Abstract
Lack of rigorous reproducibility and validation are significant hurdles for scientific development across many fields. Materials science, in particular, encompasses a variety of experimental and theoretical approaches that require careful benchmarking. Leaderboard efforts have been developed previously to mitigate these issues. However, a comprehensive comparison and benchmarking on an integrated platform with multiple data modalities with perfect and defect materials data is still lacking. This work introduces JARVIS-Leaderboard, an open-source and community-driven platform that facilitates benchmarking and enhances reproducibility. The platform allows users to set up benchmarks with custom tasks and enables contributions in the form of dataset, code, and meta-data submissions. We cover the following materials design categories: Artificial Intelligence (AI), Electronic Structure (ES), Force-fields (FF), Quantum Computation (QC), and Experiments (EXP). For AI, we cover several types of input data, including atomic structures, atomistic images, spectra, and text. For ES, we consider multiple ES approaches, software packages, pseudopotentials, materials, and properties, comparing results to experiment. For FF, we compare multiple approaches for material property predictions. For QC, we benchmark Hamiltonian simulations using various quantum algorithms and circuits. Finally, for experiments, we use the inter-laboratory approach to establish benchmarks. There are 1281 contributions to 274 benchmarks using 152 methods with more than 8 million data points, and the leaderboard is continuously expanding. The JARVIS-Leaderboard is available at the website: https://pages.nist.gov/jarvis_leaderboard/
Similar content being viewed by others
Introduction
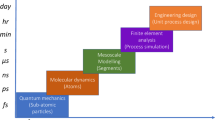
The accelerated design and characterization of materials of technological interest has been a rapidly evolving area of research in the last few decades1. Materials design requires approaches spanning a variety of length and time scales2. For atomistic design, the methods employed may include computational approaches such as density functional theory, tight-binding, and force-fields, and highly accurate approaches such as quantum Monte Carlo or quantum computations. A wide range of approaches are employed above the purely atomistic level, such as mesoscale and finite-element methods3. Similarly, experimental characterization approaches include X-ray diffraction, vibroscopy, manometry, scanning electron microscopy, and magnetic susceptibility measurements.
Moreover, data produced from these techniques can be of various types: chemical formulae, atomic/micro-structures, images, spectra, and text-documents4,5,6. The data analysis and curation methods add further complexity to benchmarking efforts, which are extremely important7,8,9,10,11,12,13,14,15,16,17,18. For example, more than 70 % of research works were shown to be non-reproducible19,20,21, and this number could be much higher depending upon the field of investigation. Although there have been significant advances in individual fields, there is an urgent need to establish a large-scale benchmark for systematic, reproducible, transparent, and unbiased scientific development.
Developing such metrology is a highly challenging task, even for one of these methods, let alone the entire galaxy of available methods. Projects and approaches, such as the materials genome and FAIR initiatives1,22, have resulted in several well-curated datasets and benchmarks. These, in turn, have led to several materials informatics applications23,24,25,26. Although electronic structure approaches such as density functional theory (DFT) tend to be more reproducible than other categories16,27, a systematic effort must be made to validate these methods and estimate the error in predictions. Hence, it is highly desirable to have a large-scale benchmarking platform in the materials science field for reproducibility and method validation.
Massive progress in fields such as image recognition/image classification (ImageNet28), protein structure prediction (AlphaFold29), large language modeling (Generative pretrained transformers (GPT))30) has been possible primarily because of well-defined benchmarks in respective fields. With regards to AI methods for structure-to-property predictions31, benchmarking efforts have enabled drastic improvements in the accuracy of predicted properties (i.e., moving away from descriptor-based predictions and including graph neural networks in the model architectures to improve accuracy).
For deterministic electronic structure methods such as DFT, extensive benchmarking of software and different DFT approximations (functionals, pseudopotentials, etc.) has led to increased reproducibility and precision in individual results and workflows27,32. Such benchmarks allow a wide community to solve problems collectively and systematically. In addition, since there already exists highly accurate models for specific tasks (i.e., energy prediction), more comprehensive evaluations of the models are required so that the performance ranking is not overfitted to one biased data source. We believe that such a universal and large-scale set of benchmarks for materials science will significantly benefit the scientific community.
To this date, several benchmarks of individual methods have already been developed. For artificial intelligence (AI) methods, there have been several benchmarks and leaderboards, such as MatBench33. MatBench provides a leaderboard for machine-learned structure-based property predictions of inorganic materials using 13 supervised machine learning tasks (thermodynamic, tensile, optical, thermal, elastic, and electronic properties) from 10 datasets (including DFT and experiment)33. Similar AI benchmarking and leaderboard platforms include MoleculeNet34, OpenCatalystProject35, sGDML36,37, mLEARN38, MatScholar39, and AtomAI40. For electronic structure methods, some of the notable benchmarks include the work by Lejaeghere et al.27, Borlido et al.41, Huber et al.42, Zhang et al.43, Tran et al.44 and several other projects45,46,47,48. Other method benchmarks include phase-field benchmarks by Wheeler et al.49, Lindsay et al.50, and microscopy benchmarks such as by Wei et al.51. A few additional benchmarking studies in materials science include refs. 52,53,54,55,56,57,58,59,60,61,62,63,64. More details on some of these benchmarking efforts are provided in later sections.
The goal of this project is to provide a more comprehensive framework for materials benchmarking than previous works. In particular, most existing efforts: 1) lack the flexibility to readily incorporate new tasks or benchmarks, which is a limitation given the continuous discovery of new materials and quantities in science, 2) are specialized towards a single modality, such as electronic structure, rather than providing a comprehensive framework that can accommodate multiple modalities, 3) offer only a limited set of tasks or properties, 4) are primarily focused on computational methods, overlooking the importance of experimental benchmarking, and 5) make adding contributions to existing platforms rather complex, creating a barrier to entry. In general, there is a need to simplify the process of user contributions to leaderboards to foster broader community engagement.
In this work, we present a user-friendly, comprehensive approach to integrate the benchmarking of both computational, experimental and data-analytics methods. The JARVIS-Leaderboard framework (https://pages.nist.gov/jarvis_leaderboard/) covers a variety of categories: Artificial Intelligence (AI), Electronic Structure (ES), Force-field (FF), Quantum Computation (QC), and Experiments (EXP). It also covers various data types, including atomic structures, spectra, images, and text. This project can be used to: (1) check the state-of-the-art methods in respective fields, (2) add a contribution model on an existing benchmark, (3) add a new benchmark, (4) compare new ideas and approaches to well-known approaches. To enhance reproducibility, we encourage each contribution to (1) be from peer-reviewed articles with an associated DOI for all contributions, models, and tools, (2) include a run script to exactly reproduce the results (especially for computational tools), (3) include a metadata file with details such as team name, contact information, computational timing and software (with software version)/hardware used in order to enhance transparency.
It is important to note differences between a typical data repository and a benchmarking platform. Some of the key distinguishing factors between a usual large data repository (such as JARVIS-DFT) and the present leaderboard effort are: 1) the leaderboard contains well-characterized/well-known samples/tasks (i.e., with digital object identifier/peer-reviewed article links) with all the scripts/metadata easily available to reproduce the results rather than just being a look-up table to find data, 2) large data repositories usually contain more variation in materials chemistry/structure and less variation of methods while the leaderboard focuses on a larger number of method comparisons.
For example, the JARVIS-DFT contains DFT data for more than 80,000 materials and millions of material properties with a few specific ES methods and hence there are only a few entries for, say, the electronic bandgap of Silicon from different methods, while the leaderboard contains electronic bandgaps for Silicon using more than 17 ES methods from various contributors. Similarly, JARVIS-ALIGNN project contains AI models for more than 80 properties/tasks of materials, i.e., just one model for a well-known property such as formation energy, while there are more than 12 methods for formation energy task in the leaderboard (as discussed later).
Furthermore, the JARVIS-leaderboard attempts to bridge together multiple categories of methods (AI, ES, FF, QC, EXP) and types of data (single properties, structure, spectra, text, etc.) with the goal of broadening benchmarking efforts across several fields of study. What differentiates the JARVIS-Leaderboard from platforms such as MatBench33, is that MatBench33 provides a handful of tasks to evaluate ML methods on larger datasets (i.e. 104 entries, most of which are from the Materials Project65). A potential drawback of this approach is that the resulting performance rankings could be biased towards the data distribution of a single source. In contrast, the JARVIS-Leaderboard covers a broader range of datasets and properties and provides a better overview of model performance.
Recently in the field of machine learning in materials science, there has been a fixation on performance metrics for newly developed models. This begs the question of whether or not benchmarking can be destructive to the development of new methods if these new methods cannot immediately outperform the previous state-of-the-art approaches. This also begs the question of whether or not benchmarking can lead to overfitting or poor generalization66,67.
Therefore, we outline how the leaderboard can also be used to identify and focus on some of the major challenges in different fields, such as: (1) how to evaluate extrapolation capability68? (2) why is it difficult to develop a reasonably good AI model with similar accuracy to electronic structure methods?, (3) how can we reduce the computational cost of higher accuracy electronic structure predictions (such as bandgaps and bandoffsets)?, (4) how do we identify examples of materials that require high-fidelity methods (beyond DFT accuracy)?, (5) how can we identify material space where methodological improvements need to be targeted?, (6) how can we establish figures of merits for mesoscale and device-scale models such as phase field and technology computer-aided design (TCAD)?, (7) how can we make atomistic image analysis quantitative rather than qualitative?, (8) and how do we develop and benchmark multi-modal models (such as text, image, video, atomic structures, etc.)69?
The JARVIS-Leaderboard is seamlessly integrated into the existing and well-established NIST-JARVIS infrastructure70,71, which hosts several datasets, tools, applications, and tutorials for materials design, motivated by the materials genome initiative1. The framework is open access to the entire materials science community for progressing the field collectively and systematically. JARVIS (Joint Automated Repository for Various Integrated Simulations)70,71 is a repository designed to automate materials discovery and optimization using classical force-field, density functional theory, machine learning calculations, and experiments. Nevertheless, the leaderboard is not limited to NIST-JARVIS infrastructure and can be linked with other external projects as well.
