Abstract
Abscisic acid (ABA) is one of the plant hormones that regulate various physiological processes, including stomatal closure, seed germination and development. ABA is synthesized mainly in vascular tissues and transported to distal sites to exert its physiological functions. Many ABA transporters have been identified, however, the molecular mechanism of ABA transport remains elusive. Here we report the cryogenic electron microscopy structure of the Arabidopsis thaliana adenosine triphosphate-binding cassette G subfamily ABA exporter ABCG25 (AtABCG25) in inward-facing apo conformation, ABA-bound pre-translocation conformation and outward-facing occluded conformation. Structural and biochemical analyses reveal that the ABA bound with ABCG25 adopts a similar configuration as that in ABA receptors and that the ABA-specific binding is dictated by residues from transmembrane helices TM1, TM2 and TM5a of each protomer at the transmembrane domain interface. Comparison of different conformational structures reveals conformational changes, especially those of transmembrane helices and residues constituting the substrate translocation pathway during the cross-membrane transport process. Based on the structural data, a ‘gate-flipper’ translocation model of ABCG25-mediated ABA cross-membrane transport is proposed. Our structural data on AtABCG25 provide new clues to the physiological study of ABA and shed light on its potential applications in plants and agriculture.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The 3D cryo-EM density maps of ABCG25 have been deposited in the Electron Microscopy Data Bank under the accession numbers EMD-35145, EMD-35146, EMD-35147, EMD-35148, EMD-35149 and EMD-35150, respectively. Coordinates for structure models have been deposited in the Protein Data Bank (PDB) under the accession codes 8I38, 8I39, 8I3A, 8I3B, 8I3C and 8I3D, respectively. The protein sequence of Arabidopsis thaliana ABCG25 is publicly available at Uniprot (https://www.uniprot.org/) with accession code Q84TH5. Source data are provided with this paper.
References
Ton, J., Flors, V. & Mauch-Mani, B. The multifaceted role of ABA in disease resistance. Trends Plant Sci. 14, 310–317 (2009).
Kavi Kishor, P. B., Tiozon, R. N. Jr., Fernie, A. R. & Sreenivasulu, N. Abscisic acid and its role in the modulation of plant growth, development, and yield stability. Trends Plant Sci. 27, 1283–1295 (2022).
Cutler, S. R., Rodriguez, P. L., Finkelstein, R. R. & Abrams, S. R. Abscisic acid: emergence of a core signaling network. Annu. Rev. Plant Biol. 61, 651–679 (2010).
Kuromori, T., Seo, M. & Shinozaki, K. ABA transport and plant water stress responses. Trends Plant Sci. 23, 513–522 (2018).
Nambara, E. & Marion-Poll, A. Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 56, 165–185 (2005).
Seo, M. & Koshiba, T. Complex regulation of ABA biosynthesis in plants. Trends plant Sci. 7, 41–48 (2002).
Kang, J. et al. PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc. Natl Acad. Sci. USA 107, 2355–2360 (2010).
Kuromori, T. et al. ABC transporter AtABCG25 is involved in abscisic acid transport and responses. Proc. Natl Acad. Sci. USA 107, 2361–2366 (2010).
Kanno, Y. et al. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc. Natl Acad. Sci. USA 109, 9653–9658 (2012).
Zhang, H. et al. A DTX/MATE-type transporter facilitates abscisic acid efflux and modulates ABA sensitivity and drought tolerance in Arabidopsis. Mol. Plant 7, 1522–1532 (2014).
Kang, J. et al. Abscisic acid transporters cooperate to control seed germination. Nat. Commun. 6, 8113 (2015).
Anfang, M. & Shani, E. Transport mechanisms of plant hormones. Curr. Opin. Plant Biol. 63, 102055 (2021).
Do, T. H. T., Martinoia, E., Lee, Y. & Hwang, J. U. 2021 update on ATP-binding cassette (ABC) transporters: how they meet the needs of plants. Plant Physiol. 187, 1876–1892 (2021).
