Abstract
Genome engineering for materials synthesis is a promising avenue for manufacturing materials with unique properties under ambient conditions. Biomineralization in diatoms, unicellular algae that use silica to construct micron-scale cell walls with nanoscale features, is an attractive candidate for functional synthesis of materials for applications including photonics, sensing, filtration, and drug delivery. Therefore, controllably modifying diatom structure through targeted genetic modifications for these applications is a very promising field. In this work, we used gene knockdown in Thalassiosira pseudonana diatoms to create modified strains with changes to structural morphology and linked genotype to phenotype using supervised machine learning. An artificial neural network (NN) was developed to distinguish wild and modified diatoms based on the SEM images of frustules exhibiting phenotypic changes caused by a specific protein (Thaps3_21880), resulting in 94% detection accuracy. Class activation maps visualized physical changes that allowed the NNs to separate diatom strains, subsequently establishing a specific gene that controls pores. A further NN was created to batch process image data, automatically recognize pores, and extract pore-related parameters. Class interrelationship of the extracted paraments was visualized using a multivariate data visualization tool, called CrossVis, and allowed to directly link changes in morphological diatom phenotype of pore size and distribution with changes in the genotype.
Similar content being viewed by others
Introduction
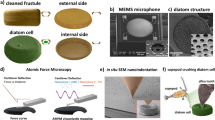
Living organisms can construct complex three-dimensional structures from inorganic materials at ambient temperature, pressure, and pH in aqueous conditions, yet the complexity and precision of this synthesis rival modern industrial methods. Understanding and exploiting biomineral formation processes will enable the design of next-generation materials for photonics,1,2 sensing,3,4,5 filtration, and drug delivery.6 One of the best-studied examples of biomineralization is in diatoms, unicellular algae that use silica to construct micron-scale cell walls with nanoscale features, yielding a multiscalar structure ranging over eight orders of magnitude.7 A model diatom, Thalassiosira pseudonana, like all diatoms, has a cylindrical cell wall—a frustule made from silica. The top and bottom of the cylinder, termed valves, are connected by a series of circular girdle bands. Within each valve there is an array of ribs, pores, and rimoportulae (Fig. 1). Altogether the diatom morphology is a complex, functional three-dimensional agglomeration of nanosacle to microscale structures exceeding the complexity possible with current synthetic approaches.8,9
SEM analysis of wild type T. pseudonana diatom. a A 45° view of an individual diatom. Published under a Creative Commons Attribution 4.0 International License from ref. 47 Scale bar is 1 µm. b Top view of a diatom valve (scale bar is 1 µm) and (c) valve morphology (scale bar is 300 nm). b, c Two different examples of a diatom valve
The pores in diatom valves are of particular importance for both diatom survival in the environment, as well as potential industrial applications. The porous architecture of their exoskeleton enables attractive optical properties, such as light harvesting, confinement, and selective optical transmission.10,11 Moreover, despite high porosity, diatom silica exhibits remarkably high mechanical stability, which is important for filtration applications.12,13,14 Controllably changing the pores thus empowers an array of properties, and in turn applications of diatom systems for a wide variety of tasks.
Diatom frustules, including the pores, are assembled through the concerted action of dozens of proteins encoded in the diatom genome. These proteins modulate silica precipitation and sintering to generate specific structures. New techniques in multi-omic characterization have identified many genes that may be involved in biomineralization, based on coordinated expression during cell growth and division.15,16 Validation of these predictions, however, is challenging, requiring disrupting expression of each putative frustule formation gene followed by identification of any resulting changes in morphology. While in certain cases changes to frustules may be obvious,17 often structure variation requires more rigorous analysis. Specifically, the recognition of persistent (but a priori unknown) features is a significant challenge. Further complicating this is the fact that isogenic diatom populations have substantial natural variation in morphological properties,18,19 which will render the task more difficult with regards to generalization. Therefore, to controllably modify diatom frustule structure through targeted genetic modifications, a robust genotype to phenotype linking methodology needs to be developed.
