Abstract
Parkinson’s disease (PD) is the second most common neurodegenerative disease in the United States. Decades before motor symptoms manifest, non-motor symptoms such as hyposmia and rapid eye movement (REM) sleep behavior disorder are highly predictive of PD. Previous immune profiling studies have identified alterations to the proportions of immune cells in the blood of clinically defined PD patients. However, it remains unclear if these phenotypes manifest before the clinical diagnosis of PD. We utilized longitudinal DNA methylation (DNAm) microarray data from the Parkinson’s Progression Marker’s Initiative (PPMI) to perform immune profiling in clinically defined PD and prodromal PD patients (Prod). We identified previously reported changes in neutrophil, monocyte, and T cell numbers in PD patients. Additionally, we noted previously unrecognized decreases in the naive B cell compartment in the defined PD and Prod patient group. Over time, we observed the proportion of innate immune cells in PD blood increased, but the proportion of adaptive immune cells decreased. We identified decreases in T and B cell subsets associated with REM sleep disturbances and early cognitive decline. Lastly, we identified increases in B memory cells associated with both genetic (LRRK2 genotype) and infectious (cytomegalovirus seropositivity) risk factors of PD. Our analysis shows that the peripheral immune system is dynamic as the disease progresses. The study provides a platform to understand how and when peripheral immune alterations occur in PD and whether intervention at particular stages may be therapeutically advantageous.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD) is an age-related neurodegenerative disorder with multisystemic manifestations characterized histopathologically by the loss of dopaminergic (DA) neurons in the substantia nigra pars compacta1. PD-related proteinopathy is characterized by the identification of aggregates of alpha-synuclein protein and alpha-synuclein immunoreactive Lewy Bodies in brain regions near and distal to the sites of neurodegeneration2. The diagnostic criteria for PD rely on the clinical determination of bradykinesia, in conjunction with rest tremor or rigidity, while ruling out other potential causes3,4. Before the onset of motor symptoms and years before clinical diagnosis, non-motor symptoms such as REM sleep behavior disorder (RBD) and hyposmia are common5,6,7. Patients who experience these symptoms but do not exhibit clear motor deficits are considered to be in the prodromal phase of PD7.
Inflammation is an easily identified histopathological component of PD8. Current data support the role of inflammation in the pathogenesis of PD citing microglial activation9, the identification of genetic variation in genes such as HLA-DR10,11, and evidence of immune infiltration in the brain parenchyma12,13,14,15. Interactions between the peripheral immune system and PD are poorly understood but analysis of post-mortem tissue, cerebrospinal fluid (CSF), and blood have provided evidence of complex interactions between peripheral, local, innate, and adaptive immune responses during the neurodegenerative process16.
The innate immune system provides an early-stage line of defense against infectious agents or damage to other cells in the body. Broadly, these cells recognize damaged neurons in the PD brain, clear debris, and relay the signal to the periphery17. Microglia are the central nervous system (CNS) resident innate immune cells, while myeloid (neutrophils, basophils, eosinophils, and monocytes) and natural killer (NK) cells are those in the blood. Microglia detect PD-related damage-associated molecular patterns (DAMPs) such as alpha-synuclein proteinopathy or neuronal damage and relay signals to peripheral immune cells17. However, the role of peripheral immune cells in initiating or propagating inflammatory damage in the CNS of PD patients is not entirely known. Neutrophils can increase the permeability of the blood-brain barrier (BBB), giving all immune cells increased access to the CNS18. Evidence for neutrophil activity in the PD brain is based on the observed expression of myeloperoxidase19. Basophils and eosinophils are specialized in responses to allergens and parasitic infections; some associations have been reported between asthma or allergies to plants, pollen, or antibiotics and an increased risk of PD20,21. Monocytes are phagocytes that can clear debris and recruit immune cells. In PD, monocytopenia, increased phagocytic behavior, and decreased viability and cytokine production upon ex vivo stimulation have been observed22,23,24. Lastly, NK cells recognize non-specific biomarkers and have been thought to aid in the clearance of alpha-synuclein and communication with dying neurons25. Others have noted that the activity and numbers of NK cells increase as PD progresses22,26,27,28,29.
The adaptive immune system provides a secondary line of defense against antigens and produces a memory response facilitated by T and B cells. Recent work has suggested both pathogenic and neuroprotective roles of the adaptive immune cells in various neurodegenerative contexts30. T cells have been identified within the substantia nigra of PD patients31,32,33. Additionally, PD patients possess CD4+ T cells that are primed to a Th1 proinflammatory phenotype and react to epitopes of alpha-synuclein34,35,36. B cells are the antibody-producing cells of the body. In PD, antibodies specific to alpha-synuclein have been identified in patient serum and CSF37,38. Whether an inflammatory immune response is a cause or a consequence of tissue damage in neurodegenerative diseases like PD is still controversial. However, it is agreed that the CNS has a direct line of communication with the peripheral immune system, and intercepting these signals may allow us to monitor CNS disease states from the blood39.
Immune profiling on an epidemiological scale is challenging due to the lack of fresh samples that can be processed using flow/mass cytometry. Epigenomics technology has opened the potential to scale this type of data collection without sacrificing resolution. DNA methylation (DNAm) is an epigenetic marker placed on cytosine bases and is used to regulate gene expression and cell differentiation40; cell-type specific genes can be expressed and are hypomethylated, while other genes are repressed with hypermethylation of regulatory gene elements41. These durable, intrinsic lineage markers allow researchers to use DNAm data to estimate the composition of a specimen by using cell-type specific DNAm profiles42,43,44,45. This approach, known as cellular deconvolution, can be used to assess compositional differences as a potential element of the disease processes and allow for control of confounding by cellular heterogeneity across samples in epigenome-wide association studies (EWAS)46.
Several studies have already performed immune profiling of PD blood samples using conventional methods such as flow cytometry and DNAm data23,47,48,49,50,51,52,53,54. However, our current knowledge of the peripheral immune dynamics in PD is limited. Firstly, many of these studies investigated broad proportions of immune cells where details about subsets of B cells, T cells, and granulocytes may have been lost due to technical limitations23,52. Similarly, previous DNA-based assessments have utilized earlier versions of deconvolution platforms that also do not investigate some of these cell subtypes49. Secondly, many of these studies have assessed immune dysregulation in clinically defined PD cases; nonetheless, we know that CNS damage may begin decades before motor symptom onset55. There is a significant gap in our understanding of when precisely the peripheral immune perturbations manifest concerning the onset of neurological damage rather than a clinical diagnosis of PD.
To fill this gap, we utilized the data available from the Parkinson’s Progression Markers Initiative (PPMI) to study the early immunological perturbations during PD. The PPMI is a large PD case-control study implemented to identify PD biomarkers56,57. The PPMI collected a battery of longitudinal data, including whole blood DNA data, from PD patients, healthy participants, and a group of prodromal PD patients58. Our goal is to perform an in-depth analysis of cellular composition using the PPMI data with a deconvolution library developed and validated by our laboratory that extends to 12 cell types42. We can assess how early the immunological perturbations manifest in the PD progression timeline by analyzing the prodromal PD patients in parallel to the clinically defined PD patients. We hypothesize that immunological dysregulation will manifest alongside the earliest clinically observable events, and thus, we will see peripheral immunological perturbations in prodromal PD patients.
Results
Patient data
Participants selected for this analysis were assigned to one of three patient groups (Table 1). The healthy control group (HC) were neurologically healthy participants of similar demographics to the case groups. The Parkinson’s disease group (PD) consisted of participants diagnosed with PD less than two years before the screening visit and were naive to PD-specific medications. The prodromal PD group (Prod) consisted of participants who exhibited symptoms indicative of early stages of PD or tested positive for PD genetic risks alleles (Table 2). After enrollment, blood DNAm and clinical data were collected annually.
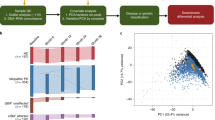
Raw bulk DNAm data were downloaded from the PPMI database and analyzed (Fig. 1). After the quality control assessment, we included 2184 samples in our study (Supplementary Table 1). The FlowSorted.BloodExtended.EPIC package was used to deconvolute each blood sample, i.e., calculate the proportions of immune cells42: this includes basophils (Bas), naive and memory B cells (Bnv, Bmem), naive and memory CD4+T cells (CD4nv, CD4mem), naive and memory CD8+T cells (CD8nv, CD8mem), eosinophils (Eos), monocytes (Mono), neutrophils (Neu), natural killer cells (NK), and T regulatory cells (Treg). These values were then adjusted to account for each cell type’s limit of detection (LoD, see Methods). Because 98.5% of samples were below the LoD for Tregs, these cells were excluded from all subsequent analyses (Supplementary Fig. 1).
All available DNAm data files were downloaded from PPMI, but after our quality control analyses, we included 2184 samples in the study. Using the DNAm data, we estimated the proportions of immune cells in each blood sample and performed different epidemiological analyses, denoted by the figure number in the abbreviated causal diagram.
We noticed two batch effects present in the dataset (Supplementary Fig. 2). The first was associated with the DNAm data processing batch; the second was associated with the run date of the array. Each of these variables was transformed into binary variables and they were included as covariates in all subsequent analyses. Sensitivity analyses confirmed our findings were valid in a subset of data that did not include these batch effects (data not shown).