Since its creation in 2017, JARVIS has had over 50,000 users worldwide, over 45 JARVIS-associated articles have been published, and over 80,000 materials currently reside in the database. As these numbers continue to multiply, significant effort on external outreach to the materials science community has been an additional goal of JARVIS, with several events (https://jarvis.nist.gov/events/) such as the Artificial Intelligence for Materials Science (AIMS) and Quantum Matters in Materials Science (QMMS) workshops and hands-on JARVIS-Schools, which have had hundreds of participants throughout the last few years. Based on the level of success and support from the community with regard to the existing JARVIS infrastructure, we believe that the integration of the JARVIS-Leaderboard will have a similar level of engagement and success, with a growing number of contributors from all over the world (in government, academia and industry) and in different sub-fields of materials science.
Results and discussion
Leaderboard overview
At the homepage, information regarding the number of methods, benchmarks, contributions, and datapoints are provided. A snapshot of the homepage with various categories is shown in Fig. 1a. Clicking on one of the entries (or searching in the ’Search’ box) such as “formation_energy_peratom” opens a new tab with available contributions. This new tab consists of 1) a description of the benchmark, 2) a plot of various available contributions (as shown in Fig. 1b), 3) explicit table for the plot (as shown in Fig. 1c). For each contribution, links are provided to the submitted data (in .csv.zip format), reference benchmark data (in JSON file), a shell script to reproduce the contribution (run.sh file) and metadata file (metadata.json). The metadata file contains details about the team name, the electronic mail address of the contributor(s), DOI number, software (with software version), hardware, instrument, computational timing and other relevant details of a benchmark.
a Homepage sanpshot showing list of categories and number of available contributions at the time of writing, b an example AI regression model benchmark for formation energy with several contributions. The methods are sorted based on the mean absolute error (MAE) values. Lower MAE values indicate higher accuracy, c explicit table for the plot in panel b.Links to individual csv.zip (AI-SinglePropertyPrediction-formation_energy_peratom-dft_3d-test-mae.csv.zip), json.zip (dft_3d_formation_energy_peratom.json.zip), shell script (run.sh) and detailed info (metadata.json) files are provided to help enhance reproducibility. Such results plots and tables are available for each benchmark in the leaderboard.
There are several categories for the benchmarks, including AI, ES, QC, FF, and EXP and their combinations. Some example contributions and a summary table are also provided on the webpage to help a user navigate through the project. The summary table breaks down the available information into categories and sub-categories of different methodologies.
JARVIS-Leaderboard is an evolving project, so additions to the project are anticipated, welcome, and easy to make. We show a general flowchart for adding a new benchmark to the leaderboard in Fig. 2. The user can populate the reference dataset (with well-defined data splits) used for a specific benchmark (e.g., for 2D exfoliation energies in JARVIS-DFT dataset using an AI method: “AI-SinglePropertyPrediction-exfoliation_energy-dft_3d-test”). AI benchmarks have pre-defined training/validation/test identifiers and target data in a corresponding json.zip file, while other methods have only reference test set for evaluation because they do not require model training like an AI method does. For most benchmarks in the leaderboard, experimental data is used as the reference data.
The jarvis_populate_data.py scripts generate a benchmark dataset. A user can apply their method, train models, or run experiments on that dataset and prepare a csv.zip, a metadata.json file, and other files in a new folder in the contributions directory. The contributions can be locally checked by the user using jarvis_server.py script. Then the folder can be uploaded to a user’s GitHub account by the automated jarvis_upload.py script involving several GitHub uploading steps. The administrators of the JARVIS-Leaderboard at NIST will verify the contributions and then finally, it will become part of the leaderboard website.
There is a helper script jarvis_populate_data.py to generate a benchmark dataset. A user can apply their method, train models, or run experiments on that dataset and prepare a csv.zip file, a metadata.json file, and also if possible, a conda environment.yaml/Nix/Dockerfile and a run.sh file. This step helps to reproduce the benchmark. These files are kept in a folder with the name of the folder as the team name and can be uploaded to a user’s GitHub account by the automated jarvis_upload.py script. This script automatically forks the parent usnistgov/jarvis_leaderboard repo for the user, adds the team-name folder with its files in that forked repo, runs a few minimal sanity checks on the new contribution, and then makes a pull request to the parent repo. The contribution addition and automated testings are carried out using GitHub actions. The administrators of the JARVIS-Leaderboard at NIST will verify the contributions and then finally, it will become part of the leaderboard website.
This project is available on GitHub at: https://github.com/usnistgov/jarvis_leaderboard. The administrators of the JARVIS-Leaderboard at NIST will fully oversee the upload of contributions and benchmarks. A tree structure of the repo is shown in Fig. 3. There are two main directories in the repo: (1) benchmarks (reference) and (2) leaderboard contributions (for various leaderboard entries), as shown by the green highlighted boxes in Fig. 3.
There are two main directories in the repo: (1) benchmarks (reference) and (2) leaderboard contributions (for various leaderboard entries). In the “benchmarks” directory, there are folders for the AI, ES, QC, FF, and EXP categories. Within them, there are sub-folders for specific sub-categories. In the “contributions” directory there is a collection of folders that consists of .csv.zip, metadata.json files, and optionally a Dockerfile and run.sh file for available contributions from each method. The csv.zip file contains entries of identifier (id) and corresponding prediction values as obtained by the corresponding model/method. These test identifiers (such as JVASP-1408) must match the test set ids in the json.zip file in the benchmarks folder for the metric measurements to work.
The “benchmarks” directory has folders for the AI, ES, QC, FF, and EXP categories. Within them, there are sub-folders for specific sub-categories such as (1) SinglePropertyPrediction (where the output of a model/experiment is one single number for an entry), (2) SinglePropertyClass (where the output is class-ids, i.e., 0,1,.. instead of floating values), (3) ImageClass (for multi-class image classification), (4) TextClass (for multi-label text classification), (5) MLFF (machine learning force-field), (6) Spectra (for multi-value data) and (7) EigenSolver (for Hamiltonian simulation). In each of these sub-folders, there are .json.zip files with well-defined reference datasets and available properties as also available in the JARVIS-Tools package https://pages.nist.gov/jarvis/databases/. To avoid storage of large files in the GitHub repo, the actual datasets are part of JARVIS-Tools and are stored in the Figshare repository with specific DOIs and version numbers.
Next, in the “contributions” directory, there is a collection of folders that consist of .csv.zip, metadata.json files, and optionally a Dockerfile and run.sh file. The csv.zip file contains identifier (id) entries and corresponding prediction values obtained by the corresponding model/method. These test identifiers (such as JVASP-1408 in Fig. (3)) must match the test set IDs in the json.zip file in the benchmarks folder for the metric measurements to work. Each of the csv.zip files must contain six components in the filename to place the contribution in the appropriate webpage. The components are the categories (such as AI), sub-categories (such as ImageClass), property (such as bravais_lattice), dataset-name (such as stem_2d_image as available in the JARVIS-Tools database page), and data-split. For entries in the AI category, the data is in train-validation-test splits (using a fixed random number generator). For the current leaderboard format, we report the performance accuracy in the test set only. These files can be easily edited with common text editors. Each contribution folder (e.g. alignn-model) consists of one or several csv.zip files corresponding to each benchmark (such as for formation energies, bandgap, etc.).
Model-specific details are kept in the metadata.json file with required keys such as model_name, project_url, team_name and an email address. Users can keep other data such as the uncertainty, time taken, and instrument/software/hardware used in the metadata file as well. For computational models, the run.sh script can be used to reproduce the contributions completely as a single command line script or job submission script. If a method requires additional steps or details beyond a simple command line script, a user can upload a README file containing the additional details. For enhanced reproducibility, we also optionally allow users to include a Dockerfile and an ipython/Google-colab notebook for each benchmark. These notebooks can be used to run the contributions in the Google-cloud without downloading anything locally.
In addition, there is a “docs” directory in the JARVIS-leaderboard. The docs folder consists of a directory structure that is similar to the benchmarks folder with categories names (AI, ES, etc.), and sub-categories (such as SinglePropertyPrediction, ImageClass etc.) with markdown (.md) files that will be converted automatically into corresponding html pages for the website. For each benchmark (i.e., json.zip file), a corresponding docs entry (i.e., md file) should be present. A new benchmark must be associated with a peer-reviewed article and a DOI, in order to have trust in the reference benchmark data. A new benchmark must also be verified by the JARVIS-Leaderboard administrators.
As mentioned above, there already exist several other materials science-specific benchmarks. We compare some of these benchmarks in Table 1 based on the categories that are included. We find that there is no single, large-scale benchmark encompassing the various fields as in the JARVIS-Leaderboard. Also, the data format, metadata, and website for these different leaderboards vary significantly. Hence, having a uniform way to compare different methods would greatly help the materials community.
Benchmarks
The benchmarks consist of experimental data, density functional theory, or numerical solutions that are well-known and have already been published in peer-reviewed articles or books. A benchmark should be considered the “ground truth” for a particular task. Therefore, it is mandatory to have a digital object identifier (DOI) for each benchmark from a peer-reviewed article. There can be multiple contributions from different models or experiments for a benchmark, e.g., contributions from various DFT functionals in predicting the electronic bandgap of silicon with respect to experimental data. Typically, for electronic structure (ES) method-based contributions, the benchmarks are experimental data; for artificial intelligence (AI) methods, they are the test split; for force-field (FF) methods, they are electronic structure data; for quantum computation (QC), they are analytical results; and for experiments (EXP), they are other experiments. Currently, we have more than 270 benchmarks in the leaderboard. The JARVIS-Leaderboard flexible and dynamic nature allows the addition of new benchmarks as well.
