Abstract
Why are materials with specific characteristics more abundant than others? This is a fundamental question in materials science and one that is traditionally difficult to tackle, given the vastness of compositional and configurational space. We highlight here the anomalous abundance of inorganic compounds whose primitive unit cell contains a number of atoms that is a multiple of four. This occurrence—named here the rule of four—has to our knowledge not previously been reported or studied. Here, we first highlight the rule’s existence, especially notable when restricting oneself to experimentally known compounds, and explore its possible relationship with established descriptors of crystal structures, from symmetries to energies. We then investigate this relative abundance by looking at structural descriptors, both of global (packing configurations) and local (the smooth overlap of atomic positions) nature. Contrary to intuition, the overabundance does not correlate with low-energy or high-symmetry structures; in fact, structures which obey the rule of four are characterized by low symmetries and loosely packed arrangements maximizing the free volume. We are able to correlate this abundance with local structural symmetries, and visualize the results using a hybrid supervised-unsupervised machine learning method.
Similar content being viewed by others
Introduction
Computational materials discovery is a fast-growing discipline leading to innovation in many fields. Within a specific technological sector (i.e., communications, renewable energies, medical), the choice of material is critical for the long-lasting success of the given product. Therefore, it is important—and of fundamental interest—to efficiently identify materials’ structural and energetic characteristics through materials’ data analysis to select structures for innovative applications. The emerging field of materials informatics has demonstrated its potential as a springboard for materials development, alongside first-principles techniques such as density-functional theory (DFT)1,2. The increase in computational power, together with large-scale experimental3 and computational high-throughput studies4, is paving the way for data-intensive, systematic approaches to classify materials’ features and to screen for optimal experimental candidates. In addition, the collection of statistical methods offered by machine learning (ML) has accelerated these efforts, both within fundamental and applied research5,6,7,8,9,10.
However, the success of these endeavours is ultimately limited by the quality and diversity of the data serving as the underlying data source. Understanding the space of materials spanned by a dataset is integral to data-driven materials searches or machine-learning workflows. Thus, when anomalous correlations arise in datasets, it is useful to understand and investigate the origins, and potential implications, of such peculiarities. We use here the name rule of four (RoF) to describe the unusually high relative abundance of structures with primitive unit cells containing a multiple of 4 atoms. This occurrence is explored within two different databases of inorganic crystal structures: the Materials Project (MP)11 database, which contains crystal structures that have been relaxed with first-principles calculations starting from experimental databases or from structure-prediction methods, and the Materials Cloud 3-dimensional crystal structures ‘source’ database (MC3D-source); this latter combines experimental structures from the crystallographic open database (COD)12,13,14,15, the inorganic crystal structures database (ICSD)16 and the materials platform for data science (MPDS). Note that for the ICSD and COD, occasionally some theoretically predicted structures can also be present, see section I in the supplementary information for more details. Figure 1 is a visual representation of this striking abundance, while Table 1 demonstrates the RoF by comparing the relative abundance of structures with primitive unit cells made up of multiple of 3, 4, 5, 6 and 7 atoms.
The two datasets (the Materials Project (MP)11 and the Materials Cloud 3-dimensional crystal structures ‘source’ database (MC3D-source)) contain a disproportionate amount (coloured in red) of compounds with a primitive unit cell containing multiples of 4 atoms. nRoF characterises the number of structures in the datasets that obey the rule of four, while nnon−RoF the ones that do not. The distributions are normalised.
Within the context of this study, we will label a structure that belongs to the subset of structures with a unit cell size multiple of four as a RoF structure, and one that does not belong to the subset as a non-RoF structure. In Fig. 1 the x axis is capped at 100 atoms to best represent the RoF, as respectively 97.51% and 91.00% of structures in the MP and in the MC3D-source databases contain 100 atoms or less (the largest cell in the MP database contains 296 atoms, while the MC3D-source one contains 4986 atoms).
