Abstract
The sensitivity of thin-film materials and devices to defects motivates extensive research into the optimization of film morphology. This research could be accelerated by automated experiments that characterize the response of film morphology to synthesis conditions. Optical imaging can resolve morphological defects in thin films and is readily integrated into automated experiments but the large volumes of images produced by such systems require automated analysis. Existing approaches to automatically analyzing film morphologies in optical images require application-specific customization by software experts and are not robust to changes in image content or imaging conditions. Here, we present a versatile convolutional neural network (CNN) for thin-film image analysis which can identify and quantify the extent of a variety of defects and is applicable to multiple materials and imaging conditions. This CNN is readily adapted to new thin-film image analysis tasks and will facilitate the use of imaging in automated thin-film research systems.
Similar content being viewed by others
Introduction
Film morphology optimization is important for reducing the detrimental impacts of defects on the performance of thin-film devices such as photovoltaics1,2 and light-emitting diodes3. Images of thin films carry information about common morphological defects, such as cracking4 and dewetting5,6, which are controlled by film synthesis conditions. While automated experiments can generate images of thin films synthesized under numerous distinct conditions, existing approaches to automatically analyzing film morphologies in such images typically require application-specific customization by software experts and are not robust to changes in image content or imaging conditions7,8. Here we present a versatile convolutional neural network (CNN) for thin-film image analysis which can identify and quantify the extent of a variety of defects, is applicable to multiple materials and imaging conditions, and is readily adapted to new thin-film image analysis tasks.
The severity of film defects such as thickness variations, cracks, precipitates, or dewetting can often be identified by the naked eye or with optical microscopy4,9,10. For this reason, rapid, non-destructive optical inspection of thin films is often carried out in the place of more expensive, more destructive, or more time-consuming methods such as stylus profilometry, atomic force microscopy, or electron microscopy. Quantitative defect analysis enables researchers to identify potentially subtle trends in film morphology as a function of experimental conditions. Researchers frequently perform quantitative image analyses using semi-manual software tools, such as measuring film coverage with ImageJ11, or surface roughness with Gwyddion12. Semi-manual analysis becomes impractical, however, when applied to high-throughput experiments or high-speed manufacturing where images of thin films are generated at high frequency or in large numbers. In such cases, automated image analysis is necessary. Automated analyses of images of thin-film materials and devices are often performed using image-processing algorithms that are specific to the material, morphology, and imaging modality of interest13. An example of this type of approach is the matrix-based analysis of orientational order in AFM images of P3HT nanofibers7. This traditional type of computer vision includes application-specific feature extraction subroutines with numerous adjustable parameters, such as imaging condition-dependent thresholds, which can make them difficult to adapt to new applications14, such as new materials, morphologies, or imaging modalities. Here, we describe a new approach to image-based thin-film defect quantification which uses a CNN to overcome many limitations of previous approaches. The CNN we developed for this purpose, which we call DeepThin, quantifies the extent of several types of common morphological defects (e.g., particles, cracks, scratches, and dewetting) in images of thin films. We show that DeepThin works with different imaging modalities (darkfield imaging and brightfield microscopy), different magnifications and different materials (a small-molecule organic glass and a metal oxide) and can readily be retrained to detect new defect types.
CNNs are a family of machine learning algorithms that have been applied to image classification15, feature detection16, image segmentation17, and object recognition problems18. CNNs have achieved classification accuracy comparable to human experts in computer vision challenges19,20. The performance and robustness demonstrated by CNNs make them appealing for thin-film defect analysis. An additional benefit of CNNs is that they can be easily trained using examples provided by a domain expert (e.g., a materials scientist) rather than through involved algorithm customization by a computer vision expert14. CNNs are an established approach to electron microscopy image analysis tasks, with examples in the materials sciences including mechanical property estimation21, nanoparticle segmentation22, nanostructure classification23, and the study of atomic-scale defects24,25,26. However, CNNs have only recently been applied for the analysis of optical images of thin films in two highly-application-specific ways: classifying the corrosion conditions under which surface films formed on metal surfaces27,28 and determining the thickness of exfoliated 2D crystals29. The DeepThin CNN reported here is a general-purpose CNN for classifying or quantifying common morphological defects in optical images of thin films.