Each entry in the benchmark dataset consists of a unique identifier. Most of these datasets are integrated into JARVIS-Tools databases page already (but not limited by it), with an associated JARVIS ID number (JID) and are backed up in Figshare, Google Drive and NIST-internal storage systems. The number of entries can vary from a few (which is especially applicable for experimental and high-accuracy computational methods, where generating a very large dataset is not feasible in terms of time and resources) to hundreds of thousands of entries in a dataset.
An overview of the dataset can be found in Fig. 4. Considering all possible entries in the dataset, we have close to 7 million datapoints. For example, an atomic structure can have multiple properties calculated, such as bandgaps and formation energies, among other properties. We find the JARVIS-DFT-3D dataset to have the largest number of entries. Considering unique systems, we can find the distribution in Fig. 4b). In this case, qe-tb (fitting dataset for ThreeBodyTB.jl72) is one of the largest datasets available in the leaderboard. Note that these datasets contain all varieties of data modalities, such as atomic structure, images, spectra, and text. In Fig. 5, we show the fractional distribution of periodic table elements in the entire dataset. We find that the most common elements are C, N, O, Cu, which is similar to the natural abundance of these elements.
a all entries in leaderboard, b entries with unique identifiers. Note that one identifier (such as JVASP-1002 for silicon) can have multiple properties (such as bandgap, bulk modulus etc.). A script to generate this figure is also provided on the leaderboard website as the leaderboard is continuously evolving.
Experimental results are uploaded as benchmarks (i.e. what is regarded as the reference). In the absence of experimental data, high-fidelity computational methods can be used as a reference. If there are multiple experimental measurements available in the literature, each can be individually added as separate benchmarks (i.e., different json.zip files to distinguish one benchmark from another) and users can submit contributions for each of them. As time and the materials science field progresses, certain experimental data may need to be revisited (i.e. more accurate measurements in the future or results are reported that contradict previous experimental data). As a response to this, separate reference (experimental) benchmarks can be added, and users will be able to plot and compare the evolution of these benchmarks over time.
In addition, Leaderboard users can raise an issue on GitHub pertaining to reference benchmarks. The administrators will also upload a README file, which contains additional information about the experiments conducted, including associated DOI, the experimental conditions and provide details if additional experiments conducted on the same material/property exist in the literature. The experimental conditions described in the README file can be important when comparing the reference benchmark to calculated results, which may be in different conditions than the experiment (i.e., the bandgap of a material is never measured at 0 K, as DFT predicts).
Contributions to the leaderboard in the form of user-submitted experimental data can be compared with previous experiments, electronic structure methods or other numerical results. ES-based contributions are benchmarked against experimental results and can be compared with other ES methods. QC data can be compared with classical computation data or exact analytical results. For FF, contributions can be compared to DFT (or other ES data) or high-level interatomic potential benchmark suites (specifically for MLFFs)38. For AI, a test dataset is used. Unlike other methods, AI methods can have both “train” and “test” datasets, while others have only “test” sets in the corresponding dataset. For AI methods, if the “train” dataset is not provided and only “test” is given, the benchmark can be used for checking extrapolation behavior such as vacancy formation energy benchmarks.
Analysis of benchmarks
Presently, the leaderboard has 5 categories, 10 sub-categories, 152 methods, 274 benchmarks, 1281 contributions and 8714228 datapoints. In this section, we show a few of the hundreds of example analyses that can be carried out using the available benchmarks and contributions. In Fig. 6, we show the MAE of the AI-computed formation energy and ES-computed bandgap for Si for a variety of contributions in the leaderboard. In Fig. 6(a) we see the comparison of 12 AI models (each AI model had a well-defined 80:10:10 split for training, validation and testing respectively from the JARVIS-3D database) and find the kgcnn_coGN73 has the highest accuracy/lowest error, followed by Potnet74, Matformer75 and ALIGNN76,77 models. This can be attributed to the fact that as we include more structural information and use deep-learning methods rather than descriptor methods, we get improvement in accuracy.
a Artificial intelligence (AI) formationenergy for test set with 5572 materials in JARVIS-DFT 3D dataset, b electronic structure (ES) Si (JARVIS-DFT ID:JVASP-1002) bandgap, c classical force-field (FF) based Voigt bulk modulus of Si and d machine learning force-field (MLFF) based forces for Si. We provide Jupyter/Google colab notebooks to easily plot such comparisons for all available benchmarks. Also, similar analysis figures for all the available benchmarks are available in the Supplementary Information (Supplementary Figs. 1–298). As a note, these plots are a current snapshot of the leaderboard, and it is possible that new and more accurate models will be developed and added here in the future.
Similarly, in Fig. 6(b) we compare the bandgap of Si using several methods and find GLLB-sc78 calculated with GPAW79 to yield the lowest error, while G0W080 (VASP81,82), GW080 (VASP81,82), TBmBJ83,84 (VASP), and DMC85 (QMCPACK86) methods follow. This can be attributed to the inclusion of the discontinuity potential (GLLB-sc78) or kinetic energy density (TBmBJ83,84) in the density functional or incorporating many-body physics (G0W080, GW080, DMC85) into the methodology, which can lead to improved accuracy for bandgap prediction. Also, similar methods such as PBE87 data from Open Quantum Materials Database (OQMD)88,89, AFLOW90 and Materials Project65 compare well with each other.
In Fig. 6(c) we compare how several classical FFs compute the Voigt bulk modulus of Si. In Fig 6(d) we compare several MLFF models for the forces of Si. We compare various pretrained MLFFs and other MLFFs we specifically trained on the MLEARN38 dataset (PBE-based DFT data). We see that ALIGNN-FF91 and MatGL92 perform similarly for the prediction of forces. Fig. 6(d) provides a comprehensive comparison of MLFFs that are trained and tested on the same dataset and pretrained models that were trained elsewhere. The comparisons are presented in tabular form for all the benchmarks on the leaderboard website. We have provided tools and notebooks in the leaderboard GitHub repository that can be used for making such plots for all the available benchmarks and contributions. A collection of such figures for method comparison is available in the supplementary information (Supplementary Figures 1-298). We have also added interactive plots for such comparisons on the website. These tools can aid in identifying examples of materials that require high-fidelity methods beyond the accuracy of DFT in order to understand their underlying properties. In addition, these tools can be used to validate electronic structure methods and provide insight for error estimation.
The leaderboard has a large number of benchmarks and can enable a more comprehensive comparison of different methods for better revealing their respective advantages and limitations. For instance, neural networks outperform descriptor-based models by a large degree in all of the 10 regression tasks in the latest Matbench33 leaderboard. To check if this is also the case for 44 regression benchmarks in the current JARVIS leaderboard, we compare the performance of the best descriptor-based model to that of the best neural network. As shown in Fig. 7, the best neural network outperform the best descriptor-based model in 34 tasks, but only 14 out of 44 (32 %) tasks see a performance difference by more than 20 %. This indicates that descriptor-based models are still competitive with respect to neural networks, especially considering their better interpretability and orders of magnitude lower training cost66,67. Notably, the best descriptor-based model is found to outperform the best neural network in 10 tasks including those with 104-105 training data, opening up interesting questions and potential direction to further model improvement. For instance, the inferior performance of neural networks in the regression tasks for the heat capacity and hMOF data may be related to the recently revealed incapability of graph neural networks in capturing periodicity93.
The benchmark name and the corresponding best performing neural network are indicated in the left and right y axis, respectively. For all the considered AI benchmarks, the best descriptor-based model is the tree-based model using Magpie185 and Voronoi-tessellation201 features. As a disclaimer, these plots are a current snapshot of the leaderboard, and it is possible that new and more accurate models will be developed in the future.
Analysis of error metrics
Although a metric such as the MAE can be useful to compare methods for a specific benchmark, it is difficult to compare across different methods, since MAE values can differ substantially. Hence, we use the mean absolute deviation (MAD, computed with respect to the average value of the training data as a baseline/random-guess model) to MAE ratio for both AI and ES single-property-prediction categories. Mean absolute deviation values act as a baseline/random-guessing model for the benchmark and contributed models should have MAE performance better than MAD values. We show the MAD/MAE ratios for AI and ES benchmarks in Fig. 8. We find that the MAD/MAE values range from 2 to 50. MAD/MAE values close to 1 suggest low predictive power. We observe that quantum properties such as the bandgap have lower MAD/MAE than classical quantities (quantities that do not require quantum mechanical simulations) such as total energy or bulk modulus. Interestingly, such trends for classical vs. quantum quantities are observed for both the AI and ES approaches.
Interactive view of benchmarks and contributions
In addition to making bar plots as shown in Figs. 6 and 8, the raw data available in benchmarks and contributions can be presented in various other forms such as scatter plots, bandstructures, adsorption spectra, and diffraction spectra. In Fig. 9, we show example comparisons of different methods for AI, ES, QC and EXP categories, including (a) formation-energy-per atom model using AI, (b) bulk modulus predictions using ES, (c) electronic bandstructure of Al using QE with different quantum circuits94, (d) CO2 capture for zeolite at several labs in round-robin fashion95. In Fig. 9a), we find that formation energy is one of the easiest quantities to train AI models and even simple chemistry-only-based models can perform reasonably well (i.e., cfid_chem). Including more structural features (such as bond angles and dihedral angles) and using deep learning models (such as graph neural network vs descriptor-based models) further help improve accuracy. Similarly, for ES example for predicting bulk modulus, we find irrespective of DFT based method used, they are in relatively close agreement with experimental bulk modulus data as shown in Fig. 9b). In Fig 9c), we find that the selection of a quantum circuit is critically important for predicting electronic band structures well. Here, we used 6 different quantum94 circuits and found the SU(2)96 circuit to compare well with classical computer-based electronic bandstructures. This can be attributed to various entanglements captured in the SU(2)96 circuits that may be missing in other circuits. Finally, for experimental inter-laboratory/round-robin type measurements of the zeolite CO2 isotherm, we find excellent agreement across different labs95.
a formation-energy-per atom model using AI for JARVIS-DFT 3D dataset with 5572 materials in the test set, (b) bulk modulus predictions using ES methods for 21 materials, c electronic bandstructure of Aluminum using QC methods with different quantum circuits on a coarse k-point mesh, d CO2 capture for zeolite (ZSM-5) at several labs in inter-laboratory/round-robin fashion.
