Abstract
Photoreceptor proteins utilise chromophores to sense light and trigger a biological response. The discovery that adenosylcobalamin (or coenzyme B12) can act as a light-sensing chromophore heralded a new field of B12-photobiology. Although microbial genome analysis indicates that photoactive B12-binding domains form part of more complex protein architectures, regulating a range of molecular–cellular functions in response to light, experimental evidence is lacking. Here we identify and characterise a sub-family of multi-centre photoreceptors, termed photocobilins, that use B12 and biliverdin (BV) to sense light across the visible spectrum. Crystal structures reveal close juxtaposition of the B12 and BV chromophores, an arrangement that facilitates optical coupling. Light-triggered conversion of the B12 affects quaternary structure, in turn leading to light-activation of associated enzyme domains. The apparent widespread nature of photocobilins implies involvement in light regulation of a wider array of biochemical processes, and thus expands the scope for B12 photobiology. Their characterisation provides inspiration for the design of broad-spectrum optogenetic tools and next generation bio-photocatalysts.
Similar content being viewed by others
Introduction
Photoreceptor proteins are ubiquitous in nature and regulate a large range of biological processes in response to light. They have become essential components for optogenetic applications, where they can be fused to different output domains to provide light control of various cellular functions1. The number of known photoreceptor families in biology has recently been expanded2 by the discovery of a new superfamily of B12 photoreceptors that repurpose coenzyme B12 or adenosylcobalamin (AdoCbl) for light sensing. Cobalamins are complex cobalt-containing tetrapyrrole molecules, in which various forms differ in the nature of the upper axial ligand (e.g., methyl, cyano, hydroxyl or adenosyl) that is ligated to the central Co atom. For a long time, cobalamin has only been known as a widespread organometallic cofactor to many thermally-driven enzymes that catalyse a wide range of processes essential to living organisms, including humans3. The discovery of CarH, the canonical AdoCbl photoreceptor, revealed B12 can support a light-mediated repression of carotenoid biosynthetic genes4. In the dark, the AdoCbl-bound CarH tetramer specifically binds to DNA and blocks transcription. On exposure to light, the photolabile Co-C bond of AdoCbl is cleaved, which results in the release of free 4’,5’-anhydro-adenosine4,5. B12 photochemistry triggers structural changes culminating in the formation of a bis-His ligated light state of CarH and the dissociation of the tetramer, together with the concomitant release from DNA and ultimately transcription initiation4. CarH has already been used in several green light-dependent biotechnological applications, such as in the formation of light-responsive hydrogels for drug delivery6,7,8,9, light-activated technology devices10 and the regulation of mammalian gene expression11. However, the optogenetic potential of B12 photoreceptors would be significantly enhanced by broadening the wavelength range to include the more penetrative red/far-red region of the spectrum. More complex putative B12 photoreceptors have since been identified in genomes, but lack functional characterisation12,13. Sequence analysis reveals many CarH-like homologues form considerably more complex protein architectures, with the B12-binding domain fused to additional enzyme or chromophore-binding domains12. Based on these previous findings, we describe a new subfamily of B12-dependent photoreceptors, consisting of a B12 binding domain fused to a globin-like (or photoglobin6) domain. The hypothesis that these coupled photoglobin and B12-binding domains may act as a light-sensing regulatory bundle13 provides exciting opportunities to develop new optogenetic tools. We now demonstrate that these proteins can simultaneously bind B12 and the linear tetrapyrrole biliverdin (BV), and are sensitive to light across the entire visible spectrum (Fig. 1). Hence, we propose these form a new family of photocobilins (photoactive-cobalamin-bilin, Pcob) that can mediate light-regulation of a range of processes.
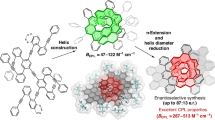
Schematic showing the structures of the adenosylcobalamin (AdoCbl or coenzyme B12) and biliverdin (BV) chromophores in photocobilin, together with the wavelength range over which they absorb. The photocobilins are found as both standalone proteins or fused to other domains (a diguanylate cyclase or DGC domain is shown). The nature of the upper R ligand in B12 is responsible for the different B12 analogues (R=adenosyl, AdoCbl; = CH3, methylcobalamin; = OH, hydroxcobalamin; =CN, cyanocobalamin). The protein (surface is coloured blue and orange by chains) and absorbance spectrum shown in the figure is from the SasPcob protein with bound AdoCbl and BV under dark conditions.
