Abstract
The adenosine A3 receptor (A3AR), a key member of the G protein-coupled receptor family, is a promising therapeutic target for inflammatory and cancerous conditions. The selective A3AR agonists, CF101 and CF102, are clinically significant, yet their recognition mechanisms remained elusive. Here we report the cryogenic electron microscopy structures of the full-length human A3AR bound to CF101 and CF102 with heterotrimeric Gi protein in complex at 3.3-3.2 Å resolution. These agonists reside in the orthosteric pocket, forming conserved interactions via their adenine moieties, while their 3-iodobenzyl groups exhibit distinct orientations. Functional assays reveal the critical role of extracellular loop 3 in A3AR’s ligand selectivity and receptor activation. Key mutations, including His3.37, Ser5.42, and Ser6.52, in a unique sub-pocket of A3AR, significantly impact receptor activation. Comparative analysis with the inactive A2AAR structure highlights a conserved receptor activation mechanism. Our findings provide comprehensive insights into the molecular recognition and signaling of A3AR, paving the way for designing subtype-selective adenosine receptor ligands.
Similar content being viewed by others
Introduction
The adenosine receptor subfamily of G protein-coupled receptors consists of four subtypes: A1, A2A, A2B, and A31,2. These receptors are activated by the endogenous ligand, adenosine, to transduce downstream signals that mediate a number of important physiological and pathological functions including immunomodulation, energy balance, cardiac function, and neuroprotection3,4,5. The gene of A3AR was firstly cloned in 19916 and characterized as a subtype of adenosine receptor in 19931. It is expressed in various tissues including the brain, heart, lungs, liver, kidneys, and immune cells7. A3AR participates in regulating cardiac function, vasodilation, inhibition of inflammation, protection against ischemia-reperfusion injury, and suppression of oxidative stress. Additionally, A3AR is highly expressed in several tumor types, making it as a promising therapeutic target for suppressing cancer cell proliferation7,8,9.
A1AR and A3AR preferentially couple to the inhibitory G protein (Gi), leading to the suppression of adenylate cyclase activity and a reduction in intracellular cyclic AMP levels, contrasting with the stimulatory G protein (Gs) signaling triggered by A2AAR and A2BAR activation2. The structure of adenosine has inspired the design of various agonists and antagonists targeting A3AR, particularly for cancer, inflammation, and pain management10. Studies highlight that alterations at the N6 position of the purine ring and the 5’-N position of the ribose group enhance the potency and selectivity of A3AR agonists11,12,13. Notably, N6-methyladenosine (m6A), a methylated adenosine metabolite, emerged as a potent A3AR agonist14. CF101 and CF102 are representatives of such modification strategy with similar nucleoside core structure and only one chloro-substituent difference, both demonstrate high affinity and selectivity for A3AR15,16,17. These effective orally compounds have shown promise in disrupting key signaling pathways in cancer and inflammatory cells10. CF101 has demonstrated efficacy in Phase III trials for psoriasis and rheumatoid arthritis, while CF102 is being evaluated for hepatocellular carcinoma and non-alcoholic steatohepatitis18,19. The broad expression of adenosine receptors poses challenges in designing subtype-selective compounds20,21. The lack of structural information for A3AR, unlike other adenosine receptor subtypes, limits our understanding of its specific signaling mechanisms and impedes structure-based drug design.
Here, we present the cryo-EM structures of A3AR bound to the Gi protein in the presence of CF101 and CF102. These structures reveal the mechanisms of ligand recognition and activation in A3AR, providing valuable insights for designing effective, targeted therapies for conditions like cancer and inflammation.
Results and discussion
Overall structures of the complexes
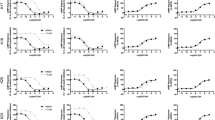
CF101 and CF102 are A3AR agonists that contain modifications to the ribose and adenine moieties, which confer their potent binding to A3AR. Specifically, CF101 and CF102 have a 5’-N-methylcarboxamide substitution on the ribose group and a N6-(3-iodobenzyl) substitution on the adenine base (Fig. 1a). These combined modifications result in significantly higher A3AR potency compared to the endogenous A3AR agonist adenosine. To ensure the specificity of our experiments in the context of HEK293 cells, which are known to express high levels of A1AR, A2AAR, and A2BAR but not A3AR, we employed NanoBiT association assays. These assays were crucial in determining the selectivity of CF101 and CF102 for A3AR, as opposed to other adenosine receptor subtypes (Fig. 1b–d). While adenosine activated four subtypes with similar micromolar potencies, CF101 and CF102 displayed strong potency of ~3 nM on A3AR but had weak or negligible response on other subtypes of adenosine receptors.
a Chemical structures of the adenosine, CF101 and CF102 are provided, highlighting modifications at the 5’-N-methylcarboxamide in the ribose group, as well as the N6 and C2 positions of the adenosine group. The atom numbering is indicated in blue. CF101, is also named IB-MECA and N6-(3-iodobenzyl)adenosine-5’-N-methyluronamide. CF102, is also named Cl-IB-MECA and 2-chloro-N6-(3-iodobenzyl)adenosine-5’-N-methyluronamide. NanoBiT association assays monitoring ligand activity on adenosine receptors for adenosine (b), CF101 (c) and CF102 (d), respectively. Data shown are mean ± S.E.M. of three independent experiments (n = 3). Source data are provided as a Source Data file. Cryo-EM map (e) and model (f) of the CF101-A3AR-Gi complex, with inset showing CF101 density. The density map in the inset is shown at 0.232 threshold. Cryo-EM map (g) and model (h) of the CF102-A3AR-Gi complex, with inset showing CF102 density. The density map in the inset is shown at 0.17 threshold. Subunits are colored as indicated.