Methods
The JARVIS-Leaderboard aims to provide a comprehensive framework covering a variety of length and time-scale approaches2 to enable realistic materials design. In this section, we provide a brief overview of the methods that are currently available in the leaderboard. In this work we use the terms categories, sub-categories, methods, benchmarks, and contributions often, so we define them as follows.
Currently, there are five main “categories” in the leaderboard: Artificial Intelligence (AI), Electronic Structure (ES), Force-field (FF), Quantum Computation (QC), and Experiments (EXP). Each category is divided into “sub-categories”, a list of which is provided on the website. These sub-categories include single-property-prediction, single-property-classification, atomic force prediction, text classification, text-token classification, text generation, image classification, image segmentation, image generation, spectra-prediction, and eigensolver. These sub-categories are highly flexible and new categories can be easily added. “Benchmarks” are the reference data (in the form of json.zip file, discussed later) used to calculate performance metrics for each specific contribution. “Methods” are a set of precise specifications for evaluation against a benchmark. For example, within the ES category, density functional theory (DFT) performed with the specifications of the Vienna Ab initio Simulation Package (VASP)81,82, Perdew-Burke-Ernzerhof (PBE)87 functional and PAW81,82 pseudopotentials (VASP-PBE-PAW) is a method. Similarly, within the AI category, descriptor/feature-based models with specifications of MatMiner97 chemical features and the LightGBM98software is a method. “Contributions” are individual data (in the form of csv.zip files) for each benchmark computed with a specific method. Each contribution files consist of six components: category (e.g. AI), sub-category (e.g. SinglePropertyPrediction), property (e.g. formation energy), dataset (e.g. dft_3d), data-split (e.g. test), metric (e.g. mae).
Electronic structure
Electronic structure approaches cover short-length scales and short-time scales with high-fidelity. There are a variety of ES methodologies such as such as tight-binding72,99,100, density functional theory (DFT)101, quantum Monte Carlo85, dynamical mean field theory102 and many-body perturbation theory (Green’s function with screened Coulomb potential, GW methods)80. For each of the methodologies, there are a number of specifications to completely describe a method including the exact software, exchange-correlation functional, pseudopotential, and other relevant parameters. Example methods used in this work are given in Table 2.
Each method in the ES category can have a variety of contributions. For example, using a specific method, one can calculate various properties such as bandgaps, formation energies, bulk moduli, solar cell efficiencies, and superconducting transition temperatures as well as spectral quantities such as dielectric functions. While there are more than 400 approximate exchange-correlation functionals proposed in DFT literature103, currently, we have OptB88vdW104, Opt86BvdW105, LDA106, PBE87, PBEsol107, GLLB-sc78, TBmBJ83,84, SCAN108, r2SCAN109, HSE06110, in the leaderboard. We use converged k-points and cut-offs as available in the JARVIS-DFT database111. We have used the Vienna Ab initio Simulation Package (VASP)81,82, ABINIT112,113,114, GPAW79 and Quantum Espresso (QE)115 as DFT software packages, but other packages can be easily added as well. In addition, we use VASP81,82 to perform GW calculations including “single-shot” G0W0 and self-consistent GW0 methods80. Other ES approaches include tight-binding (TB)99 and quantum Monte Carlo (QMC)85. For TB, we use the recently developed ThreeBodyTB.jl code72 along with the Wannier90116 code, while the QMCPACK86 code is used for diffusion Monte Carlo (DMC)85 calculations.
Force-field
Force fields can be used in molecular dynamics and Monte Carlo simulations for studying larger time and length scales compared to electronic structure methods. Traditional force fields are developed for specific chemical systems and applications and may not be transferable to other uses. It is important to check the validity of an FF before using it in a particular application. Moreover, the development of FFs is a cumbersome task. Examples of typical FFs include embedded-atom method (EAM) potentials117 (i.e. Al099.eam.alloy for aluminum system118), Lennard Jones (LJ)119 for 2D liquids, reactive empirical bond order (REBO)120 for Si, and classical, atomistic force fields for biomolecular systems121,122. Recently, machine learning force fields (MLFF)123,124,125,126,127,128 have become popular because of their higher accuracy and ease of development (such as SNAP129 FFs). Nevertheless, early generations of MLFFs were also developed for specific types of chemistry and applications. Very recently, several MLFFs have been developed that can be used to simulate any combination of periodic table elements. Some of these FFs include M3GNET92, ALIGNN-FF91, and CHGNet130. In the leaderboard, we include benchmarks for energies, forces, and stress tensors for both specific systems and universal datasets.
Traditional FFs are available in LAMMPS131, while MLFFs are integrated into the Atomic Simulation Environment (ASE)132 package. Some of these MLFFs are now available in LAMMPS and other large-scale MD codes. In addition to static quantities, FFs can be used for Monte Carlo simulations, such as CO2 adsorption in metal-organic frameworks (MOFs)133 using the RASPA134 code. In addition to energy, force, and stress, we also have FF benchmarks for classical properties such as the bulk modulus. For biomolecular systems, GROMACS135 is commonly used, and we present here free energy differences and conformational state population benchmarks for three model peptides136,137,138.
Artificial intelligence
Recently artificial intelligence methods have become popular for materials prediction across all lengths and time scales. We currently have benchmarks for four types of data used as input for the AI models: (1) atomic structure, (2) spectra, (3) images, and 4) text. AI techniques can be used for both forward prediction and inverse design. For atomic structure datasets, we use DFT datasets such as JARVIS-DFT70,71, Materials Project (MP)65, Tight binding three-body dataset (TB3)72, Quantum-Machine 9 (QM9)139,140. For spectral data, we use either DFT-based spectra of, for example, electron or phonon density of states (DOS), Eliashberg functions, or numerical XRD spectra. For images, we have simulated and experimental scanning transmission electron microscope (STEM) and scanning tunneling microscopy (STM) images for 2D materials. For text data, we have used the publicly available arXiv dataset.
Currently, we have models for feature-based/tabular models (such as RandomForest141, Gradient boosting141, Linear regression141), graph-based models (such as ALIGNN76,77, SchNet142, CGCNN143, M3GNET92, AtomVision144, ChemNLP145) as well as transformers (such as OPT146, GPT30, and T5147). These models use popular AI code bases, including PyTorch148, scikit-learn141, TensorFlow149, LightGBM98, JAX150, and HuggingFace151. These models are used for a variety of properties such as formation energies, electron bandgaps, phonon spectra, forces, text data etc.
Quantum computation
Quantum chemistry is one of the most promising applications of quantum computations152. Quantum computers with relatively few logical qubits can potentially exceed the performance of much larger classical computers because the size of Hilbert space increases exponentially with the number of electrons in the system. Predicting the energy levels of a Hamiltonian is a typical and fundamentally important problem in quantum chemistry. We use Hamiltonian simulations with quantum algorithms and compare it with classical solvers. Determination of appropriate quantum circuit for a specific QC problem is a challenging task. For example, we use the tight-binding Hamiltonians for electrons and phonons in JARVIS-DFT and evaluate the electron bandstructures using quantum algorithms (such as variational quantum eigen solver (VQE)153 and variational quantum deflation (VQD)154) and with different quantum circuits (such as PauliTwo design96 and SU(2)96 circuits). We primarily use the Qiskit96 software in this work through the JARVIS-Tools/AtomQC94 interface, but other packages such as Tequila155, Circq156, and Pennylane157,158 can also be easily integrated. In addition to studying algorithm and circuit architecture dependence, the leaderboard can be used for studying the noise-levels in quantum circuits across different quantum computers, which is a key issue hindering quantum computer commercialization. Currently, we are only using statevector simulators for the quantum algorithms available in the Qiskit96 library.
Experiments
Although experimental results for material properties and spectra are referenced in comparison to computational methods (within the JARVIS-Leaderboard and other leaderboards such as MatBench33), we dedicated a portion of the JARVIS-Leaderboard to experimental benchmarking. Benchmarking experiments essentially boils down to the comparison of different experiments for the same desired result/s. A systematic way to perform this benchmarking is through round-robin testing159. This is an inter-laboratory test performed independently several times, which can involve multiple scientists and a variety of methods and equipment. This approach has been applied successfully for a range of materials science applications95,160,161,162,163, but many more of such experiments are still needed. Specifically in the JARVIS-Leaderboard, we include experimental round-robin results for manometric measurements of CO2 adsorption95. It is important to note that the experimental results included in the leaderboard are for well-characterized materials with well-defined properties and phenomena that can be easily reproduced (in contrast to replication attempts of variable experiments, such as the recent attempt to synthesize room temperature superconductors164,165,166,167). Some of the experiments we used for benchmarking purposes are XRD, magnetometry, vibroscopy, and scanning electron microscopy (SEM) and transition electron microscopy (TEM). We purchase the samples from industrial vendors with available identifiers such as CAS-number. We also carried out XRD for MgB2 (a superconducting material) to verify its crystal structure before carrying out magnetometry measurements to determine the transition temperature. This measurement was compared with numerical XRD data. Magnetometry measurements for superconductors were also conducted to compare their superconducting transition temperatures with respect to predicted or experimentally available values168. Strain-stress measurements were done for Kevlar for failure analysis168. We have several instruments, such as Bruker D8, Titan, Quantum design PPMS and FAVIMAT in the leaderboard currently.
