Abstract
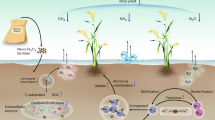
Plant nitrogen (N)-use efficiency (NUE) is largely determined by the ability of root to take up external N sources, whose availability and distribution in turn trigger the modification of root system architecture (RSA) for N foraging. Therefore, improving N-responsive reshaping of RSA for optimal N absorption is a major target for developing crops with high NUE. In this study, we identified RNR10 (REGULATOR OF N-RESPONSIVE RSA ON CHROMOSOME 10) as the causal gene that underlies the significantly different root developmental plasticity in response to changes in N level exhibited by the indica (Xian) and japonica (Geng) subspecies of rice. RNR10 encodes an F-box protein that interacts with a negative regulator of auxin biosynthesis, DNR1 (DULL NITROGEN RESPONSE1). Interestingly, RNR10 monoubiquitinates DNR1 and inhibits its degradation, thus antagonizing auxin accumulation, which results in reduced root responsivity to N and nitrate (NO3−) uptake. Therefore, modulating the RNR10-DNR1-auxin module provides a novel strategy for coordinating a desirable RSA and enhanced N acquisition for future sustainable agriculture.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Raw RNA-seq data are deposited at the National Center for Biotechnology Information’s Sequence Read Archive (SRA) (accession number PRJNA986304). Proteomic identification and ubiquitination proteomics data are deposited at ProteomeXchange (accession number PXD043400). Source data are provided with this paper.
References
Jia, Z., Giehl, R. & von Wirén, N. Nutrient-hormone relations: driving root plasticity in plants. Mol. Plant 15, 86–103 (2022).
Liu, H., Liu, Q., Gao, X. & Fu, X. Role of nitrogen sensing and its integrative signaling pathways in shaping root system architecture. Front. Agr. Sci. Eng. 9, 316–332 (2022).
Masclaux-Daubresse, C. et al. Nitrogen uptake, assimilation and remobilization in plants: challenges for sustainable and productive agriculture. Ann. Bot. 105, 1141–1157 (2010).
Kiba, T. & Krapp, A. Plant nitrogen acquisition under low availability: regulation of uptake and root architecture. Plant Cell Physiol. 57, 707–714 (2016).
Ma, W. et al. Auxin biosynthetic gene TAR2 is involved in low nitrogen-mediated reprogramming of root architecture in Arabidopsis. Plant J. 78, 70–79 (2014).
Liu, Y. & von Wirén, N. Integration of nutrient and water availabilities via auxin into the root developmental program. Curr. Opin. Plant Biol. 65, 102117 (2022).
Shao, A. et al. The auxin biosynthetic TRYPTOPHAN AMINOTRANSFERASE RELATED TaTAR2.1-3A increases grain yield of wheat. Plant Physiol. 174, 2274–2288 (2017).
Tromas, A. et al. Auxin-binding protein 1 is a negative regulator of the SCF (TIR1/AFB) pathway. Nat. Commun. 4, 2496 (2013).
Zhang, L. et al. Function of histone H2B monoubiquitination in transcriptional regulation of auxin biosynthesis in Arabidopsis. Commun. Biol. 4, 206 (2021).
Nakagawa, T. & Nakayama, K. Protein monoubiquitylation: targets and diverse functions. Genes Cells 20, 543–562 (2015).
Zhang, S. et al. Natural allelic variation in a modulator of auxin homeostasis improves grain yield and nitrogen use efficiency in rice. Plant Cell 33, 566–580 (2021).
Xi, Z. Y. et al. Development of a wide population of chromosome single-segment substitution in the genetic background of an elite cultivar of rice (Oryza sativa L.). Genome 49, 476–484 (2006).
Jain, M. et al. F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol. 143, 1467–1483 (2007).
Kahloul, S. et al. Structural, expression and interaction analysis of rice SKP1-Like genes. DNA Res. 20, 67–78 (2012).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Zheng, X. et al. Genomic signatures of domestication and adaptation during geographical expansions of rice cultivation. Plant Biotech. J. 20, 16–18 (2022).
Chen, D. et al. Parkin mono-ubiquitinates Bcl-2 and regulates autophagy. J. Biol. Chem. 285, 38214–38223 (2010).
