normalized using Pearson residuals55. Then, n out of N genes were selected to be analyzed in the next step.
Binarization
The normalized expression matrix X was binarized by applying the expression threshold of 0, which means that expression values greater than 0 are set to 1, and 0 otherwise56,57. The outcome of the binarization was saved to Xb, which is a matrix of dimension n × m.
Labeling
Labels were assigned for each cell in Xb, such that the label is a string vector composed of the binarized expression of the n genes in a cell. Thus, a label is the activation state of a gene in the corresponding cells.
Activation ratios
Activation ratios were computed for each gene as the percentage of cells expressing that gene in Xb. Then, the n rows in Xb were ordered decreasingly by the activation ratio and were labeled as g0, g1, ⋯, gn−1.
Observed distribution
We computed the percentage of occurrences of each label within the m cells to obtain the observed distribution pobs. The percentage of label \({{\rm{|}}0{\rm{\rangle }}}_{n}\) in pobs was set to 0, and the rest of the distribution was rescaled to sum to 1. The rationale for setting the \({{\rm{|}}0{{\rangle }}}_{n}\) probability to 0 is that only cells with expression values for at least one of n genes are informative. Sparsity is a common characteristic of scRNA-seq data because of the dropout event that occurs during the sequencing process. Figure 2a shows the steps just described in the workflow of the quantum-classical framework.
Initialization of the parameter \({\mathbf{\theta}}\)
The non-diagonal elements \({\theta }_{k,{p}}\) corresponding to c-Ry gates were initialized to 0. The diagonal elements \({\theta }_{k,{k}}\) corresponding to Ry gates were initialized to \(2\,\cdot \,{{\rm{sin }}}^{-1}\sqrt{{{\rm{act}}}_{k}}\), where \({{\rm{act}}}_{k}\) is the activation ratio for the kth gene. The rationale for the formula is that, independently on each qubit, the probability of observing 1 is the activation ratio of the corresponding gene after the Lenc layer. Algorithm 2 illustrates the initial setup of the workflow.
Algorithm 2
Initial setup of the workflow
Require: Normalized scRNA-seq matrix X
1: Select n gene from X
2: Xb = Binarized matrix X for the selected n genes
3: Label each cell in Xb of dimension n × m
4: for \(k=0,\,1,\,\cdots ,{n}-1\) then
5: \({{\rm{act}}}_{k}={\#{\rm{cells}}}_{k}/m\), where \({\#{\rm{cells}}}_{k}\) is the number of cells expressing the k gene
6: end for
7: for each label \({\bf{x}}\in {\left\{\mathrm{0,1}\right\}}^{n}\) then
8: \({p}_{{\bf{x}}}^{{\rm{obs}}}={\#{\rm{cells}}}_{{\bf{x}}}/m\), where \({\#{\rm{cells}}}_{{\bf{x}}}\) is the number of cells having \({\bf{x}}\) as label
9: end for
10: Rescale pobs
11: \({\mathbf{\theta}}\) = Create an n × n matrix of all elements 0
12: for \(k=0,\,1,\,\cdots ,{n}-1\) then
13: for \(p=0,\,1,\,\cdots ,{n}-1\) then
14: if \(k=p\) then
15: \({\theta }_{k,{k}}=2\,\cdot\, {{\rm{sin }}}^{-1}\sqrt{{{\rm{act}}}_{k}}\)
16: end if
17: end for
18: end for
19: return initial parameter \({\mathbf{\theta}}\), pobs
Measuring the output register of the qscGRN model
We measured the output register \({{|}}{{\psi}}_{{\rm{out}}}{{\rangle}}\) to obtain the output distribution pout of observing the basis states. The probability of the state \({{{|}}0{{\rangle}}}_{n}\) in pout was set to 0, and the rest of the distribution was rescaled to sum to 1.