Metrics used
We use several metrics in the leaderboard depending on the “sub-categories” mentioned above. We use mean absolute error (MAE), accuracy (acc), multi-mae (L1 norm of multi-dimensional data), recall-oriented understudy for gisting evaluation (ROUGE) for the single property prediction, single property classification, spectra/eigensolver/atomic forces and textGen/text summary subcategories respectively. As the user contributes their data to compare against the reference data (benchmarks), other complementary metrics (such as those available in the sklearn.metrics library) can be easily calculated as the raw contribution data is also made available through the website. For the sake of readability and ease of use, we primarily employ the metrics mentioned above. For single property prediction, there is only scalar values per column in the csv.zip file with id and prediction separate by comma., For spectra, force-prediction and other multi-value quantities (i.e. with multiple prediction values per id) we concatenate the array and separate by semicolon (to avoid comma convention in csv files). The benchmark data is also stored in a similar format. We provide tools to convert these csv.zip files into json or other file formats if needed. We also provided notebooks to visualize the data through Jupyter/Colab notebooks. In addition, we plan to eventually add metrics for timing, uncertainty, development cost and other details.
Data availability
Multiple datasets used in this work are available at the Figshare repository: https://figshare.com/authors/Kamal_Choudhary/4445539. Index and usage guidelines are provided at https://pages.nist.gov/jarvis/databases/.
Code availability
JARVIS-Leaderboard package mentioned in the article can be found at https://github.com/usnistgov/jarvis_leaderboard.
References
Ward, C. H. & Warren, J. A. Materials genome initiative: materials data (US Department of Commerce, National Institute of Standards and Technology, 2015).
Callister, W. D. et al. Fundamentals of materials science and engineering, Vol. 471660817 (Wiley London, 2000).
Chen, L.-Q. Phase-field models for microstructure evolution. Annu. Rev. Mat. Res. 32, 113–140 (2002).
Agrawal, A., Gopalakrishnan, K., & Choudhary, A. Materials image informatics using deep learning, in Handbook on Big Data and Machine Learning in the Physical Sciences: Volume 1. Big Data Methods in Experimental Materials Discovery, series and number World Scientific Series on Emerging Technologies, edited by (WorldScientific, 2020) pp. 205–230.
Choudhary, K. et al. Recent advances and applications of deep learning methods in materials science. npj Comp. Mat. 8, 59 (2022).
Audus, D. J. et al. Artificial intelligence for materials, in https://doi.org/10.1142/9789811265679_0023Artificial Intelligence for Science, Chapter 23, pp. 413–430.
Camerer, C. F. et al. Evaluating the replicability of social science experiments in Nature and Science between 2010 and 2015. Nat. Hum. Behav. 2, 637–644 (2018).
Fanelli, D. Is science really facing a reproducibility crisis, and do we need it to? Proc. Nat. Acad. Sci. 115, 2628–2631 (2018).
Sun, Z. et al. Are we evaluating rigorously? Benchmarking recommendation for reproducible evaluation and fair comparison. Proc. 14th ACM Conf. on Recomm. Sys. (2020).
Amrhein, V., Korner-Nievergelt, Fränzi & Roth, T. The earth is flat (p> 0.05): significance thresholds and the crisis of unreplicable research. PeerJ 5, e3544 (2017).
Grimes, DavidRobert, Bauch, C. T. & Ioannidis, JohnP. A. Modelling science trustworthiness under publish or perish pressure. Roy. Soc. Open Sci. 5.1, 171511 (2018).
Allen, G. I., Gan, L., & Zheng, L. Interpretable Machine Learning for Discovery: Statistical Challenges and Opportunities. Ann. Rev. Stat. and App. 11 (2023).
Prager, E. M. et al. Improving transparency and scientific rigor in academic publishing. J. Neuro. Res. 97, 377–390 (2019).
Papadiamantis, A. G. et al. Metadata stewardship in nanosafety research: community-driven organisation of metadata schemas to support FAIR nanoscience data. Nanomat 10, 2033 (2020).
Hao-Nan, Z. & Rubio-González, C. On the reproducibility of software defect datasets, 2023 IEEE/ACM 45th International Conference on Software Engineering (ICSE) IEEE (2023).
Lehtola, S. & Marques, M. Reproducibility of density functional approximations: How new functionals should be reported. J. Chem. Phys. 159, 114116 (2023).
Sayre, F. & Riegelman, A. The reproducibility crisis and academic libraries. Coll. Res. Lib. 79, 2 (2018).
Papadiamantis, A. G., Ward, L. & Hattrick-Simpers, J. Metadata stewardship in nanosafety research: Community-driven organisation of metadata schemas to support FAIR nanoscience data. Dig. Disc. 3, 281–286 (2024).
Park, J., Howe, J. D. & Sholl, D. S. How reproducible are isotherm measurements in metal–organic frameworks? Chem. Mat. 29, 10487–10495 (2017).
Baker, M. 1,500 scientists lift the lid on reproducibility. Nature 533, 452–454 (2016).
Hutson, M. Artificial intelligence faces reproducibility crisis. Science 359, 725–726 (2018).
Wilkinson, M. D. et al. The fair guiding principles for scientific data management and stewardship. Sci. Data 3, 1–9 (2016).
Agrawal, A. & Choudhary, A. Perspective: Materials informatics and big data: Realization of the “fourth paradigm” of science in materials science. APL Mat. 4, 053208 (2016).
Rickman, J., Lookman, T. & Kalinin, S. Materials informatics: From the atomic-level to the continuum. Acta Mat. 168, 473–510 (2019).
Agrawal, A. & Choudhary, A. Deep materials informatics: Applications of deep learning in materials science. MRS Comm. 9, 779–792 (2019).
Gupta, V., Liao, W.-k, Choudhary, A. & Agrawal, A. Evolution of artificial intelligence for application in contemporary materials science. MRS Comm. 13, 754–763 (2023).
Lejaeghere, K. et al. Reproducibility in density functional theory calculations of solids. Science 351, aad3000 (2016).
Russakovsky, O. et al. Imagenet large scale visual recognition challenge. Int. J. Comp. Vis. 115, 211–252 (2015).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589 (2021).
Brown, T. et al. Language models are few-shot learners. Adv. Neur. Info Proc. Sys. 33, 1877–1901 (2020).
Zhang, X. et al. Artificial intelligence for science in quantum, atomistic, and continuum systems. Preprint at https://arxiv.org/abs/2307.08423 (2023).
Bosoni, E. et al. How to verify the precision of density-functional-theory implementations via reproducible and universal workflows. Nat. Rev. Phys. 6, 45–58 (2024).
Dunn, A., Wang, Q., Ganose, A., Dopp, D. & Jain, A. Benchmarking materials property prediction methods: the matbench test set and automatminer reference algorithm. npj Comp. Mat. 6, 138 (2020).
Wu, Z. et al. Moleculenet: a benchmark for molecular machine learning. Chem. Sci. 9, 513–530 (2018).
Chanussot, L. et al. Open catalyst 2020 (oc20) dataset and community challenges. ACS Catal. 11, 6059–6072 (2021).
Chmiela, S. et al. Machine learning of accurate energy-conserving molecular force fields. Sci. Adv. 3, e1603015 (2017).
Chmiela, S., Sauceda, H. E., Poltavsky, I., Müller, K.-R. & Tkatchenko, A. sgdml: Constructing accurate and data efficient molecular force fields using machine learning. Comp. Phys. Comm. 240, 38–45 (2019).
Zuo, Y. et al. Performance and cost assessment of machine learning interatomic potentials. J. Phys. Chem. A 124, 731–745 (2020).
Weston, L. et al. Named entity recognition and normalization applied to large-scale information extraction from the materials science literature. J. Chem. Inf. Model. 59, 3692–3702 (2019).
Ziatdinov, M., Ghosh, A., ChunYin(Tommy), W. & Kalinin, S. V. AtomAI framework for deep learning analysis of image and spectroscopy data in electron and scanning probe microscopy. Nat. Mach. Intel. 4, 1101–1112 (2022).
Borlido, P. et al. Large-scale benchmark of exchange–correlation functionals for the determination of electronic band gaps of solids. J. Chem. Theor. Comp. 15, 5069–5079 (2019).
Huber, S. P. et al. Common workflows for computing material properties using different quantum engines. npj Comp. Mat. 7, 136 (2021).
Zhang, G.-X., Reilly, A. M., Tkatchenko, A. & Scheffler, M. Performance of various density-functional approximations for cohesive properties of 64 bulk solids. N. J. Phys. 20, 063020 (2018).
Tran, R. et al. The Open Catalyst 2022 (OC22) Dataset and Challenges for Oxide Electrocatalysts. ACS Catal. 13, 3066–3084 (2023).
Jurečka, P., Šponer, J., Černy`, J. & Hobza, P. Benchmark database of accurate (mp2 and ccsd (t) complete basis set limit) interaction energies of small model complexes, dna base pairs, and amino acid pairs. Phy. Chem. Chem. Phys. 8, 1985–1993 (2006).
Brauer, B., Kesharwani, M. K., Kozuch, S. & Martin, J. M. The s66 × 8 benchmark for noncovalent interactions revisited: Explicitly correlated ab initio methods and density functional theory. Phys. Chem. Chem. Phys. 18, 20905–20925 (2016).
Mata, R. A. & Suhm, M. A. Benchmarking quantum chemical methods: Are we heading in the right direction? Angew. Chem. Int. Ed. 56, 11011–11018 (2017).
Taylor, D. E. et al. Blind test of density-functional-based methods on intermolecular interaction energies. J. Chem. Phys. 145, 124105 (2016).
Wheeler, D. et al. Pfhub: the phase-field community hub. J. Open Res. Soft. 7, 29 (2019).
Lindsay, A. D. et al. 2.0 - MOOSE: Enabling massively parallel multiphysics simulation. SoftwareX 20, 101202 (2022).