One potential method that has been pivotal of late is the use of artificial neural networks (NNs) that can learn abstract features from large datasets, negating the need for hand-crafted features. These deep NNs have been instrumental in the recent dramatic improvement of automatic image classification and speech recognition, and have also been applied to scientific domains, such as fitting potential energy surfaces,20,21,22,23 fundamental understating of phase transitions,24,25,26 processing of the atomically resolved images27,28 as well as fitting multiparametric empirical models.29,30,31,32 Classical image processing approaches work, but data generation rates are replacing them with more automated unsupervised and supervised learning methods.33 Artificial NNs, however, excel at classification problems and can be very efficient. NNs employ a “learning by example” approach that optimizes parameters by training on labeled data. Hence, if there are systematic differences between classes, and the network has presented those cases in the training sets, a NN can detect and label features in real/validation data. Given that diatoms exhibit a diverse set of frustule morphologies, a NN is a reasonable choice of a classification scheme to detect the genotype to phenotype translation.
In this work, we investigated wild type and genetically modified T. pseudonana to capture the interplay between the changing genotype and the expressed phenotype, as gene manipulation34 could enable these organisms to be used as a direct source of specifically tailored nanostructured and microstructured materials.35 We modified the genotype by knocking down genes we suspected to be involved with frustule formation and characterized the phenotype by scanning electron microscopy (SEM). We used image processing33,36 and machine learning classification algorithms (artificial NNs)37,38 to screen for genes that affect diatom phenotype and to distinguish diatoms with wild type and modified morphologies. With regard to inspected Thaps3_21880 modification, we demonstrated a NN that can identify wild and modified diatoms with 94% accuracy. To explain the apparent efficiency of NN-based classification, class activation maps (CAMs) were used to highlight the image regions used by the network.39 It was found that pores are the defining features separating wild-type diatoms from one specific knockdown strain. We then created a separate neural net to focus specifically on pores and to extract their parameters. This automated feature extraction process allowed us to correlate the genetic modification with diatom morphology. Our approach identifies the changes in valve structure that result from a given genetic modification, offering biological insight into the biomineralization process.
Results and discussion
Gene modification and testing
We identified protein Thaps3_21880 as a potential frustule biosynthetic enzyme based on structural features and coordinated expression during silicization.16 To perturb gene expression in T. pseudonana, we synthesized an antisense RNA containing the first 427 nucleotides of the associated gene and expressed this construct from a heterologous plasmid. Two independent clonal knockdown lines were selected for nourseothricin resistance, and then confirmed by polymerase chain reaction (PCR) (Supplementary materials Fig. S1 and Methods section for details).40
Identification of morphological changes
A successful gene knockdown does not necessarily result in any morphological change in the diatom frustule and, furthermore, in those cases where the morphology is altered, the precise change is difficult to predict. Therefore, a robust method is needed for determining whether a population of diatoms has a variant phenotype and, if so, the details of the variation. SEM was used to visualize the surface morphology of the modified and nonmodified samples, and a typical wild-type diatom is depicted in Fig. 1b, with the details of the structure highlighted in Fig. 1c (see Supplementary materials, Figs. S3 and S4 for more examples of wild diatoms and the example images of modified diatoms). The images reveal a complex feature-rich structure making it difficult for the untrained eye to separate the wild and the modified diatoms, let alone quantify physical changes. Therefore, we used image processing and machine learning strategies to automatically classify wild and modified phenotypes directly from images. Classical image processing techniques proved to be cumbersome in trying to process multiple images since subtle contrast variation required significant manual parameter adjustment and precluded automation. We, therefore, decided to use a NN. Labeled training sets of wild (labeled as zero) and modified (labeled as one) diatom images were prepared in the following way. Each image was spliced into subsets of 89 × 89-pixel frames. Each frame underwent a series of geometric transformations, such as horizontal and vertical mirroring, rotation, and change in contrast to expand the training set. The NN architecture is shown in Fig. 2a (see Methods section for details). The validation test for this NN showed 94% class identification accuracy. We also justified the NN by only training the net on images of wild diatoms and validated on the separate set containing wild diatoms. As a result, the accuracy was ~50%, which was expected.
a Schematic of an artificial NN to distinguish between wild and modified diatoms (NN 1); scale bar is 300 nm; b–e Class activation maps explaining operation of NN 1. b, d Regions of the diatom images that trigger wild activation class (AC); c, e Show regions of corresponding images that trigger modified AC. b–e Scale bar is 50 nm
This NN allowed us to automatically screen for changes in images between the modified and wild genotypes, and indirectly screen genes that modify the structure of diatoms. However, NNs are often thought of as a “black box” approach to solving a problem—it is difficult to ascertain what features drive the NNs to make decisions. To gain insights into what image aspects made a significant contribution to classification, we modified this NN, as described in Methods section, in order to peer into the layers using CAMs. Figure 2b, c shows the same frame of the diatom image, which corresponds to the modified class and was identified by the NN. Figure 2b highlights the regions that trigger wild activation class (AC), while Fig. 2c emphasizes the regions that, for the NN, indicate modified AC. As shown by the colorbar, a whiter color corresponds to higher AC intensity. Figure 2d, e also displays an identical frame of another diatom image, corresponding to the wild class. CAMs revealed that the NN focuses on pores as one of the main features to distinguish wild and modified diatoms (for additional examples of CAMs for NN, please, see Fig. S2 in Supplementary Materials). In this way, we identified a specific gene connected to the changes in pores of diatoms.
