Abstract
A novel computational methodology for large-scale screening of MOFs is applied to gas storage with the use of machine learning technologies. This approach is a promising trade-off between the accuracy of ab initio methods and the speed of classical approaches, strategically combined with chemical intuition. The results demonstrate that the chemical properties of MOFs are indeed predictable (stochastically, not deterministically) using machine learning methods and automated analysis protocols, with the accuracy of predictions increasing with sample size. Our initial results indicate that this methodology is promising to apply not only to gas storage in MOFs but in many other material science projects.
Similar content being viewed by others
Introduction
Metal–organic frameworks (MOFs) or porous coordination polymers are a rapidly growing family of hybrid inorganic–organic nanoporous materials, which belong to the category of coordination polymers.1,2,3 These relatively new materials consist of a three-dimensional periodic network, constructed from molecular building blocks, such as metal clusters and organic linkers (Fig. 1). The possible combinations of these numerous building blocks under different topologies result is an almost unlimited number of potential MOFs!
Since their discovery4 MOFs have attracted significant scientific attention due to their extraordinary properties. As “skeleton” materials, they pose very large pores and outstanding apparent surface area. If we were able to unwrap the surface of only one gram of these “very empty” materials, we could cover the area of a football court! These unique characteristics of the MOFs made them excellent candidates for catalysis and gas storage applications.
MOFs have shown exceptional performance in gas storage and separation. Both useful and harmful gases can be absorbed in their pores in very large amounts. The storage of hydrogen,5 methane,6,7 carbon dioxide,8,9 ammonia,10,11 hydrogen sulfide,11,12 etc. have been intensively investigated in MOFs and many new MOFs have been designed for that purpose.13 Some MOFs pose today the world record value for the storage of several gases.14,15
The two most studied gases in MOFs are H2 and CO2. The reasons are obvious: the 1st could solve the energy transferring problem, while the 2nd is responsible for global warming. There are several review papers presenting collections of MOFs and their youngest brothers—covalent organic frameworks,16,17 zeolitic imidazolate frameworks18,19 etc., with very high performance in gas storage.
Together with the experimental work, there is also a substantial theoretical support in the field. Theory undoubtedly plays a significant role in the development of the field, mainly in two ways: by explaining the experimental results20 and by leading the experiments.21 In the literature, there are several methodologies investigating the gas storage problem in MOFs. There are accurate ab initio quantum chemical approaches,22 computational light and fast classical Monte Carlo and molecular dynamics techniques23 and “multi-scale” methods that try to combine both.22 All of them address specific MOFs, either synthesized earlier, or designed for a specific application following chemical intuition.
Lately, a completely different computational approach appeared based on a large-scale screening of hypothetical MOFs.24,25 This new approach firstly generates all conceivable MOFs from a given chemical library of building blocks ending up in thousands of combinations. Then, a low computational cost screening is taking place using Monte Carlo classical techniques, ending up in the most promising candidates for a specific application.
Both general approaches have advantages and disadvantages in investigating existing MOFs and designing novel architectures. The 1st, that uses ab initio techniques and based in quantum theory is very accurate and the results are in most cases unquestionable. But unfortunately, since the computational cost is very high (approximately a week/MOF in a typical computer) it can be applied in very limited systems.
On the other hand, the large-scale screening with cost effective computational methods can monitor rapidly any MOF (approximately a minute/MOF in a typical computer) and thus thousands can be evaluated in the same period of time. But, as the authors state, “chemical intuition is absence and low-level computations can lead to unreal results or miss important species in the screening” .24
The ultimate computational technique should have the accuracy of the ab initio methods and the speed of the classical approaches, strategically combined with the chemical intuition of the researcher. But is this possible? “Can the computer learn chemistry?”