Zhang, Y. et al. ABA homeostasis and long-distance translocation are redundantly regulated by ABCG ABA importers. Sci. Adv. 7, eabf6069 (2021).
Park, Y. et al. Spatial regulation of ABCG25, an ABA exporter, is an important component of the mechanism controlling cellular ABA levels. Plant Cell 28, 2528–2544 (2016).
Kuromori, T. et al. Overexpression of AtABCG25 enhances the abscisic acid signal in guard cells and improves plant water use efficiency. Plant Sci. 251, 75–81 (2016).
Lee, J. Y. et al. Crystal structure of the human sterol transporter ABCG5/ABCG8. Nature 533, 561–564 (2016).
Sun, Y. Y. et al. Molecular basis of cholesterol efflux via ABCG subfamily transporters. Proc. Natl Acad. Sci. USA 118, e2110483118 (2021).
Skarda, L., Kowal, J. & Locher, K. P. Structure of the human cholesterol transporter ABCG1. J. Mol. Biol. 433, 167218 (2021).
Taylor, N. M. I. et al. Structure of the human multidrug transporter ABCG2. Nature 546, 504–509 (2017).
Jackson, S. M. et al. Structural basis of small-molecule inhibition of human multidrug transporter ABCG2. Nat. Struct. Mol. Biol. 25, 333–340 (2018).
Manolaridis, I. et al. Cryo-EM structures of a human ABCG2 mutant trapped in ATP-bound and substrate-bound states. Nature 563, 426–430 (2018).
Orlando, B. J. & Liao, M. ABCG2 transports anticancer drugs via a closed-to-open switch. Nat. Commun. 11, 2264 (2020).
Melcher, K. et al. A gate-latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature 462, 602–608 (2009).
Miyazono, K. et al. Structural basis of abscisic acid signalling. Nature 462, 609–614 (2009).
Yin, P. et al. Structural insights into the mechanism of abscisic acid signaling by PYL proteins. Nat. Struct. Mol. Biol. 16, 1230–1236 (2009).
Noriyuki, N. et al. Structural mechanism of abscisic acid binding and signaling by dimeric PYR1. Science 326, 1373–1379 (2009).
Pawela, A., Banasiak, J., Biala, W., Martinoia, E. & Jasinski, M. MtABCG20 is an ABA exporter influencing root morphology and seed germination of Medicago truncatula. Plant J. 98, 511–523 (2019).
Kuromori, T., Sugimoto, E. & Shinozaki, K. Brachypodium BdABCG25 is a homolog of Arabidopsis AtABCG25 involved in the transport of abscisic acid. FEBS Lett. 595, 954–959 (2021).
Cao, M. J. et al. Combining chemical and genetic approaches to increase drought resistance in plants. Nat. Commun. 8, 1183 (2017).
Chen, J. Molecular mechanism of the Escherichia coli maltose transporter. Curr. Opin. Struct. Biol. 23, 492–498 (2013).
Locher, K. P. Mechanistic diversity in ATP-binding cassette (ABC) transporters. Nat. Struct. Mol. Biol. 23, 487–493 (2016).
Rees, D. C., Johnson, E. & Lewinson, O. ABC transporters: the power to change. Nat. Rev. Mol. Cell Biol. 10, 218–227 (2009).
Thomas, C. & Tampe, R. Structural and mechanistic principles of ABC transporters. Annu. Rev. Biochem. 89, 605–636 (2020).
Rempel, S., Stanek, W. K. & Slotboom, D. J. ECF-type ATP-binding cassette transporters. Annu. Rev. Biochem. 88, 551–576 (2019).
Thomas, C. et al. Structural and functional diversity calls for a new classification of ABC transporters. FEBS Lett. 594, 3767–3775 (2020).