Wei, J. et al. Benchmark Tests of Atom Segmentation Deep Learning Models with a Consistent Dataset. Micro Microanal. 29, 552–562 (2023).
Ren, J. et al. Diligent102: A photometric stereo benchmark dataset with controlled shape and material variation, in Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) (2022) pp. 12581–12590.
Li, M. et al. Multi-view photometric stereo: A robust solution and benchmark dataset for spatially varying isotropic materials. IEEE Trans. Im. Proc. 29, 4159–4173 (2020).
Henderson, A. N., Kauwe, S. K. & Sparks, T. D. Benchmark datasets incorporating diverse tasks, sample sizes, material systems, and data heterogeneity for materials informatics. Data Brief. 37, 107262 (2021).
Fung, V., Zhang, J., Juarez, E. & Sumpter, B. G. Benchmarking graph neural networks for materials chemistry. npj Comp. Mat. 7, 84 (2021).
Cecen, A., Dai, H., Yabansu, Y. C., Kalidindi, S. R. & Song, L. Material structure-property linkages using three-dimensional convolutional neural networks. Acta Mat. 146, 76–84 (2018).
Baird, S. G., Issa, R. & Sparks, T. D. Materials science optimization benchmark dataset for multi-objective, multi-fidelity optimization of hard-sphere packing simulations. Data Brief. 50, 109487 (2023).
Chen, L., Tran, H., Batra, R., Kim, C. & Ramprasad, R. Machine learning models for the lattice thermal conductivity prediction of inorganic materials. Comp. Mat. Sci. 170, 109155 (2019).
Tian, S. et al. Quartet protein reference materials and datasets for multi-platform assessment of label-free proteomics. Genome Bio. 24, 202 (2023).
Fu, N. et al. Materials transformers language models for generative materials design: a benchmark study. Preprint at https://arxiv.org/abs/2206.13578 (2022).
Meredig, B. et al. Can machine learning identify the next high-temperature superconductor? examining extrapolation performance for materials discovery. Mol. Syst. Des. Eng. 3, 819–825 (2018).
Lejeune, E. Mechanical mnist: A benchmark dataset for mechanical metamodels. Ext. Mech. Lett. 36, 100659 (2020).
Clement, C. L., Kauwe, S. K. & Sparks, T. D. Benchmark aflow data sets for machine learning. Int. Mat. Manufact. Innov. 9, 153–156 (2020).
Varivoda, D., Dong, R., Omee, S. S. & Hu, J. Materials property prediction with uncertainty quantification: A benchmark study. Appl. Phys. Rev. 10, 021409 (2023).
Jain, A. et al. Commentary: The materials project: A materials genome approach to accelerating materials innovation. APL Mat. 1, 011002 (2013).
Li, K., DeCost, B., Choudhary, K., Greenwood, M. & Hattrick-Simpers, J. A critical examination of robustness and generalizability of machine learning prediction of materials properties. npj Comp. Mat. 9, 55 (2023).
Li, K. et al. Exploiting redundancy in large materials datasets for efficient machine learning with less data. Nat. Commun. 14, 7283 (2023).
Choudhary, K. & Sumpter, B. G. Can a deep-learning model make fast predictions of vacancy formation in diverse materials? AIP Adv. 13 (2023).
Vuorio, R., Sun, S.-H., Hu, H. & Lim, J. J. Multimodal model-agnostic meta-learning via task-aware modulation, in Advances in Neural Information Processing Systems, Vol. 32, (eds H. Wallach, H. Larochelle, A. Beygelzimer, F. d’Alché-Buc, E. Fox, & R. Garnett) (CurranAssociates, Inc., 2019). https://proceedings.neurips.cc/paper_files/paper/2019/file/e4da3b7fbbce2345d7772b0674a318d5-Paper.pdf.
Choudhary, K. et al. The joint automated repository for various integrated simulations (jarvis) for data-driven materials design. npj Comp. Mat. 6, 173 (2020).
Wines, D. et al. Recent progress in the JARVIS infrastructure for next-generation data-driven materials design. Appl. Phys. Rev. 10, 041302 (2023).
Garrity, K. F. & Choudhary, K. Fast and accurate prediction of material properties with three-body tight-binding model for the periodic table. Phys. Rev. Mat. 7, 044603 (2023).
Reiser, P., Eberhard, A. & Friederich, P. Graph neural networks in tensorflow-keras with raggedtensor representation (kgcnn). Soft. Imp. 9, 100095 (2021).
Lin, Y. et al. Efficient approximations of complete interatomic potentials for crystal property prediction, in Proceedings of the 40th International Conference on Machine Learning (2023).
Yan, K., Liu, Y., Lin, Y., & Ji, S. Periodic graph transformers for crystal material property prediction, in The 36th Annual Conference on Neural Information Processing Systems (2022) pp. 15066–15080.
Choudhary, K. & DeCost, B. Atomistic line graph neural network for improved materials property predictions. npj Comp. Mat. 7, 185 (2021).
Gupta, V. et al. Structure-aware graph neural network based deep transfer learning framework for enhanced predictive analytics on diverse materials datasets. npj Comp. Mat. 10, 1 (2024).
Kuisma, M., Ojanen, J., Enkovaara, J. & Rantala, T. T. Kohn-sham potential with discontinuity for band gap materials. Phys. Rev. B 82, 115106 (2010).
Enkovaara, J. et al. Electronic structure calculations with gpaw: a real-space implementation of the projector augmented-wave method. J. Phys.: Cond. Matt. 22, 253202 (2010).
Onida, G., Reining, L. & Rubio, A. Electronic excitations: density-functional versus many-body green’s-function approaches. Rev. Mod. Phys. 74, 601–659 (2002).
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169 (1996).
Kresse, G. & Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Comp. Mat. Sci. 6, 15–50 (1996).
Tran, F. & Blaha, P. Importance of the kinetic energy density for band gap calculations in solids with density functional theory. J. Phys. Chem. A 121, 3318–3325 (2017).
Rai, D. P., Ghimire, M. P. & Thapa, R. K. A dft study of bex (x = s, se, te) semiconductor: Modified becke johnson (mbj) potential. Semicond 48, 1411–1422 (2014).
Foulkes, W. M. C., Mitas, L., Needs, R. J. & Rajagopal, G. Quantum Monte Carlo simulations of solids. Rev. Mod. Phys. 73, 33–83 (2001).
Kim, J. et al. Qmcpack: an open source ab initio quantum monte carlo package for the electronic structure of atoms, molecules and solids. J. Phys.: Cond. Matt. 30, 195901 (2018).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Saal, J. E. et al. Materials design and discovery with high-throughput density functional theory: The open quantum materials database (oqmd). JOM 65, 1501–1509 (2013).
Kirklin, S. et al. The open quantum materials database (oqmd): assessing the accuracy of dft formation energies. npj Comp. Mat. 1, 15010 (2015).
Curtarolo, S. et al. Aflow: An automatic framework for high-throughput materials discovery. Comp. Mat. Sci. 58, 218–226 (2012).
Choudhary, K. et al. Unified graph neural network force-field for the periodic table: solid state applications. Dig. Disc. 2, 346–355 (2023).
Chen, C. & Ong, S. P. A universal graph deep learning interatomic potential for the periodic table. Nat. Comp. Sci. 2, 718–728 (2022).
Gong, S., Xie, T., Shao-Horn, Y., Gomez-Bombarelli, R., & Grossman, J. C. Examining graph neural networks for crystal structures: limitations and opportunities for capturing periodicity. Preprint at https://arxiv.org/abs/2208.05039 (2022).
Choudhary, K. Quantum computation for predicting electron and phonon properties of solids. J. Phys.: Cond. Matt. 33, 385501 (2021).
Nguyen, H. G. T. et al. A reference high-pressure co2 adsorption isotherm for ammonium zsm-5 zeolite: results of an interlaboratory study. Adsorption 24, 531–539 (2018).
IBM Quantum, https://quantum-computing.ibm.com (2021).
Ward, L. et al. Matminer: An open source toolkit for materials data mining. Comp. Mat. Sci. 152, 60–69 (2018).
Ke, G. et al. Lightgbm: A highly efficient gradient boosting decision tree. Adv. Neur. Info Proc. Sys. 30, 3146–3154 (2017).
Harrison, W. A. Electronic structure and the properties of solids: the physics of the chemical bond (Courier Corporation, 2012).
Garrity, K. F. & Choudhary, K. Database of wannier tight-binding hamiltonians using high-throughput density functional theory. Sci. Data 8, 106 (2021).
Martin, R. M. Electronic structure: basic theory and practical methods (Cambridge University Press, 2020).
Kotliar, G. et al. Electronic structure calculations with dynamical mean-field theory. Rev. Mod. Phys. 78, 865 (2006).
Lehtola, S., Steigemann, C., Oliveira, M. J. & Marques, M. A. Recent developments in libxc—a comprehensive library of functionals for density functional theory. SoftwareX 7, 1–5 (2018).
Klimeš, J., Bowler, D. R. & Michaelides, A. Chemical accuracy for the van der waals density functional. J. Phys.: Cond. Matt. 22, 022201 (2009).
Klimeš, J. C. V, Bowler, D. R. & Michaelides, A. Van der waals density functionals applied to solids. Phys. Rev. B 83, 195131 (2011).
Hohenberg, P. & Kohn, W. Inhomogeneous electron gas. Phys. Rev. 136, B864–B871 (1964).
Perdew, J. P. et al. Restoring the density-gradient expansion for exchange in solids and surfaces. Phys. Rev. Lett. 100, 136406 (2008).
Sun, J., Ruzsinszky, A. & Perdew, J. P. Strongly constrained and appropriately normed semilocal density functional. Phys. Rev. Lett. 115, 036402 (2015).
Furness, J. W., Kaplan, A. D., Ning, J., Perdew, J. P. & Sun, J. Accurate and numerically efficient r2scan meta-generalized gradient approximation. J. Phys. Chem. Lett. 11, 8208–8215 (2020).