Automatic recognition of relevant features and extraction of their parameters
Identification of pores emphasizes the importance of pore-related parameters. Thus, a new NN 2 was generated and trained to focus on these image characteristics. First, pores were recognized in several raw images of diatoms. As a result, matching pairs of the original images and corresponding binary images containing only pores were generated and used as a training set for NN 2. Similar to previous NN, training images were cut into 46 × 46-pixel frames, which underwent identical geometric and contrast treatment to enlarge the training set. It was important to perform this image segmentation before loading the frames into the neural net, as the contrast variation within a given image often precluded the use of typical filtering approaches. Each frame was processed by the NN 2 (the architecture is shown in Fig. 3a; see Methods section for details), built in a way that its output would be a pixel-wise classification map corresponding to the input image.
a Schematic of an artificial NN to recognize pores of the diatoms (NN 2). b Frame of an original diatom image that is loaded into the NN 2. c Pixel-wise classification map of the NN 2 corresponding to raw image (b). d Result of the application of the NN 2 to the image of a diatom (red circles are recognized pores); scale bar is 300 nm
Figure 3b, c shows pixel-matching images, where Fig. 3b is an original frame of a diatom image that is loaded into the NN 2, and Fig. 3c is an output image of the NN 2. Obtained NN-processed image matches with the raw image and closely resolves the presented pores. Scanning across each image, the pores can be determined, as shown in Fig. 3d, where the raw image is overlapped with the contours of NN-identified pores (red circles). Since the CAMs pointed to pores as the features NN 1 used to separate wild and modified diatoms, we wanted to provide a quantitative analysis of multiple pore-related parameters to capture this change. Using NN-processed images, we extracted the following parameters: density of pores, mean area of pores, and the percentage of area occupied by the pores relative to the total area of the valve captured in an image. Additionally, pore area distribution was extracted and fitted with a Gaussian distribution, which yielded two more parameters, Gaussian value µ and Gaussian sigma ς, producing a total of five parameters. To minimize the influence of outliers (such as small features that NN could recognize as pores), threshold for pore size was set to be larger than three pixels.
The values of four of these parameters were plotted against the values of the fifth parameter (mean area of pores) to test if different regions of the parameter space produce distinguishable types of behavior (Fig. 4). Figure 4a reveals that the best separation between wild and modified diatoms can be established through density of pores against the mean area of pores. Figure 4b, c shows that the separation of these two classes of diatoms can also be shown through the plots of expected Gaussian value and Gaussian sigma as a function of mean area, respectively. Finally, no significant difference between classes is observed in relation to the ratio of pore area to total area parameter (Fig. 4d).
It is also helpful to plot the extracted parameters together to visualize parameter intercorrelation. We explored class interrelationship using a multivariate data visualization tool called CrossVis. CrossVis represents an evolution of both the EDEN and MDX visual analysis systems41,42 (see Methods section for details). For clarity, the expected Gaussian value µ was removed since its values are identical to the mean area of pores (Fig. 4b). Parameter values are graphed as polylines in a parallel coordinate plot,43 and each vertical axis represents one of the pore-related parameters, as shown in Fig. 5. Parameters of modified diatoms are colored orange, and parameters of wild diatoms are blue. Vertical columns are arranged so the parameters showing better separation between classes are on the left-hand side of the Type column, and the parameters with lesser separation are on the right-hand side. Certain statistics are graphically represented in the interior boxes of each column. Namely, thin blue and orange stripes within each column capture the mean-centered standard deviation of a given parameter of wild and modified diatoms, respectively. Similar to Fig. 4, this statistical display shows that there is little separation between classes based on the ratio of the pore area to total area and Gaussian sigma ς, but diatoms can be clearly classified based on the mean area and density of pores. The distribution of each parameter is also displayed along the vertical axis as shaded histogram bins (light grey is for modified diatoms; dark gray is for wild diatoms).