Before delving into an extensive analysis, we rule out that the RoF is simply an artefact of how structures are mathematically described, or of how this description is curated and processed for storage in the aforementioned databases (III A). We then decide to probe the RoF more deeply and attempt to understand its origins and impact. First, we examine the RoF with respect to traditional materials science metrics, including energies and symmetries, and uncover that the RoF is largely correlated with loosely-packed polyatomic systems (III B, III C). We then use symmetry-adapted machine learning techniques to relate the RoF to local atomic environments and determine that it has only little implications for formation energy (III D). We finally manage to correctly classify the RoF by only providing the algorithm with information on local structural symmetry rather than a global one (III D). Although we explored many meaningful avenues to rationalize the rule’s existence and emergence, a full explanation of the anomalous distribution is still missing. Since the most plausibile causes have been explored, the present work serves also as a reference for future research on the topic.
Results
Within this study, we make sure that the data is sufficiently diverse for the training set to cover the whole design space17 by procuring the structural data from open and FAIR repositories18,19,20; the same analytical workflow is applied to two different databases of bulk, crystalline, stoichiometric compounds. One database is the Materials Project, which contained 83 989 data entries obtained via high-throughput DFT calculations as of 10/18/2018, corresponding to the mp all 20181018 dataset retrieved with the matminer.datasets module21. The other data source, the MC3D-source, contains 79 854 unique structures extracted from the MPDS, ICSD and COD, which have been curated via an AiiDA22 workflow, as explained in Section I of the SI.
Primitive unit cell
When materials structure datasets are prepared, it is standard procedure to ‘primitivise’ unit cells, i.e., to reduce the unit cell to its minimum volume. As many conventional unit cells contain exactly four times the number of atoms that would be found in their respective primitive unit cell, it could be expected that misclassifying conventional unit cells as primitive ones could lead to an artificial emergence of the RoF. Both the MP and MC3D-source databases obtain the primitive unit cell using the spglib software23. When primitivizing the structure, one needs to set the symprec tolerance parameter, which allows for slight deviations in the atomic positions stemming from thermal motion or experimental noise. To rule out that the primitivization is the source of the emergence of the RoF, we show in Fig. 2 that changing the symprec (1E-8 to 1E-1Å) parameter has little effect on the RoF distribution, converting around 1% of RoF structures into non-RoF ones. It is only when one increases the symprec to unreasonably large values (close to 1 Å) that the slope changes—this is expected, as using such a large tolerance effectively considers sites with the same element that should be different as identical, producing primitive unit cells with a reduced number of sites, but which no longer correctly describe the structure. Encouraged by these results, we proceed with a more extensive analysis.
The black and green lines correspond to structures in the MP and MC3D-source datasets, respectively. At typical symmetrization parameters, there is little to no change in the number of RoF structures (roughly 1% of RoF structures go to non-RoF). At larger symmetrization parameters (≈1 Å), this increases to roughly 6% based upon the large deviations allowed in considering sites as symmetrically equivalent.
Formation energy
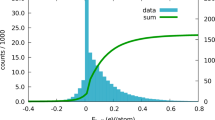
We first test whether the RoF is correlated with stability with respect to elemental phases, as this would provide a straightforward explanation for the phenomenon. To test this assumption, we analyze the information contained in the MP dataset, namely the formation energy per atom within each compound. This is the energy of the compound with respect to standard states (elements), normalized per atom. For example, for Fe2O3 the formation energy is [E(Fe2O3)–2E(Fe)–(3/2)E(O2)]/5. It is computed at a temperature of 0 K and a pressure of 0 atm. This quantity is often a good approximation for formation enthalpy at ambient conditions, where a negative formation energy implies stability with respect to elemental compounds.
Our initial results provide no evidence of a correlation between RoF compounds and their formation energy, as shown in Fig. 3. Nevertheless, it does appear that structures obeying the RoF have a longer positive tail of large formation energies, seen towards the bottom right of the figure.
However, this result can be misleading—it does not take into consideration the large variance in structural composition across the database—and we must aim to compare the energies of similar structures within the RoF and non-RoF subsets, as we will do in later sections.
Correlation with symmetry descriptors
The crystal symmetries of compounds—defined by the set of symmetry operations that, when performed, leave the structure unchanged—are captured in crystals by their space groups and point groups. Higher symmetry space groups inherit the symmetry operations of their ‘parent’ point groups; for example, cubic space groups inherit the one-fold, two-fold, and four-fold rotational symmetries (for the interested reader, the concept of inherited symmetry is enumerated nicely in Fig. 1.5 of the book chapter by Hestenes24). Figure 4 shows histograms of inherited symmetries and their relative abundance within each of the two sets (RoF in red and non-RoF in blue). The point groups are ordered from the ones with the least number of symmetry operations (bottom) to the highest order ones (top). Symmetry groups that are equally represented in both sets (i.e. 1-rotation, since all compounds are invariant to the simplest symmetry) have tails of equal length, whereas symmetries seen in a larger percentage of RoF structures have a red tail to the right of the histogram.