Smoothing p obs and p out
Laplace smoothing was used to reshape \({p}^{{\rm{obs}}}\) and \({p}^{{\rm{out}}}\) to distributions \({\hat{p}}^{{\rm{obs}}}\) and \({\hat{p}}^{{\rm{out}}}\) respectively. These smoothed distributions were computed as \({\hat{p}}^{i}=\frac{{\#{\rm{ocu}}}^{i}+\alpha }{m+{2}^{n}\bullet \alpha }\), where \(i\in \left\{{\rm{out}},{\rm{obs}}\right\}\), α is the smoothing parameter being typically 1 and \({\#{\rm{ocu}}}^{i}\) is the number of occurrences in the distribution \({p}^{i}\). In other words, \({p}^{i}=\tfrac{{\#{\rm{ocu}}}^{i}}{m}\) is the original distribution.
Loss function
The loss function consists of KL and constrain terms, named as LKL and Lcons, were defined as
where \({\mathbf{\theta}}\) is the parameter in the qscGRN model and \({\left\{\mathrm{0,1}\right\}}^{n}\) is the n Cartesian power of the set \(\left\{\mathrm{0,1}\right\}\). Thus, the loss function was defined as
where \(\lambda\) is a dynamic coefficient that rescales Lcons to the same order of magnitude than LKL. In summary, the LKL term fits the output distribution pout in the observed distribution pobs. Meanwhile, the Lcons constraints any parameter in \({\mathbf{\theta}}\) to not get close to \({\rm{\pi }}/2\).
Optimization of the parameter \({\mathbf{\theta}}\)
The optimization was achieved by minimizing iteratively the loss function to a threshold value of 2n × 1e − 4 using a modified-gradient descent algorithm with a learning rate lr of 0.05. Otherwise, the optimization was performed for a pre-defined iterations t. Then, the parameter \({\mathbf{\theta}}\) in the iteration \(s+1\) was defined as
where \({\nabla }^{\text{T}}\) is the transpose of the gradient of loss function, allowing to keep the parameter \({\mathbf{\theta}}\) as a symmetric matrix. The diagonal parameters \({\theta }_{k,{k}}\) were not trained during optimization under the assumption that these parameters encode the binarized scRNA-seq matrix given as an input to the quantum framework. Algorithm 3 illustrates the optimization of parameter \({\mathbf{\theta}}\) and Fig. 2b shows details of the optimization in the workflow. Our work is also integrated to Qiskit—an open source library for working with quantum computers—that simulates a noisy quantum circuit using Aer Simulator backend with default parameters.
Algorithm 3
Optimization of parameter \({\mathbf{\theta}}\)
Require: Initial parameter \({{\mathbf{\theta}}}_{0}\), pobs
1: \({\hat{p}}^{{\rm{obs}}}={\mathrm{smooth}}({p}^{{\rm{obs}}})\)
2: for \(s=0,\,1,\,\cdots ,t-1\) then
3: qscGRN = Constructed quantum circuit using \({{\mathbf{\theta}}}_{s}\)
4: Measure output register and obtain \({p}^{{\rm{out}}}\)
5: \({\hat{p}}^{{\rm{out}}}={\mathrm{smooth}}({p}^{{\rm{out}}})\)
6: \(\text{loss}=L\left({{\mathbf{\theta}}}_{s}\right)\)
7: if loss < loss_threshold then
8: return \({{\mathbf{\theta}}}_{s}\)
9: end if
10: Compute gradient \(\nabla L\left({{\mathbf{\theta}}}_{s}\right)\)
11: \({{\mathbf{\theta}}}_{s+1}={{\mathbf{\theta}}}_{s}-{\rm{lr}}\,\cdot\, \frac{\nabla L\left({{\mathbf{\theta}}}_{s}\right)+{\nabla}^{\text{T}}L\left({{\mathbf{\theta}}}_{s}\right)}{2}\)
12: end for
13: return \({{\mathbf{\theta}}}_{t}\)
Recovery of gene regulatory network
We removed non-diagonal parameters in \({\mathbf{\theta}}\) that had an absolute value less than \(\tfrac{{\rm{\pi }}}{180\,\cdot \,2}\) because no significant rotation was performed by the corresponding c-Ry gate. Next, we used the remaining parameter in \({\mathbf{\theta}}\) to construct the adjacency matrix of the biological GRN, which is a weighted symmetric network. Figure 2c shows this last step in the workflow of the quantum-classical framework.