Heyd, J., Scuseria, G. E. & Ernzerhof, M. Hybrid functionals based on a screened coulomb potential. J. Chem. Phys. 118, 8207–8215 (2003).
Choudhary, K. & Tavazza, F. Convergence and machine learning predictions of monkhorst-pack k-points and plane-wave cut-off in high-throughput dft calculations. Comp. Mat. Sci. 161, 300–308 (2019).
Gonze, X. et al. Recent developments in the abinit software package. Comp. Phys. Comm. 205, 106–131 (2016).
Romero, A. H. et al. Abinit: Overview, and focus on selected capabilities. J. Chem. Phys. 152, 124102 (2020).
Gonze, X. et al. The abinit project: Impact, environment and recent developments. Comp. Phys. Comm. 248, 107042 (2020).
Giannozzi, P. et al. Quantum espresso: a modular and open-source software project for quantum simulations of materials. J. Phys.: Cond. Matt. 21, 395502 (2009).
Mostofi, A. A. et al. An updated version of wannier90: A tool for obtaining maximally-localised wannier functions. Comp. Phys. Comm. 185, 2309–2310 (2014).
Daw, M. S. & Baskes, M. I. Embedded-atom method: Derivation and application to impurities, surfaces, and other defects in metals. Phys. Rev. B 29, 6443–6453 (1984).
Choudhary, K. et al. Evaluation and comparison of classical interatomic potentials through a user-friendly interactive web-interface. Sci. Data 4, 160125 (2017).
Jones, J. E. & Chapman, S. On the determination of molecular fields.—i. from the variation of the viscosity of a gas with temperature. Proc. Roy. Soc. Lond. Ser. A, Contain. Pap. A Math. Phys. Character 106, 441–462 (1924).
Tersoff, J. New empirical approach for the structure and energy of covalent systems. Phys. Rev. B 37, 6991–7000 (1988).
Case, D. A. et al. The amber biomolecular simulation programs. J. Comp. Chem. 26, 1668–1688 (2005).
Huang, J. et al. Charmm36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Novoselov, I., Yanilkin, A., Shapeev, A. & Podryabinkin, E. Moment tensor potentials as a promising tool to study diffusion processes. Comp. Mat. Sci. 164, 46–56 (2019).
Drautz, R. Atomic cluster expansion for accurate and transferable interatomic potentials. Phys. Rev. B 99, 014104 (2019).
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B 87, 184115 (2013).
Zhang, L., Han, J., Wang, H., Car, R. & Weinan, E. Deep potential molecular dynamics: A scalable model with the accuracy of quantum mechanics. Phys. Rev. Lett. 120, 143001 (2018).
Botu, V. & Ramprasad, R. Adaptive machine learning framework to accelerate ab initio molecular dynamics. Int. J. Quant. Chem. 115, 1074–1083 (2015).
Smith, J. S. et al. Automated discovery of a robust interatomic potential for aluminum. Nat. Comm. 12, 1257 (2021).
Chen, C. et al. Accurate force field for molybdenum by machine learning large materials data. Phys. Rev. Mat. 1, 043603 (2017).
Deng, B. et al. CHGNet as a pretrained universal neural network potential for charge-informed atomistic modelling. Nat. Mach. Intell. 5, 1031–1041 (2023).
Thompson, A. P. et al. LAMMPS - a flexible simulation tool for particle-based materials modeling at the atomic, meso, and continuum scales. Comp. Phys. Comm. 271, 108171 (2022).
Larsen, A. H. et al. The atomic simulation environment—a python library for working with atoms. J. Phys.: Cond. Matt. 29, 273002 (2017).
Choudhary, K. et al. Graph neural network predictions of metal organic framework co2 adsorption properties. Comp. Mat. Sci. 210, 111388 (2022).
Dubbeldam, D., Calero, S., Ellis, D. E. & Snurr, R. Q. Raspa: molecular simulation software for adsorption and diffusion in flexible nanoporous materials. Mol. Sim. 42, 81–101 (2016).
Páll, S. et al. Heterogeneous parallelization and acceleration of molecular dynamics simulations in GROMACS. J. Chem. Phys. 153 (2020).
Tsai, S.-T., Smith, Z. & Tiwary, P. Sgoop-d: Estimating kinetic distances and reaction coordinate dimensionality for rare event systems from biased/unbiased simulations. J. Chem. Theor. Comp. 17, 6757–6765 (2021).
Mehdi, S., Wang, D., Pant, S. & Tiwary, P. Accelerating all-atom simulations and gaining mechanistic understanding of biophysical systems through state predictive information bottleneck. J. Chem. Theor. Comp. 18, 3231–3238 (2022).
Wang, D. & Tiwary, P. State predictive information bottleneck. J. Chem. Phys. 154, 134111 (2021).
Ruddigkeit, L., van Deursen, R., Blum, L. C. & Reymond, J.-L. Enumeration of 166 billion organic small molecules in the chemical universe database gdb-17. J. Chem. Info Model. 52, 2864–2875 (2012).
Ramakrishnan, R., Dral, P. O., Rupp, M. & von Lilienfeld, O. A. Quantum chemistry structures and properties of 134 kilo molecules. Sci. Data 1, 1–7 (2014).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Schütt, K. T., Sauceda, H. E., Kindermans, P.-J., Tkatchenko, A. & Müller, K.-R. Schnet – a deep learning architecture for molecules and materials. J. Chem. Phys. 148, 241722 (2018).
Xie, T. & Grossman, J. C. Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys. Rev. Lett. 120, 145301 (2018).
Choudhary, K., Gurunathan, R., DeCost, B. & Biacchi, A. J. Atomvision: A machine vision library for atomistic images. J. Chem. Info Model. 63, 1708–1722 (2023).
Choudhary, K. & Kelley, M. L. ChemNLP: A Natural Language-Processing-Based Library for Materials Chemistry Text Data. J. Phys. Chem. C. 127, 17545–17555 (2023).
Zhang, S. et al. Opt: Open pre-trained transformer language models. Preprint at http://arxiv.org/abs/2205.01068 (2022).
Raffel, C. et al. Exploring the limits of transfer learning with a unified text-to-text transformer. Preprint at http://arxiv.org/abs/1910.10683 (2020).
Paszke, A. et al. Pytorch: An imperative style, high-performance deep learning library. Preprint at http://arxiv.org/abs/1912.01703 (2019).
Abadi, M. et al. https://www.tensorflow.org/ TensorFlow: Large-scale machine learning on heterogeneous systems, (2015), software available from tensorflow.org
Bradbury, J. et al. http://github.com/google/jax JAX: composable transformations of Python+NumPy programs, (2018).
Wolf, T. et al. Huggingface’s transformers: State-of-the-art natural language processing. Preprint at http://arxiv.org/abs/1910.03771 (2020).
Nielsen, M. A. & Chuang, I. L. Quantum computation and quantum information. Phys. Today 54, 60 (2001).
Peruzzo, A. et al. A variational eigenvalue solver on a photonic quantum processor. Nat. Comm. 5, 4213 (2014).
Higgott, O., Wang, D. & Brierley, S. Variational Quantum Computation of Excited States. Quantum 3, 156 (2019).
Kottmann, J. S. et al. Tequila: a platform for rapid development of quantum algorithms. Quantum Sci. Tech. 6, 024009 (2021).
Developers, C. https://doi.org/10.5281/zenodo.7465577 Cirq, (2022), See full list of authors on Github: https://github.com/quantumlib/Cirq/graphs/contributors.
Bergholm, V. et al. Pennylane: Automatic differentiation of hybrid quantum-classical computations. Preprint at http://arxiv.org/abs/1811.04968 (2022).
Arrazola, J. M. et al. Differentiable quantum computational chemistry with pennylane. Preprint at http://arxiv.org/abs/2111.09967 (2023).
Pierson, R. H. & Fay, E. A. Guidelines for interlaboratory testing programs. Anal. Chem. 31, 25A–49A (1959).
Lowhorn, N. D. et al. Round-robin studies of two potential seebeck coefficient standard reference materials, in 2007 26th International Conference on Thermoelectrics pp. 361–365, https://doi.org/10.1109/ICT.2007.4569495 (2007).
Moylan, S., Brown, C. U. & Slotwinski, J. Recommended protocol for round-robin studies in additive manufacturing. J. Test. Eval. 44, 1009–1018 (2016).
Brown, C. U. et al. Interlaboratory study for nickel alloy 625 made by laser powder bed fusion to quantify mechanical property variability. J. Mat. Eng. Perf. 25, 3390–3397 (2016).
Alleno, E. et al. Invited Article: A round robin test of the uncertainty on the measurement of the thermoelectric dimensionless figure of merit of Co0.97Ni0.03Sb3. Rev. Sci. Inst. 86, 011301 (2015).
Jiang, Y. et al. \({{{{\rm{Pb}}}}}_{9}{{{\rm{Cu}}}}{({{{{\rm{PO}}}}}_{4})}_{6}{({{{\rm{OH}}}})}_{2}\): Phonon bands, localized flat-band magnetism, models, and chemical analysis. Phys. Rev. B 108, 235127 (2023).
Lee, S., Kim, J.-H. & Kwon, Y.-W. The first room-temperature ambient-pressure superconductor Preprint at http://arxiv.org/abs/2307.12008 (2023).
Guo, K., Li, Y. & Jia, S. Ferromagnetic half levitation of lk-99-like synthetic samples. Sci. China Phys., Mech. ; Astro 66, 107411 (2023).
Kumar, K., Karn, N. K., Kumar, Y. & Awana, V. P. S. Absence of superconductivity in LK-99 at ambient conditions. Preprint at http://arxiv.org/abs/2308.03544 (2023).
Engelbrecht-Wiggans, A. et al. Effects of temperature and humidity on high-strength p-aramid fibers used in body armor. Text. Res. Journ. 90, 2428–2440 (2020).