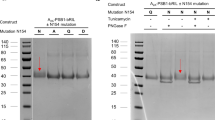
We used HiBiT tether approach to stabilize the full-length A3AR-G protein complexes, as it has been used for many GPCR structural studies22,23,24 (Supplementary Fig. S1). The large NanoLuc domain (LgBiT) and small high affinity fragment (HiBiT) was fused at the C-terminal of A3AR and Gβ, respectively. Meanwhile, A3AR used in this study had an N-terminal thermostabilized apocytochrome b562RIL (BRIL) fusion to enhance its expression, which is co-expressed with G protein subunits and scFv16, an antibody fragment that is used to further stabilize the receptor G protein complex. For the CF101-A3AR-Gi complex, data from 20,779 movies comprising 271,323 particles was used to determine the structure at 3.29 Å resolution (Supplementary Fig. S2, Supplementary Table S1). For the CF102-A3AR-Gi complex, data from 13,581 movies yielding 283,561 particles was used to determine the structure at a resolution of 3.19 Å (Supplementary Fig. S3, Supplementary Table S1). The structures of the CF101/CF102-A3AR-Gi complexes revealed that the ligands occupy the orthosteric binding pocket, with the core structures modeled clearly into the cryo-EM density at the center of the receptor transmembrane helical domain (TMD) (Fig. 1e–h).
The structures showed the canonical seven-transmembrane architecture for A3AR, with the intracellular domains occupied by the α5 helix of Gαi for Gi coupling. The density maps enabled modeling of most of the structures, except for A3AR N-terminus residues M1-L8, third intracellular loop N211-Y222, C-terminus V301-E318, and the alpha-helical domain of Gαi. The extracellular loop M151-S165 was also less defined but the backbone could be established (Supplementary Fig. S4). Aside from these regions, the models were well-resolved. Overall, the two agonist-bound complexes were highly similar, with 0.593 Å root mean square deviation (RMSD) for the whole receptor.
Binding mode of CF101/CF102 in A3AR orthosteric site
The A3AR agonists CF101 and CF102 bind at conserved orthosteric pocket forms by ECL2, TM3, TM5, TM6 and TM7, akin to the endogenous ligand adenosine bound to other adenosine receptor subtypes (Fig. 2a, b). However, the orientations of the modified 3-iodobenzyl moieties differ between CF101 and CF102. The adenine core mediates conserved receptor interactions commonly seen in other adenosine receptors23,25,26. Notably, the adenine pyrimidine forms π-stacks against F45.52, and the F45.52A mutation greatly affected the ability of CF101/CF102 to induce the receptor activation in the NanoBiT association assay (Fig. 2c–f, Supplementary Table S2). Additionally, 2’ and 3’ hydroxyl group in ribose and purine group form hydrogen bonds with polar side chains at positions 3.36, 6.55 and 7.43, which are conserved and critical for recognition of nucleoside ligands by adenosine receptors (Fig. 2c–f, Supplementary Table S2).
Detailed interactions between A3AR and CF101 (a) or CF102 (b) from the membrane plane. Residues involved in ligand interaction are colored blue and pink in two complexes, respectively. Black dashed lines indicate hydrogen bonds. Dose-response curves of mutants of A3AR induced by CF101 (upper panels, c, e) or CF102 (lower panels, d, f) using NanoBiT assay. Data shown are mean ± S.E.M. of three independent experiments (n = 3). Source data are provided as a Source Data file.
The ligand binding pocket is mainly comprised of hydrophobic residues, including position 3.33, 5.38, 5.47, 6.48, 6.51 and 7.39, which form hydrophobic contacts that are important for CF101/CF102 potencies (Fig. 2c–f, Supplementary Table S2). Alanine mutations at these positions severely reduced agonists’ ability to induce receptor activation. His3.37 and Ser5.42 participate van der Waals contacts with the bound ligands, their alanine mutations also affected activity (Fig. 2c–f, Supplementary Table S2). To confirm the functional data, cAMP accumulation assays were carried out to assess the agonist activity (Supplementary Fig. S5, Supplementary Table S3). The results from the NanoBiT association assay and cAMP accumulation assay were consistent. The side chains from M1745.35 and L2647.35 in the receptor form hydrophobic interactions with the 3-iodobenzyl group extended from the N6 position of the adenosine base of CF101. In contrast, the corresponding group of CF102 is surrounded by V169ECL2 and L2647.35 from the receptor. Alanine mutations on these residues did not significantly affect the potency of the compounds on A3AR (Supplementary Fig. S6, Supplementary Table S2), suggesting that the 3-iodobenzyl substituents may exist alternative states at the receptor extracellular domains. This demonstrates that the N6 position may accommodate various substituted groups through distinct conformations in the A3AR pocket.
CF102 is a 2-chloro derivative of CF101. In CF102-bound A3AR, Y151.35 is situated near the 2-chloro group of CF102 (Supplementary Fig. S7a). The Y151.35A mutation in A3AR abolished the agonist activity of both CF101 and CF102 (Fig. 2c, d). However, the Y151.35F mutant only slightly impacted the potency of CF101 and CF102 (Supplementary Fig. S7b, c). Y151.35 forms extensive π-π contact with Y2657.36 in TM7. The Y2657.36A mutant also affected the receptor’s ability to bind CF101 or CF102 (Supplementary Fig. S7). This implies that Y151.35 likely plays a critical role in maintaining the stability and structural integrity of A3AR, thus affecting both CF101 and CF102 binding to the receptor. Additionally, modifications at the 2-position of adenosine tend to be well tolerated for A3AR binding16, whether incorporating a small or large substituent, or even linking it to the N6 moiety to form a macrocycle27. Elucidation of these subtle ligand and receptor interaction variations thus provides molecular insight into the conformational adaptability and binding poses governing molecular recognition by A3AR.