Class interrelationship visualization using CrossVis v.2.1.2. Parameters of modified diatoms are colored orange, and parameters of wild diatoms have a blue color. Histogram bins show the distribution of each parameter, and they have a light grey color for modified diatoms, and a dark gray color for wild diatoms
Figure 5 also facilitates a visual correlation between presented parameters shown with blue and orange lines connecting neighboring vertical axis. Namely, each line connects values of the parameters extracted from the same diatom image. On the one hand, some parameters, such as Gaussian sigma and the ratio of pore area to total area, do not seem to show any correlation between each other. On the other hand, looking at the links between density of pores and mean area of pores, it is observed that these two parameters are negatively correlated within each diatom class (higher density corresponds to smaller mean area and vice versa), and the links have opposite directions for wild and modified diatoms. This behavior suggests that higher density of pores in wild diatoms is compensated by the smaller area of these pores, which is opposite to the trend shown by modified diatoms. In turn, this trend explains why the ratio of pore area to total area is very similar for both classes.
This pore-based multifactor analysis confirmed that, while there is an overlap between parameters of wild and modified diatoms, the proposed and investigated algorithm allowed for their separation and established a correlation between realized gene modification and alteration of frustule morphology. Moreover, it was shown that the performed gene alteration mainly affects such pore parameters as their area and density, increasing the former and decreasing the latter, while keeping the ratio of pore area to total area of frustule constant.
In this work, we have illustrated how machine learning can aid in biomaterial research using diatoms. After gene knockdown in T. pseudonana diatoms, supervised machine learning was successfully applied to screen for genes that altered the structure of the diatoms and separate those with modified morphology. This yielded a process where morphological changes were connected to a specific gene alteration. In the case of Thaps3_21880 modification, one artificial NN was developed to distinguish wild and modified diatoms based on the SEM images of frustules, resulting in 94% accuracy. Then CAMs visualized physical changes that allowed the NNs to separate diatom strains, subsequently establishing a specific gene that controls pores. Another artificial NN was created to process image data, automatically recognize pores, and extract pore-related parameters. In turn, multifactor analysis and visualization of these parameters emphasized the alteration in density and area of pores. An important advantage of the presented approach is that it allows for automated screening of the genes that modify diatom morphology, and it enables recognition and analysis of features, such as pores, whose changes might be indistinguishable with the naked eye.
Methods
Culture and growth conditions
T. pseudonana (CCMP1335) stock cultures were maintained in NEPC (North East Pacific Culture) medium (http://www3.botany.ubc.ca/cccm/NEPCC/esaw.html). The cultures were grown in a temperature controlled incubator at 16 °C under a 14 h:10 h (light:dark) photoperiod at 100 μmol photon m−2 s−1 of photosynthetically active radiation.
Genetic manipulations
An antisense construct containing the reverse complement of the first 427 nucleotides of Thaps3_21880 was synthesized de novo and cloned into pMHL00940 immediately following the NAT coding sequence (GenScript, Piscataway, NJ). This construct was transformed into T. pseudonana by microparticle bombardment using a Bio-Rad Biolistic PDS-1000/He particle delivery system, as described previously.40 Briefly, exponentially grown cells were harvested and 1 × 108 cells were plated onto an NEPC agar plate lacking antibiotics. A nuclear transformation was performed by bombarding plasmid coated tungsten beads (1.1 μm diameter, Bio-Rad M-17) at 1100 lb/in2 under vacuum at a distance of 8 cm onto cells plated on the agar plate. After bombardment, NEPC medium was added to plates, which wasincubated under a 14 h:10 h (light:dark period) for 24 h. After 24 h, the plasmid bombarded diatom cells were plated on NEPC bacto-agar plate with 100 µL/mL of the antibiotic nourseothricin. The resistant colonies were transferred into 24-well plates with 2 ml of NEPC medium. To confirm construct integration, cells from well plates were screened by PCR using primers NAT_SeqF (5′-AAGGTGTTCCCCGACGACGAATC-3′) and 21880_SeqR (5′-TATGAGCATGTCTTTGCCACTCAGAC-3′), as described previously.40
Sample preparation for SEM imaging
One milliliter of diatom culture (2 × 106 cells/ml) was collected by centrifugation at 5000 rpm for 5 min and rinsed once with deionized water. The diatom frustules were cleaned with 1 mL concentrated sulfuric acid and boiled in a water bath for 10 min. After cooling, we added 20 mg KNO3, then boiled again for 10 min. Samples were then washed three times with deionized water using centrifugation. The diatom frustules were spotted on silicon wafer and air dried in hood for approximately 10 min.44 While these harsh cleaning methods may affect the fine structures of diatom pores, they were necessary to remove residual organic matter and allow clear imaging.