From Fig. 4, the relative abundance of non-RoF structures in the high symmetry point groups emerges, while on the contrary most RoF structures in both databases are grouped in the lowest symmetry point groups (2, m, 2/m, mm2, 222 and mmm), which generally contain a relative abundance of them apart from one exception (the MC3D-source presents. a slightly higher relative abundance of non-RoF structures in the mm2 point group). This analysis shows how 4-fold symmetry is not a determining descriptor to classify the phenomenon.
The the lack of higher symmetry groups in RoF compounds could be correlated by a heterogeneous composition of atoms; this heterogeneity can be quantified by counting the number of atomic species (Nspecies) (first column of Fig. 5, in logarithmic scale) composing the structures: from this analysis we see that RoF materials are mostly composed of 4 or more elements (statistics start being less reliable after Nspecies = 8), while non-RoF structures present a larger abundance of simpler composition, containing more often 1, 2, or 3 elements. When looking at chemical composition, the hypothesis of the RoF emerging from signatures associated to a specific element have been ruled out. For example, when selecting from both datasets only structures containing Si (the first direct candidate having a typical coordination number of 4), the RoF still emerges with a probability of 59.07% for the MP dataset and 57.32% for the MC3D-source one. These statistics are not sufficiently divergent from the results in Table 1.
Different geometric properties of each compound are analysed for the (a) MC3D-source and (b) MP databases. From left to right, the plots represent the normalised distribution of the number of elemental species (Nspecies), the relative abundance of small (NS) to large (NL) atomic radii (x), the ratio between smallest (RS) and largest (RL) atomic radii (α) and the packing fraction (PF) for compounds with a unit cell size between 0 and 100 atoms. All of the results are plotted for the two sets, RoF (red) and non-RoF (blue), with the probability normalized to each set.
Another property that emerges from our analysis and is more evident in the MP dataset (second column of Fig. 5(b)) is the relative abundance of smaller atomic radii within RoF compounds, as often defined by the parameter \(x=\frac{{N}_{s}}{{N}_{s}+{N}_{L}}\)25, where NS and NL are the counts, in a given structure, of the smallest and largest radii respectively. When considering this parameter, we focus our attention on the MC3D-source dataset, which contains bigger and more complex structures, where the divergence between the smallest and largest atoms is more considerable. In fact, the first peak in Fig. 1 for the MP is very large, hinting at the implicit bias of computational studies, where larger structures are often avoided based upon computational cost of calculations. The abundance of small atomic radii in RoF compounds of the MC3D-source dataset (higher x parameter) partly explains the lower symmetries that characterise them, as more atoms will be inserted as ‘interstitial’ elements in a given structure, characterising the ‘imperfections’ that eventually contribute in lowering the overall structural symmetry of point groups analyzed in Fig. 4.
In general, the symmetry type of atomic crystal systems is strictly linked to packing mechanisms26,27,28. While the mathematical problem of sphere packing is not hard to pose (Kepler conjecture), it was historically difficult to prove29, and the complexity of its solution rises exponentially with polydispersity30.
Despite this, a qualitative analysis of RoF configurations shows that they contain chemical elements whose size variance is greater compared to the variance in the non-RoF population.
This size variance is quantified by the parameter \(\alpha =\frac{{R}_{S}}{{R}_{L}}\) (where RL is the radius of the largest atomic radius and RS of the smallest one), namely the ratio between the smallest and the biggest atomic radii within each compound (third column of Fig. 5). For the same reason as above, in the context of this parameter the results of the MC3D-source dataset are considered more relevant. RoF compounds from the MC3D-source exhibit a greater standard deviation between largest and smallest atoms, with the α parameter presenting a peak at around 0.35; this finding suggests the presence of very small radii filling the interstitial spaces, which contribute to keeping the symmetry of RoF compounds low, as was previously highlighted by the analysis of the x paramter.