Comparing the cellular proportions of PD and Prod groups to HC
We performed regression analyses to estimate the effect of PD on the proportion of each immune cell. Our regression analyses were adjusted for the patient’s age at their baseline visit, sex, and batch effects. We conducted two parallel analyses using cross-sectional and longitudinal approaches. This dual strategy captures inter-timepoint dynamics across linear cross-sectional models at each study time point. We then utilized a longitudinal model to have a well-powered estimate for the overall effect of having PD on the immune proportion. Radar plots show the average difference in cell proportion between the PD group and the HC group at the BL sample (Fig. 2a) and at the final major study timepoint (Fig. 2b). We performed a single-sample t-test for each effect size to test if the estimate was non-zero and generate a p-value. We used a linear mixed effects model for the longitudinal approach, including a random effect for each patient to account for the repeated measures and time as an additional variable (Fig. 2c).
Plotted is the effect size and 95% confidence interval for having PD compared to HC in our regression models for each cell type. a–c plot the effect size as derived from the models, whereas (d–f) plot the variance normalize mean difference. a, b, d, e were derived using linear models, whereas (c, f) were derived using mixed effect regression models. Models were adjusted for age, sex, and batch variables. Only p values <0.05 are denoted (regression model t test; df = 375 for (a, d); df = 342 for (b, e); df = 385.3 for (c, f)). HC healthy control group, PD Parkinson’s disease group, Bas basophils, Eos eosinophils, Neu neutrophils, Bnv B-cells naïve, Bmem B-cells memory, CD4nv helper CD4+ T-cells naïve, CD4mem helper CD4+ T-cells memory, Treg CD4+ T regulatory cells, CD8nv cytotoxic CD8+ T-cells naïve, CD8mem cytotoxic CD8+ T-cells memory, Mono monocytes, NK natural killer cells, df degrees of freedom.
We anticipated that comparing the linear effect sizes from these models would visually inflate the changes to abundant cell types such as Neu. To calibrate the effect sizes, we divided each by the standard deviation of the cell proportion in the HC group (Fig. 2d–f). These values represent the mean difference in cell proportion in the units of the number of standard deviations away from the healthy population (Eq. II, see Methods). This data representation allows us to visualize small yet significant changes in lowly abundant cell types, such as Bnv cells.
Consistent across both cross-sectional and longitudinal methods, we found that the PD group had lower proportions of CD4mem and Bnv and higher proportions of Neu. We also identified a decrease in Eos (\({\hat{\beta }}_{Z}\) = -0.48, p = 0.015) and Mono (\({\hat{\beta }}_{Z}\) = -0.61, p = 0.0057) proportions in the longitudinal model. At the baseline visit, we found a decrease in Mono proportions (\({\hat{\beta }}_{Z}\) = -0.79, p = 0.0023); however, it was no longer significant at the Year 3 time point (\({\hat{\beta }}_{Z}\) = -0.45, p = 0.082).
We employed the same dual approaches to compare the cellular composition of the Prod group to HC (Fig. 3). We first looked at yearly time points in the study (Fig. 3a, b) in addition to a longitudinal model (Fig. 3c). At the baseline time point, we observed a decrease in Bnv (\({\hat{\beta }}_{Z}\) = -0.42, p = 0.047) and Mono (\({\hat{\beta }}_{Z}\) = -0.62, p = 0.031) (Fig. 3d). Although not statistically significant, we observed an increase in Bmem cells (\({\hat{\beta }}_{Z}\) = 0.19, p = 0.21). At the Year 3 time point, we found a decrease in CD4mem cells (\({\hat{\beta }}_{Z}\) = -1.2, p = 0.050) (Fig. 3e). Looking at the longitudinal model, we observe an overall reduction in Bnv cells (\({\hat{\beta }}_{Z}\) = -0.42, p = 0.027) (Fig. 3f). We also identified strong trends in Bmem (\({\hat{\beta }}_{Z}\) = 0.21, p = 0.15) and Mono (\({\hat{\beta }}_{Z}\) = -0.47, p = 0.059) in the longitudinal model.
Plotted is the effect size and 95% confidence interval for having Prod compared to HC in our regression models for each cell type. a–c plot the effect size as derived from the models, (d–f) plot the variance normalized mean difference. a, b, d, e were derived using linear models, whereas (c, f) were derived using mixed effect regression models. Models were adjusted for age, sex, and batch variables. Only p values <0.05 are denoted (regression model t test; df = 252 for (a, d); df = 197 for (b, e); df = 262.6 for (c, f)). HC healthy control group, Prod prodromal Parkinson’s disease group, Bas basophils, Eos eosinophils, Neu neutrophils, Bnv B-cells naïve, Bmem B-cells memory, CD4nv helper CD4+ T-cells naïve, CD4mem helper CD4+ T-cells memory, Treg CD4+ T regulatory cells, CD8nv cytotoxic CD8+ T-cells naïve, CD8mem cytotoxic CD8+ T-cells memory, Mono monocytes, NK natural killer cells, df degrees of freedom.
We directly compared the PD and Prod groups to each other to identify changes in cell composition that may differ between pre-motor deficit PD and clinically defined PD (Supplementary Fig. 3). We identified that the increased levels of Neu is specific to the clinically defined PD patients (\({\hat{\beta }}_{Z}\) = 2.2, p = 0.0032 in the longitudinal model). We also observe marginal decreases in CD4T cell populations in the PD group compared to the Prod group.
Because we obtained different results across cross-sectional time points, we assessed how the cell proportions changed over time in each group and determined if they changed at different rates using stratified longitudinal models (Fig. 4). Interestingly, we observed a decrease in CD8nv cells over time in all groups (HC \({\hat{\beta }}_{{time}}\) = -0.060, p = 0.0013; Prod \({\hat{\beta }}_{{time}}\) = -0.034, p = 0.0098; PD \({\hat{\beta }}_{{time}}\) = -0.032, p = 2.7E-4). For the Prod group, we also observed an increase in Bas (\({\hat{\beta }}_{{time}}\,\)= 0.046, p = 1.6E-5), and a decrease in Bnv (\({\hat{\beta }}_{{time}}\,\)= -0.054, p = 0.0031) and CD4nv cells (\({\hat{\beta }}_{{time}}\) = -0.13, p = 0.0022). For the PD group, we observed a decrease in Bnv (\({\hat{\beta }}_{{time}}\) = -0.067, p = 8.1E-6), CD4nv (\({\hat{\beta }}_{{time}}\,\)= -0.079, p = 0.019), CD4mem (\({\hat{\beta }}_{{time}}\) = -0.18, p = 5.4E-4), and CD8mem (\({\hat{\beta }}_{{time}}\) = -0.12, p = 0.023) over time, while we saw an increase in Mono (\({\hat{\beta }}_{{time}}\,\)= 0.063, p = 0.052) and Neu (\({\hat{\beta }}_{{time}}\) = 0.49, p = 9.4E-4).
Plotted is the effect size and 95% confidence interval for the effect of time on the level of each cell type in the mixed effect longitudinal models. We stratified the data and ran a model for each patient group to identify a potential interaction. Each model was adjusted for age, sex, and batch variables. Only p values <0.05 are denoted (regression model t test; df = 243.7 for HC; df = 867.7 for PD; df = 507.0 for Prod). HC healthy control group, PD Parkinson’s disease group, Prod prodromal Parkinson’s disease group, Bas basophils, Eos eosinophils, Neu neutrophils, Bnv B-cells naïve, Bmem B-cells memory, CD4nv helper CD4+ T-cells naïve, CD4mem helper CD4+ T-cells memory, Treg CD4+ T regulatory cells, CD8nv cytotoxic CD8+ T-cells naïve, CD8mem cytotoxic CD8+ T-cells memory, Mono monocytes, NK natural killer cells, df degrees of freedom.
To formally test if the rate of change in the PD or Prod group is unique from the HC group, we performed likelihood ratio tests between models that did and did not include an interaction term for case status and time. A significant result implies that allowing the model to fit unique trends over time for each group (interaction model) better represents the data compared to a model that tries to fit one trend for the entire dataset (additive model). Comparing PD to HC, the likelihood ratio test was significant for Bas (p = 0.0097), Bmem (p = 0.036), and CD8nv (p = 0.046). Comparing Prod to HC, the likelihood ratio test was significant for Bmem (p = 0.016), Bnv (p = 0.035), and CD8nv (p = 0.036).
Lastly, we assessed whether the immune profile is associated with clinical measures of disease activity. Using seven clinical scores and two DaT imaging variables (Table 3), we assessed the relationship between PD symptomology and the proportions of immune cells (Fig. 5). At the baseline visit, we identified that patients in the highest quartile of sleep disturbances had lower proportions of CD8mem (\({\hat{\beta }}_{Z}\,\)= -0.24, p = 0.038) and Bmem (\({\hat{\beta }}_{Z}\,\)= -0.045, p = 0.015) compared to those in the lowest quartile (Fig. 5a). We identified decreased proportions of CD4nv (\({\hat{\beta }}_{Z}\,\)= -0.27, p = 0.043) when comparing patients in the highest quartile for cognitive dysfunction (low MoCA) to those in the lowest quartile (Fig. 5b). We also identified an association between Bmem proportions and scores on STAI (\({\hat{\beta }}_{Z}\) = 0.0063, p = 0.017) and GDS (\({\hat{\beta }}_{Z}\,\)= 0.048, p = 0.017) (Supplementary Fig. 4), but no other associations were found at baseline (Supplementary Fig. 4, Supplementary Fig. 5).