The role of ECL3 in A3AR subtype selectivity
CF101 and CF102 show high selectivity on A3AR rather than other subtypes (Fig. 1c, d). Analysis the sequence of adenosine receptors reveals strong conservation within TMs, while the extracellular loops diverge among subtypes (Supplementary Fig. S8). ECL1 shows relatively distant from the orthosteric site. The residue F16845.52 in ECL2 of adenosine receptors provides critical π-π interactions with both agonists and antagonists binding to these receptors. However, A3AR possesses a shorter ECL3 than other subtypes (Fig. 3a). The shorter ECL3 may rigidify A3AR to minimize its conformational changes for ligand binding (Fig. 3b).
a sequence alignment of ECL3 among adenosine receptors. The disulfide bond was shown as green linker. b Superimposed structures of adenosine receptors reveal that A3AR has the shortest ECL3. The residues in A3AR are shown in pink. The residues formed disulfide bond on ECL3 in A1AR were shown in green. Other TMs were omitted. c–e Assessing the effects of adenosine, CF101, and CF102 on A1AR, A2AAR, and A2BAR, along with their corresponding mutants containing the swapped ECL3 from the A3AR using NanoBiT assays. The results were from three independent experiments. Data shown are mean ± S.E.M. of three independent experiments (n = 3). Source data are provided as a Source Data file. f, g Assessing the effects of CF101 and CF102 on A3AR and its mutants with flexible ECL3 using NanoBiT assays. Data shown are mean ± S.E.M. of three independent experiments (n = 3). Source data are provided as a Source Data file.
To assess the role of ECL3 in A3AR, we engineered chimeric receptors by grafting ECL3 from A3AR onto the backbones of other adenosine receptors. NanoBiT assays were performed to test the binding abilities of adenosine, CF101, and CF102 to wide-type or chimeric adenosine receptors (Fig. 3c–e). The result showed that the three chimeric adenosine receptors did not show increased binding ability to the endogenous ligand adenosine (Fig. 3c, Supplementary Table S4). However, the three ECL3-chimeric receptors gained the ability to bind CF101 and CF102 with increased efficacy or potency (Fig. 3d, e, Supplementary Table S3). These findings suggest that ECL3 could serve as a structural factor mediating the selective recognition CF101 and CF102 by A3AR. The significance of ECL3’s length and amino acid composition in A3AR’s ligand binding was further investigated through mutations. We mutated the original four ECL3 residues of A3AR to GGGS or (GGGS)2 that has the same length as ECL3 of A2BAR. Neither mutant above showed any binding ability to CF101 and CF102 (Fig. 3f, g), suggesting that both the specific length and the unique amino acid sequence of ECL3 play critical roles in the selective binding of ligands to A3AR, underscoring the complexity of ligand-receptor interactions in this context.
The proximity of the ECL3 to the N6 position in adenosine is likely a crucial factor in the selectivity of A3AR for N6-modified adenosine derivatives, as indicated by structure-activity relationship (SAR) studies15,16. Substituents at the N6 position, whether too small or overly bulky, can adversely affect the potency and affinity of ligands for A3AR. This relationship underscores the importance of ECL3 in ligand recognition, as the N6 position extends into A3AR’s binding pocket near ECL3. Understanding these intricate structural interactions is key for discerning the selectivity mechanisms of structurally similar ligands at different adenosine receptors.
Residues in binding pocket across adenosine receptors
Among adenosine receptors, A3AR stands out with the lowest sequence identity compared to other subtypes. This distinction is particularly evident in the orthosteric binding pocket (Fig. 4a), where A3AR’s unique residues at specific positions contribute to its selective ligand binding. Notably, positions 3.32, 3.37, 5.42, 5.47, 6.52, and 6.58 feature different amino acids in A3AR compared to A1, A2A, and A2B receptors (Fig. 4b, Supplementary Fig. S9). Mutations at these positions to their counterparts in other subtypes were conducted to evaluate their impact on CF101 and CF102 binding and activity. NanoBiT assays and cAMP accumulation assays were utilized to cross-confirm the effects of the mutations (Fig. 4c, d, Supplementary Fig. S10, Supplementary Tables S2 and 3).
a Aligning the residues in the orthosteric binding pocket among the adenosine receptors. The conserved residues were colored in blue, and stars were used as markers. The unique residues in A3AR, distinct from other adenosine receptors subtypes were colored in orange, while residues in corresponding positions in other subtypes were colored in green. All residues were annotated based on GPCR Ballesteros-Weinstein numbering scheme. b In the superposition of adenosine receptors, the unique residues in A3AR, distinct from those in other adenosine receptors, are represented as yellow spheres. c, d Effects of CF101/CF102 on A3AR mutants containing swapped residues from other adenosine receptors by NanoBiT assay. Data shown are mean ± S.E.M. of three independent experiments (n = 3). Source data are provided as a Source Data file. e–i The binding cavities of the adenosine receptors were generated in PyMOL and depicted in gray. In A3AR, a subpocket is formed by His3.37, Ser5.42, and Ser6.52, while these positions are conserved as Gln3.37, Asn5.42, and His6.52 in other adenosine receptor subtypes (His, H; Ser, S; Gln, Q; Asn, N). In h and i, dashed lines depict the hydrogen bonds between His3.37 and Ser5.42. The names of the receptors and their associated PDB codes23,26,45 are indicated below each model.