However, there is no overarching evidence in the distributions of the x and α parameters that allows us to confirm a correlation between the emergence of the RoF and the abundance of ‘intestitial’ elements.
The packing fraction (PF), defined as as \({PF}=\frac{{V}_{{tot},{atoms}}}{{V}_{{cell}}}\) (where V tot,atoms is the total volume of all atoms composing the structure, and V cell is the unit cell’s volume) is another related property of sphere packing26,27. This quantity is noticeably lower (with peaks at values around 0.1–0.2) for RoF structures, as can be seen in the last column of Fig. 5a, b, pointing away from packing arguments as the cause of this database anomaly. The sharp red peaks in PF might characterise disordered compounds such as porous materials, which have been determined to be outliers for the MC3D-source dataset.
Employing symmetry-adapted descriptors for further insight
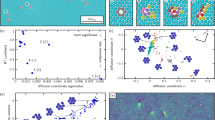
Up until this point we have employed classical techniques for analyzing crystal structures; these have offered little to no insights on the origins of the RoF anomalous distribution, but have allowed us to exclude packing arguments and symmetrical global descriptors as features that make the RoF emerge. Here, we therefore turn to more modern data- driven techniques. In the field of atomistic modelling, it has been common, albeit non-trivial, to represent crystal structures through symmetrized density correlations9,31,32 in order to predict broad swaths of materials properties. Here, we represent the compounds using the Smooth Overlap of Atomic Positions (SOAP)31, a popular ML representation for structure-energy relations that contains information on the average three-body local environment for atomic arrangements. SOAP vectors provide an avenue for a statistical analysis on local environments, offering a robust framework through which we can explore and visualize the chemical and configuration space of the materials studied33. We use two parameterizations of SOAP vectors, detailed in Section II of the Supplementary Information: one that uses separate channels to represent different chemical species and another that ignores the chemical identities in order to highlight the geometry of the local symmetry. The former, from hereon called the species-tagged representation, is necessary in energetic analysis, as similar geometry symmetries can correspond to wildly different energetics given the elements present; however, this representation is computationally cumbersome (roughly 100 000 sparse features for each compound, from which we take a diverse subset of 2 000 features). Thus, in later analyses where the chemical identities play a smaller role, it is beneficial and conceptually more straightforward to use the more lightweight, latter representation (roughly 80 features for each compound), hereon called the species-invariant representation.
Earlier, we noted that simply presenting a histogram of RoF and non-RoF energetics did not provide any specific understanding of the RoF; it might be more insightful to compare the energies of chemically similar structures. To determine whether the RoF structures exhibit lower energy than structurally-similar non-RoF ones, we use Principal Covariates Regression (PCovR)34,35, a ML method which constructs a latent space projection to explore the correlation between stability and local symmetries within the dataset by expanding regression models to incorporate information on the structure of the input data, as implemented in the scikit-matter library36,37. In this mixing model, the projection is weighted towards the property of interest using a mixing parameter (of which a more extensive explanation is given in Supplementary Fig. 1), and, where the input linearly correlates with the target property, the resulting embedding will reflect this property along the first component, with subsequent components representing orthogonal dimensions in structure space. In our case the PCovR is always trained on the species-tagged SOAP vectors and their formation energies. We plot the first two principal covariates in Fig. 6. The first principal covariate is strongly correlated to the energetic descriptor, as can be seen in Fig. 6, where in the lower plots we have coloured each point in the projection by their RoF classification (left) and formation energy (right). However, the second covariate (and all significant subsequent covariates, see Supplementary Figs. 2-4) fail to separate the datasets into two distinct populations corresponding to this phenomenom. This implies that for structurally similar compounds, there is no significant difference in energy between RoF and non-RoF samples. We also see little difference in the spread of RoF versus non-RoF structures in the latent space, as shown by the kernel density probability map in the upper panel of Fig. 6. Further principal covariates for the same PCovR representation are plotted in Supplementary Figs. 2-4, as well as other relevant energetic descriptors (the energy above the convex hull energy, i.e., the envelope connecting the lowest energy compounds in the chemical space, and the band gap energy), in order to show how these targets yield similar results. Thus the RoF is neither correlated with the energetics, nor are RoF lower in formation energy when compared to chemically similar non-RoF ones.