Pavlopoulos, E. et al. Neuralized1 activates CPEB3: a function for nonproteolytic ubiquitin in synaptic plasticity and memory storage. Cell 147, 1369–1383 (2011).
Sun, H. et al. Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat. Genet. 46, 652–656 (2014).
Li, S. et al. Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 560, 595–600 (2018).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Wang, S. et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 44, 950–954 (2012).
He, Y. et al. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 11, 1210–1213 (2018).
Huang, X. et al. Natural variation at the DEP1 locus enhances grain yield in rice. Nat. Genet. 41, 494–497 (2009).
Wang, S. et al. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat. Genet. 47, 949–954 (2015).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol. 146, 368–376 (2008).
Wang, F. et al. Biochemical insights on degradation of Arabidopsis DELLA proteins gained from a cell-free assay system. Plant Cell 21, 2378–2390 (2009).
Wu, K. et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 367, eaaz204 (2020).
Zhao, Q. et al. A plant-specific in vitro ubiquitination analysis system. Plant J. 74, 524–533 (2013).
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Ho, C. H., Lin, S. H., Hu, H. C. & Tsay, Y. F. CHL1 functions as a nitrate sensor in plants. Cell 138, 1184–1194 (2009).
Loqué, D. et al. Additive contribution of AMT1;1 and AMT1;3 to high-affinity ammonium uptake across the plasma membrane of nitrogen-deficient Arabidopsis roots. Plant J. 48, 522–534 (2006).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Bandelt, H. J., Forster, P. & Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Layer, R. M. et al. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 15, R84 (2014).
Acknowledgements
We thank G. Xu (Nanjing Agricultural University) for critical suggestions. This research was supported by the National Key Research and Development Program of China (No. 2021YFF1000400 to Shan Li), Jiangsu Province Key Research and Development Program (No. BE2022335-2 to Shan Li), the National Natural Science Foundation of China (No. 32122065 to Shan Li), the Jiangsu Natural Science Foundation (No. BK20200540 to Shan Li), Fundamental Research Funds for the Central Universities (No. KJYQ2022001 to Shan Li) and Jiangsu Collaborative Innovation Center for Modern Crop Production. Work in N.P.H.’s laboratory was supported by the BBSRC-Newton ‘Rice’ Initiative (grant no. BB/N013611/1 to N.P.H.) and also by BBSRC Response Modes grant no. BB/S013741/1 to N.P.H.
Author information
Authors and Affiliations
Contributions
Shan Li, Y.H. and Z.J. conceived and designed this study. Y.H. performed most of the experiments; Y.H., Y. Tao, S. Wei, S.K. and C.S. conducted map-based cloning. Y.H., S.Z., Y.F., Y.Q., P.J. Shunqi Li, X.L., and Y. Tian constructed transgenic rice plants and performed field experiments and biochemical experiments. Y.H. and Z.J. conducted phylogenetic analysis. W.J. performed bioinformatic analysis. S. Wang provided SSSL lines. Z.J. and Shan Li wrote the manuscript. Z.J., Q.S., N.P.H., S. Wang and Shan Li revised the manuscript. All authors discussed the results and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Nicolaus von Wirén and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 N-dependent developmental plasticity responses of indica and japonica rice plants.
a, 21-day-old rice plants grown at LN (0.375 mM NH4NO3) versus HN (1.25 mM NH4NO3) supply. Scale bar, 10 cm. b, c, The absolute value of total length (b) and total area (c) of visible roots of 10-day old indica and japonica varieties.
Extended Data Fig. 2 RNR10 was identified by fine-scale mapping, and its sequence diverges between indica and japonica.
a, Successive maps of the candidate gene, RNR10, using indicated recombination break points and linked DNA markers to an ~ 3.3 kb segment flanked by the markers P241 and P242 on chromosome 10. The gene structure of RNR10 is shown underneath, where the thick black bars represent the protein-coding sequence, and the start and stop codons are labelled as ATG and TGA, respectively. The 3496-bp SV and 604-bp deletion in the RNR10IRAT261 promoter relative to the RNR10 start ATG (nucleotide 1), are shown. b, Agarose gel electrophoresis of PCR amplified products from RNR10 promoter, open reading frame and protein coding region, respectively. c, d, Agarose gel electrophoresis of PCR amplified products using primer pairs flanking the 3496-bp segment in the promoter region (c) and the 604-bp segment in the open reading frame (d) from our collection of 12 indica and 12 japonica varieties. e, Agarose gel electrophoresis shows that there is no detectable difference in the size of the protein coding region. f, The differences in RNR10 transcript level between indica and japonica varieties. Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. Data are mean ± s.e.m. (n = 3 biologically independent samples).