All radar plots represent variance normalized mean difference in cell proportion with 95% confidence intervals. a, b show relationships between specific clinical variables and the proportions of each cell type in which PD patients in the most symptomatic quartile are compared to PD patients in the least symptomatic quartile. c, e are heatmaps of the mean value of each clinical score in each cluster identified by the clustering approach. All scores were mean scaled and centered before clustering, color scale shows scaled Z-score. d, f show the comparison of group 2 to group 1 immune cell profiles. Models were adjusted for age, sex and batch variables. Only p values <0.05 are denoted (regression model t test; df = 271 for (a); df = 268 for (b); df = 269 for (d); df = 230 for (f)). Bas basophils, Eos eosinophils, Neu neutrophils, Bnv B-cells naïve, Bmem B-cells memory, CD4nv helper CD4+ T-cells naïve, CD4mem helper CD4+ T-cells memory, Treg CD4+ T regulatory cells, CD8nv cytotoxic CD8+ T-cells naïve, CD8mem cytotoxic CD8+ T-cells memory, Mono monocytes, NK natural killer cells, AI asymmetry index, NP1PTOT Movement Disorder Society Unified Parkinson’s Disease Rating Scale Part 1 (total score), NP2PTOT Movement Disorder Society Unified Parkinson’s Disease Rating Scale Part 2 (total score), ESS_tot Epworth Sleepiness Scale (total score), MCATOT Montreal Cognitive Assessment (total score), STAI_tot State-Trait Anxiety Inventory (total score), REM_tot REM Sleep Behavior Questionnaire (total score), GDS_tot Geriatric Depression Scale (total score), df degrees of freedom.
We performed k-means clustering on the clinical variable data at baseline and at the year 2 timepoint which cluster the patients into two groups (Fig. 5c, d). Group 2 exhibited worse clinical scores across all variables; thus, we interpreted that the clustering split patients into “more symptomatic” (Group 2) and “less symptomatic” (Group 1). We compared the immune cell proportions of Group 2 to Group 1 at baseline (Fig. 5e) and at the year 2 time point (Fig. 5f). Although these findings were not statistically significant, we observed that more symptomatic patients had higher proportions of Bmem at baseline (\({\hat{\beta }}_{Z}\) = 0.18, p = 0.11), increased proportions of Neu at year 2 (\({\hat{\beta }}_{Z}\) = 1.27, p = 0.32), and decreased proportions of Mono at year 2 (\({\hat{\beta }}_{Z}\) = -0.55, p = 0.072). We tested for a potential interaction between cluster group and time to determine if the rate of change of immune cells in each group was unique to each cluster. For all 11 cell types, the test was significant: Bas (p = 6.9E-04), Eos (p = 0.0035), Neu (p = 0.022), Mono (p = 0.0048), Bmem (p = 0.0016), Bnv (p = 0.0042), NK (p = 0.0043), CD4nv (p = 0.0040), CD8nv (p = 9.9E-04), CD8mem (p = 0.022) and CD4mem (p = 0.010).
Comparing the DNAm profiles of PD and Prod groups to HC
We performed cross-sectional and longitudinal epigenome-wide association studies (EWAS) to identify which CpGs were differentially methylated between PD and HC and between Prod and HC (Fig. 6). These models were adjusted for age at the baseline visit, sex, batch effects, and Neu, Mono, Bnv, CD4mem, and Eos proportions. A differentially methylated CpG (DMC) is defined when the average difference in methylation level (Δβ) is greater than |0.1| and the adjusted p-value to maintain a false discovery rate of 5% is still significant.
In (a, c), PD patients are compared to HC while in (b, d), prodromal PD patients are compared to HC. a, b show volcano plots highlighting differentially methylated CpGs (DMCs). False discovery rate was controlled at 5% using the BH procedure to generate adjusted p values (p values generated from empirical Bayes moderated regression model t test). c, d show bar plots comparing the DMCs identified in each cross-sectional model compared to the longitudinal model. The input to each EWAS analyses was the top 100,000 CpGs with the largest Δβ.
We did not identify any DMCs between PD and HC in the cross-sectional models; however, we found 1 DMC in the longitudinal model (cg13102742, Fig. 6a). Considering only the number of statistically significant CpGs, regardless of the Δβ, we still observe no DMCs in the cross-sectional models but many DMCs in the longitudinal model (Fig. 6c).
Surprisingly, we observed much more epigenetic dysregulation in the Prod group. Our longitudinal model identified 13 hypermethylated and 15 hypomethylated DMCs between Prod and HC (Fig. 6b, Supplementary Figs. 6-8). Hypermethylated DMCs were associated with PLEKHA1, VASN, CORO7, DENND4C, NUP93, PYROXD2, L3HYPDH, ACOT1, HEATR4, STK4. Hypomethylated DMCs were associated with SPATA4, LENG8-AS1, LENG8, BHLHE40, GLIPR2, EPB41L5, PKD1L2, KIRREL3, ERICH1, RAD51B, LSG1, SHANK2, and PCNX1. We identified DMCs in each cross-sectional model but vastly more in the longitudinal model (Fig. 6d). Most of the identified DMCs lie within 50 base-pairs of previously identified lowly-penentrant SNPs (Supplementary Table 2).
Lastly, we performed similar EWAS analyses comparing the PD group to the Prod group (Supplementary Fig. 9). We identified that two CpGs (cg18845950 and a CpG near SHANK2) were hypermethylated in the PD patients. These two CpGs were hypomethylated when comparing Prod to HC suggesting that these molecular alterations are indeed specific to the Prod group samples.
PD risk factors have varying effects on the immune cell proportions present in the blood
Previous findings have shown a genetic influence on the peripheral immune cell proportions in PD patients23,59. Using the baseline sample data, we compared the immune cell composition of LRRK2 risk allele positive (LRRK2+) PD patients with the major allele of LRRK2 (LRRK2-) (Fig. 7a). We identified higher proportions of Bmem (\({\hat{\beta }}_{Z}\) = 0.29, p = 0.021) but lower proportions of Bnv (\({\hat{\beta }}_{Z}\,\)= -0.40, p = 0.043) in LRRK2+ PD patients. Similarly, we repeated this analysis with LRRK2+ and LRRK2- Prod patients but did not identify these differences (Fig. 7b). We also performed this analysis for GBA risk allele patients. However, as anticipated we were underpowered to test for this, given the low number of GBA+ patients (Supplementary Fig. 10).
All radar plots represent variance normalized mean difference in cell proportion with 95% confidence intervals. a, b compare the immune cell proportions between LRRK2 risk allele positive patients to LRRK2 risk allele negative patients, within the PD group (a) or Prod group (b). c–f Is in reference to DNAm derived CMV serostatus. c shows the number of predicted seropositive and seronegative patients in each group. d–f compare the immune cell proportions in CMV seropositive to CMV seronegative participants in each group. Models were adjusted for age, sex, and batch variables. Only p values <0.05 are denoted (regression model t test; df = 291 for (a); df = 169 for (b); df = 292 for (d); df = 169 for (e); df = 78 for (f)). HC healthy control group, PD Parkinson’s disease group, Prod prodromal Parkinson’s disease group, Bas basophils, Eos eosinophils, Neu neutrophils, Bnv B-cells naïve, Bmem B-cells memory, CD4nv helper CD4+ T-cells naïve, CD4mem helper CD4+ T-cells memory, Treg CD4+ T regulatory cells, CD8nv cytotoxic CD8+ T-cells naïve, CD8mem cytotoxic CD8+ T-cells memory, Mono monocytes, NK natural killer cells, df degrees of freedom.
We also assessed how environmental risk factors, such as viral infections, contributed to changes in the immune cell composition of the blood60. We utilized a DNAm-based prediction model to predict cytomegalovirus (CMV) seropositivity. We did not identify an association between CMV seropositivity and having PD (Fig. 7c). However, we observed very large differences in immune cell composition between individuals who were predicted to be CMV seropositive (CMV + ) and those who were predicted to be seronegative (CMV-) (Fig. 7d–f). Across all three patient groups, we find CMV+ subjects to have elevated CD8mem (HC \({\hat{\beta }}_{Z}\,\)= 6.1, p = 4.2E-9; Prod \({\hat{\beta }}_{Z}\) = 6.7, p = 1.5E-21; PD \({\hat{\beta }}_{Z}\) = 7.6, p = 5.8E-41) and decreased CD4nv (HC \({\hat{\beta }}_{Z}\) = -2.3, p = 4.5E-4; Prod \({\hat{\beta }}_{Z}\) = -1.5, p = 0.013; PD \({\hat{\beta }}_{Z}\) = -1.5, p = 6.8E-5). We identified a decrease in Neu in CMV + PD (\({\hat{\beta }}_{Z}\) = -4.3, p = 2.5E-5) and Prod (\({\hat{\beta }}_{Z}\,\)= 0.29, p = 0.021) subjects that is not observed in CMV + HC. Similarly, we saw only changes in Bnv (\({\hat{\beta }}_{Z}\,\)= -0.60, p = 1.8E-4) and Bmem (\({\hat{\beta }}_{Z}\) = 0.25, p = 0.048) proportions in the CMV + PD subjects but not the other groups. Lastly, a decrease in NK cells in CMV + HC (\({\hat{\beta }}_{Z}\) = -1.3, p = 0.058) was not observed in CMV + PD or CMV+ Prod subjects. Together these data support that both genetic and environmental risk factors of PD also disrupt the proportions of peripheral immune cells in the blood of PD and Prod patients.