The model is regressed on the formation energy per atom. The three plots contain the same data, represented on the top through a kernel density probability distribution (the RoF subset is coloured in red and the non-RoF one in blue), coloured according to the subset classification (lower left) and according to the formation energy per atom (lower right). The plot on the top is generated with the seaborn.kdeplot() function.
The linear correlation between the average local symmetries and the RoF is not particularly strong (a logistic regression on the SOAP vectors results in an accuracy on the order of 0.6, as listed in Table 1 of the SI); thus, we turn to non-linear classifications to understand if the RoF is potentially correlated with these local neighbourhoods. We ignore the species information to focus solely on the average local symmetries. We build a Random Forest (RF) classification38 on both datasets, first varying the interaction cutoff that defines the local environment (see Fig. 7). We see a plateau in accuracy on the test set at 87% as we consider local environments of 4.0 Å, suggesting that differentiating local symmetries occur within the first two neighbour shells, also supported by the high false positive (FP) rate at small cutoff radii. From the learning curve on the 4.0 Å escriptors (inset), we see that the classification has a positive learning rate, although shows little saturation despite the large training set. This result implies that local features are sufficient for the ML model to pick up the complexity of the datasets and to predict with good probability the correct classification. We report the accuracy on the test set achieved by other classification algorithms in Section V of the SI. To visualise the stoichiometry of materials falling into one of the two categories, the interested reader is referred to our Materials Cloud Archive39 entry, from which a json.gz file for each database can be downloaded. This can be uploaded on the Chemiscope40 web interface to visualise a 2D plot of the first principal covariates. The chemical composition of each dataset structure can be inspected by clicking on the dots composing the scatter plot. The colouring can be done according to different parameters, of which the most insightful one is the classification outcome.
Here we use the species-invariant 3-body SOAP vectors to build a random forest ensemble classifier. Test set accuracy, represented on the y axis, saturates at ~4.0 A, with little additional gain at larger cutoff radii. Below the figure we show the table of true positive (TP), true negative (TN), false positive (FP), and false negative (FN) results, showing that the classifier is unable to differentiate RoF and non-RoF structures at lower cutoff radii, leading to a high false positive (FP) rate. Inset in the upper figure is a learning curve for a cutoff radius of 4.0 Å, which shows a positive learning rate, albeit no saturation, an indication that secondary effects beyond the local environments play a role (or, more unlikely, that the dataset is not sufficiently large).
Discussion
Through an extensive investigation, in this work we highlight and analyze the anomalous abundance of inorganic compounds whose primitive unit cell contains a number of atoms that is a multiple of four, a property that we name rule of four (RoF) and that is observed in both experimental and DFT-generated structure databases. Here, we:
-
highlight the rule’s existence, especially notable when restricting oneself to mostly experimentally known compounds;
-
explore its possible relationship with established energetic descriptors, namely formation energies, and utilise hybrid ML methods combining regression and principal component analysis to surprisingly rule out the possibility that the relative abundance has the (expected) effect of stabilising compounds, bringing them to a lower energy state;
-
conclude, through a global structural composition analysis of point groups and packing fractions, that the overabun- dance does not either correlate with high-symmetry structures, but rather to low symmetries and loosely packed arrangements maximising the free volume;
-
predict, with an accuracy of 87% the association to the rule of four of a compound by providing a random forest classification algorithm with local structural descriptors (the smooth overlap of atomic positions) only, eventually highlighting the importance of local symmetry rather than global one for the emergence of the rule of four.
This analysis constitutes a valuable reference for further systematic studies targeting the classification of materials’ features with specific ML approaches in order to screen for optimal experimental candidates. Moreover, the study provides a starting point for future investigations on the rule’s emergence, given that a fully satisfactory explanation of such anomalous distribution is as yet lacking.
Methods
Reduction to the primitive cell
All the structures in both databases are reduced to the primitive cell using the find primitive function of the spglib23 package, varying the symprec value in the range of 1E − 8 to 1 Å.