In this communication, we go one step further and we introduce “machine learning” for predicting gas adsorption in MOFs. Machine Learning is not new in the scientific community. Machine learning is the subfield of artificial intelligence that studies methods that can automatically learn interesting patterns from data. It overlaps with statistics in scope. The latter is a sub-branch of mathematics and so it traditionally favors problems with mathematical analytic solutions and arguably focuses less on automation and computation. The fields of statistics, machine learning, pattern recognition, and data mining are gradually merging into the emerging field of data science. Machine learning techniques have been successfully applied in a variety of domains. Examples of such applications include: recommender systems,26 automatic speech recognition,27 real-time face detection,28 identification of protein-protein interactions in yeast,29 peptide identification from tandem mass spectra,30 prediction of brain maturity from fMRI data,31 prediction of diffuse large B-cell lymphoma32 and hepatocellular carcinoma33 from gene expression data.
Results
In our study, we carefully collected from the literature 100 MOFs that have been synthesized in different laboratories around the world and their CO2 and H2 storage properties were accurately measured in specific thermodynamic conditions. For the H2 we gather data from different MOFs measured at 1 bar, 77 K. For the CO2 a small pressure and temperature range (0.99–1.20 bar in 293–313 K) was unavoidable since the 100 selected experiments were not in identical thermodynamic conditions. Nevertheless, we believe this small deviation will not affect the findings of our investigation, which is the testing of a new methodology based in machine learning.
With those we construct a small database (also called “dataset”) containing their key structural parameters like organic linker, metal cluster and functional groups. Across all selected MOFs, 62 organic linkers, 18 metal clusters and 12 functional groups were identified. Each such structural property can be encoded as a binary parameter (also called variable or feature), indicating its presence or absence in a specific MOF. A matrix representation was used to represent each dataset, with one row for each MOF (100 in total) and one column for each structural feature (92 in total). The datasets are provided as Supplementary Material.
A Machine Learning algorithm then learns a function (also called “model”) f that approximates the storage properties of MOFs given their structural parameters (Fig. 2). The hope is that the model f can generalize and accurately predict an approximate value for a new material, never seen before, represented with the values of its structural parameters. If the predicted property for the new material is not in the desired level, there is no point in synthesizing the material and vice-versa. Thus, an accurate predictive model can guide the synthesis of new materials and experimentation.
For our computational experiments, we used a customized version of the Just Add Data v0.6 tool (JAD Bio; Gnosis Data Analysis; www.gnosisda.gr). JAD has also been recently successfully applied to the prediction of proteins to periplasmic or cytoplasmic given their mature amino acid sequence.34 Though, a totally different problem, the application shows the ability of the automated pipeline to learn patterns from data that generalize to new data. JAD employs a fully-automated machine learning pipeline for producing a model from a dataset and an estimate of its predictive performance on new, unseen MOFs. The latter is especially important, as the main goal is to create a model that is able to perform well on new data, rather than the data used for producing it.
We performed a computational experiment to validate the predictive performance estimation of the model provided by JAD on new MOFs. For each original dataset, sub-samples (i.e., subsets of MOFs) were randomly selected (without replacement) of size 40, 50, …, 100%, and a model was learned using JAD from each such sub-dataset. In other words, we simulate the scenario where only a portion of the original data is available, the tool is run to learn a predictive model on them, and subsequently more data is collected prospectively upon which the predictive performance of the model is estimated. JAD outputs a predictive performance estimation derived from its input dataset. This is compared against the remaining data held out to obtain a second performance estimation, simulating the application of the predictive models on new MOFs. The above was repeated 100 times for each sub-sample size. This experiment is used only as a verification of the whole procedure and the estimation of its variance.
Discussion
It can be clearly seen in Fig. 3 that the error decreases with increased sample size. For both datasets, the mean absolute error on the test set is lower than that estimated on the training set, suggesting that the data analysis pipeline used by JAD does not provide optimistic estimations; on the contrary, it provides slightly conservative performance estimates. This allows us to have confidence in the performance estimation provided by the tool of the final models trained on the full dataset. Note that, even when 100% of the MOFs are used by JAD there is still some variance on the estimated error, which is the reason why all experiments were repeated 100 times. The (conservative) estimated mean absolute error of the models (when 100% of the available sample is used for training) is 4.07 g _CO2/g_MOF and 0.47 g_H2/g_MOF.