For our final analysis, we wanted to assess if there were changes in immune cell composition in the Prod subjects that converted to PD. For a small subset of patients, the PPMI collected DNAm data on patients before and after their PD diagnosis (n = 20, Supplementary Fig. 11). We compared their pre and post-conversion samples and identified an increase in CD4nv post-conversion (Fig. 8a). We also performed an EWAS analysis to identify possible DMCs associated with conversion to PD (Fig. 8b). Although no DMC surpassed our Δβ threshold, the model identified three statistically significant DMCs (cg13892688, SLC6A15, KATNAL1).
a compares the samples from patients after their clinical diagnosis of PD to those before their diagnosis of PD. The values represented are variance normalized mean differences in cell proportion with 95% confidence intervals. P values were generated from regression model t tests; df = 71.5). b shows the EWAS comparing the same set of samples. (p values generated from empirical Bayes moderated regression model t test) and the false discovery rate was controlled at 5%. The input to each EWAS analyses was the top 100,000 CpGs with the largest Δβ. Bas basophils, Eos eosinophils, Neu neutrophils, Bnv B-cells naïve, Bmem B-cells memory, CD4nv helper CD4+ T-cells naïve, CD4mem helper CD4+ T-cells memory, Treg CD4+ T regulatory cells, CD8nv cytotoxic CD8+ T-cells naïve, CD8mem cytotoxic CD8+ T-cells memory, Mono monocytes, NK natural killer cells, df degrees of freedom.
Discussion
From longitudinally collected blood DNAm data in the PPMI database, we performed an immune profiling study on clinically defined PD patients, prodromal PD patients and matched neurologically healthy participants. We sought to validate others’ findings about the relative quantities of peripheral immune cells in PD and determine at which point along the disease course these immunological perturbations occur.
We identified significant perturbations to the proportions of innate immune cells in well-established PD. Comparing PD to HC, we found an increase in neutrophils and a decrease in monocytes and eosinophils. In PD patients, there is an increase in neutrophils and monocytes over time, with a consistent trend in the Prod group as well. We observed a slight decrease in monocytes in more symptomatic PD patients. Lastly, although we observed changes in the innate immune cells at the baseline PD timepoint, there is not a strong change in the innate immune cell compartment as patients transition from prodromal PD to PD, except for a slight increase in NK cells.
There are conflicting results on whether the proportions of neutrophils change in the blood of PD patients, with some papers stating no changes to healthy controls, while others report higher neutrophil proportions or a higher neutrophil to lymphocyte ratio (NLR)23,50,61,62,63. Neutrophils are notoriously fragile and can easily be underrepresented in single-cell RNA sequencing or flow cytometry if the sample processing is not correct, which may explain why this result is mixed across studies64,65,66,67. Here, we leverage DNAm cell deconvolution strategies to analyze neutrophils in which we identified increases in the proportions of neutrophils in PD patients. In Alzheimer’s disease (AD), it is well established that the release of neutrophil extracellular traps (NET) in the vasculature of the brain can increase the permeability of the blood-brain barrier which facilitates chronic inflammation68. Our results support the notion that these cells may be involved in similar pathogenic mechanisms across chronic neuroinflammatory diseases including PD.
In concordance with others’ findings, we also reproduced a decrease in monocytes in PD patients22,23,24. It was interesting as well that we found this effect to be stronger and more correlated to symptom severity at later rather than earlier time points, which is supported in the literature by known trends in activity and phenotype as PD progresses69. Pigment-laden macrophages are observed in the degenerating mesencephalon in PD but it remains unclear to what degree peripheral monocytes contribute to this population typically thought to arise from CNS resident microglia. A decrease in circulating monocytes may result from increases in monocyte diapedesis into the CNS in response to tissue damage. The analysis described here provides an important starting point for studies designed to further subset peripheral immune cells like monocytes to contextualize their states concerning CNS damage.
Taking these findings together, we conclude that perturbations to the innate cell compartment manifest when PD is highly symptomatic and developed, whereas at very early stages of the disease, such as during the prodromal stage or very close to diagnosis, we see smaller changes in the relative proportions of innate immune cells.
We found that perturbations to the adaptive immune landscape occur at slightly different periods compared to innate immune perturbations. We identified a strong decrease in CD4+ memory T cells and naive B cells in clinically defined and prodromal PD. In PD patients, we see a significant decrease in many adaptive immune cell populations over time, including all naive lymphocytes and memory T cells. In the prodromal group, there is a significant decrease in the naïve lymphocytes but not a significant trend in the memory compartments. Symptomology changes correlated more with proportions of adaptive immune cells than innate cells. For instance, proportions of all memory lymphocyte populations were associated with REM sleep disturbances, which matches previous findings about the relationships of peripheral T cell phenotypes associated with this phenotype70. In addition, higher proportions of CD4+ naive cells were identified in patients with less cognitive decline and we found a slight increase in B memory cells in those who were more symptomatic at baseline. B memory cells were also higher in PD patients with more symptoms of depression and anxiety. Strikingly, when we compared samples of prodromal PD patients before and after their transition to clinically defined PD, we see a significant increase in CD4+ naive cells after diagnosis.
Across multiple studies, there is a decrease in T cells in the periphery, primarily naive T cell and CD4+ T cell populations23,50,54,71,72. Some studies also report a decrease in naive B cells and an increase in memory B cells within the peripheral blood, which matches the results of our analysis as well54,73. One single-cell RNA sequencing study identified an increase in isotype-switched (IgG and IgA) proportions in PD blood further supporting our results that the B cell compartment in PD patients is more antigen-experienced and shifted to more mature B cell phenotypes74.
Together, these data suggest that the perturbations to the peripheral adaptive immune profile are linked with early or acute disease events in PD. We believe that in “preclinical” PD, an adaptive immune response may be initiated against a range of antigens, possibly emanating from the gut, stress related to environmental insult, to CNS antigens related to early-stage neurodegenerative processes. Our findings are consistent with “systemic” PD, demonstrating that levels of various T and B cell populations correlated with non-motor symptoms such as sleep disturbances and cognitive ability. We observe another increase in the CD4+ naive T cell levels in patients right after they transition to a clinical diagnosis of PD. We hypothesize that the initial presentation of motor symptoms is associated with increases in neurodegenerative antigens and possibly the dissemination of protienopathic stressors such as seeding competent alpha synuclein modulating T cell state and leading to more naive T cells responding to or being recruited to the CNS34.
However, as degeneration progresses and there is sustained CNS inflammation, we observe larger changes to the innate immune system. In clinically defined PD patients, there is a significant decrease in monocyte and increase in neutrophil proportions compared to healthy controls. However, over time the proportions of both are increasing. For monocytes, we suspect that the decreased level in the periphery compared to controls may be due to an increased infiltration into the CNS, consistent with work done in rodents75,76. The levels of monocytes may be increasing over time as more CNS damage takes place and recruits more innate immune cells. The increase in neutrophils we suspect are involved in opening of the BBB to recruit more immune cells to the CNS, as others have described BBB instability in PD patients77. Similarly, BBB instability has been associated with neutrophil activity in multiple sclerosis (MS) and Alzheimer’s disease78,79. As the levels of monocytes and neutrophils are increasing in PD patients over time, we see a decrease in many lymphocyte population levels over time. Compared to healthy controls, PD patients exhibit a decrease in CD4+ memory cells and naive B cells. Because the levels of these cells are lower and they are decreasing over time, it can be interpreted in two different manners: either (1) that there is migration of these cells to the CNS, causing a decrease of their levels in the periphery or (2) there is a dampening of the adaptive immune response as a protective mechanism to mitigate the risk of CNS autoimmunity. Current hypotheses pose that chronic inflammatory signaling can cause a shift from adaptive to innate immunity as the predominating inflammatory mechanism during the progression of CNS disorders80,81,82,83,84,85,86,87. Further functional studies of these cell types from early PD patients are required to test these hypotheses.
We performed longitudinal EWAS analyses using (1) a cross-sectional EWAS analysis at each major study timepoint or (2) a longitudinal EWAS analysis by adding a random effect into the model, which will account for the longitudinally in the dataset. We found that in all of our comparisons (PD vs. HC, Prod vs. HC, and PD vs. Prod), more DMCs were called in our longitudinal models compared to the cross-sectional models. We believe that in the longitudinal modeling approach, within-patient variation is better accounted for via the random effect, and thus this increases the power of hypothesis testing.
When comparing PD vs. HC, we found a single differentially methylated CpG, cg13102742, previously associated with schizophrenia88. We saw much more of an epigenetic shift when comparing Prod vs. HC. When visualizing the distribution of DNAm values at these positions, we noticed methylation patterns that may be indicative of underlying genetic variation, since we observe a trimodal distribution centered at 0, 0.5, and 1 for many of these DMCs and that these DMCs lie close to lowly-penetrant SNPs. Our findings show that the DNA methylome seems to be more deviated from HC in the Prod group compared to the PD group. This may suggest that the DNAm perturbations are more associated with earlier disease events, in other words, the initial immune responses to alpha-synuclein.
There has been a strong body of research investigating the genetic and inheritable risk factors for developing PD identifying risk variants to glucocerebrosidase (GBA) and leucine-rich repeat kinase 2 (LRRK2) among others89,90. LRRK2 variants with pathogenic associations with PD account for 5–13% of familial PD cases and 1–5% of sporadic PD91. Variation in this protein function is thought to disrupt kinase cascades and alpha-synuclein clearance in the brain91. Interestingly, LRRK2 may also act on the neuro-immune axis in PD and other chronic inflammatory diseases92,93,94. Others have shown a potential interaction between that the LRRK2 genotype and PD case status when looking at the proportions of immune cells23. We show that LRRK2+ PD patients had lower proportions of naive B cells and higher proportions of memory B cells. It has been shown that LRRK2 is expressed in B cells and mice lacking LRRK2 have a more naive B cell repertoire95. Because this B cell trend was not observed in the Prod cohort, we hypothesize that the inflammatory-mediated genetic risk of LRRK2 variants may affect the chronic immune responses in defined PD, rather than the first presence of neuronal debris or alpha-synuclein. We repeated this analysis comparing GBA positive and negative PD patients, however, in this dataset only 20 patients were GBA allele positive, which greatly limited our statistical power. Although the trends were not significant, we did observe many of the trends to be in the same direction as the LRRK2+ vs. LRRK2- comparison. Similarly, others have stated an effect of PD-related GBA mutations on B cell biology96, thus, more research is needed to further understand how this peripheral immune activity mediates the effect of these genetic alleles on PD risk.