We found that changing the leucine at 3.32 to valine, similar to other subtypes, had no significant effect on the activity of CF101 and CF102 in NanoBiT assay, likely due to their comparable hydrophobic properties (Fig. 4c, d). However, mutations at positions 5.47 and 6.58 altered the receptor activation, indicating the importance of side chain length at these positions for ligand binding (Fig. 4c, d).
Furthermore, the hydrogen bond formation between H953.37 and S1815.42 in A3AR, which was absent in other subtypes, appears critical (Fig. 4e–i). Mutations H953.37Q and S2475.42N significantly impacted CF101 and CF102 activities (Fig. 4c, d), highlighting the importance of these residues in ligand-receptor interaction. The mutation of S2476.52 to histidine also reduced ligand activity, suggesting the influence of steric and electronic properties of the side chains (Fig. 4c, d, Supplementary Table S2).
Residues H953.37, S1815.42 and S2476.52 form a unique sub-pocket in A3AR to accommodate the 5’-N-methylcarboxamide from the ribose (Fig. 4h, i, Supplementary Fig. S11). The mutational results implicate this sub-pocket might serve as a structural determinant for stabilizing CF101 and CF102 in A3AR versus other subtypes. Our results above with NanoBiT assay were replicated with traditional cAMP accumulation assays (Fig. 4c, d, Supplementary Fig. S10), further demonstrating that how minor sequence variations in receptors can significantly influence their conformations and ligand binding specificity.
Activation mechanisms of A3AR
Structural comparisons between active, agonist-bound A3AR complexes and an inactive, antagonist-bound A2AAR structure (PDB ID: 4EIY)28 reveal classical hallmarks of conformational changes associated with GPCR activation29,30. Notably, the A3AR structures exhibit an outward movement of TM6 compared to inactive A2AAR, shifting 11.6 Å based on measurements of residue Glu6.30 at Cα atoms in receptors (Fig. 5a). Additional rearrangements of activation include inward movements of TM1 and TM7 and an upward shift of TM3 in A3AR relative to inactive A2AAR (Fig. 5b–d).
a, b Superposition of active A3AR-CF101/CF102 complexes (blue/pink) with inactive A2AAR-ZM241385 complex (gray, PDB ID 4EIY). Comparison of extracellular (c) and cytoplasmic (d) views of active A3AR and inactive A2AAR. e–h Conformational changes in conserved motifs, including the toggle switch, PIF, DRY and NPxxY, upon CF101/CF102 binding to A3AR relative to inactive state of A2AAR-ZM241385. Arrows indicate movement directions. In e, The sub-pocket in A3AR is formed by residues at position 3.37, 6.52 and 6.52. The residues at these positions from both A3AR and A2AAR were labeled in green.
Detail structural analysis also provide potential mechanism of ligand induced A3AR activation.
A unique sub-pocket formed by H3.37, S5.42 and S6.52 residues confers selectivity over other adenosine receptor subtypes (Fig. 5e). This facilitates deeper binding of CF101/CF102 compared to A2AAR antagonists, enabling engagement with conserved motifs like the W6.48 “toggle switch”. Propagation through P5.50I3.40F6.44, D3.49R3.50Y3.51, and N7.49P7.50xxY7.53 motifs transduces rearrangements (Fig. 5f–h), while limited ECL3 flexibility likely assists selective activation. By elucidating the structural transitions from inactive to active A3AR, our findings provide molecular insights connecting specialized agonist recognition to downstream signaling activation.
G protein coupling of adenosine receptors
Adenosine receptors exhibit differential G protein coupling preferences that correlate with distinct conformational orientations of the associated G proteins23,25,26. Structural alignment reveals A3AR-Gi shares better overlay with A1AR-Gi versus A2A/A2BAR-Gs (Fig. 6a). The analogous Gi-binding modes of A3AR and A1AR contrast A2A/A2BAR’s Gs-coupling preferences, consistent with sequence and functional profiles. Notably, TM6 positioning facilitates differential G protein accommodation, 3.1 Å inward shift enables A1/A3AR-Gi versus A2A/A2BAR-Gs binding (Fig. 6b). Additionally, α5 helix and αN of Gi protein tilt orient differently between complexes, induced by receptors’ hydrophobic and polar residue interactions (Fig. 6c, d). The α5 helix of Gαs subunits in A2AAR-Gs displays an 8.6 Å displacement relative to its orientation in A3AR-Gi complexes based on measurements of the Cα atom of GαH5.03 (Fig. 6c). The αN helix of Gαi exhibits a 3.3 Å tilt compared to Gs when measuring the Cα of GαHN.39.
a Comparison of adenosine receptor and Gα protein conformations in A1/A3AR-Gi and A2A/A2BAR-Gs complexes. Omitting the Gβ and Gγ subunits. The PDB codes for A1AR, A2AAR and A2BAR are 7LD4, 6GDG and 8HDP, respectively. b Conformational comparison of TM6 in adenosine receptors, with reference to the toggle switch residue W6.48 in TM6 of receptor. c, d Conformational comparison of the α5 helix and αN helix in G protein among adenosine receptor-G protein complexes. Arrows indicate movement directions. e, f Distinguishing residues on TM3 and ICL2 in adenosine receptor subtypes that participate in G protein coupling are highlighted. Components of adenosine receptors-G protein complexes are colored as indicated.