Single-cell transcriptomic data
The scRNA-seq data used in this study was generated from lymphoblastoid cell lines (LCLs), which are widely used cell line systems derived from human primary B cells. The single cell sequencing libraries were prepared using the 10x Genomics platform. Information about the experimental procedure and the acquisition of sequence data is provided in reference to our original study31. The data set has been deposited to the Gene Expression Omnibus (GEO) database and can be accessed with accession number GSE126321. To increase the number of cells in this study, we merged our data set with another LCL scRNA-seq data set downloaded from the GEO database with accession number GSE158275 ref. 32. The data matrices were pre-processed using scGEAToolbox58 and combined to produce the final matrix, which contains expression counts of 9,905 genes of 28,208 cells. The matrix was then normalized using the Pearson residuals method55. Normalized expression values of six genes: IRF4, REL, PAX5, RELA, PRDM1, and AICDA in the NF-κB signaling pathway, were extracted. The 6-gene expression matrix with dimensions of 6 × 28,208 was binarized and used as the input of our qscGRN analysis. The biological regulatory relationships between these genes, called the baseline model of GRN, were obtained from the previously established B-cell differentiation circuit model33,34.
Data availability
The scRNA-seq data analyzed in the current study is available in the NCBI GEO database under accession numbers GSE126321 and GSE158275.
Code availability
The processed data and the source code implementation of the qscGRN package are provided in the GitHub repository at https://github.com/cailab-tamu/QuantumGRN/. The repository also includes tutorials written in Python language.
References
Huynh-Thu, V. A. & Sanguinetti, G. Gene regulatory network inference: an introductory survey, in Gene Regulatory Networks: Methods and Protocols, Sanguinetti G. & Huynh-Thu V. A., Editors, Springer: New York, NY, 1–23 (2019).
Hecker, M., Lambeck, S., Toepfer, S., van Someren, E. & Guthke, R. Gene regulatory network inference: data integration in dynamic models-a review. Biosystems 96, 86–103 (2009).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Chan, T. E., Stumpf, M. P. H. & Babtie, A. C. Gene regulatory network inference from single-cell data using multivariate information measures. Cell. Syst. 5, 251–267.e3 (2017).
Osorio, D., Zhong, Y., Li, G., Huang, J. Z. & Cai, J. J. scTenifoldNet: a machine learning workflow for constructing and comparing transcriptome-wide gene regulatory networks from single-cell data. Patterns 1, 100139 (2020).
Yang, Y. et al. scTenifoldXct: a semi-supervised method for predicting cell-cell interactions and mapping cellular communication graphs. Cell. Syst. 14, 302–311.e4 (2023).
Kim, S. ppcor: an R package for a fast calculation to semi-partial correlation coefficients. Commun. Stat. Appl. Methods 22, 665–674 (2015).
Huynh-Thu, V. A., Irrthum, A., Wehenkel, L. & Geurts, P. Inferring regulatory networks from expression data using tree-based methods. PLoS One 5, e12776 (2010).
Kotiang, S. & Eslami, A. A probabilistic graphical model for system-wide analysis of gene regulatory networks. Bioinformatics 36, 3192–3199 (2020).
Lahdesmaki, H., Hautaniemi, S., Shmulevich, I. & Yli-Harja, O. Relationships between probabilistic Boolean networks and dynamic Bayesian networks as models of gene regulatory networks. Signal Process. 86, 814–834 (2006).
Friedman, N., Linial, M., Nachman, I. & Pe’er, D. Using Bayesian networks to analyze expression data. J. Comput. Biol. 7, 601–620 (2000).
Shmulevich, I., Dougherty, E. R., Kim, S. & Zhang, W. Probabilistic Boolean Networks: a rule-based uncertainty model for gene regulatory networks. Bioinformatics 18, 261–274 (2002).
Delgado, F. M. & Gomez-Vela, F. Computational methods for gene regulatory networks reconstruction and analysis: a review. Artif. Intell. Med. 95, 133–145 (2019).