Many PD risk factors are related to environmental exposures such as infections. Since there may be an association between PD and viral infections, including herpes viruses97,98, we wished to test this association in the PPMI data by predicting cytomegalovirus (CMV) seroprevalence from the DNAm data99. Although the relationship between seroprevalence for CMV and PD risk is not entirely clear60,100, recent literature has shown there may be disrupted T cell and myeloid populations in CMV seropositive PD patients101,102. Firstly, we did not identify a significant association between CMV seroprevalence and having PD. However, CMV infection has been tied to various diseases; thus, this our estimate is susceptible to selection bias by over-selecting HC who did not contract CMV or are resilient to CMV-related conditions. In our HC population, we found CMV positive participants had higher proportions of CD8+ memory T cells, with decreases in NK cells, CD8+, and CD4+ naive T cells. We found these perturbations in the other groups; however, the effect was weakest in the HC group and stronger in PD patients. However, CMV positive PD and Prod patients have lower proportions of neutrophils compared to CMV negative patients and this was not observed in the HC group. Furthermore, the PD CMV positive patients exhibited a decrease in naive B cells and an increase in B memory cells. These alterations were in the same direction as what was observed with the effect LRRK2 genotype on the peripheral immune cell proportions. Together these data suggest that the way the genetic and environmental PD risk factors affect the peripheral immune system may follow overlapping mechanisms and that combinations of genetic and environmental risk may be synergistic concerning the effects on inflammation. Our findings that B cell perturbations are found in the Prod population and are associated with genetic and infectious risk factors of PD provide a rationale for the further study of B cells in early PD.
Although this study is in line with previous literature, there are certain limitations. Firstly, we anticipate residual confounding in our results due to medication use and the presence of other comorbidities. Although all patients were medication naive at baseline, some patients did go on to receive treatment during follow-up. Secondly, our analyses consider proportions of cell types rather than actual counts. Third, we are limited by the availability of deconvolution libraries to resolve certain subsets of leukocytes, such as immature from mature neutrophils or helper T-cell subtypes. As reference libraries for these cell types are developed, these analyses can be repeated to gain said resolution. Fourth, in our longitudinal analyses, we used the patient visit ID coding system to determine the number of months away from baseline the sample was taken place. This “binning” of the data will introduce a slight nondifferential misclassification bias in the data concerning the true timing of the sample. Fifth, we observed a batch effect that could not be explained by known technical or biological variables. Although we adjust for this in our analysis and performed sensitivity analyses, it remains unclear as to what is the source of this variation. Lastly, the direction of our hypothesis testing was to test if there was an effect of having PD on the proportion of a particular immune cell. Thus, we are assuming that having PD is causal to the change in immune cell proportion. There is potential for reverse causality, in which the change in immune cell proportion is casual to having PD. Until we understand more about the underlying biology of inflammation and neuroinflammation in PD, the direction of this hypothesis testing is left up to debate.
By utilizing blood DNAm from the PPMI cohort, we were able to employ cellular deconvolution to predict the proportions of immune cells and conduct a longitudinal immune profiling study. We established that in prodromal PD, the immune landscape is significantly different in age and sex-matched healthy individuals. This suggests that the peripheral immune system is responding to the earliest presence of CNS antigens (alpha-synuclein and/or CNS debris). Our findings suggest that the change in the peripheral adaptive immune profile is altered with earlier disease events. However, as the patients accrue more clinical burden and neurodegeneration accelerates, we observe dynamic shifts of innate and memory populations, which may be representative of a shift toward a chronic inflammatory state.
Methods
PPMI patient selection and data collection
For detailed patient recruitment and study protocols, please visit the PPMI consortium publications56,57, the PPMI website (https://www.ppmi-info.org/study-design/research-documents-and-sops), and ClinicalTrials.gov (NCT01141023, registered June 10, 2010). In brief, all participants gave informed consent before enrollment. Participants were organized into one of three groups (Table 1): Parkinson’s disease (PD), healthy control (HC), or prodromal PD (Prod). The PD group consists of people with clinically defined PD4. Individuals were excluded if they had received their diagnosis more than two years before screening or received, or expected to receive, treatment for PD (levodopa, dopamine agonists, MAO-B inhibitors, or amantadine) in the initial six months of follow-up. The HC group comprises age- and sex-matched individuals without significant neurological disorders and naive to PD-related medications. Lastly, the Prod group includes people with an elevated risk of progressing to PD. Individuals were excluded from this group if they met the criteria of clinically defined PD. However, a combination of symptomology (hyposmia or REM sleep behavior disorder) or genetic background (first-degree biological relative or specific gene variants) excluded these individuals from being in the HC group (Table 2). The consensus committee determined the assignment of each patient to a group through PPMI. All three groups of patients had a baseline sample (BL) in which basic demographic information and medical history were recorded. These patients were followed longitudinally with 1-4 visits per year for clinical evaluation and biospecimen collection. The following data were included in our analyses:
Whole blood DNAm data
Genomic DNA from blood collected at study visits was isolated (AutoGen Flex DNA extraction), quantified (Qubit 2.0 Fluorometer), bisulfite converted (Zymo EZ-96 DNA Methylation Kit), and loaded onto Infinium MethylationEPIC BeadChip arrays (Illumina, v1.0). The arrays were read with the Illumina iScan System to develop raw .idat files which contain DNAm data for over 850,000 CpG sites in the human genome.
Dopamine Transporter (DaTSCAN) Single-photon emission computerized tomography (SPECT)
The DaTScan SPECT imaging103 was done according to the PPMI imaging protocol. This imaging modality allows for visualizing DA presynaptic neuron loss in the striatum. At least two nuclear medicine experts analyzed all images for interpretation. Raw data were reconstructed and processed according to the stated methods for calculating the striatal binding ratio (SBR). In brief, regions of interest were placed on 9-slice averaged images to select the left and right putamen and caudate. The occipital cortex was used as a reference. The SBR was calculated as the ratio of the signal intensity of the target region to the reference region minus one. The asymmetry index (AsI) is calculated as [(R-L)/(R + L)]*100 where R and L represent the SBR of the left and right putamen or caudate (Table 3).
Clinical evaluations
For this analysis, we used data on seven clinical exams (MDS-UPDRS1, MDS-UPDRS2, ESS, MoCA, STAI, REM, and GDS) to assess possible relationships with different sets of PD-related symptoms (Table 3).
DNAm data preprocessing
Data preprocessing and analysis were done in R Studio (v4.2). All available DNAm data as of July 2022 were downloaded from the PPMI web interface in raw .idat format. Patient data were matched to available DNAm data. Patient visits annotated to multiple arrays was excluded. Additionally, if any sample had missing sex or age information, it was also excluded. The remaining data were analyzed using a combination of functions from various R packages, including minfi [v1.42.0]104,105, sesame [v1.14.2]106, ENmix [v1.32.0]107,108, and ewastools [v1.7]109. We utilized normal-exponential out-of-band correction with linear dye-bias normalization to derive a beta value for every array position for every sample110.
DNAm data quality control and filtering
We first employed a sample-wise filtering approach to determine each array’s quality and remove data that fails quality control assessments. Each sample had to pass three criteria to be included in the analysis: (1) the median beta value of the sample had to lie within three inter-quartile ranges of the set of all median beta values in the study, (2) the predicted sex, as estimated by the minfi package, matches the reported sex, (3) all quality control probe signals are within the appropriate regions as identified by the ENmix and ewastools software. The number of samples and their reason for exclusion are listed in Supplementary Table 1.
Next, we employed a probe-wise filtering approach to remove poor-quality probe data within each sample. Detection p-values were used to determine if the intensity signal for each probe was significantly above the background control probes (p < 0.05). If the probe fails, its beta value was masked as missing. In addition, probes located on the X or Y chromosome, near single nucleotide polymorphisms (SNPs), or in regions of known array cross-reactivity were masked, allowing for a final set of 743,840 CpGs of data to remain in the study for analysis. For each CpG, a β value was calculated, representing the methylation level at that position.
Calculation of DNAm-derived variables
Blood immune cell proportions
The blood immune cell fractions were calculated using the FlowSorted.BloodExtended.EPIC software42. These values were then adjusted for their limit of detection (LoD) values as previously described111. Any value below the cell type-specific LoD was replaced by the LoD/\(\surd 2\). After replacing the LoD, the remaining cell types were scaled, so the sum of all cell proportions was constrained to 1.
Cytomegalovirus (CMV) seropositivity prediction
CMV seropositivity was calculated with a previously published elastic net prediction model with the output delivered as a binary seropositive or seronegative for CMV99.