Furthermore, different adenosine receptors induced a variation in the N-terminal helix (αN) tilt of the Gα protein (Fig. 6d). The residue at position 34.51 (L/L/L/V, the residue in A1/A2A/A2B/A3-AR) in receptors is conserved as a hydrophobic residue that forms hydrophobic interactions with the G protein by inserting into the cleft between αN and α5 of the Gα protein (Supplementary Fig. S12a). Besides, sequence alignment of adenosine receptors showed that residues at positions 3.53 (R/A/A/R) and 34.55 (M/G/S/R) revealed different preferences in different subtypes (Fig. 6e, f, Supplementary Fig. S12a). The longer side chains in Gi-coupled A1AR and A3AR likely triggered more noticeable translocations in the αN and α5 helix to accommodate the Gαi protein. The main chains from R3.53 and P34.50 in A1AR formed a polar interaction with N348 in Gαi protein (Supplementary Fig. S12b). The side chain of R3.53 and H4.39 in A3AR formed a polar interaction with the side chain of N347 and E28 in Gαi protein, respectively (Supplementary Fig. S12c). Both of the complexes of A2AAR/A2BAR-Gs, H41 in Gαs formed a polar interaction with the main chain from the receptor’s ICL2 (Supplementary Fig. S12d, e). Together, these findings reveal that preferred Gi-coupled adenosine receptors adopt conserved Gi protein-binding conformations that differ distinctly from those of Gs-coupled adenosine receptor subtypes.
In summary, we have determined cryo-EM structures of the A3AR bound to selective agonists CF101 and CF102 with heterotrimeric Gi protein. Despite the conserved binding of the core adenosine moiety, the structures revealed differences in the orientations of the N6 substituted groups in CF101 and CF102. We have identified ECL3 and key sub-pocket residues His3.37, Ser5.42 and Ser6.52 that confer selectivity over other adenosine receptor subtypes by structural and mutational studies. Comparison to an inactive A2AAR structure provided insight into the conformational changes associated with A3AR activation and G protein coupling. By elucidating the molecular mechanisms governing ligand recognition, signaling, and subtype selectivity, the experimentally determined A3AR structures significantly advance our fundamental understanding of this important drug target. The findings pave the way for structure-guided design of selective ligands targeting adenosine receptors subtypes for the treatment of cancer, inflammation, and other diseases.
Methods
Construct design
The full-length gene coding human A3AR was synthesized (Synbio) and subcloned into pFastBac vector using ClonExpress II one step cloning kit (Vazyme Biotech, C112). A hemagglutinin signal peptide and thermostabilized apocytochrome b562RIL (BRIL) were fused at the N-terminal of A3AR to enhance receptor expression. To enhance complex stability, a NanoBiT tethering approach was used where an LgBiT domain was fused to the C-terminal of the receptor22. A dual maltose-binding protein was linked after LgBiT through a tobacco etch virus protease site (TEV site) for further cleavage. A dominant-negative mutant of bovine Gαi containing G203A/A326S31 mutations was generated to stabilize the heterotrimeric Gαiβγ protein. Rat Gβ1 was fused with a HiBiT at C-terminal for structural complementation of LgBiT to form a NanoBiT. The single-chain variable fragement scFv16 was applied to bind the Gαiβγ protein for stabilization32. Gαi, Gβ1-HiBiT, Gγ, and scFv16, were cloned into pFastBac vector (Supplementary Fig. 1a), respectively.
Protein expression and purification
The recombinant A3AR, Gαi, Gβ1-HiBiT, Gγ, and scFv16 were co-expressed in Trichoplusia ni High Five insect cells using the Bac-to-Bac baculovirus expression system. High Five cells were co-infected with the baculovirus at a cell density of 3.5 × 106 cells per milliliter. Fourty-eight hours later, the infected cells were harvested and stored at −80 °C until used.
For the purification of the CF101-A3AR-Gi complex, cells pellets were thawed and resuspended in Buffer A (100 mM NaCl, 20 mM HEPES, pH 7.5) supplemented with protease inhibitor cocktail (TargetMol, C0001). Cells were lysed by dounce homogenization (Sigma-Aldrich, D9188) followed by centrifugation to remove unsoluble materials. The pellets were resuspended in Buffer B (100 mM NaCl, 10 %(v/v) glycerol, 20 mM HEPES, pH 7.5) supplemented with 10 mM MgCl2, 5 mM CaCl2, 0.2 mM Tris-(2-carboxyethyl)phosphine (TCEP, Hampton Research, HR2-801) and protease inhibitor cocktail. We formed the complexes by rotating the samples at room temperature for 1 h after addition of 25 mUnit/mL apyrase and 10 μM CF101 (MedChemExpress, HY-13591). After incubation, the sample was solublized in 0.5 %(w/v) lauryl maltose neopentylglycol (LMNG, anatrace, NG310) and 0.1%(w/v) cholesteryl hemisucinate (CHS, anatrace, CH210) for 3 h at 4 °C. The supernatant was clarified by centrifugation at 100,000× g for 40 min. The supernatant was incubated with dextrin beads 6FF (Smart-Lifesciences, SA02601L) for 3 h at 4 °C. The beads were loaded onto a gravity column and washed with 20 column volumes of Buffer C (100 mM NaCl, 2 mM MgCl2, 10 μM CF101, 0.2 mM TCEP, 0.01 %(w/v) LMNG, 0.002 %(w/v) CHS, 20 mM HEPES, pH 7.5). The complex was eluted with Buffer C supplemented with 10 mM maltose and further concentrated using 100 kDa molecular weight cut-off concentrator. TEV protease was added to the concentrated protein at 4 °C overnight to cleave dual maltose binding protein from fusion protein. After digestion, sample was loaded onto Superdex 200 Increase 10/300 GL column (Cytiva, 28-9909-44) with Buffer D (100 mM NaCl, 2 mM MgCl2, 10 μM CF101, 0.1 mM TCEP, 0.00075 %(w/v) LMNG, 0.00025 %(w/v) glyco-diosgenin, 0.0002 %(w/v) CHS, 20 mM HEPES, pH 7.5). The desired fractions were pooled and concentrated to 5–8 mg/mL for cryo-EM sample preparation. The purification procedures of CF102-A3AR-Gi complex were almost the same as in CF102-A3AR-Gi complex preparation, while the CF101 compounds was replaced by CF102 (TargetMol, T6884).