SEM imaging of diatoms
Dried diatoms on Si wafers were imaged with a Merlin Field Emission SEM (Carl Zeiss, Oberkochen, Germany) operating at a base pressure of 2.8 × 10−6 mbar. Images were acquired using an in-lens detector with an accelerating voltage of 1 kV, probe current of 100 pA, cycle time of 14.4 s, 10 line averages, and resolution of 1024 × 768. Images were collected of diatom valves with a 2–3 µm field of view, depending on the individual size of the diatom, to capture the arrays of ribs and pores. Diatoms in various orientations were observed with SEM and only those with valves that were enact, unwrinkled, and approximately parallel to the Si wafer were included in this study, such as in Fig. 1b, c.
Computational methods
A series of images of wild and modified diatoms, that were used as training sets, were analyzed using Python 3.6 libraries, such as numpy, scipy, matplotlib, opencv, and scikit-image.45 Conversion from pixels to nanometers was based on image scale bar, which was programmed to be automatically recognized by the Python code. Both artificial NNs were created using Keras library46 with a TensorFlow backend in Python 3.6.
Training dataset for NN 1
We have selected 29 images of wild diatoms and 29 images of modified ones. A random selection would first determine the type of the particular image to be added into training set, then the specific image of that type would be also randomly drawn. This ensures that the presence of both types of diatoms is equal. Otherwise, the classifier becomes biased to output the type numerically dominant in the dataset. Image augmentation was done in the following manner. All the images were rescaled using resize function from opencv to have the same pixel/nm value of 3. A 250 × 250 px cutaways were randomly selected from a randomly selected image. Vertical and horizontal mirroring were applied at random. The rotation of the cutaway was done using opencv functions getRotationMatrix2D and warpAffine. We have applied non-linear adjustment of contrast for the training set to incorporate possible artifacts from SEM images. Two cases with equal probabilities were created: e(a*I) and (1 − e(−a*I)) where a is a random value between 0 and 1 and I is the intensity in the pixel. The resulting arrays were normalized to [0, 1]. We have intentionally avoided rescaling as we expected that size of the pores may be characteristic to the diatom type, while SEM contrast, rotation, and mirroring were expected to be varying within the type.
Artificial NN to distinguish between wild and modified diatoms (NN 1)
NN 1 architecture, shown in Fig. 2a, consists of two convolutional layers with a max pooling layer in between, and then a dense (fully connected) layer. The max pooling layer was added to increase the “connectedness” of the convolutional filters. Rectified linear units (ReLUs) were used as activation functions in these layers. The last layer consists of a two-unit dense layer with softmax activation, so that the classification outputs would sum to one, respecting the choices available. It is this layer that provides the estimate of which class the input image belongs. Optimization of the network was performed with the Adam optimizer utilizing the cross-entropy metric, and NN was trained on a randomly selected subset of images, where 80% were a training set, and 20% were a validation set. The validation test, which predicts the class of the labeled images, showed that NN can determine the class of the diatom with 94% accuracy.
The performance of created neural net (NN 1) was additionally validated to confirm that the NN differentiates between wild and modified diatoms based on actual patterns in the morphology and not based on noise or artifacts. We took only images of wild diatoms, randomly split them into two training sets with different labels, and retrained NN 1. Because both sets consist only of images of wild diatoms, NN should not be able to learn from these training sets and separate them if it works properly. As a result, accuracy of the training was ca. 50%, which was expected and validated the performance of NN 1.
Modification of NN 1 to obtain CAMs
CAMs allowed determination of the image regions used by NN 1 to distinguish between wild type and modified diatoms. However, in order to apply the concept of CAMs, next-to-last layer of NN 1 had to be changed from dense layer to global average pooling layer. This modification did not affect the accuracy of NN 1, but allowed to peer within the layers of the NN and extract CAMs.