Statistical analysis: comparing immune cell compositions
We utilized linear regression models to test the effect of a binary variable, \({X}_{Z}\), on the composition of immune cells at a single cross-sectional timepoint. Let \({y}_{i}\) represent the proportion of cell type \(i\). For every cell type, the cross-sectional regression model at a given timepoint is represented as Eq. I where \(\alpha\) is the intercept term, \(\varepsilon\) is the error term, and \(X\) is the covariate matrix, consisting of a binary variable for sex, a continuous variable for the age of the patient at their baseline visit, and two binary batch effect variables. An estimate for \({\hat{\beta }}_{{Z}_{i}}\) and 95% confidence intervals were calculated for each cell type and plotted on the radar plots. P-values were generated for each estimate by performing a regression model t-test to determine if estimate was significantly nonzero. For the variance-normalized radar plots, the estimate and confidence intervals for \({\hat{\beta }}_{{Z}_{i}}\) was divided by the standard deviation of \({y}_{i}\) in the reference group (Eq. II). We adapted this approach for longitudinal data by introducing a random effects term to Eq. I by adding a random effect for each patient (\({u}_{{subject}}\)) and time as a continuous fixed effect (Eq. III). To identify if there was an interaction between time and \({X}_{Z}\) for each cell type, we performed likelihood ratio tests using the lmtest and lmerTest software, comparing the models generated by Eq. III and Eq. IV to test the hypothesis that adding the interaction better fits the data112,113.
Statistical Analysis: K-means clustering of clinical data
We performed k-means clustering using the cluster software114, including the following clinical variables: MDS-UPDRS1, MDS-UPDRS2, ESS, MoCA, GDS, STAI, REM, putamen AsI, and caudate AsI. Values were mean-centered and scaled before clustering. Decisions about the optimal number of clusters to divide the data were determined using the fviz_nbclust and clusGap visualizations.
Statistical analysis: longitudinal epigenome-wide association studies (EWAS)
EWAS analysis was done using the minfi and limma software115. To identify differentially methylated CpGs (DMCs) between two groups, we utilized the same regression approach as in Eq. I, but instead \({y}_{i}\) represents the DNAm level at CpG \(i\). DNAm proportions (β values) were converted to M-values by computing the logit of the β values in base 2. We subset the EWAS analyses to the top 100,000 CpGs with the largest mean difference in β values (Δβ) between the two groups. An empirical Bayes method was used to normalize CpG-wise residual variance. P-values for each CpG were generated using regression model t-tests to determine if the coefficient is nonzero. P-values were adjusted for multiple hypothesis testing using the Benjamini-Hochberg procedure to maintain a false discovery rate of 5%. To perform a longitudinal EWAS analysis, we utilize the duplicateCorrelation functionality in the limma software to produce models equivalent to Eq. II for differential methylation testing, including the random effect to adjust for repeated measurements.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Data used in the preparation of this article were obtained from the Parkinson’s Progression Markers Initiative (PPMI) database (www.ppmi-info.org/access-data-specimens/download-data), RRID:SCR_006431. For up-to-date information on the study, visit www.ppmi-info.org.
Code availability
All R scripts used to preprocess and analyze the data are available on our laboratory’s GitHub page (https://github.com/SalasLab/PPMI_DNAm_Analysis) and on Zenodo116.
References
Latif, S. et al. Dopamine in Parkinson’s disease. Clin. Chim. Acta 522, 114–126 (2021).
Surmeier, D. J., Obeso, J. A. & Halliday, G. M. Selective neuronal vulnerability in Parkinson disease. Nat. Rev. Neurosci. 18, 101–113 (2017).
Postuma, R. B. et al. MDS clinical diagnostic criteria for Parkinson’s disease. Mov. Disord. 30, 1591–1601 (2015).
Gelb, D. J., Oliver, E. & Gilman, S. Diagnostic criteria for Parkinson disease. Arch. Neurol. 56, 33–39 (1999).
Chen, H. et al. Olfaction and incident Parkinson disease in US white and black older adults. Neurology 89, 1441–1447 (2017).
Tekriwal, A. et al. REM sleep behaviour disorder: prodromal and mechanistic insights for Parkinson’s disease. J. Neurol. Neurosurg. Psychiatry 88, 445–451 (2017).
Postuma, R. B. & Berg, D. Prodromal Parkinson’s disease: the decade past, the decade to come. Mov. Disord. 34, 665–675 (2019).
Marogianni, C. et al. Neurodegeneration and inflammation: an interesting interplay in Parkinson’s disease. Int J. Mol. Sci. 21, 8421 (2020).
Tansey, M. G. et al. Inflammation and immune dysfunction in Parkinson disease. Nat. Rev. Immunol. 22, 657–673 (2022).
Saiki, M. et al. Association of the human leucocyte antigen region with susceptibility to Parkinson’s disease. J. Neurol. Neurosurg. Psychiatry 81, 890–891 (2010).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014).
Zhang, X. et al. Immune profiling of Parkinson’s disease revealed its association with a subset of infiltrating cells and signature genes. Front Aging Neurosci. 13, 605970 (2021).
Evans, F. L., Dittmer, M., de la Fuente, A. G. & Fitzgerald, D. C. Protective and regenerative roles of T cells in central nervous system disorders. Front Immunol. 10, 2171 (2019).
McGeer, P. L., Itagaki, S., Boyes, B. E. & McGeer, E. G. Reactive microglia are positive for HLA-DR in the substantia nigra of Parkinson’s and Alzheimer’s disease brains. Neurology 38, 1285–1291 (1988).
Orr, C. F., Rowe, D. B., Mizuno, Y., Mori, H. & Halliday, G. M. A possible role for humoral immunity in the pathogenesis of Parkinson’s disease. Brain 128, 2665–2674 (2005).
Tansey, M. G. & Romero-Ramos, M. Immune system responses in Parkinson’s disease: early and dynamic. Eur. J. Neurosci. 49, 364–383 (2019).
Heneka, M. T., Kummer, M. P. & Latz, E. Innate immune activation in neurodegenerative disease. Nat. Rev. Immunol. 14, 463–477 (2014).
Santos-Lima, B., Pietronigro, E. C., Terrabuio, E., Zenaro, E. & Constantin, G. The role of neutrophils in the dysfunction of central nervous system barriers. Front Aging Neurosci. 14, 965169 (2022).
Gellhaar, S., Sunnemark, D., Eriksson, H., Olson, L. & Galter, D. Myeloperoxidase-immunoreactive cells are significantly increased in brain areas affected by neurodegeneration in Parkinson’s and Alzheimer’s disease. Cell Tissue Res. 369, 445–454 (2017).
Heilbron, K. et al. The Parkinson’s phenome-traits associated with Parkinson’s disease in a broadly phenotyped cohort. NPJ Parkinsons Dis. 5, 4 (2019).
Cheng, C. M. et al. Risk of developing Parkinson’s disease among patients with asthma: a nationwide longitudinal study. Allergy 70, 1605–1612 (2015).
Tian, J. et al. Specific immune status in Parkinson’s disease at different ages of onset. NPJ Parkinsons Dis. 8, 5 (2022).
Muñoz-Delgado, L. et al. Peripheral inflammatory immune response differs among sporadic and familial Parkinson’s disease. NPJ Parkinsons Dis. 9, 12 (2023).
Nissen, S. K. et al. Alterations in blood monocyte functions in Parkinson’s disease. Mov. Disord. 34, 1711–1721 (2019).
Earls, R. H. & Lee, J. K. The role of natural killer cells in Parkinson’s disease. Exp. Mol. Med. 52, 1517–1525 (2020).
Niwa, F., Kuriyama, N., Nakagawa, M. & Imanishi, J. Effects of peripheral lymphocyte subpopulations and the clinical correlation with Parkinson’s disease. Geriatr. Gerontol. Int. 12, 102–107 (2012).
Cen, L. et al. Peripheral lymphocyte subsets as a marker of Parkinson’s disease in a Chinese population. Neurosci. Bull. 33, 493–500 (2017).
Mihara, T. et al. Natural killer cells of Parkinson’s disease patients are set up for activation: a possible role for innate immunity in the pathogenesis of this disease. Parkinsonism Relat. Disord. 14, 46–51 (2008).
Menees, K. B. & Lee, J. K. New insights and implications of natural killer cells in Parkinson’s disease. J. Parkinsons Dis. 12, S83–S92 (2022).
Mayne, K., White, J. A., McMurran, C. E., Rivera, F. J. & de la Fuente, A. G. Aging and neurodegenerative disease: is the adaptive immune system a friend or foe? Front Aging Neurosci. 12, 572090 (2020).
McGeer, P. L., Itagaki, S., Akiyama, H. & McGeer, E. G. Rate of cell death in parkinsonism indicates active neuropathological process. Ann. Neurol. 24, 574–576 (1988).
Brochard, V. et al. Infiltration of CD4+ lymphocytes into the brain contributes to neurodegeneration in a mouse model of Parkinson disease. J. Clin. Invest 119, 182–192 (2009).
Sommer, A. et al. Th17 Lymphocytes Induce Neuronal Cell Death in a Human iPSC-Based Model of Parkinson’s Disease. Cell Stem Cell 23, 123–131.e126 (2018).
Sulzer, D. et al. T cells from patients with Parkinson’s disease recognize α-synuclein peptides. Nature 546, 656–661 (2017).
Lindestam Arlehamn, C. S. et al. α-Synuclein-specific T cell reactivity is associated with preclinical and early Parkinson’s disease. Nat. Commun. 11, 1875 (2020).
Kustrimovic, N. et al. Parkinson’s disease patients have a complex phenotypic and functional Th1 bias: cross-sectional studies of CD4+ Th1/Th2/T17 and Treg in drug-naive and drug-treated patients. J. Neuroinflammation 15, 205 (2018).