Results and discussion
Photocobilin proteins exhibit spectral changes upon light activation
We have identified and characterised two types of photocobilins, one from Saccharothrix syringae (SasPcob) that represents a standalone photocobilin photoreceptor and a more complex protein that consists of a photocobilin diguanylate cyclase (DGC) fusion from Acidimicrobiaceae bacterium (AbDPcob) that is representative of a photocobilin domain fused to an enzyme domain (Fig. 1 and Supplementary Figs. 1 and 2). Both proteins are able to bind AdoCbl and BV, although there are significant differences in BV binding between the two proteins (Figs. 2a, b, 3). In SasPcob, the ternary AdoCbl-BV complex is readily formed when both chromophores are provided. AdoCbl binding allosterically affects BV affinity, with the BV dissociation constant decreasing ~10-fold from Kd ~ 7 µM to ~0.7 µM in the presence of AdoCbl (Fig. 3 and Supplementary Fig. 3). In the full-length AbDPcob, full occupancy of AdoCbl is observed in the absence of BV, but the binding of both chromophores simultaneously is diminished compared to the SasPcob (Fig. 3 and Supplementary Fig. 3). The weaker binding and lower occupancy levels (~20%) of BV in AbDPcob suggest that evolutionary pressures may have reduced the requirement for BV and resulted in a different role for the globin-like domain in these more complex fusion proteins. Despite these differences, both the standalone photocobilin (SasPcob) and diguanylate cyclase fusion (AbDPcob) show similar light-sensing behaviour (Fig. 2c, d and Supplementary Fig. 4a, b). Illumination of AdoCbl-only bound photocobilin with red light (660 nm LED) has no effect (Fig. 2 and Supplementary Fig. 4), whereas illumination with green light (530 nm LED) elicits spectral changes similar to those observed previously with CarH14. These include the formation of new absorbance features at 356 nm and 500 nm, together with the disappearance of the absorbance band at approximately 570 nm. In contrast, the BV-only bound species does not display any obvious spectral changes after illumination with green or red light (Fig. 4a, b). However, the ternary AdoCbl-BV-photocobilin complexes respond to both green and red light, with excitation of the BV chromophore leading to spectral changes associated with Co-C bond cleavage in the B12 cofactor (Fig. 2 and Supplementary Fig. 4). Hence, the two chromophores are optically coupled and excitation of either can trigger AdoCbl photoconversion, thereby expanding light sensitivity to cover most of the visible spectrum. The relative efficiency of conversion to the light state is higher for the direct AdoCbl excitation route (Supplementary Fig. 5). The ability to respond to red light is lost when AdoCbl is replaced with methylcobalamin, which contains a smaller methyl group as upper axial ligand (Fig. 4c−h). This indicates that the upper adenosyl ligand is required for optical coupling and cleavage of the upper axial C-Co bond. However, the exact mechanism of coupling between the 2 chromophores is currently not clear and will require further characterisation.
a absorbance spectra of SasPcob with either BV, AdoCbl or both bound. b absorbance spectra of AbDPcob with either BV, AdoCbl or both bound. Difference absorbance spectra of photocobilin photoreceptors after illumination with green light (c) or red light (d) for SasPcob. Interaction of BV and AdoCbl molecules in SasPcob in dark (e) and light (f) state. The Rossmann fold region is coloured blue, four-helix-bundle region in green and BV binding region in red. The protein main scaffold is shown as cartoon and coloured as above. The opposite monomer is shown as grey cartoon. Main residues involved in the binding are shown as sticks. The AdoCbl and BV molecules are shown as pink and yellow sticks, and the upper ligand of AdoCbl is shown as cyan sticks, respectively. The N and O atoms in the sticks are coloured blue and red, respectively. The cobalt and water molecules as pink and red spheres, respectively. The polar interactions are shown as yellow dash and labelled with distance (angstrom).