Results
Training dataset and model development
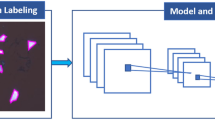
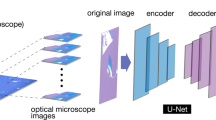
To develop and validate DeepThin (Fig. 1), we first created a dataset of 2600 darkfield images of organic semiconductor thin films (each 4000 × 3000 pixels) exhibiting varying extents of cracking and dewetting due to differences in film composition and annealing conditions. These films were deposited by spin-coating, annealed, and imaged by a flexible robotic platform equipped with a darkfield photography system (see “Methods” section, ref. 30). The images in this darkfield dataset were labeled with respect to the extent of dewetting and of cracking by materials scientists with expertise in thin-film materials research (see “Methods” section). Labeling was on a subjective integer scale from zero (no defects observed) to ten (extremely defected) for both defect types. This labeled dataset was first randomly divided into training, validation, and test sets to facilitate the development of a CNN for image-based thin film defect analysis. These datasets were then augmented by applying rotations and mirroring to the labeled images to obtain a total of 17,374 labeled images. Finally, some images of non-defected films were removed to improve the balance of the datasets, with the final breakdown as detailed in Table S1. This balancing was done to avoid biasing the model towards labeling images as non-defected. We evaluated the suitability of several state-of-the-art CNNs architectures31,32 for this task before choosing to develop a new architecture for DeepThin (Methods) inspired by the VGG16 CNN (see “Methods” section). We trained DeepThin using five-fold cross-validation and the Adam optimizer (see “Methods” section). After this optimization, DeepThin quantified the extent of cracking, and of dewetting, in each of the darkfield dataset images with >93 % accuracy (Table S2). With the largest possible quantification error being 10, the root-mean-square-errors between the model’s scores and the ground truth were 0.086609 for crack quantification and 0.090362 for dewetting quantification.
a Scheme for applying DeepThin to score the extent of cracking (Scrack) and dewetting (Sdewet): (1) thin film sample is created, for example by spin-coating. (2) A darkfield photograph of the sample is taken. (3) The image is subdivided into n patches (4) The DeepThin convolutional neural network (architecture shown) is applied to each patch (5) Scores for the extent of cracking and dewetting are computed by averaging the scores of all n patches. The structure of DeepThin is shown (bottom). The numbers below each layer indicate the output data size of each convolution or fully connected layer. Conv Convolution layer, Pool Pooling layer, ReLU Rectified Linear Units layer, FC fully connected layer. b Example images of organic thin films from the testing portion of the darkfield dataset with varying extents of dewetting, ordered by the scores assigned to them by DeepThin. Scale bar, 500 µm.
Model validation against a known, monotonic morphological trend
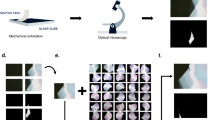
To further validate DeepThin we carried out an experiment where an organic semiconductor film was imaged as it underwent thermally-activated dewetting9 (Fig. 2a). This experiment provided a series of images in which the extent of dewetting was known to increase monotonically with respect to time. We then used DeepThin to quantify the extent of dewetting in each image. The resulting dewetting scores also increased monotonically with respect to time (Fig. 2b), showing that DeepThin can correctly order a set of images of thin films based on a one-dimensional trend in the film morphology.
a The experimental setup used for capturing a series of images of a thin film with an extent of dewetting which increases monotonically in time. b Dewetting score assigned to images of the thin film sample as a function of heating time. The extent of dewetting increased monotonically with time throughout the experiment as seen in the images from 10, 30, 50, 70, and 90 s into the experiment. Similarly, the dewetting score reported by DeepThin also increases monotonically. Scale bar in image, 10 mm.