Extended Data Fig. 3 The root and shoot phenotypes of the NIL plants.
a, The ratio [(LN-HN)/HN] of total length of visible roots. b, The absolute value of total length of visible roots. c, The ratio [(LN-HN)/HN] of total area of visible roots. d, The absolute value of total area of visible roots. e-i, Mature plant height (e), the number of tillers per plant (f), the number of secondary branches per panicle (g), the number of grains per panicle (h), and grain yield per plant (i) of the NIL plants. j, The ratio [(LN-HN)/HN] of root to shoot biomass ratio. k, l, The changes in the ratios [(LN-HN)/HN] of root biomass (k) and shoot biomass (l) over time. m, n, The absolute values of root biomass (l) and shoot biomass (m). a, c, e-i, P values were generated from two-sided Student’s t tests. b, d, j-l, Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. a-d, j-n, Data are mean ± s.e.m. (n = 3 biologically independent samples). e, Data are mean ± s.e.m. (n = 16 biologically independent samples). f-i, Data are mean ± s.e.m. (n = 12 biologically independent samples).
Extended Data Fig. 4 The higher expression of RNR10 driven by the RNR10IRAT261 promoter in the HJX74 background resulted in reduced root growth and NO3− uptake.
a, Morphology of mature HJX74 and HJX74/pRNR10IRAT261::RNR10-GFP plants. Scale bar, 10 cm. b, Root RNR10 transcript abundance. Transcript abundance was measured relative to HJX74 (set to 1). c, Root RNR10-GFP protein abundance. HSP82 severs as loading control. The pictures of western blots represent one of the three experiments performed independently with similar results. d, Root systems of HJX74 and HJX74/pRNR10IRAT261::RNR10-GFP plants. Scale bar, 10 cm. e, The ratio [(LN-HN)/HN] of total length of visible roots. f, The absolute value of total length of visible roots. g, The ratio [(LN-HN)/HN] of total area of visible roots. h, The absolute value of total area of visible roots. i, The ratio [(LN-HN)/HN] of root to shoot biomass ratio. j, The absolute value of the ratio of root to shoot biomass. k-n, Mature plant height (k), the number of tillers per plant (l), the number of grains per panicle (m), and grain yield per plant (n). b, e, g, i, k-n, P values were generated from two-sided Student’s t tests. f, h, j, Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. b, e-j, Data are mean ± s.e.m. (n = 3 biologically independent samples). k, Data are mean ± s.e.m. (n = 16 biologically independent samples). l-n, Data are mean ± s.e.m. (n = 12 biologically independent samples).
Extended Data Fig. 5 RNR10 transcript abundance, RNR10 protein accumulation and RSA of ZH11, ZH11/pAct::RNR10-Flag and ZH11/rnr10 plants.
a, RNR10 mRNA abundance in ZH11/pAct::RNR10-Flag-1, ZH11/pAct::RNR10-Flag-2 and ZH11/rnr10 plants, relative to ZH11 (set to 1). b, Comparison of RNR10 protein abundance detected by an anti-DDDDK-tag antibody in ZH11 versus ZH11/pAct::DNR1-Flag. HSP82 severs as loading control. c, Gene structure of the RNR10 gene showing the location of the CRISPR/Cas9-generated 1-bp deletion in the ZH11/rnr10 mutant. d, Comparison of RNR10 protein abundance detected by an anti-RNR10 antibody in ZH11 versus ZH11/rnr10. e-g, The absolute values of total length of visible roots (e), total area of visible roots (f) and root to shoot biomass ratio(g). a, e-g, Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. Data are mean ± s.e.m. (n = 3 biologically independent samples). The experiments in b and d were repeated independently at least three times with similar results.
Extended Data Fig. 6 Phylogenetic relationship among the rice F-box proteins and their homologous genes, and amino acid sequence alignment of the FBA subfamily.
a, Phylogenetic tree of RNR10 and its homologous genes, constructed in MEGA11 using the Neighbour-Joining method. RNR10 is indicated by the star. b, Alignment of the protein sequences of members in the FBA subfamily. Black represents identical amino acids. The numbers indicate the positions of the amino acids.