Artificial NN to recognize pores of the diatoms (NN 2)
NN 2 architecture, shown in Fig. 3a, consists of a convolutional layer with a max pooling layer, followed by two convolutional layers, and an up-sampling layer. An up-sampling layer was added to bring the output image to a higher resolution that would match the resolution of the input image. ReLUs were used as activation functions in these layers. The raw images were normalized to [0, 1], and the thresholding value of 0.5 was used to generate binary images for the NN 2. The last layer consisted of a convolutional layer with only one filter and a sigmoid activation, so that its output would be a pixel-wise classification map corresponding to the input image.
Data visualization
Class interrelationship between diatom parameters was inspected using a multivariate data visualization tool called CrossVis. The CrossVis system is a visual analytics system, which combines interactive data visualization and statistical analytics techniques. The system represents an evolution of both the EDEN41 and MDX42 systems. CrossVis includes a number of enhancements to the popular parallel coordinate information visualization technique.43 As shown in Fig. 6, the parallel coordinates plot yields a two-dimensional plot of multivariate data by representing each N-dimensional tuple with points on N parallel axes, which are joined with a polyline. CrossVis extends the basic parallel coordinates plot with human directed interactions and graphical representations of summary statistics. Users can select parameter ranges of interest by using the mouse to drag a rectangular region on the vertical axes. As shown in Fig. 5, within the interior of each vertical axis, the mean and mean-centered standard deviation range (95% confidence interval) are shown, where the rectangular box is the range and the horizontal line intersecting the box is the mean value. Correlations between parameters (vertical axes) are apparent when two axes are side-by-side. If the polylines cross to form an X-shaped pattern, the two parameters are negatively correlated—as one parameter increases in value the other decreases (see the polyline crossings for the two leftmost axes in Fig. 5). If the polylines have few crossings, the two parameters are positively correlated—as one parameter increases in value, the other also increases. CrossVis also allows categorical parameters (see the middle Type axis in Fig. 5) to be represented, which is an extension of the standard parallel coordinate plot. The display of categorical parameters enables study of the wild vs. modified classes in the current work. Users can select a class by clicking on the appropriate box for the categorical axis. For more details on the parallel coordinate-based techniques, the reader is urged to consult the related references.41,42,43
Data availability
All data used in this manuscript are available from the authors on request.
Code availability
Full code is available from the authors on request.
Change history
13 August 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
De Tommasi, E. et al. UV-shielding and wavelength conversion by centric diatom nanopatterned frustules. Sci. Rep. 8, 16285 (2018).
Jeffryes, C., Solanki, R., Rangineni, Y., Wang, W., Chang, C.-H. & Rorrer, G. L. Electroluminescence and photoluminescence from nanostructured diatom frustules containing metabolically inserted germanium. Adv. Mater. 20, 2633–2637 (2008).
Bismuto, A., Setaro, A., Maddalena, P., De Stefano, L. & De Stefano, M. Marine diatoms as optical chemical sensors: a time-resolved study. Sens. Actuat B-Chem 130, 396–399 (2008).
Gale, D. K., Gutu, T., Jiao, J., Chang, C.-H. & Rorrer, G. L. Photoluminescence detection of biomolecules by antibody-functionalized diatom biosilica. Adv. Funct. Mater. 19, 926–933 (2009).
Selvaraj, V., Muthukumar, A., Nagamony, P. & Chinnuswamy, V. Detection of typhoid fever by diatom-based optical biosensor. Environ. Sci. Pollut. R 25, 20385–20390 (2018).
Delalat, B. et al. Targeted drug delivery using genetically engineered diatom biosilica. Nat. Commun. 6, 1–11 (2015).
Ghobara, M. et al. On Light and diatoms: a photonics and photobiology review. In: Diatoms: Fundamentals and Applications (eds Seckbach, J. & Gordon, R.) (Wiley-Scrivener, 2019).
Hildebrand, M., Holton, G., Joy, D. C., Doktycz, M. J. & Allison, D. P. Diverse and conserved nano- and mesoscale structures of diatom silica revealed by atomic force microscopy. J. Microsc. 235, 172–187 (2009).
Hildebrand, M. et al. Nanoscale control of silica morphology and three-dimensional structure during diatom cell wall formation. J. Mater. Res. 21, 2689–2698 (2006).
De Tommasi, E., Gielis, J. & Rogato, A. Diatom frustule morphogenesis and function: a multidisciplinary survey. Mar. Genom. 35, 1–18 (2017).
Pawolski, D., Heintze, C., Mey, I., Steinem, C. & Kröger, N. Reconstituting the formation of hierarchically porous silica patterns using diatom biomolecules. J. Struct. Biol. 204, 64–74 (2018).