Zhang, P. Structure and mechanism of energy-coupling factor transporters. Trends Microbiol. 21, 652–659 (2013).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Hagn, F., Nasr, M. L. & Wagner, G. Assembly of phospholipid nanodiscs of controlled size for structural studies of membrane proteins by NMR. Nat. Protoc. 13, 79–98 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr D 74, 531–544 (2018).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Bao, Z. et al. Structure and mechanism of a group-I cobalt energy coupling factor transporter. Cell Res. 27, 675–687 (2017).
Acknowledgements
We thank the Center for Excellence in Molecular Plant Sciences core facility centre for mass spectrometry analysis, confocal analysis and diagnostic cryo-EM analysis. The two ABA mimics/derivatives were kind gifts from professor Y. Zhang of Center for Excellence in Molecular Plant Sciences. We thank H. Zhao and X. Zhang at the cryo-EM centre of Fudan University and M. Zhang at the cryo-EM centre of the Chinese Academy of Sciences interdisciplinary Research Center on Biology and Chemistry for their technical assistance on cryo-EM data collection. This work was supported by grants from the National Natural Science Foundation of China (32025020 and 32230050 to P.Z., 31970146 to Z.C. and 32100961 to X.Z.), the Chinese Academy of Sciences (XDB27020103 to P.Z.).
Author information
Authors and Affiliations
Contributions
X.H., and X.Z. designed and performed the bulk of the experiments. X.H. and N.A. carried out protein expression and purification, sample preparation, biochemical analysis and transport assay. X.Z. and X.H. carried out cryo-EM data collection and structure determination supervised by Z.C. and P.Z. M.Z. contributed to grid sample preparation and diagnostic cryo-EM analysis. M.M. contributed to protein purification and MST analysis. Y.Y., L.J. and Y.W. contributed to transport assay and mass spectrometry analysis. P.Z., X.H. and X.Z. wrote the manuscript with inputs from other authors. P.Z. conceived the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Youngsook Lee, Xiaochun Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and ATPase activity of AtABCG25.
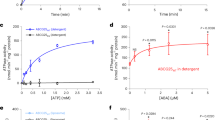
a-c, Gel filtration profile of a Superdex-200 column and Coomassie-blue-stained SDS-PAGE analysis of AtABCG25 wild type and mutant in buffer containing 0.05% digitonin. Independent experiments have been repeated at least three times with similar results. d, Gel filtration profile of a Superdex-200 column and Coomassie-blue-stained SDS-PAGE analysis of AtABCG25 E232Q mutation in nanodisc. Independent experiments have been repeated at least three times with similar results. e-f, Effect of substrate addition on the ATPase activity of AtABCG25 purified in buffer containing 0.05% digitonin (e) and reconstituted into nanodiscs (f). Data are mean ± s.e.m. (In 0.05% digitonin, n = 4 for protein incubate with DMSO or 0, 1, 2 and 5 μM ABA, n = 3 for protein incubate with 10, 20 and 50 μM ABA. In nanodiscs, n = 3 for protein incubate with DMSO or 2, 10, 50 μM ABA, n = 4 for protein incubate with 0, 1, 5, and ABA).
Extended Data Fig. 2 Transport activity assay of AtABCG25 in Xenopus oocytes.
Procedure of the transport activity assay. b, GFP fluorescence indicates the expression of genes encoding N-terminal GFP tagged AtABCG25 wild type and mutations on the membrane of Xenopus oocytes. Water was used as a control. The oocytes used for transporter assay analysis were selected based on the GFP fluence level, which indicates the protein expression amount of AtABCG25. Independent experiments have been repeated three times with similar results. Bar=2 μm.
Extended Data Fig. 3
Cryo-EM analysis of AtABCG25. a, The cryo-EM data analysis pipeline of AtABCG25. The examples of AtABCG25inward-ABA and AtABCG25outward are shown. b-g, Representative micrograph, 2D class averages and gold-standard Fourier shell correlation (FSC) curves for AtABCG25CHS-apo (b), AtABCG25CHS+ABA (c), AtABCG25inward-apo (d), AtABCG25inward-ABA (e), AtABCG25outward (f) and AtABCG25nanodisc (g).