Estimated mean absolute error as a function of the sample-size percentage used for training the model. The average estimations over 100 repetitions of the computational experiment and the 2.5th and 97.5th percentiles of the performance estimation by JAD are shown. In addition, the average of the estimation on the hold-out set is shown too. The metric shown is the Mean Absolute Error. Results for both gases CO2 and H2 are shown in each column, respectively. Values are grams of absorbed gas per grams of material
Internally, in order to provide with a conservative predictive performance estimate JAD also (repeatedly) holds out some of the input data and evaluates predictions of the model learnt on the remaining data on the held out set. These are called “out-of-sample” predictions. JAD also outputs the out-of-sample predictions on the input MOFs (see “Methods” for a detailed description). Figure 4 shows the average predicted values over 100 repeated executions of JAD on the complete set of MOFs. There is a clear positive relationship between the predicted and actual values (Pearson correlation of 0.68 and 0.61 for CO2 and H2, respectively), although the ones on the lower and upper ranges are overestimated and underestimated respectively. It is expected that predictions would further improve if more MOFs were used for training the models.
Predicted gas adsorption values by JAD for each MOF. Each point is the average predicted value over 100 executions of JAD on the complete sets of MOFs. Predicted values are obtained on hold-out data, and not on MOFs used for training the models. The red curve shows the smoothed predictions. Values are grams of absorbed gas per grams of material
To gauge the practical significance of the results we check the accuracy of the model in filtering in practically useful and promising-to-synthesize MOFs. Specifically, we use the model to select the MOFs that are above a given threshold t, which is deemed of practical significance. We then measure how many predictions are indeed above the threshold. We used all predictions of JAD using the complete datasets and across all 100 repetitions. The results are shown in Table 1. The thresholds have been pre-selected and not optimized for performance through repeated trials. For example, for threshold 16 (gr_CO2/gr_MOF) for a researcher employing the model to synthesize a MOF with predicted CO2 above the threshold would expect 86% chance that the MOF actually exhibits a CO2 retention above 16 (gr_CO2/gr_MOF). In contrast, if the model is not employed for predictions, random guessing has a chance of about 21% to synthesize high retention MOFs.
Overall, the results demonstrate that:
(a) the chemical properties of MOFs are predictable (stochastically, not deterministically)
(b) the accuracy of predictions increases with sample size
(c) modern machine learning algorithms and automated predictive analytics pipelines can learn predictive models that generalize to unseen MOFs, and guide experiments in material discovery by predicting properties of new materials.
Limitations
Despite the promising results, both the theory and the scope of the computational experiments have limitations that need to be considered. First, we note that the performance estimation is correct when employed on new MOFs coming from the same statistical distribution as the MOFs employed during training. The collection of MOFs used for training is a selection from a “universe” of possible MOFs. If new MOFs are selected for testing with a different probability, the performance of predictions may increase or decrease.
Another limitation of the approach is that the models cannot be expected to generalize to unseen linkers or metals. This is a restriction of the data representation, rather than machine learning in general. If for example, the representation of a MOF included not just the presence or absence of a linker or metal, but in a finer detail, chemical properties of the linker or metal, then potentially a model could learn to predict gas storage in new MOFs employing new linkers not seen in the training test.
Methods
The problem that we study, that is, to predict the gas adsorption properties of MOFs given their structural properties, is called “supervised” learning in the field of machine learning. In “supervised learning”, the computer learns from labeled historical examples where the outcome is known, to generalize and make predictions on future data where the outcome is unknown.
Just Add Data (JAD Bio; Gnosis Data Analysis PC; www.gnosisda.com) is an automated tool that produces a supervised machine learning model and an estimate of its predictive performance. For regression problems (i.e., when the outcome is a continuous value, like CO2 and H2 adsorption), JAD employs state-of-the-art machine learning algorithms, such as “random forest regression” (RF),35 “support vector regression” (SVR)36 using both polynomial and Gaussian kernels and “ridge linear regression”37 although the list is continuously being enriched. In the computational experiments, out of 100 repetitions on the full set of MOFs, RF and ridge regression were selected two times each and SVR 96 times for the H2 data, while for the CO2 data only SVR models were selected. This happens due to the random split of the data in the cross-validation procedure. Overall however, the results are fairly robust.