Aitken, Z. H., Luo, S., Reynolds, S. N., Thaulow, C. & Greer, J. R. Microstructure provides insights into evolutionary design and resilience of Coscinodiscus sp. frustule. Proc. Natl Acad. Sci. USA 113, 2017–2022 (2016).
Hamm, C. E. et al. Architecture and material properties of diatom shells provide effective mechanical protection. Nature 421, 841–843 (2003).
Leon, S. D. & Markus, J. B. Influence of geometry on mechanical properties of bio-inspired silica-based hierarchical materials. Bioinspir. Biomim. 7, 036024 (2012).
Hildebrand, M. & Lerch, S. J. L. Diatom silica biomineralization: parallel development of approaches and understanding. Semin. Cell Dev. Biol. 46, 27–35 (2015).
Shrestha, R. P. et al. Whole transcriptome analysis of the silicon response of the diatom Thalassiosira pseudonana. BMC Genomics 13, 499–515 (2012).
Tesson, B., Lerch, S. J. L. & Hildebrand, M. Characterization of a new protein family associated with the silica deposition vesicle membrane enables genetic manipulation of diatom silica. Sci. Rep. 7, 13457 (2017).
Amato, A., Orsini, L., D’Alelio, D. & Montresor, M. Life cycle, size reduction patterns, and ultrastructure of the pennate planktonic diatom Pseudo-nitzschia delicatissima (Bacillariophyceae). J. Phycol. 41, 542–556 (2005).
Hense, I. & Beckmann, A. A theoretical investigation of the diatom cell size reduction–restitution cycle. Ecol. Model. 317, 66–82 (2015).
Dawes, R., Thompson, D. L., Guo, Y., Wagner, A. F. & Minkoff, M. Interpolating moving least-squares methods for fitting potential energy surfaces: computing high-density potential energy surface data from low-density ab initio data points. J. Chem. Phys. 126, 184108 (2007).
Handley, C. M. & Popelier, P. L. Potential energy surfaces fitted by artificial neural networks. J. Phys. Chem. A 114, 3371–3383 (2010).
Manzhos, S., Wang, X., Dawes, R. & Carrington, T. Jr. A nested molecule-independent neural network approach for high-quality potential fits. J. Phys. Chem. A 110, 5295–5304 (2006).
Blank, T. B., Brown, S. D., Calhoun, A. W. & Doren, D. J. Neural-network models of potential-energy surfaces. J. Chem. Phys. 103, 4129–4137 (1995).
Carrasquilla, J. & Melko, R. G. Machine learning phases of matter. Nat. Phys. 13, 431 (2017).
van Nieuwenburg Evert, P. L., Liu, Y.-H. & Huber Sebastian, D. Learning phase transitions by confusion. Nat. Phys. 13, 435–439 (2017).
Broecker, P., Carrasquilla, J., Melko, R. G. & Trebst, S. Machine learning quantum phases of matter beyond the fermion sign problem. Sci. Rep. 7, 8823 (2017).
Ziatdinov, M. et al. Deep learning of atomically resolved scanning transmission electron microscopy images: chemical identification and tracking local transformations. ACS Nano 11, 12742–12752 (2017).
Ziatdinov, M., Maksov, A. & Kalinin, S. V. Learning surface molecular structures via machine vision. npj Comput. Mater. 3, 31 (2017).
Yin, F., Mao, H. J., Hua, L., Guo, W. & Shu, M. S. Back propagation neural network modeling for warpage prediction and optimization of plastic products during injection molding. Mater. Des. 32, 1844–1850 (2011).
Yun, S. Y., Namkoong, S., Rho, J. H., Shin, S. W. & Choi, J. U. A performance evaluation of neural network models in traffic volume forecasting. Math. Comput. Model. 27, 293–310 (1998).
Gupta, V. K. et al. Prediction of capillary gas chromatographic retention times of fatty acid methyl esters in human blood using MLR, PLS and back-propagation artificial neural networks. Talanta 83, 1014–1022 (2011).
Lee, W. Y., Park, G. G., Yang, T. H., Yoon, Y. G. & Kim, C. S. Empirical modeling of polymer electrolyte membrane fuel cell performance using artificial neural networks. Int. J. Hydrog. Energ. 29, 961–966 (2004).