Cryo-EM data collection
Cryo-EM grids were prepared with the Vitrobot Mark IV plunger (FEI) set to 8 °C and 100% humidity. Three-microliters of the CF101-A3AR-Gi complex were applied to glow- discharged Quantifoil R1.2/1.3 holey carbon grids. The sample was incubated for 10 s on the grids before blotting for 3.5 s (double-sided, blot force 1) and flash-frozen in liquid ethane immediately. The same conditions were used for the CF102-A3AR-Gi complex sample.
For CF101-A3AR-Gi complex, three datasets comprising 20,779 movies were collected on a Titan Krios equipped with a Gatan K3 direct electron detection device at 300 kV with a magnification of 105,000 corresponding to a pixel size 0.824 Å. Image acquisition was performed with EPU Software (FEI Eindhoven, Netherlands). We collected a total of 36 frames accumulating to a total dose of 50 e− Å−2 over 2.5-s exposure.
For CF102-A3AR-Gi complex dataset, two datasets totaling 13,581 movies were collected on a Titan Krios equipped with a Gatan K3 detector at 300 kV with a magnification of 105,000 and pixel size of 0.824 Å, using EPU Software (FEI Eindhoven, Netherlands). Thirty-six frames were collected over a 2.5-s exposure to a dose of 50 e− Å−2.
Image processing
MotionCor2 was used to perform the frame-based motion-correction algorithm to generate drift-corrected micrograph for further processing, and CTFFIND4 provided estimation of contrast transfer function (CTF) parameters33,34.
For the CF101-A3AR-Gi dataset, the previously resolved structure of BAY 60-6583-A2BAR-Gs23 was used as a reference for automatic particle picking in RELION 3.035. Particle picking and extraction yielded 4,550,294 particles after 2D classification clearance, which were imported into CryoSPARC36. Four rounds of 2D classification selected 1,267,837 particles, followed by two rounds of 3D heterogenous refinement giving 982,833 particles. After two additional rounds of 2D classification and two rounds of heterogenous refinement, 271,323 particles were refined to a structure at 3.29 Å global resolution using non-uniform refinement (Supplementary Fig. 2).
For CF102-A3AR-Gi complex dataset, the BAY 60-6583-A2BAR-Gs structure23 was again used for reference-based particle picking. 4,090,959 and 4,833,382 particles were autopicked and extracted from Dataset 1 and Dateset 2, respectively. For Dataset 1, two rounds of 2D classification were used to separate out 1,070,085 particles. Masked 3D classification on the receptor part was used to separate out 175,747 particles that resulted to a clearer density of A3AR. For Dataset 2, two rounds of 2D classification and two rounds of 3D classification were performed to separate out 246,392 particles. After clearance, the remained particles from two datasets were combined and subjected to alignment-free 3D classification. 283,561 particles were remained and transferred in CryoSPARC36. One round of heterogenous refinement yielded a final 102,581 particles were refined to a structure at 3.19 Å global resolution using non-uniform refinement (Supplementary Fig. 3).
Model building
An A3AR structure predicted by AlphaFold2 was used as the starting reference models for receptors building37. Structures of Gαi, Gβ, Gγ and the scFv16 were derived from PDB entry 7EZH38 were rigid body fit into the density. All models were fitted into the EM density map using UCSF Chimera39 followed by iterative rounds of manual adjustment and automated rebuilding in COOT40 and PHENIX41, respectively. The model was finalized by rebuilding in ISOLDE42 followed by refinement in PHENIX with torsion-angle restraints to the input model. The final model statistics were validated using Comprehensive validation (cryo-EM) in PHENIX and provided in the supplementary information, Supplementary Table 1. All structural figures were prepared using Chimera39, Chimera X43, and PyMOL (Schrödinger, LLC.).
NanoBiT assay
To monitor G protein interaction with A1AR, A2AAR, A2BAR or A3AR upon agonist stimulation, a NanoLuc-based NanoBiT enzyme complementation assay was used as previously described44. The C terminus of A1AR, A2AAR or A2BAR was fused to SmBiT, while LgBiT was fused to the N terminus of miniG proteins. The C terminus of A3AR was fused with LgBiT, and the SmBiT was fused to the N terminus of miniG proteins. HEK293 cells were seeded at 4 × 104 cells/well on 96-well plates and co-transfected with adenosine receptor-SmBiT and LgBiT-G protein plasmid. After 24 h, cells were replaced with 40 μL fresh culture medium without fetal bovine serum. Ten microliter Nano-Glo Live Cell reagent was added followed the manufacturer’s protocol (Promega, N2011), and incubated at 37 °C, 5 % CO2 for 5 min. Another 25 μL culture medium containing various concentrations of compounds were added and incubated at room temperature for 10 min before measuring bioluminescence using EnVision multiplate reader (PerkinElmer).
cAMP accumulation assays
HEK293 cells expressing wide-type (WT) or mutant A3AR were harvested and resuspended in DMEM containing 500 μM 3-isobutyl-1-methylxanthine (IBMX) at a density of 2 × 105 cells/ mL. Cells were then plated onto 384-well assay plates at 1000 cells/ 5 μL/ well. Another 5 μL buffer containing 1 μM Forskolin and various concentrations of test compounds were added to the cells. After incubation at room temperature for 15 min, intracellular cAMP level was tested by a LANCE Ultra cAMP kit (PerkinElmer, TRF0264) and EnVision multiplate reader according to the manufacturer’s instructions.