Resolution of a two-dimensional film-morphology response surface
To illustrate the applicability of our method to thin film optimization, we used DeepThin to resolve a 2-dimensional film-morphology response surface in a set of experiments where both film composition and processing were varied. Following our previous work30, thin films of spiro-OMeTAD doped with varying amounts of FK102 Co(III) TFSI salt and annealed for varying durations were prepared and then imaged using a robotic platform (see “Methods” section). These experiments provided an array of images exhibiting morphological trends as a function of both film composition and processing. The analysis of these images using DeepThin automatically provided a response surface quantifying the extent of dewetting as a function of the film composition and annealing time (Fig. 3). From this surface, two trends can readily be identified: (i) the extent of dewetting increased as the dopant-to-spiro-OMeTAD molar ratio increased from 0 to 0.4, then decreased at higher dopant levels; (ii) longer annealing times produced more dewetted films across all dopant-to-spiro-OMeTAD ratios, with the exception of undoped films which did not exhibit dewetting regardless of annealing time. The ability to automatically obtain composition-processing-morphology response surfaces such as the one shown in Fig. 3 using rapid, inexpensive, and non-destructive imaging is a benefit of our approach.
Applicability of DeepThin to multiple materials, defect types and imaging modalities
To demonstrate the versatility of DeepThin, we next applied it to a different imaging modality (bright-field microscopy) at different magnifications (×5 and ×20), to additional defect types (scratches, particles, and thickness non-uniformities) and to films of a different material (a metal oxide) (Fig. 4). For these demonstrations, three new image datasets were manually obtained using a bright-field microscope (see “Methods” section): a set of 129 images of organic semiconductor films at ×5 magnification and two sets of images of TiOx films (81 images at ×5 magnification and 82 at ×20 magnification). These microscope images, originally 1024 × 768 pixels, were divided into 100 × 100 pixels patches, manually labeled based on the types of defects present and then subjected to reflections and rotations to obtain augmented datasets of adequate size for retraining and testing DeepThin (Tables S3–S5). These images were classified based on the presence or absence of cracking, dewetting, and additional defect types not considered in the darkfield dataset originally used for model development (scratches, particles, and thickness non-uniformities). After a separate retraining for each of the three microscopy datasets, DeepThin was able to accurately detect the presence or absence of the five labeled defect types (cracks, dewetting, particles, scratches, and thickness non-uniformities) wherever they appeared in the datasets for the different materials and magnifications (Fig. 4). The model could classify a given brightfield image as having no defects or any combination of one or more defects. Confusion matrices, which highlight the model’s ability to discriminate between defect-types, are also provided in Tables S6–S8. These results demonstrate that CNNs such as DeepThin may be applied to a broad scope of thin-film materials, defect morphologies, and imaging conditions.
In the top row, the accuracy for reproducing the human-labeled scores for the extent of cracking and the extent of dewetting in an image is shown. In all other rows, the accuracy for correctly classifying the images based on the presence or absence of different morphological defects is shown. Empty cells in the figure are associated with defects that were not present in the datasets or were not labeled for this study. Images in each row have the same scale. Scale bars from top to bottom are 500 µm, 50 µm, 50 µm, and 12.5 µm.
Benchmarking against concrete defect detection literature
To assess the performance of DeepThin for defect detection in a domain other than thin films, we benchmarked DeepThin against previously reported state-of-the-art algorithms for crack detection33 and segmentation34,35 in images of concrete and road surfaces. To benchmark the road-surface crack detection ability of DeepThin, we used the training and testing datasets provided by Zhang and coworkers33 to first retrain and then test DeepThin. The accuracy statistics given in Table S9, architecture comparison given in Table S10, and the receiver operating characteristic (ROC) curves in Fig. S1 show that DeepThin outperforms the three crack detection algorithms used by Zhang and coworkers33. Next, we benchmarked DeepThin against the road-surface crack segmentation algorithms described by Shi et al.34 and Fan et al.35 using the 118 image CFD dataset provided by Shi et al. We used 72 of these images for training and 46 images for testing as was done by Shi et al. and again found that DeepThin again achieves state-of-art performance (Tables S11 and S12). The state-of-the-art performance of DeepThin in these benchmarks likely arises from our use of a CNN architecture based on one of the best available (VGG16) and further optimized for crack detection. These results suggest that DeepThin may also have utility in areas of materials science other than thin films.
Discussion
We have shown that a CNN can accurately identify several different types of morphological defects in images of organic and inorganic thin films acquired under a variety of imaging conditions. The versatility of this approach to defect detection arises due to the ease with which it can be adapted to new defect types using labeled example images. The labeling of images containing examples of materials defects provides a straightforward mechanism for materials scientists to encode their domain expertise into an image analysis algorithm. As this example-based process for algorithm customization does not require software engineering expertise, we expect CNN-based approaches to material defect analysis to increase the accessibility of automated image analysis to the materials science community. Our DeepThin CNN provides the ability to rapidly and automatically identify trends in film morphology arising from manipulations of composition and process variables. We anticipate that capabilities of this kind, particularly in combination with automated experimentation, will accelerate thin film materials science research by facilitating the optimization of materials in design spaces where the morphological response to the experimental parameters is initially unknown.