Extended Data Fig. 4 Local resolution estimation of AtABCG25 and densities in the substrate-binding cavity.
a-f, Local resolution estimation of AtABCG25CHS-apo (a), AtABCG25CHS+ABA (b), AtABCG25inward-apo (c), AtABCG25inward-ABA (d), AtABCG25outward (e) and AtABCG25nanodisc (f). The colour represents the local resolution in Å. g-h, Density maps in the substrate-binding cavity of AtABCG25CHS-apo (g) and AtABCG25CHS+ABA(h). i-j, Density maps in the substrate-binding cavity of AtABCG25inward-apo (i) and AtABCG25inward-ABA (j). Density maps processed under C1 and C2 symmetry were both shown. (Map contour level = 5 σ.).
Extended Data Fig. 5 Representative densities of AtABCG25.
Cryo-EM density of representative segments superimposed with the atomic model in AtABCG25inward-apo (a), AtABCG25inward-ABA (b), AtABCG25outward (c). (Map contour level = 5 σ.).
Extended Data Fig. 6 Structure comparison of AtABCG25 and human ABCG subfamily members.
Overall structure and substrate binding site of AtABCG25 (a), HsABCG1 (b), HsABCG2 (c) and HsABCG5/ABCG8 (d) in inward-facing conformation. The corresponding substrate in the cavity is labelled, indicated the substrate binding model is different in ABCG subfamily members.
Extended Data Fig. 7 Configuration of ABA in ABCG25.
a-b, ABA (2-cis-4-trans) configurations in ABCG25 and ABA receptors were shown in separate view (a) and in superimposition view (b). PDB codes of ABA receptors are shown below. c, Comparison of (+)ABA and (-)ABA. The dashed circle indicated the difference. d, Substrate binding site of (+)ABA superimposed with (-)ABA. The dashed circle shows the conflict between (-)ABA and Thr552.
Extended Data Fig. 8 Multiple sequence alignment of the transmembrane domain of AtABCG25.
Invariant residues are highlighted with red box; conserved residues are highlighted by yellow box. Residues involved in substrate binding are indicated with green circles, and residues play key roles during substrate translocation are indicated with red stars. Secondary structures of AtABCG25 transmembrane domain are shown on the top. At: Arabidopsis thaliana, Uniprot: Q84TH5; Bd: Brachypodium distachyon, Gene ID: BRADI_4g24120; Mt: Medicago truncatula, Gene ID: MTR_1g093990; Zm: Zea mays, Gene ID: ZEAMMB73_Zm00001d053049; Os: Oryza sativa, Gene ID: OSNPB_110177400; Gm: Glycine max, Gene ID: GLYMA_10G217300.
Extended Data Fig. 9 Interaction between TMDs and NBDs in AtABCG25.
a, Salt bridges and hydrogen bonds between CnH, CpH and E-helix. b, Multiple sequence alignment of (a) among AtABCG25 and Homo sapiens ABCG families. Conserved residues are highlighted by purple box. c-d, Cation-pi interaction between residue Arg463 from CpH of one protomer and residue Phe203 from the opposing protomer induced by ATP-binding. Residue Arg463 from CpH of one protomer and residue Phe203 from the opposing protomer fall apart in the absence of ATP (c), and form cation-pi interaction (d) due to ATP-binding to the NBD.
Supplementary information
Supplementary Information
Cryo-EM data collection, refinement and validation statistics.
Source data
Source Data Fig. 1
Statistical analysis source data.
Source Data Fig. 2
Statistical analysis source data.
Source Data Fig. 5
Statistical analysis source data.
Source Data Extended Data Fig. 1
Statistical analysis source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, X., Zhang, X., An, N. et al. Cryo-EM structure and molecular mechanism of abscisic acid transporter ABCG25. Nat. Plants 9, 1709–1719 (2023). https://doi.org/10.1038/s41477-023-01509-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01509-7
This article is cited by
-
Structural basis for abscisic acid efflux mediated by ABCG25 in Arabidopsis thaliana
Nature Plants (2023)