Cell-surface expression assay
The same constructs were used in the cell-surface expression assays, NanoBiT assays, and cAMP measurements. A human influenza hemagglutinin tag (HA-tag) was fused to the N-terminus of the adenosine receptor and mutant gene sequences in the pcDNA3.0 vector constructs used across the various assays. HEK293 cells were transfected with wild type (WT) or adenosine receptor mutants and then were seeded at 4 × 104 cells/well on 96-well plates. After 24 h, cells were washed with PBS buffer, fixed with 4 %(w/v) paraformaldehyde for 15 min, and blocked with 2 %(w/v) bovine serum albumin (BSA) for 1 h. Next, cells were incubated with the polyclonal anti-HA antibody (diluted at a ratio of 1:1,000, Sigma-Aldrich, H6908) overnight at 4 °C, followed by 1 h with horseradish peroxidase (HRP)-conjugated anti-rabbit antibody (diluted at a ratio of 1:10,000, Cell Signaling, 7074S) at room temperature. After washing, 50 μL tetramethylbenzidine (Sigma, T0440) was added for 30 min before stopping the reaction with 25 μL 3,3,5,5 - tetramethylbenzidine (TMB) substrate stop solution (Beyotime, P0215). Absorbance at 450 nm was measured on a FlexStation III microplate reader (Molecular Devices).
Statistical analysis
All functional study data were analyzed in Prism 8 (GraphPad) and presented as means ± S.E.M. from at least three independent experiments. Concentration-response curves were evaluated with a three-parameter logistic equation. pEC50 values were calculated using the sigmoid three-parameter equation. Significance was determined by one-way ANOVA followed by multiple comparisons test, and *P < 0.05 vs. wild-type (WT) was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support this study are available from the corresponding authors upon request. The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-37985 (A3AR-CF101-Gi complex) and EMD-37986 (A3AR-CF102-Gi complex). The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 8X16 [https://doi.org/10.2210/pdb8X16/pdb] (A3AR-CF101-Gi complex) and 8X17 [https://doi.org/10.2210/pdb8X17/pdb] (A3AR-CF102-Gi complex). Previously published structures can be accessed via accession codes 7LD4, 6GDG, 8HDP and 4EIY. Source data are provided with this paper.
References
Salvatore, C. A., Jacobson, M. A., Taylor, H. E., Linden, J. & Johnson, R. G. Molecular cloning and characterization of the human A3 adenosine receptor. Proc. Natl Acad. Sci. USA 90, 10365–10369 (1993).
Sheth, S., Brito, R., Mukherjea, D., Rybak, L. P. & Ramkumar, V. Adenosine receptors: expression, function and regulation. Int. J. Mol. Sci. 15, 2024–2052 (2014).
Borea, P. A. et al. The A3 adenosine receptor: history and perspectives. Pharmacol. Rev. 67, 74–102 (2015).
Jacobson, K. A. et al. A(3) Adenosine Receptors as Modulators of Inflammation: From Medicinal Chemistry to Therapy. Med. Res. Rev. 38, 1031–1072 (2018).
Hauser, A. S. et al. GPCR activation mechanisms across classes and macro/microscales. Nat. Struct. Mol. Biol. 28, 879–888 (2021).
Meyerhof, W., Muller-Brechlin, R. & Richter, D. Molecular cloning of a novel putative G-protein coupled receptor expressed during rat spermiogenesis. FEBS Lett. 284, 155–160 (1991).
Fishman, P., Bar-Yehuda, S., Liang, B. T. & Jacobson, K. A. Pharmacological and therapeutic effects of A3 adenosine receptor agonists. Drug Discov. Today 17, 359–366 (2012).
Fishman, P., Bar-Yehuda, S., Madi, L. & Cohn, I. A3 adenosine receptor as a target for cancer therapy. Anticancer Drugs 13, 437–443 (2002).
Marwein, S., Mishra, B., De, U. C. & Acharya, P. C. Recent Progress of Adenosine Receptor Modulators in the Development of Anticancer Chemotherapeutic Agents. Curr. Pharm. Des. 25, 2842–2858 (2019).
Fishman, P. Drugs Targeting the A3 Adenosine Receptor: Human Clinical Study Data. Molecules 27, 3680 (2022).
Jacobson, K. A. Adenosine A3 receptors: novel ligands and paradoxical effects. Trends Pharmacol. Sci. 19, 184–191 (1998).
Jacobson, K. A. et al. Medicinal chemistry of the A3 adenosine receptor: agonists, antagonists, and receptor engineering. Handb. Exp. Pharmacol. 193, 123–159 (2009).
Barkan, K. et al. Pharmacological characterisation of novel adenosine A(3) receptor antagonists. Sci. Rep. 10, 20781 (2020).
Ogawa, A. et al. N(6)-methyladenosine (m(6)A) is an endogenous A3 adenosine receptor ligand. Mol. Cell 81, 659–674.e7 (2021).
Gallo-Rodriguez, C. et al. Structure-activity relationships of N6-benzyladenosine-5’-uronamides as A3-selective adenosine agonists. J. Med. Chem. 37, 636–646 (1994).