Scalar global descriptors
The symmetry of compounds is investigated by looking at space groups and point groups. subgroup of symmetry operations over which the space group is invariant. With a total number of 32 point groups, it is easier to convey the symmetric properties of the vast variety of compounds via their point group rather than their space groups; while space groups uniquely identify geometric properties, point groups identify symmetry classes and reduce the parameter space to a lower degree when investigating the symmetries of all compounds. The point groups are calculated through the spglib23 and seekpath41 packages for the MC3D-source database, while we used the SymmetryAnalyzer pymatgen module—which also relies on the spglib package developed by Togo and Tanaka23—to find the symmetry operators and point groups for the MP dataset, at a symprec of 0.3 Å. As concerns packing mechanisms, we extend the conventions employed by Hopkins25 to n-elements packing and employ the α, PF and x parameters. In structures with FCC and HCP symmetry, the maximum packing fraction is 0.74. α = 1 denotes unary compound. Conversely, when α 0 the compounds contain elements whose atomic radii distribution presents a wider spread.
Local symmetry descriptors and ML pipeline
We adopt the following ML pipeline to study local symmetries and energetic effects. First, the atomic representation of each compound is obtained with SOAP vectors (see section II of the Supplementary Information), computed with the librascal library33. The SOAP features are then averaged within each compound, and the representations from the two datasets are normalised simultaneously. We then select a diverse subset of 2 000 features through Furthest Point Sampling (FPS) algorithm36,37,42,43, efficiently reducing the dataset size without losing important information. For Sec.III D, we perform a linear ridge regression with 4-fold cross-validation—which optimises the regularisation parameter to prevent overfitting—on the formation energies data retrieved from the MP database to ascertain the accuracy of the model. Table 2 illustrates the RMSE and the uncertainty in units of eV of the predicted energetic quantities.
Compared to results in the literature, which achieve an accuracy in formation energy prediction of 0.173 eV (Automatminer44) and 0.0332 eV (Crystal Graph Convolutional Neural Networks45), the accuracy of 0.4002 eV is sufficient for this study, since the aim of our study is not to find the most efficient way to predict energies, but rather to provide a sufficient regression prediction to employ in PCovR analysis. We use the species-invariant SOAP vectors to classify the RoF phe- nomenon using scikit-learn’s46 RandomForestClassifier algorithm47, which accepts binary labels as target properties (RoF or non-RoF) and outputs a probability between 0 and 1 for each compound to fall into the RoF subset. Training and testing set constitute respectively 90 and 10% of the whole dataset. Our random forest classification comprises 100 random decision trees. This classifier performs better in our case compared to Support Vector Machine (SVM) and Logistic Regression (LR) classifiers, signifying a need for a stochastic model.
Data availability
The full dataset employed for the analysis can be downloaded from the Materials Cloud Archive39, where the MC3D- source data is only provided in SOAP format as the experimental structures can not be released due to licensing constraints. Its DFT–relaxed counterpart is available at: https://archive.materialscloud.org/record/2022.38. Instead, we provide the full list of structure IDs for each database, including the version of the database upon the time of extraction.
Code availability
The codes to reproduce the results and figures can be found at: https://github.com/epfl-theos/r4-project. As the MC3D- source structure data cannot be made publicly available due to licensing contraints, the repository contains example data from a reduced random subset of the publicly available MP dataset in order to test run a preliminary analysis.
Change history
13 May 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41524-024-01284-9
References
Hohenberg, P. & Kohn, W. Inhomogeneous electron gas. Phys. Rev. 136, B864–B871 (1964).
Marzari, N., Ferretti, A. & Wolverton, C. Electronic-structure methods for materials design. Nat. Mater. 20, 736–749 (2021).
Suram, S. K., Haber, J. A., Jin, J. & Gregoire, J. M. Generating information-rich high-throughput experimental materials genomes using functional clustering via multitree genetic programming and information theory. ACS Comb. Sci. 17, 224–233 (2015).
Curtarolo, S. et al. The high-throughput highway to computational materials design. Nat. Mater. 12, 191–201 (2013).
Vasudevan, R. K. et al. Materials science in the artificial intelligence age: high-throughput library generation, machine learning, and a pathway from correlations to the underpinning physics. MRS Commun. 9, 821–838 (2019).
Pilania, G., Wang, C., Jiang, X., Rajasekaran, S. & Ramprasad, R. Accelerating materials property predictions using machine learning. Sci. Rep. 3, 2810 (2013).