Recently it has been shown experimentally on a variety of problems that random forests and support vector machines outperform other algorithms on average,38 and thus these were deemed sufficient choices for the moment. We note however that the best algorithm depends on the problem (see the no free lunch theorem for machine learning,39 and thus there may be better suited algorithms for the problems considered in this work. A high-level overview of the pipeline used by JAD is shown in Fig. 5. A detailed description of the pipeline follows.
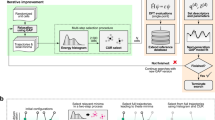
Schematic representation of the analysis pipeline employed by JAD. Based on the type of data and its size, the tool determines a set of combinations of tuning hyper-parameter values to try, called configurations. Hyper-parameters are depicted as tuning sliders. The data are partitioned to K-folds and for each fold and configuration a predictive model is trained. These are evaluated on the held-out folds and the average performance of each configuration is estimated. Based on the best configuration found a final model is produced on all data. The estimate of the best configuration is optimistic (see ref. 44 for an explanation); the optimism is removed using a bootstrap procedure before it is returned in a similar fashion as in ref. 44
All those algorithms require the user to set a number of parameters (called “hyper-parameters” in this context) that determine their behavior, and whose optimal values are problem-dependent. Results can greatly vary depending on correctly tuning the values of the hyper-parameters. The hyper-parameters are depicted as sliders in Fig. 5. Their optimal values cannot be found analytically; their values must be found by trial-and-error. JAD uses the statistical properties of the input data (such as the number of training examples and number of features) to determine a set of hyper-parameter combinations (called “configuration” hereafter) to try. In order to find the best algorithm and hyper-parameter configuration and to learn a final model, JAD uses the “K-fold cross-validation protocol”,40 described next.
The K-fold cross-validation protocol splits the data into K non-overlapping approximately equal-sized sets (called “folds”) of MOFs. In this work, we used K = 5. Each of them is held-out for testing purposes and the rest are used for training. It proceeds by keeping each fold out once, training models using all configurations on the remaining K-1 folds, and estimating their performance on the held-out fold. The held-out test sets are used to simulate the application of the models on new MOFs. In the end, K performance estimates are computed for each configuration, and the one with the best average performance is selected as the best configuration. A final model is produced by applying the best configuration on the complete set of MOFs. Note that, the predictions obtained for each MOF when in the hold-out test set can also be returned, and can be used to get an idea of how the final model would perform on unseen MOFs; these are the predictions used for Fig. 4 and Table 1.
Unfortunately, the estimation of this best performing configuration is optimistically biased because numerous models have been tried.41 The optimistic estimation is equivalent to the problem of multiple testing in statistics, called “the multiple induction problem in Machine Learning”.42 The problem is created because the test sets (folds) are employed multiple times, once for each configuration tried. The optimism problem has been noted both theoretically as well as experimentally. JAD estimates the bias of the performance using a bootstrap method,43,44 and removes it to return the final performance estimate.
Data availability
The 2 databases of the 100 MOFs (one for CO2 and one for H2 gases) used for this study are available as Supplementary Material.
Change history
08 November 2017
A correction to this article has been published and is linked from the HTML version of this article.
References
Furukawa, H., Cordova, K. E., O’Keeffe, M. & Yaghi, O. M. The chemistry and applications of metal-organic frameworks. Science 341, 1230444 (2013).
Farruseng, D. Metal-Organic Frameworks: Applications from Catalysis to Gas Storage (Wiley-VCH Verlag & Co. KGaA, 2011).
Kitagawa, S. & Matsuda, R. Chemistry of coordination space of porous coordination polymers. Coord. Chem. Rev. 251, 2490–2509 (2007).
Meek, S. T., Greathouse, J. A. & Allendorf, M. D. Metal-organic frameworks: a rapidly growing class of versatile nanoporous materials. Adv. Mater. 23, 249–267 (2011).