Methods
Robotic platform for film deposition, annealing, and imaging
Deposition, annealing, and darkfield imaging of all the organic thin films included in the database were performed using a flexible robotic platform configured for thin-film experiments described in detail in ref. 30. Briefly, the robotic platform consists of a multi-purpose robotic arm that can handle fluids and planar glass substrates, as well as a variety of other modules that enable other tasks to be performed. The modules relevant to this study include: trays of stock solutions and mixing vials which enable the formulation of spin-coating inks with various compositions; a spin-coater for depositing inks on substrates to form thin films; an annealing station for variable-time annealing of thin films; a darkfield imaging station for imaging the thin films.
Materials
Toluene (ACS grade) was purchased from Fisher Chemical, and was used without further purification. Acetonitrile (≥99.9%), 2-propanol (≥99.5%), acetone (≥99.5%), 4-tert-Butylpyridine (96%), Spiro-MeOTAD (99%), FK 102 Co(III) TFSI salt (98%, SKU 805203-5G), and Zinc di[bis(trifluoromethylsulfonyl)imide] (Zn(TFSI)2, 95%) were purchased from Sigma Aldrich and were used without any further purification. Extran 300 Detergent was purchased from Millipore Corporation. Titanium(IV) 2-ethylhexanoate (97%) was purchased from Alfa Aesar and was used without any further purification. White glass microscope slides (3″ × 1″ × 1 mm) were purchased from VWR International. Fused silica wafers (100 mm diameter, 500 µm thickness, double-side polished) were purchased from University Wafer.
Fused silica wafers and microscope slides were cleaned prior to thin film deposition. A solution of 1% v/v Extran 300 in deionized water was prepared. The substrates were sonicated successively in the diluted Extran 300, deionized water, acetone, and 2-propanol. Before each sonication step, the substrates were rinsed in the following solvent. Substrates were stored submersed in 2-propanol. Prior to use, the substrates were dried with filtered, compressed air and inspected by eye for defects.
Organic thin film deposition
Stock solutions of spiro-OMeTAD, FK102 Co(III) TFSI salt, Zn(TFSI)2, and 4-tert-butylpyridine were prepared at 50 mg mL−1 in 1:1 v/v acetonitrile/toluene. These stock solutions were combined using the robotic platform described above to form 150 µL of ink. 100 µL of ink was deposited by the robotic platform onto a microscope substrate rotating at 1000 rpm; rotation was maintained for 60 s following ink injection. The resulting thin films were then annealed for 0 to 250 s using a custom forced air annealer (an aluminum enclosure around heat gun, Model 750 MHT Products, Inc.). All of these procedures are described in more detail in ref. 30.
Metal oxide thin film deposition
Amorphous titanium oxide films were prepared by manual spincoating. The samples were prepared by pipetting 100 µL of Titanium(IV) 2-ethylhexanoate solution (0.1 M, 2-propanol) onto cleaned fused silica wafers rotating at 3000 rpm; rotation was maintained for 30 s following ink injection. The resulting samples were irradiated with deep ultraviolet light (Atlantic Ultraviolet G18T5VH/U lamp – 5.8 W 185/254 nm, ~2 cm from the bulb, atmospheric conditions) for 15 min. After irradiation, the samples were transparent and highly refractive.
Robotic darkfield imaging
All darkfield images taken with the robot were captured with a FLIR Blackfly S USB3 (BFS-U3-120S4C-CS) camera using a Sony 12.00 MP CMOS sensor (IMX226) and an Edmund Optics 25 mm C Series Fixed Focal Length Imaging Lens (#59-871). The C-mount lens was connected to the CS-mount camera using a Thorlabs CS- to C-Mount Extension Adapter, 1.00”-32 Threaded, 5 mm Length (CML05). The sample was illuminated from the direction of the camera using an AmScope LED-64-ZK ring light. For imaging, the lens was opened to f/1.4, and black flocking paper (Thorlabs BFP1) was placed 10 cm behind the sample.