Thiyagalingam, J. et al. https://github.com/stfc-sciml/sciml-bench Scimlbench: A benchmarking suite for ai for science, (2021).
Brown, N., Fiscato, M., Segler, M. H. & Vaucher, A. C. Guacamol: benchmarking models for de novo molecular design. J. Chem. Info Model. 59, 1096–1108 (2019).
Chen, G. et al. Alchemy: A quantum chemistry dataset for benchmarking ai models. Preprint at https://arxiv.org/abs/1906.09427 (2019).
Khatib, M. E.& de Jong, W. A. Ml4chem: A machine learning package for chemistry and materials science. Preprint at https://arxiv.org/abs/2003.13388 (2020).
Broccatelli, F., Trager, R., Reutlinger, M., Karypis, G. & Li, M. Benchmarking accuracy and generalizability of four graph neural networks using large in vitro adme datasets from different chemical spaces. Mol. Info 41, 2100321 (2022).
Johnson, R. D. et al. Nist computational chemistry comparison and benchmark database. http://srdata.nist.gov/cccbdb (2006).
Prandini, G., Marrazzo, A., Castelli, I. E., Mounet, N. & Marzari, N. Precision and efficiency in solid-state pseudopotential calculations. npj Comp. Mat. 4, 72 (2018).
Karls, D. S. et al. The openkim processing pipeline: a cloud-based automatic material property computation engine. J. Chem. Phys. 153, 064104 (2020).
Hale, L. M., Trautt, Z. T. & Becker, C. A. Evaluating variability with atomistic simulations: the effect of potential and calculation methodology on the modeling of lattice and elastic constants. Model. Sim. Mat. Sci. Eng. 26, 055003 (2018).
Choudhary, K. et al. High-throughput assessment of vacancy formation and surface energies of materials using classical force-fields. J. Phys.: Cond. Matt. 30, 395901 (2018).
Cross, A. W., Bishop, L. S., Sheldon, S., Nation, P. D. & Gambetta, J. M. Validating quantum computers using randomized model circuits. Phys. Rev. A 100, 032328 (2019).
Tomesh, T. et al. Supermarq: a scalable quantum benchmark suite, in 2022 IEEE International Symposium on High-Performance Computer Architecture (HPCA) (IEEE, 2022) pp. 587–603.
Häse, F. et al. Olympus: a benchmarking framework for noisy optimization and experiment planning. Mach. Learn.: Sci. Technol. 2, 035021 (2021).
Aldeghi, M., Häse, F., Hickman, R. J., Tamblyn, I. & Aspuru-Guzik, A. Golem: an algorithm for robust experiment and process optimization. Chem. Sci. 12, 14792–14807 (2021).
Hattrick-Simpers, J. R. et al. An inter-laboratory study of zn–sn–ti–o thin films using high-throughput experimental methods. ACS Comb. Sci. 21, 350–361 (2019).
Choudhary, K., DeCost, B. & Tavazza, F. Machine learning with force-field-inspired descriptors for materials: Fast screening and mapping energy landscape. Phys. Rev. Mat. 2, 083801 (2018).
Ward, L., Agrawal, A., Choudhary, A. & Wolverton, C. A general-purpose machine learning framework for predicting properties of inorganic materials. npj Comp. Mat. 2, 16028 (2016).
Wexler, R. B., Gautam, G. S., Stechel, E. B. & Carter, E. A. Factors governing oxygen vacancy formation in oxide perovskites. J. Am. Chem. Soc. 143, 13212–13227 (2021).
Jha, D. et al. Elemnet: deep learning the chemistry of materials from only elemental composition. Sci. Rep. 8, 17593 (2018).
Jha, D. et al. Enhancing materials property prediction by leveraging computational and experimental data using deep transfer learning. Nat. Comm. 10, 1–12 (2019).
Gupta, V. et al. Cross-property deep transfer learning framework for enhanced predictive analytics on small materials data. Nat. Comm. 12, 6595 (2021).
Gupta, V., Liao, W.-k., Choudhary, A., & Agrawal, A. Pre-activation based representation learning to enhance predictive analytics on small materials data, in 2023 International Joint Conference on Neural Networks (IJCNN) (IEEE, 2023) pp. 1–8.
Jha, D. et al. Irnet: A general purpose deep residual regression framework for materials discovery, in Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, edited by (2019) pp. 2385–2393.
Jha, D. et al. Enabling deeper learning on big data for materials informatics applications. Sci. Rep. 11, 4244 (2021).
Jha, D., Gupta, V., Liao, W.-k, Choudhary, A. & Agrawal, A. Moving closer to experimental level materials property prediction using ai. Sci. Rep. 12, 1–9 (2022).
Gupta, V., Liao, W.-k., Choudhary, A., & Agrawal, A. Brnet: Branched residual network for fast and accurate predictive modeling of materials properties, in Proceedings of the 2022 SIAM international conference on data mining (SDM) (SIAM, 2022) pp. 343–351.
Gupta, V., Peltekian, A., Liao, W.-k, Choudhary, A. & Agrawal, A. Improving deep learning model performance under parametric constraints for materials informatics applications. Sci. Rep. 13, 9128 (2023).
Bartel, C. J. et al. A critical examination of compound stability predictions from machine-learned formation energies. Npj Comput. Mat. 6, 97 (2020).
Gasteiger, J., Groß, J., & Günnemann, S. Directional message passing for molecular graphs, in International Conference on Learning Representations (ICLR) (2020).
Gasteiger, J., Giri, S., Margraf, J. T., & Günnemann, S. Fast and uncertainty-aware directional message passing for non-equilibrium molecules, in Machine Learning for Molecules Workshop, NeurIPS (2020).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the amber ff99sb protein force field. Proteins: Struct., Func., Bioinfo. 78, 1950–1958 (2010).
Van Der Spoel, D. et al. Gromacs: fast, flexible, and free. J. Comp. Chem. 26, 1701–1718 (2005).
Ward, L. et al. Including crystal structure attributes in machine learning models of formation energies via voronoi tessellations. Phys. Rev. B 96, 024104 (2017).
Acknowledgements
K.C., D.W., K.F.G., A.F., A.J.B., M.W., and F.T. thank the National Institute of Standards and Technology for funding, computational, and data-management resources. This work was performed with funding from the CHIPS Metrology Program, part of CHIPS for America, National Institute of Standards and Technology, U.S. Department of Commerce. K.C. thanks the computational support from XSEDE (Extreme Science and Engineering Discovery Environment) computational resources under allocation number TG-DMR 190095. Contributions from K.C. were supported by the financial assistance award 70NANB19H117 from the U.S. Department of Commerce, National Institute of Standards and Technology. J.T.K., K.S., P.G. and P.R.C.K. were supported by the U.S. Department of Energy, Office of Science, Basic Energy Sciences, Materials Sciences and Engineering Division, as part of the Computational Materials Sciences Program and Center for Predictive Simulation of Functional Materials. A.F. and P. G. were supported by the Center for Nanophase Materials Sciences, which is a US Department of Energy, Office of Science User Facility at Oak Ridge National Laboratory. AHR thanks the Supercomputer Center and San Diego Supercomputer Center through allocation DMR140031 from the Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS) program, which is supported by National Science Foundation grants #2138259, #2138286, #2138307, #2137603, and #2138296. AHR also recognizes the support of West Virginia Research under the call Research Challenge Grand Program 2022 and NASA EPSCoR Award 80NSSC22M0173. N.M. and A.M. acknowledge support from the U.S. Department of Energy through the LANL LDRD Programs under grant no. 20210036DR and 20220814PRD4, respectively. V.G. and A.A. were supported by NIST award 70NANB19H005 and NSF award CMMI-2053929. S.H.W. especially thanks to the NSF Non-Academic Research Internships for Graduate Students (INTERN) program (CBET-1845531) for supporting part of the work in NIST under the guidance of K.C. A.M.K. acknowledges support from the School of Materials Engineering at Purdue University under startup account F.10023800.05.002. P.F. acknowledges support by the Federal Ministry of Education and Research (BMBF) under Grant No. 01DM21001B (German-Canadian Materials Acceleration Center).
Please note certain equipment, instruments, software, or materials are identified in this paper in order to specify the experimental procedure adequately. Such identification is not intended to imply the recommendation or endorsement of any product or service by NIST, nor is it intended to imply that the materials or equipment identified are necessarily the best available for the purpose.
This manuscript has been authored by UT-Battelle, LLC, under contract DE-AC05-00OR22725 with the US Department of Energy (DOE). The publisher acknowledges the US government license to provide public access under the DOE Public Access Plan (https://www.energy.gov/doe-public-access-plan). The Los Alamos National Laboratory is operated by the Triad National Security, LLC, for the National Nuclear Security Administration of U.S. Department of Energy (Contract No. 89233218CNA000001).
Author information
Authors and Affiliations
Contributions
K.C. conceived of the idea, developed the workflow, designed several benchmarks and oversaw the project. K.C., D.W., K.L., K.F.G. and V.G. wrote the first draft of the manuscript. K.C., D.W., K.L., K.F.G., V.G., A.H.R., J.T.K., K.S., A.F., R.W., A.M.K., K.Y., Y.L., P.R., A.M., S.H.W., E.B., A.D.R., T.D.R., A.J.B., F.T. uploaded benchmarks and contributions to the leaderboard. All authors contributed in editing and revising the manuscript. List of contributors to the GitHub repository for this work is also available at: https://github.com/usnistgov/jarvis_leaderboard/graphs/contributors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Choudhary, K., Wines, D., Li, K. et al. JARVIS-Leaderboard: a large scale benchmark of materials design methods. npj Comput Mater 10, 93 (2024). https://doi.org/10.1038/s41524-024-01259-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-024-01259-w
This article is cited by
-
Setting standards for data driven materials science
npj Computational Materials (2024)