Suh, M. P., Park, H. J., Prasad, T. K. & Lim, D.-W. Hydrogen storage in metal-organic frameworks. Chem. Rev. 112, 782–835 (2012).
Furukawa, H. et al. Ultrahigh porosity in metal-organic frameworks. Science 329, 424–428 (2010).
He, Y., Zhou, W., Qian, G. & Chen, B. Methane storage in metal-organic frameworks. Chem. Soc. Rev. 43, 5657 (2014).
Li, J.-R. et al. Carbon dioxide capture-related gas adsorption and separation in metal-organic frameworks. Coord. Chem. Rev. 255, 1791–1823 (2011).
Lu, X. et al. Strategies to enhance CO2 storage and separation based on engineering adsorbent materials. J. Mater. Chem. A 3, 12118–12132 (2015).
Morris, W., Doonan, C. J. & Yaghi, O. M. Postsynthetic modification of a metal-organic framework for stabilization of a hemiaminal and ammonia uptake. Inorg. Chem. 50, 6853–6855 (2011).
DeCoste, J. B. & Peterson, G. W. Metal-organic frameworks for air purification of toxic chemicals. Chem.Rev. 114, 5695–5727 (2014).
Khan, N. A., Hasan, Z. & Jhung, S. H. Adsorptive removal of hazardous materials using metal-organic frameworks (MOFs): a review. J. Hazard. Mater. 244-245, 444–456 (2013).
Zhang, M., Bosch, M., Gentle III, T. & Zhou, H. C. Rational design of metal-organic frameworks with anticipated porosities and functionalities. CrysEngComm 16, 4069 (2014).
Mason, J. A., Veenstra, M. & Long, J. R. Evaluating metal-organic frameworks for natural gas storage. Chem. Sci. 5, 32–51 (2014).
Nandi, M. & Uyama, H. Exceptional CO2 adsorpbing materials under different conditions. Chem. Rec. 14, 1134–1148 (2014).
Ding, S.-Y. & Wang, W. Covalent organic frameworks (COFs): from design to applications. Chem. Soc. Rev. 42, 548–568 (2013).
Zeng, Y., Zou, R. & Zhao, Y. Covalent organic frameworks for CO2 capture. Adv. Mater. 28, 2855–2873 (2016).
Chen, B., Yang, Z., Zhu, Y. & Xia, Y. Zeolitic imidazolate framework materials. Recent progress in synthesis and applications. J. Mat. Chem. A 2, 16811–16831 (2014).
Pimentel, B. R., Parulkar, A., Zhou, E.-K., Brunelli, N. A. & Lively, R. P. Zeolitic imidazolate frameworks: next-generation materials for energy-efficient gas separations. ChemSusChem 7, 3202–3240 (2014).
Lee, J. S. et al. Understanding small-molecule interactions in metal–organic frameworks: coupling experiment with theory. Adv. Mater. 27, 5785–5796 (2015).
Bernales, V. et al. Computationally guided discovery of a catalytic cobalt-decorated metal–organic framework for ethylene dimerization. J. Phys. Chem. C 120, 23576–23583 (2016).
Odoh, S. O., Cramer, C. J., Truhlar, D. G. & Gagliardi, L. Quantum-chemical characterization of the properties and reactivities of metal−organic frameworks. Chem. Rev. 115, 6051–6111 (2015).
Evans, J. D. et al. Computational chemistry methods for nanoporous materials. Chem. Mater. 29, 199–212 (2016).
Wilmer, C. E. et al. Large-scale screening of hypothetical metal–organic frameworks. Nat. Chem. 4, 83–89 (2012).
Witman, M. et al. In silico design and screening of hypothetical MOF-74 analogs and their experimental synthesis. Chem. Sci. 7, 6263–6272 (2016).
Xiaoyuan, S. & Khoshgoftaar, T. M. A Survey of collaborative filtering techniques. Adv. Artif. Intell. 4, 1–19 (2009).