Rupp, M., Tkatchenko, A., Mueller, K.-R. & von Lilienfeld, O. A. Fast and accurate modeling of molecular atomization energies with machine learning. Phys. Rev. Lett. 108, 058301 (2012).
Sanchez-Lengeling, B. & Aspuru-Guzik, A. Inverse molecular design using machine learning: generative models for matter engineering. Science 361, 360–365 (2018).
Bart´ok, A. P. et al. Machine learning unifies the modeling of materials and molecules. Sci. Adv. 3, e1701816 (2017).
De, S., Bart´ok, A. P., Cs´anyi, G. & Ceriotti, M. Comparing molecules and solids across structural and alchemical space. Phys. Chem. Chem. Phys. 18, 13754–13769 (2016).
Jain, A. et al. Commentary: The materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Vaitkus, A., Merkys, A. & Graˇzulis, S. Validation of the crystallography open database using the crystallographic information framework. J. Appl. Crystallogr. 54, 661–672 (2021).
Quir´os, M., Graˇzulis, S., Girdzijauskaite˙, S., Merkys, A. & Vaitkus, A. Using SMILES strings for the description of chemical connectivity in the crystallography open database. J. Cheminformatics 10, 23 (2018).
Merkys, A. et al. COD::CIF::Parser: an error-correcting CIF parser for the Perl language. J. Appl. Crystallogr. https://doi.org/10.1107/S1600576715022396 (2016).
Graˇzulis, S., Merkys, A., Vaitkus, A. & Okuliˇc-Kazarinas, M. Computing stoichiometric molecular composition from crystal structures. J. Appl. Crystallogr. 48, 85–91 (2015).
Levin, I. NIST Inorganic Crystal Structure Database (ICSD). National Institute of Standards and Technology Public Data Repository. https://data.nist.gov/od/id/mds2-2147 (2018)
Ajiboye, A., Abdullah-Arshah, R., Qin, H. & Isah-Kebbe, H. Evaluating the effect of dataset size on predictive model using supervised learning technique. IJSECS 1, 75–84 (2015).
Wilkinson, M. et al. The fair guiding principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Schmidt, J., Marques, M. R. G., Botti, S. & Marques, M. A. L. Recent advances and applications of machine learning in solid-state materials science. npj Comput. Mater. 5, 1–36 (2019).
Draxl, C. & Scheffler, M. NOMAD: The FAIR concept for big data-driven materials science. MRS Bull. 43, 676–682 (2018).
Pizzi, G., Togo, A. & Kozinsky, B. Matminer, access ready made datasets in one line. https://hackingmaterials.lbl.gov/matminer/#access-ready-made-datasets-in-one-line (2023).
Pizzi, G., Togo, A. & Kozinsky, B. Provenance, workflows, and crystallographic tools in materials science: AiiDa, spglib, and seekpath. MRS Bull. 43, 696–702 (2018).
Togo, A. & Tanaka, I. Spglib: a software library for crystal symmetry search. arXiv https://arxiv.org/abs/1808.01590 (2018).
Hestenes, D. in Applications of Geometric Algebra in Computer Science and Engineering (eds. Dorst, L., Doran, C. & Lasenby, J.) 3–34 (Birkh¨auser Boston, Boston, MA, 2002).
Hopkins, A. B., Jiao, Y., Stillinger, F. H. & Torquato, S. Phase diagram and structural diversity of the densest binary sphere packings. Phys. Rev. Lett. 107, 125501 (2011).
De Graaf, J., Van Roij, R. & Dijkstra, M. Dense regular packings of irregular nonconvex particles. Phys. Rev. Lett. 107, 155501 (2011).
Torquato, S. & Jiao, Y. Dense packings of the platonic and archimedean solids. Nature 460, 876–879 (2009).
Cersonsky, R. K., van Anders, G., Dodd, P. M. & Glotzer, S. C. Relevance of packing to colloidal self-assembly. Proc. Natl. Acad. Sci. USA 115, 1439–1444 (2018).
Hales, T. A proof of the kepler conjecture. Ann. Math. 162, 1065–1185 (2005).
Torquato, S. Perspective: Basic understanding of condensed phases of matter via packing models. J. Chem. Phys. 149, 020901 (2018).