a The binding ratio (chromophore:protein) of B12 and BV for SasPcob. The binding ratio was calculated based on the chromophore and protein molar concentrations. Ordinary one-way ANOVA F-test with multiple comparisons was performed, F (3, 8) = 11.3, P = 0.0021 **P < 0.01, n = 3 independent experiments. b The binding ratio of B12 and BV for AbDPcob protein. In (a, b) values are expressed as mean ± SD. Ordinary one-way ANOVA F-test with multiple comparisons was performed, F (9, 20) = 51.23, P < 0.0001, ****P < 0.0001, n = 3 independent experiments. c The absorbance increase of AdoCbl at 540 nm at increasing concentrations of SasPcob was used to calculate the binding constant (Kd) for AdoCbl to the protein. The absorbance increase of BV at 700 nm at increasing concentrations of SasPcob was used to calculate the (Kd) for BV in the absence of AdoCbl (d) and with AdoCbl pre-bound (e). f BV titration with AbDPcob. BV was kept at 3.5 μM and AbDPcob ranging from 12 to 80 μM. All measurements were repeated 3 times for data collection. All data are presented as mean values ± SDs, n = 3 independent experiments. Source data are provided in Source Data file.
a Absorbance spectra of SasPcob with only BV bound illuminated at 530 nm. b Absorbance spectra of SasPcob with only AdoCbl bound illuminated at 660 nm. Difference absorbance spectra of SasPcob with methylcobalamin bound after illumination with green (c) or red (d) light. Difference absorbance spectra of SasPcob with methylcobalamin and BV bound after illumination with green (e) or red (f) light. Difference absorbance spectra of AbDPcob with methylcobalamin and BV bound after illumination with green (g) or red (h) light. A dark spectrum was collected prior to any illumination and used as the blank in each case. The up and down arrows indicate the increase and decrease in absorbance, respectively.
X-ray crystallography provides a structural rationale for photoactivation in photocobilins
To provide a structural rationale for the photocobilin photo-sensing properties, we determined the crystal structures of the dark (PDB 8J2W) and light-adapted states (PDB 8J2X) of SasPcob, as well as the dark state of the isolated AbDPcob photoreceptor domain (AbPcob, PDB 8J2Y), at 1.7 Å;, 2.0 Å; and 2.3 Å;, respectively (Figs. 2e, f, 5a−d, Table 1 and Supplementary Fig. 6). Unfortunately, we were unable to obtain crystals of the full length AbDPcob protein or the light state of AbPcob. Despite the low sequence identity (~34%), the structures of the two photocobilin proteins are similar, with an RMSD value of 1.63 Å for 464 C-alphas after structural alignment. Both the SasPcob and AbPcob proteins contain the W-(10)x-EH and E/DxH motifs (Fig. 5e) in the B12-binding domain, which is found in other light dependent CarH-like B12 photoreceptors15. The arrangement and dimerisation of the B12 binding domains, which consist of a Rossmann fold and associated four-helix bundle, is similar to the corresponding CarH dimer core module (RMSD = 1.37 for 237 C-alphas), with the dark state SasPcob and AbPcob forming head-to-tail dimers4 (Fig. 5d, e). AdoCbl binding is also reminiscent of CarH, with the dimethylbenzimidazole tail embedded in the Rossmann fold (Figs. 2e, f, 6a), while the conserved His108 (SasPcob numbering) provides the lower axial ligand to the Co atom of the B12 cofactor (i.e., base-off His-on ligation, Fig. 7a−c). The adenosine binding pocket is strikingly similar, with the ribose moiety stacking with the conserved Trp61 residue and forming a network of polar interactions linking across the dimer interface via the conserved Glu71-His72 motif (Figs. 2e, f, 6a). The Pcob specific C-terminal five-helix globin domain does not contribute directly to the dimer interface, and is positioned such that it forms interactions with both of the B12-binding domains (i.e., Rossmann fold and four-helix bundle). This arrangement effectively shields much of the corrin ring edge that is normally solvent exposed when bound by the canonical CarH domain.