Akhtar, R. S. et al. Measurements of auto-antibodies to α-synuclein in the serum and cerebral spinal fluids of patients with Parkinson’s disease. J. Neurochem. 145, 489–503 (2018).
Smith, L. M., Schiess, M. C., Coffey, M. P., Klaver, A. C. & Loeffler, D. A. α-Synuclein and anti-α-synuclein antibodies in Parkinson’s disease, atypical Parkinson syndromes, REM sleep behavior disorder, and healthy controls. PLoS ONE 7, e52285 (2012).
De Luca, C., Colangelo, A. M., Alberghina, L. & Papa, M. Neuro-immune hemostasis: homeostasis and diseases in the central nervous system. Front Cell Neurosci. 12, 459 (2018).
Moore, L. D., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38, 23–38 (2013).
Suelves, M., Carrió, E., Núñez-Álvarez, Y. & Peinado, M. A. DNA methylation dynamics in cellular commitment and differentiation. Brief. Funct. Genomics 15, 443–453 (2016).
Salas, L. A. et al. Enhanced cell deconvolution of peripheral blood using DNA methylation for high-resolution immune profiling. Nat. Commun. 13, 761 (2022).
Titus, A. J., Gallimore, R. M., Salas, L. A. & Christensen, B. C. Cell-type deconvolution from DNA methylation: a review of recent applications. Hum. Mol. Genet. 26, R216–R224 (2017).
Muse, M. E., Carroll, C. D., Salas, L. A., Karagas, M. R. & Christensen, B. C. Application of novel breast biospecimen cell type adjustment identifies shared DNA methylation alterations in breast tissue and milk with breast cancer risk factors. Cancer Epidemiol. Biomark. Prev. 32, 550–560 (2023).
Zhang, Z. et al. HiTIMED: hierarchical tumor immune microenvironment epigenetic deconvolution for accurate cell type resolution in the tumor microenvironment using tumor-type-specific DNA methylation data. J. Transl. Med. 20, 516 (2022).
Teschendorff, A. E. & Zheng, S. C. Cell-type deconvolution in epigenome-wide association studies: a review and recommendations. Epigenomics 9, 757–768 (2017).
Schaffner, S. L. & Kobor, M. S. DNA methylation as a mediator of genetic and environmental influences on Parkinson’s disease susceptibility: Impacts of alpha-Synuclein, physical activity, and pesticide exposure on the epigenome. Front Genet. 13, 971298 (2022).
Henderson, A. R. et al. DNA methylation and expression profiles of whole blood in Parkinson’s disease. Front Genet. 12, 640266 (2021).
Henderson-Smith, A. et al. DNA methylation changes associated with Parkinson’s disease progression: outcomes from the first longitudinal genome-wide methylation analysis in blood. Epigenetics 14, 365–382 (2019).
Saunders, J. A. et al. CD4+ regulatory and effector/memory T cell subsets profile motor dysfunction in Parkinson’s disease. J. Neuroimmune Pharm. 7, 927–938 (2012).
Chen, X. et al. Evidence for peripheral immune activation in Parkinson’s disease. Front Aging Neurosci. 13, 617370 (2021).
Dommershuijsen, L. J. et al. Peripheral immune cell numbers and C-reactive protein in Parkinson’s disease: results from a population-based study. J. Parkinsons Dis. 12, 667–678 (2022).
Jiang, S., Gao, H., Luo, Q., Wang, P. & Yang, X. The correlation of lymphocyte subsets, natural killer cell, and Parkinson’s disease: a meta-analysis. Neurol. Sci. 38, 1373–1380 (2017).
Yan, Z. et al. Dysregulation of the adaptive immune system in patients with early-stage Parkinson disease. Neurol. Neuroimmunol. Neuroinflamm 8, e1036 (2021).
McKeith, I. G. et al. Research criteria for the diagnosis of prodromal dementia with Lewy bodies. Neurology 94, 743–755 (2020).
Initiative, P. P. M. The Parkinson Progression Marker Initiative (PPMI). Prog. Neurobiol. 95, 629–635 (2011).
Marek, K. et al. The Parkinson’s progression markers initiative (PPMI) - establishing a PD biomarker cohort. Ann. Clin. Transl. Neurol. 5, 1460–1477 (2018).
Roos, D. S., Klein, M., Deeg, D. J. H., Doty, R. L. & Berendse, H. W. Prevalence of prodromal symptoms of Parkinson’s disease in the late middle-aged population. J. Parkinsons Dis. 12, 967–974 (2022).
Billingsley, K. J., Bandres-Ciga, S., Saez-Atienzar, S. & Singleton, A. B. Genetic risk factors in Parkinson’s disease. Cell Tissue Res. 373, 9–20 (2018).
Bu, X. L. et al. The association between infectious burden and Parkinson’s disease: a case-control study. Parkinsonism Relat. Disord. 21, 877–881 (2015).
Ataç Uçar, C., Gökçe Çokal, B., Ünal Artık, H. A., İnan, L. E. & Yoldaş, T. K. Comparison of neutrophil-lymphocyte ratio (NLR) in Parkinson’s disease subtypes. Neurol. Sci. 38, 287–293 (2017).
Muñoz-Delgado, L. et al. Peripheral immune profile and neutrophil-to-lymphocyte ratio in Parkinson’s disease. Mov. Disord. 36, 2426–2430 (2021).
Liu, Z., Fan, Q., Wu, S., Wan, Y. & Lei, Y. Compared with the monocyte to high-density lipoprotein ratio (MHR) and the neutrophil to lymphocyte ratio (NLR), the neutrophil to high-density lipoprotein ratio (NHR) is more valuable for assessing the inflammatory process in Parkinson’s disease. Lipids Health Dis. 20, 35 (2021).
Dorward, D. A. et al. Technical advance: autofluorescence-based sorting: rapid and nonperturbing isolation of ultrapure neutrophils to determine cytokine production. J. Leukoc. Biol. 94, 193–202 (2013).
Xu, H. & Jia, J. Single-cell RNA sequencing of peripheral blood reveals immune cell signatures in Alzheimer’s disease. Front Immunol. 12, 645666 (2021).
Schafflick, D. et al. Integrated single cell analysis of blood and cerebrospinal fluid leukocytes in multiple sclerosis. Nat. Commun. 11, 247 (2020).
Diks, A. M. et al. Impact of blood storage and sample handling on quality of high dimensional flow cytometric data in multicenter clinical research. J. Immunol. Methods 475, 112616 (2019).
Pietronigro, E. C., Della Bianca, V., Zenaro, E. & Constantin, G. NETosis in Alzheimer’s Disease. Front Immunol. 8, 211 (2017).
Konstantin Nissen, S. et al. Changes in CD163+, CD11b+, and CCR2+ peripheral monocytes relate to Parkinson’s disease and cognition. Brain Behav. Immun. 101, 182–193 (2022).
De Francesco, E. et al. CD4+ T-cell transcription factors in idiopathic REM sleep behavior disorder and Parkinson’s disease. Mov. Disord. 36, 225–229 (2021).
Bhatia, D. et al. T-cell dysregulation is associated with disease severity in Parkinson’s Disease. J. Neuroinflammation 18, 250 (2021).
Baird, J. K., Bourdette, D., Meshul, C. K. & Quinn, J. F. The key role of T cells in Parkinson’s disease pathogenesis and therapy. Parkinsonism Relat. Disord. 60, 25–31 (2019).
Scott, K. M. et al. B lymphocyte responses in Parkinson’s disease and their possible significance in disease progression. Brain Commun. 5, fcad060 (2023).
Wang, P. et al. Global characterization of peripheral B cells in Parkinson’s disease by single-cell RNA and BCR sequencing. Front Immunol. 13, 814239 (2022).
Williams, G. P. et al. CD4 T cells mediate brain inflammation and neurodegeneration in a mouse model of Parkinson’s disease. Brain 144, 2047–2059 (2021).
Xu, E. et al. Pathological α-synuclein recruits LRRK2 expressing pro-inflammatory monocytes to the brain. Mol. Neurodegener. 17, 7 (2022).
Al-Bachari, S., Naish, J. H., Parker, G. J. M., Emsley, H. C. A. & Parkes, L. M. Blood-brain barrier leakage is increased in Parkinson’s disease. Front Physiol. 11, 593026 (2020).
Aries, M. L. & Hensley-McBain, T. Neutrophils as a potential therapeutic target in Alzheimer’s disease. Front Immunol. 14, 1123149 (2023).
Aubé, B. et al. Neutrophils mediate blood-spinal cord barrier disruption in demyelinating neuroinflammatory diseases. J. Immunol. 193, 2438–2454 (2014).
Furman, D. et al. Chronic inflammation in the etiology of disease across the life span. Nat. Med. 25, 1822–1832 (2019).
Weiner, H. L. A shift from adaptive to innate immunity: a potential mechanism of disease progression in multiple sclerosis. J. Neurol. 255, 3–11 (2008).
Shastri, A., Bonifati, D. M. & Kishore, U. Innate immunity and neuroinflammation. Mediators Inflamm. 2013, 342931 (2013).
Gilli, F., Li, L. & Pachner, A. R. The immune response in the CNS in Theiler’s virus induced demyelinating disease switches from an early adaptive response to a chronic innate-like response. J. Neurovirol. 22, 66–79 (2016).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
DeMaio, A., Mehrotra, S., Sambamurti, K. & Husain, S. The role of the adaptive immune system and T cell dysfunction in neurodegenerative diseases. J. Neuroinflammation 19, 251 (2022).