Belianinov, A. et al. Big data and deep data in scanning and electron microscopies: deriving functionality from multidimensional data sets. Adv. Struct. Chem. Imaging 1, 1–25 (2015).
Hildebrand, M. Prospects of manipulating diatom silica nanostructure. J. Nanosci. Nanotechnol. 5, 146–157 (2005).
Sandhage, K. H. et al. Merging biological self-assembly with synthetic chemical tailoring: the potential for 3-D genetically engineered micro/nano-devices (3-D GEMS). Int. J. Appl. Ceram. Technol. 2, 317–326 (2005).
Burch, M. J. et al. Helium ion microscopy for imaging and quantifying porosity at the nanoscale. Anal. Chem. 90, 1370–1375 (2017).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. In Proc. International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Navab, N., Hornegger, J., Wells, W. M. & Frangi, A. F.). Vol. 9351, (Springer, Cham, 2015).
Hopke, P. K. & Song, X.-H. Classification of single particles by neural networks based on the computer-controlled scanning electron microscopy data. Anal. Chim. Acta 348, 375–388 (1997).
Zhou, B., Khosla, A., Lapedriza, A., Oliva, A. & Torralba, A. Learning deep features for discriminative localization. In Proc. CVPR 2921–2929 (2016).
Shrestha, R. P. & Hildebrand, M. Evidence for a regulatory role of idatom silicon transporters in cellular silicon responses. Eukaryot. Cell 14, 29–40 (2015).
Steed, C. A., Swan, J. E., Fitzpatrick, P. J. & Jankun-Kelly, T. J. A visual analytics approach for correlation, classification, and regression analysis. In: Innovative approaches of data visualization and visual analytics (eds Mao, L. H. & Weidong, H.) (IGI Global, Hershey, Pennsylvania, USA, 2014) pp. 25–45.
Steed, C. A. et al. Big data visual analytics for exploratory earth system simulation analysis. Comput. Geosci. 61, 71–82 (2013).
Inselberg, A. The plane with parallel coordinates. Vis. Comput. 1, 69–91 (1985).
Tesson, B. & Hildebrand, M. Dynamics of silica cell wall morphogenesis in the diatom Cyclotella cryptica: substructure formation and the role of microfilaments. J Struct Biol 169, 62–74 (2010).
Van der Walt, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Chollet, F. Keras: the python deep learning library (Astrophysics Source Code Library, 2018). https://keras.io.
Javaheri, N. et al. Temperature affects the silicate morphology in a diatom. Sci. Rep. 5, 1–9 (2015).
Acknowledgements
The research was partially conducted at the Center for Nanophase Materials Sciences, which is a DOE Office of Science User Facility. Work by C.A.S. was sponsored by the Department of Energy under the Scientific Discovery through Advanced Computing RAPIDS project. Work by K.A.H. and P.K.U. was enabled through the Oak Ridge High School Math Thesis Program. The research by J.K.M., T.J.M., O.S.O., A.A.P., A.A.T., S.M. and M.H. was sponsored by the Laboratory Directed Research and Development Program of Oak Ridge National Laboratory, managed by UT-Battelle, LLC, for the U.S. Department of Energy. This paper has been authored by UT-Battelle, LLC, under Contract no. DE-AC0500OR22725 with the U.S. Department of Energy. The Department of Energy will provide public access to these results of federally sponsored research in accordance with the DOE Public Access Plan (http://energy.gov/downloads/doe-public-access-plan).
Author information
Authors and Affiliations
Contributions
A.A.T. and N.B. developed the image processing procedures, constructed and trained both the NNs. A.A.T. analyzed the data and contributed to SEM imaging. N.B. extracted the CAMs. A.A.P. established the SEM imaging procedure and performed the SEM measurements. S.M., T.J.M. and J.K.M. grew diatom cultures and performed gene modification. M.H. obtained knockdown strains for the modification of diatoms. M.A.Z. proposed use of CAMs to highlight regions of interest for NNs. K.A.H., P.K.U. and C.A.S. contributed to the image analysis and data visualization. R.V. contributed to the neural net approach. A.V.I., A.B. and O.S.O. supervised the project and analyzed the data. All authors discussed the results and co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Trofimov, A.A., Pawlicki, A.A., Borodinov, N. et al. Deep data analytics for genetic engineering of diatoms linking genotype to phenotype via machine learning. npj Comput Mater 5, 67 (2019). https://doi.org/10.1038/s41524-019-0202-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-019-0202-3