Bright-field microscopy
All brightfield images were collected using an OLYMPUS LEXT OLS 3100 microscope operating in bright-field reflection mode using ×5 and ×20 objectives.
Monotonic dewetting experiment
To collect images of an organic thin film monotonically dewetting over time, a thin film of Spiro-OMeTAD and FK102 Co(III) TFSI salt was deposited (but not annealed) using the robotic platform as described above. A camera and lightsource were positioned above the sample in the same way as they were for the robotic darkfield imaging setup. A heat gun (Model 2363333, Wagner) was positioned to heat the sample from below at a 45° degree angle so as not to obscure the black background from the camera. To perform the experiment, the heat gun was turned on high and images were acquired every second for 100 s.
Image labeling procedure to define ground truth for model development
The extent of dewetting in the dark-field images was scored by up to 3 experts on an integer scale from 0 to 9. The extent of cracking in these images was, separately, scored in the same way. In both cases, the average of the available scores was used as the ground truth. All experts used the same graphical user interface to perform the labeling. The darkfield images and the associated scores, as well as the labeling GUI can be found online (see “Data availability” section)
For the brightfield images, a vector of binary values was assigned by a single researcher to each image. Each element of the vector indicated the presence or absence of one type of defect from the following: cracks, dewetting, particles, scratches, non-uniformities. In this way, images could be labeled as having no defects, one defect (of a specified type) or more than one defect (with the types present specified). The brightfield images and the associated labels are also available online (see “Data availability” section).
Development of the DeepThin network
The DeepThin CNN architecture (Fig. 1) was developed for the thin-film image analysis tasks described here and is inspired by the VGG16 CNN architecture36. Initially, DeepThin was trained using only one convolutional layer. The model complexity was iteratively increased until the model accuracy stopped improving. We employed five-fold cross-validation37 to find a high-performance model before evaluating the model on the unseen validation data.
The input layer to DeepThin is an image with 3 RGB color channels. DeepThin has several convolutional and pooling layers as detailed in Fig. 1. The first convolutional layer uses 32 filters with a 3 × 3 × 3 kernel to convolve over the image, creating an output of size 50 × 50 × 32. Zero padding is performed so that the resulting image size is identical to the input image size. The output of the convolutional layer is passed into a ReLU activation layer. This convolutional layer is repeated, as in the VGG16 model.
Next, a maximum pooling layer of kernel size 2 × 2 is convolved over the output of the previous layer to generate a 25 × 25 × 32 output, returning the maximum value for a kernel. The two convolution layers and the pooling layer are repeated a second time. The output of the second maximum pooling layer is flattened to a 2000 × 1 vector. This output is followed by two fully connected layers of 20 neurons with ReLU activation functions and a final layer that outputs defects classes by applying a sigmoid activation function. DeepThin is trained by minimizing an error function through backpropagation using the stochastic gradient descent method. L2 (Gaussian) and Dropout regularization was used to reduce interdependent learning amongst the neurons. Regularization reduces overfitting by adding a penalty to the loss function.
DeepThin was trained using the Adam optimizer38, with an initial learning rate of 0.001 and a batch size of 100. Training loss and validation loss converged after 11 epochs.
Data availability
The labeled image datasets and a spreadsheet giving the individual expert labels for the darkfield images as well as the labeling application and the code used to develop the model are all available at https://github.com/berlinguette/ada. All other data supporting the findings of this study are available from the corresponding authors upon request.
References
Gorji, N. E. Degradation sources of CdTe thin film PV: CdCl2 residue and shunting pinholes. Appl. Phys. A 116, 1347–1352 (2014).
Feron, K., Nagle, T. J., Rozanski, L. J., Gong, B. B. & Fell, C. J. Spatially resolved photocurrent measurements of organic solar cells: Tracking water ingress at edges and pinholes. Sol. Energy Mater. Sol. Cells 109, 169–177 (2013).
Sheats, J. R. & Roitman, D. B. Failure modes in polymer-based light-emitting diodes. Synth. Met. 95, 79–85 (1998).
Ashiri, R., Nemati, A. & Sasani Ghamsari, M. Crack-free nanostructured BaTiO3 thin films prepared by sol–gel dip-coating technique. Ceram. Int. 40, 8613–8619 (2014).