Kim, H. O. et al. 2-Substitution of N6-benzyladenosine-5’-uronamides enhances selectivity for A3 adenosine receptors. J. Med. Chem. 37, 3614–3621 (1994).
Van Schaick, E. A., Jacobson, K. A., Kim, H. O., IJzerman, A. P. & Danhof, M. Hemodynamic effects and histamine release elicited by the selective adenosine A3 receptor agonist 2-Cl-IB-MECA in conscious rats. Eur. J. Pharmacol. 308, 311–314 (1996).
Suresh, R. R. et al. Design and in vivo activity of A(3) adenosine receptor agonist prodrugs. Purinergic Signal 16, 367–377 (2020).
Fishman, P. et al. The A3 adenosine receptor agonist, namodenoson, ameliorates non‑alcoholic steatohepatitis in mice. Int. J. Mol. Med. 44, 2256–2264 (2019).
Jacobson, K. A. & Gao, Z. G. Adenosine receptors as therapeutic targets. Nat. Rev. Drug Discov. 5, 247–264 (2006).
Verzijl, D. & Ijzerman, A. P. Functional selectivity of adenosine receptor ligands. Purinergic. Signal 7, 171–192 (2011).
Duan, J. et al. Cryo-EM structure of an activated VIP1 receptor-G protein complex revealed by a NanoBiT tethering strategy. Nat. Commun. 11, 4121 (2020).
Cai, H. et al. Structures of adenosine receptor A(2B)R bound to endogenous and synthetic agonists. Cell Discov. 8, 140 (2022).
Duan, J. et al. GPCR activation and GRK2 assembly by a biased intracellular agonist. Nature 620, 676–681 (2023).
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011).
Draper-Joyce, C. J. et al. Positive allosteric mechanisms of adenosine A(1) receptor-mediated analgesia. Nature 597, 571–576 (2021).
Tosh, D. K. et al. First Potent Macrocyclic A(3) Adenosine Receptor Agonists Reveal G-Protein and beta-Arrestin2 Signaling Preferences. ACS Pharmacol. Transl. Sci. 6, 1288–1305 (2023).
Jaakola, V. P. et al. The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322, 1211–1217 (2008).
Zhou, Q. et al. Common activation mechanism of class A GPCRs. Elife 8, e50279 (2019).
Mazziotta, C. et al. Cancer biology and molecular genetics of A(3) adenosine receptor. Oncogene 41, 301–308 (2022).
Liu, P. et al. The structural basis of the dominant negative phenotype of the Galphai1beta1gamma2 G203A/A326S heterotrimer. Acta Pharmacol. Sin. 37, 1259–1272 (2016).
Maeda, S. et al. Development of an antibody fragment that stabilizes GPCR/G-protein complexes. Nat. Commun. 9, 3712 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Liu, Q. et al. Ligand recognition and G-protein coupling selectivity of cholecystokinin A receptor. Nat. Chem. Biol. 17, 1238–1244 (2021).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. Recent developments in the PHENIX software for automated crystallographic structure determination. J. Synchrotron. Radiat. 11, 53–55 (2004).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D Struct. Biol. 74, 519–530 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Nehme, R. et al. Mini-G proteins: Novel tools for studying GPCRs in their active conformation. PLoS One 12, e0175642 (2017).
Garcia-Nafria, J., Lee, Y., Bai, X., Carpenter, B. & Tate, C. G. Cryo-EM structure of the adenosine A(2A) receptor coupled to an engineered heterotrimeric G protein. Elife 7, e35946 (2018).
Acknowledgements
We thank Wen Hu, Kai Wu and Qingning Yuan from the Advanced Center for Electron Microscopy (Shanghai Institute of Materia Medica, Chinese Academy of Sciences) for their technical supporting and assistance with cryo-EM dataset collection. This project was partially supported by the CAS Strategic Priority Research Program (XDB37030103 to H.E.X.); the National Natural Science Foundation of China (32301004 to H.C., 82121005 to X.X., Y.J., and H.E.X., 32130022 to H.E.X., 82330113 to X.X., 82304579 to S.G., and 32171187 to Y.J.); Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 to H.E.X.); Shanghai Municipal Science and Technology Major Project (H.E.X.); the Lingang Laboratory (LG-GG-202204-01 to H.E.X.); the National Key R&D Program of China (2022YFC2703105 to H.E.X.); the China Postdoctoral Science Foundation (2021M703341, 2023T160662 to H.C.), the Shanghai Post-doctoral Excellence Program (2021423, H.C.).
Author information
Authors and Affiliations
Contributions
H.E.X., X.X. and H.C. conceived the study. H.C. designed the expression constructs, purified the protein complexes. Y.X. and H.C. prepared the grids and performed the cryo-EM data processing and model building with the help from J.L. H.E.X., H.C. and Y.J. analyzed the structures. S.G. J.S. and Z.X. performed the functional studies under the supervision of X.X. H.C. prepared the figures and manuscript. Y.X. and S.G. contributed to manuscript preparation. H.E.X. and H.C. wrote the manuscript with input from all authors. The authors utilized Claude.ai and ChatGPT to assist with correcting grammatical errors.
Corresponding authors
Ethics declarations
Competing interests
H.E.X. is the founder and chairman of the board of Cascade Pharmaceutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Asuka Inoue and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cai, H., Guo, S., Xu, Y. et al. Cryo-EM structures of adenosine receptor A3AR bound to selective agonists. Nat Commun 15, 3252 (2024). https://doi.org/10.1038/s41467-024-47207-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-47207-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.