Zhao, M., He, W., Tang, J., Zou, Q. & Guo, F. A comprehensive overview and critical evaluation of gene regulatory network inference technologies. Brief. Bioinform. 22, 1–15 (2021).
Blencowe, M. et al. Network modeling of single-cell omics data: challenges, opportunities, and progresses. Emerg. Top. Life Sci. 3, 379–398 (2019).
Cha, J. & Lee, I. Single-cell network biology for resolving cellular heterogeneity in human diseases. Exp. Mol. Med. 52, 1798–1808 (2020).
Chen, S. & Mar, J. C. Evaluating methods of inferring gene regulatory networks highlights their lack of performance for single cell gene expression data. BMC Bioinform. 19, 232 (2018).
Pratapa, A., Jalihal, A. P., Law, J. N., Bharadwaj, A. & Murali, T. M. Benchmarking algorithms for gene regulatory network inference from single-cell transcriptomic data. Nat. Methods 17, 147–154 (2020).
Diaz, L. P. M. & Stumpf, M. P. H. Gaining confidence in inferred networks. Sci. Rep. 12, 2394 (2022).
Gyongyosi, L. & Imre, S. A survey on quantum computing technology. Comput. Sci. Rev. 31, 51–71 (2019).
Egger, D. J. et al. Quantum computing for finance: state-of-the-art and future prospects. IEEE Trans. Quantum Eng. 1, 1–24 (2020).
Fernandez-Carames, T. M. & Fraga-Lamas, P. Towards post-quantum blockchain: a review on blockchain cryptography resistant to quantum computing attacks. IEEE Access. 8, 21091–21116 (2020).
Cao, Y., Romero, J. & Aspuru-Guzik, A. Potential of quantum computing for drug discovery. IBM J. Res. Dev. 62, 6:1–6:20 (2018).
Ramezani, S. B., Sommers, A., Manchukonda, H. K., Rahimi, S. & Amirlatifi, A.. Machine Learning Algorithms in Quantum Computing: A Survey. in International Joint Conference on Neural Networks (IJCNN), Glasgow, UK, 1–8, https://doi.org/10.1109/IJCNN48605.2020.9207714, (2020).
Bauer, B., Bravyi, S., Motta, M. & Kin-Lic Chan, G. Quantum algorithms for quantum chemistry and quantum materials science. Chem. Rev. 120, 12685–12717 (2020).
Bharti, K. et al. Noisy intermediate-scale quantum algorithms. Rev. Mod. Phys. 94, 015004 (2022).
Huang, H. Y. et al. Quantum advantage in learning from experiments. Science 376, 1182–1186 (2022).
Peng, F. C., Schuurmans, D. & Wang, S. J. Augmenting naive Bayes classifiers with statistical language models. Inf. Retr. J. 7, 317–345 (2004).
Ruder, S. An overview of gradient descent optimization algorithms. Preprint at https://arxiv.org/abs/1609.04747 (2016).
Kullback, S. & Leibler, R. A. On information and sufficiency. Ann. Math. Stat. 22, 79–86 (1951).
Osorio, D., Yu, X., Yu, P., Serpedin, E. & Cai, J. J. Single-cell RNA sequencing of a European and an African lymphoblastoid cell line. Sci. Data 6, 112 (2019).
SoRelle, E. D. et al. Single-cell RNA-seq reveals transcriptomic heterogeneity mediated by host-pathogen dynamics in lymphoblastoid cell lines. eLife 10, e62586 (2021).
Roy, K. et al. A regulatory circuit controlling the dynamics of NFkappaB cRel transitions B cells from proliferation to plasma cell differentiation. Immunity 50, 616–628.e6 (2019).
Sciammas, R. et al. An incoherent regulatory network architecture that orchestrates B cell diversification in response to antigen signaling. Mol. Syst. Biol. 7, 495 (2011).
Yadav, A. et al. Identification of a ubiquitously active promoter of the murine activation-induced cytidine deaminase (AICDA) gene. Mol. Immunol. 43, 529–541 (2006).
Boi, M., Zucca, E., Inghirami, G. & Bertoni, F. PRDM1/BLIMP1: a tumor suppressor gene in B and T cell lymphomas. Leuk. Lymphoma 56, 1223–1228 (2015).