Deng, L. & Li, X. Machine learning paradigms for speech recognition: an overview, transactions on audio. Speech Lang. Process. 21, 1060–1089 (2013).
Sung, K. K., Poggio, T. Example-based learning for view-based human face detection. IEEE Trans. Pattern Anal. Mach. Intell.. 20, 39–35 (1998).
Jansen, R. et al. A bayesian networks approach for predicting protein-protein interactions from genomic data. Science 302, 449–453 (2003).
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
Dosenbach, N. U. F. et al. Prediction of individual brain maturity using fMRI. Science 329, 1358–1361 (2010).
Shipp, M. A. et al. Diffuse large B-cell lymphoma outcome prediction by gene-expression profiling and supervised machine learning. Nat. Med. 8, 68–74 (2002).
Ye, Q. H. et al. Predicting hepatitis B virus−positive metastatic hepatocellular carcinomas using gene expression profiling and supervised machine learning. Nat. Med. 9, 416–423 (2003).
Orfanoudaki, G., Markaki, M., Chatzi, K., Tsamardinos, I. & Economou, A. MatureP: prediction of secreted proteins with exclusive information from their mature regions. Sci. Rep. 7, 3263 (2017).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Smola, A. J. & Schölkopf, B. A tutorial on support vector regression. Stat. Comput. 14, 199–222 (2004).
Hoerl, A. E. & Kennard, R. W. Ridge regression: Biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
Fernández-Delgado, M., Cernadas, E., Barro, S. & Amorim, D. Do we need hundreds of classifiers to solve real world classification problems? J. Mach. Learn. Res. 15, 3133–3181 (2014).
Wolpert, D. H. The lack of a priori distinctions between learning algorithms. Neural Comput. 8, 1341–1390 (1996).
Mosteller, F., Tukey, J. in Revised Handbook of Social Psychology (eds Lindzey, G. & Aronson E.) 80–203. (Addison Wesley, 1968).
Varma, S. & Simon, R. Bias in error estimation when using cross-validation for model selection. BMC Bioinform. 2006, 7 (2006).
Jensen, D. D. & Cohen, P. R. Multiple comparisons in induction algorithms. Mach. Learn. 38, 309–338 (2000).
Efron, B. & Tibshirani, R. J. An Introduction to the Bootstrap (CRC press, 1994).
Tsamardinos, I., Greasidou, E., Tsagris, M. & Borboudakis, G. Bootstrapping the Out-of-sample Predictions for Efficient and Accurate Cross Validation. Preprint at https://arxiv.org/abs/1708.07180 (2017).
Acknowledgements
This research has been co-financed by the European Union (European Social Fund – ESF) and Greek national funds through the Operational Program “Education and Lifelong Learning” of the National Strategic Reference Framework (NSRF)—Research Funding Program: THALES. This research has been co-financed by AFOSR/EOARD under grant numberFA9550-15-1-0291.
Author information
Authors and Affiliations
Contributions
T.S., M.F. and E.K. built the data bases of the MOFs. G.B. performed the machine learning computations. G.B, I.T. and G.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history: A correction to this article has been published and is linked from the HTML version of this article.
A correction to this article is available online at https://doi.org/10.1038/s41524-017-0051-x.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Borboudakis, G., Stergiannakos, T., Frysali, M. et al. Chemically intuited, large-scale screening of MOFs by machine learning techniques. npj Comput Mater 3, 40 (2017). https://doi.org/10.1038/s41524-017-0045-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-017-0045-8
This article is cited by
-
Recent advances in metal-organic framework (MOF)-based agricultural sensors for metal ions: a review
Microchimica Acta (2024)
-
A machine learning approach utilizing DNA methylation as an accurate classifier of COVID-19 disease severity
Scientific Reports (2022)
-
Computational screening of transition metal-doped CdS for photocatalytic hydrogen production
npj Computational Materials (2022)
-
Just Add Data: automated predictive modeling for knowledge discovery and feature selection
npj Precision Oncology (2022)
-
Automated machine learning optimizes and accelerates predictive modeling from COVID-19 high throughput datasets
Scientific Reports (2021)