Sabin, K. Z. & Echeverri, K. The role of the immune system during regeneration of the central nervous system. J. Immunol. Regen. Med. 7, 100023 (2020).
Saferding, V. & Blüml, S. Innate immunity as the trigger of systemic autoimmune diseases. J. Autoimmun. 110, 102382 (2020).
He, F. et al. Exploration of mutated genes and prediction of potential biomarkers for childhood-onset schizophrenia using an integrated bioinformatic analysis. Front Aging Neurosci. 14, 829217 (2022).
Houlden, H. & Singleton, A. B. The genetics and neuropathology of Parkinson’s disease. Acta Neuropathol. 124, 325–338 (2012).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Rui, Q., Ni, H., Li, D., Gao, R. & Chen, G. The Role of LRRK2 in Neurodegeneration of Parkinson Disease. Curr. Neuropharmacol. 16, 1348–1357 (2018).
Wallings, R. L. & Tansey, M. G. LRRK2 regulation of immune-pathways and inflammatory disease. Biochem. Soc. Trans. 47, 1581–1595 (2019).
Herrick, M. K. & Tansey, M. G. Is LRRK2 the missing link between inflammatory bowel disease and Parkinson’s disease? NPJ Parkinsons Dis. 7, 26 (2021).
Cabezudo, D., Baekelandt, V. & Lobbestael, E. Multiple-hit hypothesis in Parkinson’s disease: LRRK2 and inflammation. Front Neurosci. 14, 376 (2020).
Kubo, M. et al. Leucine-rich repeat kinase 2 is a regulator of B cell function, affecting homeostasis, BCR signaling, IgA production, and TI antigen responses. J. Neuroimmunol. 292, 1–8 (2016).
Kobo, H. et al. Down-regulation of B cell-related genes in peripheral blood leukocytes of Parkinson’s disease patients with and without GBA mutations. Mol. Genet. Metab. 117, 179–185 (2016).
Leta, V. et al. Viruses, parkinsonism and Parkinson’s disease: the past, present and future. J. Neural Transm. (Vienna) 129, 1119–1132 (2022).
Caggiu, E. et al. Inflammation, infectious triggers, and Parkinson’s disease. Front Neurol. 10, 122 (2019).
Bergstedt, J. et al. The immune factors driving DNA methylation variation in human blood. Nat. Commun. 13, 5895 (2022).
Wang, H. et al. Bacterial, viral, and fungal infection-related risk of Parkinson’s disease: meta-analysis of cohort and case-control studies. Brain Behav. 10, e01549 (2020).
Vavilova, J. D. et al. Reduced immunosenescence of peripheral blood T cells in Parkinson’s disease with CMV infection background. Int J. Mol. Sci. 22, 13119 (2021).
Goldeck, D., Maetzler, W., Berg, D., Oettinger, L. & Pawelec, G. Altered dendritic cell subset distribution in patients with Parkinson’s disease: impact of CMV serostatus. J. Neuroimmunol. 290, 60–65 (2016).
Bega, D. et al. Clinical utility of DaTscan in patients with suspected Parkinsonian syndrome: a systematic review and meta-analysis. NPJ Parkinsons Dis. 7, 43 (2021).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Fortin, J. P., Triche, T. J. & Hansen, K. D. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 33, 558–560 (2017).
Zhou, W., Triche, T. J., Laird, P. W. & Shen, H. SeSAMe: reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nucleic Acids Res. 46, e123 (2018).
Xu, Z., Niu, L., Li, L. & Taylor, J. A. ENmix: a novel background correction method for Illumina HumanMethylation450 BeadChip. Nucleic Acids Res. 44, e20 (2016).
Xu, Z., Niu, L. & Taylor, J. A. The ENmix DNA methylation analysis pipeline for Illumina BeadChip and comparisons with seven other preprocessing pipelines. Clin. Epigenet. 13, 216 (2021).
Heiss, J. A. & Just, A. C. Identifying mislabeled and contaminated DNA methylation microarray data: an extended quality control toolset with examples from GEO. Clin. Epigenet. 10, 73 (2018).
Triche, T. J., Weisenberger, D. J., Van Den Berg, D., Laird, P. W. & Siegmund, K. D. Low-level processing of Illumina Infinium DNA Methylation BeadArrays. Nucleic Acids Res. 41, e90 (2013).
Bell-Glenn, S. et al. Calculating detection limits and uncertainty of reference-based deconvolution of whole-blood DNA methylation data. Epigenomics 15, 435–451 (2023).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest Package: Tests in Linear Mixed Effects Models. J. Stat. Softw. 82, 1–26 (2017).
Diagnostic Checking in Regression Relationships (R News, 2002).
Maechler, M., Rousseeuw, P., Struyf, A., Hubert, M., Hornik, K. cluster: Cluster Analysis Basics and Extensions (2022).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Code from manuscript “Immunological shifts in early-stage Parkinson’s disease identified using longitudinal DNA methylation-based cellular deconvolution of the Parkinson’s Progression Marker’s Initiative blood samples” https://doi.org/10.5281/zenodo.10054443.
Goetz, C. G. et al. Movement Disorder Society-sponsored revision of the Unified Parkinson’s Disease Rating Scale (MDS-UPDRS): scale presentation and clinimetric testing results. Mov. Disord. 23, 2129–2170 (2008).
Johns, M. W. A new method for measuring daytime sleepiness: the Epworth sleepiness scale. Sleep 14, 540–545 (1991).
Doneh, B. Epworth sleepiness scale. Occup. Med (Lond.) 65, 508 (2015).
Nasreddine, Z. S. et al. The Montreal Cognitive Assessment, MoCA: a brief screening tool for mild cognitive impairment. J. Am. Geriatr. Soc. 53, 695–699 (2005).
Spielberger, C. D., Gorsuch, R. L. & Lushene, R. E. Manual for the State-Trait Anxiety Inventory (1970).
Oei, T. P., Evans, L. & Crook, G. M. Utility and validity of the STAI with anxiety disorder patients. Br. J. Clin. Psychol. 29, 429–432 (1990).
Stiasny-Kolster, K. et al. The REM sleep behavior disorder screening questionnaire-a new diagnostic instrument. Mov. Disord. 22, 2386–2393 (2007).
Yesavage, J. A. et al. Development and validation of a geriatric depression screening scale: a preliminary report. J. Psychiatr. Res. 17, 37–49 (1982).
Tinaz, S. et al. Semiquantitative analysis of dopamine transporter scans in patients with Parkinson disease. Clin. Nucl. Med 43, e1–e7 (2018).
Shigekiyo, T. & Arawaka, S. Laterality of specific binding ratios on DAT-SPECT for differential diagnosis of degenerative parkinsonian syndromes. Sci. Rep. 10, 15761 (2020).
Acknowledgements
This study received no funding. Data used in the preparation of this article were obtained from the Parkinson’s Progression Markers Initiative (PPMI) database (www.ppmi-info.org/access-data-specimens/download-data). For up-to-date information on the study, visit www.ppmi-info.org. PPMI—a public-private partnership—is funded by the Michael J. Fox Foundation for Parkinson’s Research and funding partners, including 4D Pharma, AbbVie Inc., AcureX Therapeutics, Allergan, Amathus Therapeutics, Aligning Science Across Parkinson’s (ASAP), Avid Radiopharmaceuticals, Bial Biotech, Biogen, BioLegend, BlueRock Therapeutics, Bristol Myers Squibb, Calico Life Sciences LLC, Celgene Corporation, DaCapo Brainscience, Denali Therapeutics, The Edmond J. Safra Foundation, Eli Lilly and Company, Gain Therapeutics, GE Healthcare, GlaxoSmithKline, Golub Capital, Handl Therapeutics, Insitro, Janssen Pharmaceuticals, Lundbeck, Merck & Co., Inc., Meso Scale Diagnostics, LLC, Neurocrine Biosciences, Pfizer Inc., Piramal Imaging, Prevail Therapeutics, F. Hoffmann-La Roche Ltd and its affiliated company Genentech Inc., Sanofi Genzyme, Servier, Takeda Pharmaceutical Company, Teva Neuroscience, Inc., UCB, Vanqua Bio, Verily Life Sciences, Voyager Therapeutics, Inc. and Yumanity Therapeutics, Inc. We would like to acknowledge all persons included in the PPMI studies with particular thanks to the patients, their families, medical professionals, and research facilitators. We would also like to acknowledge the funding sources that have provided salary support to the authors: Burroughs Wellcome Fund: Big Data in the Life Sciences Fellowship (S.C.P.), Department of Defense Congressionally Directed Medical Research Program W81XWH-20-1-0778 (L.A.S.), NIGMS P20 GM104416-09/8299 (Z.Z. and L.A.S.), NIEHS 5R01ES033462 (M.H.), and NIEHS 5R01ES024745 (M.H.). In addition, we would like to thank Dr. Michael Linzey and Andrew Plese for proofreading the manuscript and providing editorial feedback.
Author information
Authors and Affiliations
Contributions
S.C.P. and L.A.S. were responsible for the study design, statistical analysis, and drafting of the manuscript. S.C.P., M.H., F.G., Z.Z., and L.A.S. aided in the interpretation of the results, collection of prior literature, and preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pike, S.C., Havrda, M., Gilli, F. et al. Immunological shifts during early-stage Parkinson’s disease identified with DNA methylation data on longitudinally collected blood samples. npj Parkinsons Dis. 10, 21 (2024). https://doi.org/10.1038/s41531-023-00626-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41531-023-00626-6