Tunyaplin, C. et al. Direct repression of prdm1 by Bcl-6 inhibits plasmacytic differentiation. J. Immunol. 173, 1158–1165 (2004).
Shapiro-Shelef, M. & Calame, K. Regulation of plasma-cell development. Nat. Rev. Immunol. 5, 230–242 (2005).
Teng, Y. et al. IRF4 negatively regulates proliferation of germinal center B cell-derived Burkitt’s lymphoma cell lines and induces differentiation toward plasma cells. Eur. J. Cell. Biol. 86, 581–589 (2007).
Sciammas, R. et al. Graded expression of interferon regulatory factor-4 coordinates isotype switching with plasma cell differentiation. Immunity 25, 225–236 (2006).
Nutt, S. L., Taubenheim, N., Hasbold, J., Corcoran, L. M. & Hodgkin, P. D. The genetic network controlling plasma cell differentiation. Semin. Immunol. 23, 341–349 (2011).
Outeiral, C. et al. The prospects of quantum computing in computational molecular biology. WIREs Comput. Mol. Sci. 11, e1481 (2021).
Marx, V. Biology begins to tangle with quantum computing. Nat. Methods 18, 715–719 (2021).
Emani, P. S. et al. Quantum computing at the frontiers of biological sciences. Nat. Methods 18, 701–709 (2021).
Cheng, H. P., Deumens, E., Freericks, J. K., Li, C. & Sanders, B. A. Application of quantum computing to biochemical systems: a look to the future. Front. Chem. 8, 587143 (2020).
Woolnough, A. P., Hollenberg, L. C. L., Cassey, P. & Prowse, T. A. A. Quantum computing: a new paradigm for ecology. Trends Ecol. Evol. 3142, 9 (2023).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Osorio, D. et al. Single-cell expression variability implies cell function. Cells 9, 14 (2019).
Tibshirani, R. Regression Shrinkage and Selection via the Lasso. J. R. Stat. Soc. Ser. B Stat. Methodol. 58, 267–288 (1996).
Dirac, P. A. M. A new notation for quantum mechanics. Math. Proc. Camb. Philos. Soc. 35, 416–418 (1939).
Meter, R. V. & Oskin, M. Architectural implications of quantum computing technologies. ACM J. Emerg. Technol. Comput. Syst. 2, 31–63 (2006).
Rieffel, E. & Polak, W. An introduction to quantum computing for non-physicists. ACM Comput. Surv. 32, 300–335 (2000).
Lee, E. A. The past, present and future of cyber-physical systems: a focus on models. Sensors 15, 4837–4869 (2015).
Vatan, F. & Williams, C. Optimal quantum circuits for general two-qubit gates. Phys. Rev. A. 69, 032315 (2004).
Lause, J., Berens, P. & Kobak, D. Analytic Pearson residuals for normalization of single-cell RNA-seq UMI data. Genome Biol. 22, 258 (2021).
Qiu, P. Embracing the dropouts in single-cell RNA-seq analysis. Nat. Commun. 11, 1169 (2020).
Bouland, G. A., Mahfouz, A. & Reinders, M. J. T. Consequences and opportunities arising due to sparser single-cell RNA-seq datasets. Genome Biol. 24, 86 (2023).
Cai, J. J. scGEAToolbox: a Matlab toolbox for single-cell RNA sequencing data analysis. Bioinformatics 36, 1948–1949 (2019).
Acknowledgements
The identification of the boundary effect in the c-Ry gate impact analysis is credited to the anonymous reviewer for whom we express our gratitude. This work was supported by the DoD grant GW200026 for J.J.C.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.J.C.; methodology, C.R. and J.J.C.; implementation of the software, C.R.; formal analysis, C.R. and J.J.C.; writing and editing, C.R. and J.J.C.; supervision, J.J.C. All authors reviewed and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Roman-Vicharra, C., Cai, J.J. Quantum gene regulatory networks. npj Quantum Inf 9, 67 (2023). https://doi.org/10.1038/s41534-023-00740-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41534-023-00740-6