Extended Data Fig. 7 RNR10 interacts with DNR1 and OSKs.
a-d, Four unique peptide sequences were identified by immunoprecipitation followed by mass spectrometry assays. e-g, Three amino acids were identified by mass spectrometric analysis. h, i, SFLC assays. cLUC-tagged RNR10 was co-transformed into tobacco leaves along with nLUC-tagged OSK1 (h) or OSK20 (i). j, k, Co-IP experiments. Flag-tagged RNR10 was co-transformed into rice protoplasts with HA-tagged OSK1 (j) or OSK20 (k). The experiments in j and k were repeated independently at least three times with similar results.
Extended Data Fig. 8 Root phenotypes and the ratio of root to shoot biomass of NIL-RNR10HJX74-DNR1HJX74, NIL-RNR10IRAT261, NIL-DNR1IRAP9, and NIL-RNR10IRAT261-DNR1IRAP9.
a, Root systems of NIL-RNR10HJX74-DNR1HJX74, NIL-RNR10IRAT261, NIL-DNR1IRAP9, and NIL-RNR10IRAT261-DNR1IRAP9 plants. Scale bar, 10 cm. b, The ratio [(LN-HN)/HN] of total length of visible roots. c, The absolute value of total length of visible roots. d, The ratio [(LN-HN)/HN] of total area of visible roots. e, The absolute value of total area of visible roots. f, 15NO3− uptake rates. g, The ratio [(LN-HN)/HN] of root to shoot biomass ratio. h, The absolute value of ratio of root to shoot biomass. b-h, Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. Data are mean ± s.e.m. (n = 3 biologically independent samples).
Extended Data Fig. 9 Gene structure, root size and above-ground phenotypes of NIL-DNR1HJX74, NIL-DNR1IRAP9, and NIL-DNR1IRAP9/rnr10.
a, Gene structure of the RNR10 gene showing the location of the CRISPR/Cas9-generated 5-bp deletion in the NIL-DNR1IRAP9/rnr10 mutant. b, c, The absolute value of total length of visible roots (b) and total area of visible roots (c). d-f, The number of tillers per plant (d), the number of secondary branches per panicle (e) and the number of grains per panicle (f) of NIL-DNR1HJX74, NIL-DNR1IRAP9 and NIL-DNR1IRAP9/rnr10. b-f, Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. b, c, Data are mean ± s.e.m. (n = 3 biologically independent samples). d-f, Data are mean ± s.e.m. (n = 12 biologically independent samples).
Extended Data Fig. 10 Effects on DNR1 and RNR10 abundance by external N, and RNR10 expression pattern.
a, b, Shoot mRNA and protein abundances of DNR1 (a) and RNR10 (b) in HJX74 grown in nutrient solutions with different N supply (0.15 N, 0.1875 mM NH4NO3; 0.3 N, 0.375 mM NH4NO3; 0.6 N, 0.75 mM NH4NO3; 1 N, 1.25 mM NH4NO3). Transcript abundance was measured relative to 0.15 N (set to 1). HSP82 severs as loading control. The experiments in a and b were repeated independently at least three times with similar results. c, Expression profile of RNR10 in root, stem, leaf, node, and panicle tissues. RNR10 transcript abundance in root was set to 1. a-c, Different letters denote significant differences (P < 0.05) from a Duncan’s multiple range test. Exact P values are listed in Supplemental Information Table 11. Data are mean ± s.e.m. (n = 3 biologically independent samples).
Supplementary information
Supplementary Table
Supplementary Tables 1–11.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Figs. 1, 3, 4, 5 and 6
Unprocessed blots and gels.
Source Data Extended Data Figs. 1, 3, 4, 5, 8, 9 and 10
Statistical source data.
Source Data Extended Data Figs. 2, 4, 5, 7 and 10
Unprocessed blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, Y., Ji, Z., Tao, Y. et al. Improving rice nitrogen-use efficiency by modulating a novel monouniquitination machinery for optimal root plasticity response to nitrogen. Nat. Plants 9, 1902–1914 (2023). https://doi.org/10.1038/s41477-023-01533-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01533-7