Choi, S.-H. & Zhang Newby, B.-M. Suppress polystyrene thin film dewetting by modifying substrate surface with aminopropyltriethoxysilane. Surf. Sci. 600, 1391–1404 (2006).
Wang, J. Z., Zheng, Z. H., Li, H. W., Huck, W. T. S. & Sirringhaus, H. Dewetting of conducting polymer inkjet droplets on patterned surfaces. Nat. Mater. 3, 171–176 (2004).
Persson, N. E., McBride, M. A., Grover, M. A. & Reichmanis, E. Automated analysis of orientational order in images of fibrillar materials. Chem. Mater. 29, 3–14 (2017).
Costa, M. F. M. Image processing. Application to the characterization of thin films. J. Phys. Conf. Ser. 274, 012053 (2011).
Reiter, G. Unstable thin polymer films: rupture and dewetting processes. Langmuir 9, 1344–1351 (1993).
Peterhänsel, S. et al. Human color vision provides nanoscale accuracy in thin-film thickness characterization. Optica 2, 627–630 (2015).
Eperon, G. E., Burlakov, V. M., Docampo, P., Goriely, A. & Snaith, H. J. Morphological control for high performance, solution-processed planar heterojunction perovskite solar cells. Adv. Funct. Mater. 24, 151–157 (2014).
Barrows, A. T. et al. Monitoring the Formation of a CH3NH3PbI3-xClx Perovskite during thermal annealing using X-ray scattering. Adv. Funct. Mater. 26, 4934–4942 (2016).
Wieghold, S. et al. Detection of sub-500-μm cracks in multicrystalline silicon wafer using edge-illuminated dark-field imaging to enable thin solar cell manufacturing. Sol. Energy Mater. Sol. Cells 196, 70–77 (2019).
O’Mahony, N. et al. Deep learning vs. traditional computer vision. Advances in Intelligent Systems and Computing, pp128–144, https://doi.org/10.1007/978-3-030-17795-9_10 (2020).
Qi, C. R. et al. Volumetric and multi-view CNNs for object classification on 3D data. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 5648–5656 (2016).
Girshick, R., Donahue, J., Darrell, T. & Malik, J. Region-based convolutional networks for accurate object detection and segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 38, 142–158 (2016).
Jaritz, M., Charette, R. D., Wirbel, E., Perrotton, X. & Nashashibi, F. Sparse and dense data with CNNs: depth completion and semantic segmentation. In 2018 International Conference on 3D Vision (3DV) 52–60 (2018).
Choutas, V., Weinzaepfel, P., Revaud, J. & Schmid, C. Potion: pose motion representation for action recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp 7024–7033 (2018).
Rastegari, M., Ordonez, V. & Redmon, J. Xnor-net: Imagenet classification using binary convolutional neural networks. In European conference on computer vision, 525–542 (2016).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition, pp 770–778 (2016).
Gallagher, B. et al. Predicting compressive strength of consolidated molecular solids using computer vision and deep learning. Mater. Des. 190, 108541 (2020).
Groschner, C. K., Choi, C. & Scott, M. C. Methodologies for successful segmentation of HRTEM Images via neural network. Preprint at https://arxiv.org/abs/2001.05022 (2020).
Matson, T., Farfel, M., Levin, N., Holm, E. & Wang, C. Machine learning and computer vision for the classification of carbon nanotube and nanofiber structures from transmission electron microscopy data. Microsc. Microanal. 25, 198–199 (2019).
Ziatdinov, M. et al. Deep learning of atomically resolved scanning transmission electron microscopy images: chemical identification and tracking local transformations. ACS Nano 11, 12742–12752 (2017).
Maksov, A. et al. Deep learning analysis of defect and phase evolution during electron beam-induced transformations in WS2. npj Comput. Mater. 5, 12 (2019).
Madsen, J. et al. A deep learning approach to identify local structures in atomic-resolution transmission electron microscopy images. Adv. Theory Simul. 1, 1800037 (2018).
Samide, A., Stoean, C. & Stoean, R. Surface study of inhibitor films formed by polyvinyl alcohol and silver nanoparticles on stainless steel in hydrochloric acid solution using convolutional neural networks. Appl. Surf. Sci. 475, 1–5 (2019).