Bart´ok, A. P., Kondor, R. & Cs´anyi, G. On representing chemical environments. Phys. Rev. B: Condens. Matter 87, 184115 (2013).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Musil, F. et al. Efficient implementation of atom-density representations. J. Chem. Phys. 154, 114109 (2021).
de Jong, S. & Kiers, H. A. Principal covariates regression: part i. theory. Chemom. Intell. Lab. Syst. 14, 155–164 (1992).
Helfrecht, B. A., Cersonsky, R. K., Fraux, G. & Ceriotti, M. Structure-property maps with Kernel principal covariates regression. Mach. Learn.: Sci. Technol. 1, 045021 (2020).
Cersonsky, R. K. et al. scikit-learn-contrib/scikit-matter: pre-release of v0.1. Zenodo https://doi.org/10.5281/zenodo.4727575. (2021)
Goscinski, A. et al. scikit-matter: A suite of generalisable machine learning methods born out of chemistry and materials science. Open Res. Europe 3, 81(2023).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Gazzarrini, E., Cersonsky, R. K., Bercx, M., Adorf, C. S. & Marzari, N. The Rule of Four: Anomalous Stoichiometries of Inorganic Compounds. https://archive.materialscloud.org/record/2023.104. (2023)
Fraux, G., Cersonsky, R. K. & Ceriotti, M. Chemiscope: interactive structure-property explorer for materials and molecules. J. Open Res. Softw. 5, 2117 (2020).
Hinuma, Y., Pizzi, G., Kumagai, Y., Oba, F. & Tanaka, I. Band structure diagram paths based on crystallography. arXiv https://arxiv.org/abs/1602.06402 (2016).
Eldar, Y., Lindenbaum, M., Porat, M. & Zeevi, Y. Y. The farthest point strategy for progressive image sampling. IEEE Trans. Image Process. 6, 1305–1315 (1997).
Cersonsky, R. K., Helfrecht, B. A., Engel, E. A., Kliavinek, S. & Ceriotti, M. Improving sample and feature selection with principal covariates regression. Mach. Learn.: Sci. Technol. 2, 035038 (2021).
Dunn, A., Wang, Q., Ganose, A., Dopp, D. & Jain, A. Benchmarking materials property prediction methods: the matbench test set and automatminer reference algorithm. npj Comput. Mater. 6, 1–10 (2020).
Ziletti, A., Kumar, D., Scheffler, M. & Ghiringhelli, L. M. Insightful classification of crystal structures using deep learning. Nat. Commun. 9, 2775 (2018).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Couronn´e, R., Probst, P. & Boulesteix, A.-L. Random forest versus logistic regression: a large-scale benchmark experiment. BMC Bioinf. 19, 270 (2018).
Acknowledgements
This work was supported by a MARVEL INSPIRE Potentials Master’s Fellowship and the MARVEL National Centre of Competence in Research (NCCR) funded by the Swiss National Science Foundation (grant agreement ID 51NF40- 182892). RKC acknowledges funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme under grant agreement No 677013-HBMAP. MB acknowledges funding by the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 824143 (European MaX Centre of Excellence “Materials design at the Exascale”). CSA acknowledges funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 760173. The authors thank the AiiDA team for giving access to the full MC3D-source dataset, and are grateful to the technical and theoretical insights offered by the members of MARVEL, THEOS, and COSMO.
Author information
Authors and Affiliations
Contributions
E.G. conducted the analysis on the data and wrote the paper in collaboration with C.S.A and R.K.C. R.K.C. instructed on how to use many of the employed computational tools, designed the computational strategy, and helped conduct the analysis and interpretation of the data. M.B. provided technical support, especially on the MC3D-source dataset and helped with the analysis and interpretation of the data. C.S.A. managed the project, wrote the paper, and helped with the analysis and interpretation of the data. N.M. supervised the project, helped with the interpretation of the results, and provided the initial suggestion for the investigation. All authors edited and reviewed the paper.
Corresponding author
Ethics declarations
Competing interests
There are no competing interests to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gazzarrini, E., Cersonsky, R.K., Bercx, M. et al. The rule of four: anomalous distributions in the stoichiometries of inorganic compounds. npj Comput Mater 10, 73 (2024). https://doi.org/10.1038/s41524-024-01248-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-024-01248-z