Samide, A. et al. Investigation of polymer coatings formed by polyvinyl alcohol and silver nanoparticles on copper surface in acid medium by means of deep convolutional neural networks. Coat. World 9, 105 (2019).
Saito, Y. et al. Deep-learning-based quality filtering of mechanically exfoliated 2D crystals. npj Comput. Mater. 5, 124 (2019).
MacLeod, B. P. et al. Self-driving laboratory for accelerated discovery of thin-film materials. Sci. Adv. 6, eaaz8867 (2020).
Gu, J. et al. Recent advances in convolutional neural networks. Pattern Recognit. 77, 354–377 (2018).
Montavon, G., Samek, W. & Müller, K.-R. Methods for interpreting and understanding deep neural networks. Digit. Signal Process. 73, 1–15 (2018).
Zhang, L., Yang, F., Zhang, Y. D. & Zhu, Y. J. Road crack detection using deep convolutional neural network. 2016 IEEE International Conference on Image Processing (ICIP), https://doi.org/10.1109/icip.2016.7533052 (2016).
Shi, Y., Cui, L., Qi, Z., Meng, F. & Chen, Z. AutomAtic Road Crack Detection Using Random Structured Forests. IEEE Trans. Intell. Transp. Syst. 17, 3434–3445 (2016).
Fan, Z., Wu, Y., Lu, J. & Li, W. Automatic pavement crack detection based on structured prediction with the convolutional neural network. Preprint at https://arxiv.org/abs/1802.02208 (2018).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. Preprint at https://arxiv.org/abs/1409.1556 (2014).
Stone, M. Cross-validatory choice and assessment of statistical predictions. J. R. Stat. Soc.: Ser. B (Methodol.) 36, 111–133 (1974).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Acknowledgements
Brightfield microscopy was performed in the Centre for High-Throughput Phenogenomics at the University of British Columbia, a facility supported by the Canada Foundation for Innovation, British Columbia Knowledge Development Foundation, and the UBC Faculty of Dentistry. We thank Pierre Chapuis for exploratory work on image analysis and Gordon Ng for assistance with the dewetting experiment. For hardware and software contributions to the robotic platform, we acknowledge Michael Elliott, Ted Haley, Karry Ocean, Alex Proskurin, Michael Rooney, and Henry Situ. The authors thank Natural Resources Canada (EIP2-MAT-001) for their financial support. C.P.B. is grateful to the Canadian Natural Sciences and Engineering Research Council (RGPIN 337345-13), Canadian Foundation for Innovation (229288), Canadian Institute for Advanced Research (BSE-BERL-162173), and Canada Research Chairs for financial support. B.P.M, F.G.L.P., T.D.M., and C.P.B. acknowledge support from the SBQMI’s Quantum Electronic Science and Technology Initiative, the Canada First Research Excellence Fund, and the Quantum Materials and Future Technologies Program. This research was enabled in part by support provided by WestGrid (www.westgrid.ca) and Compute Canada Calcul Canada (www.computecanada.ca).
Author information
Authors and Affiliations
Contributions
C.P.B. supervised the project. N.T. developed the CNN and is the guarantor. B.P.M., F.G.L.P., K.E.D., and T.D.M. used the robotic platform to create the 2600-image initial training set and to perform the doping and annealing experiments. T.D.M., K.E.D., and F.G.L.P. labeled the datasets. F.G.L.P. performed the dewetting experiment. B.P.M. performed the microscopy. F.G.L.P. created the figures with input from N.T. and B.P.M., E.P.B., B.P.M., and N.T. carried out the literature review. All authors contributed to the writing of the paper. N.T. and B.P.M. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Taherimakhsousi, N., MacLeod, B.P., Parlane, F.G.L. et al. Quantifying defects in thin films using machine vision. npj Comput Mater 6, 111 (2020). https://doi.org/10.1038/s41524-020-00380-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-020-00380-w
This article is cited by
-
Enhanced accuracy through machine learning-based simultaneous evaluation: a case study of RBS analysis of multinary materials
Scientific Reports (2024)
-
Flexible automation accelerates materials discovery
Nature Materials (2022)
-
A machine vision tool for facilitating the optimization of large-area perovskite photovoltaics
npj Computational Materials (2021)