Overall structure of the dark states of the AbPCob (a) and SasPcob (b) protein, and the light state of SasPcob (c). The Rossmann fold region is coloured blue, four-helix-bundle region in green and BV binding region in red. The AdoCbl and BV molecules are shown as pink and yellow spheres. d Structural alignment of SasPcob, AbPcob and CarH proteins (5C8A). The B12 binding region is shown as green (SasPcob), cyan (AbPcob), and magenta (CarH) ribbons. The BV binding domain is shown as yellow (SasPcob) and blue (AbPcob) ribbons. e Sequence alignment of SasPcob, AbPcob and CarH proteins. The alignment result is coloured according to sequence identity by MView56. The residues in the red box are the key residues at the dimer interface. f Structural alignment of BV binding domain of SasPcob (green) and AbPcob (cyan). The BV molecule is shown as red sticks. g Structural alignment of BV binding domains of SasPcob (green) and AbPcob (cyan) with phycobilisome proteins 1B3320 (yellow) and 3MWN21 (magenta). BV molecules and key residues involved with BV binding are shown as red (SasPcob), cyan (AbPcob), yellow (3MWN), light-blue (1B33) sticks. h Comparison of the BV chromophore in the binding pocket. Cys residues bonding with BV (Top, 1B33 and 3WMN), and residues at same position (bottom, SasPcob and AbPcob) are shown as sticks. The same colour scheme is used as in (g). The BV is shown in red for SasPcob, yellow for 1B3311 and blue for 3MWN12. i Sequence alignment of photocobilins with 3MWN and 1B33. The residues in the red box forms the BV binding motif.
AdoCbl (a) and and BV (b) binding in SasPcob. Protein residues are shown as orange sticks. Residues from the other protein chain are shown as blue sticks. AdoCbl and BV molecules are shown as yelllow and pink sticks, respectively, and water molecules as blue spheres. Structurally-relevant water molecules are shown as green spheres. Salt-bridges and their centres are shown as yellow dashes and cyan spheres, hydrophobic interactions as a blue dash and hydrogen bonds as a red dash. c–f Clustered AbPcob protein structures after MD simulations, showing possible binding pose with BV. The grey cartoon and sticks are the representations for SasPcob as comparision. BV molecule and its binding region in AbPcob are coloured as yellow sticks and red cartoon. Residues involved with BV binding are shown as red sticks. The blue dashes indicate hydrophobic interactions between BV and AbPcob protein residues. Comparison of BV binding in AbPcob (g, modelled) and SasPcob (h, crystal structure). Binding poses in cluster 6 were selected for comparison (see Supplementary Table 1 for clusters summary). BV and B12 molecules are shown as above. Residues around BV with 4 Å are shown as red (AbPcob) and grey (SasPcob) sticks.
AdoCbl binding in TtCarH (a), AbPcob (b) and SasPcob (c) are shown as line and sticks. AdoCbl molecules are shown as magenta and upper ligands as cyan. Residues involved with binding are shown as blue and green sticks. The residues in the opposite monomer are shown as grey. Overall structural changes in TtCarH (d) and SasPcob (e). The protein is shown as a cartoon. The main monomer in dark state as blue and light state as cyan. The opposite monomer as grey. The AdoCbl molecules are shown as sticks with dark state as blue and light state as cyan. Upper adenosyl group is shown as red stick. BV molecules in SasPCob are shown as grey spheres. Main residues involved in the change are shown as blue and cyan sticks for dark and light state. The arrow indicates the direction in which the protein moves, while the length of the arrow represents the scale of the movement. The circular perspective provides a more detailed observation of the dynamic motion of residues within dark and light structure. Residues in the opposite monomer are displayed as grey sticks and red dots. f Representation of photochemical changes in Pcob protein. The AdoCbl and BV molecules are shown as pink and yellow sticks, adenosyl as cyan sticks. Red, green and blue blocks represent for four helix bundle, Rossmann fold and BV binding regions in the Pcob protein.
In SasPcob, the BV is bound with a ZZZ configuration, with Asp270 forming key interactions with the co-planar A, B and C pyrrole rings (Fig. 6a, b). In comparison to other BV-binding photoreceptors, notably the bacteriophytochromes, the photocobilins lack the Cys residue that covalently links to the BV molecule and is essential for light-induced cis-trans isomerisation16,17,18 (Figs. 2e, f and 5f−i). Despite the lack of cysteinyl-linkage, the structure of the SasPcob BV-binding pocket is akin to light-harvesting phycobilisome proteins, with the BV orientation flipped by ~180° (Fig. 5g), projecting the D ring out of the globin core, as opposed to the A ring in other bilin binding proteins19,20,21. The D ring is positioned out of the ABC plane by approximately 40° due to close interactions with Trp296 (Figs. 5g and 6a, b). Although the putative BV binding globin scaffold is retained in AbPcob, the reduced BV affinity is most likely due to replacement of a key BV binding residue, Asp270 in SasPcob, with a His (Fig. 5f−i). The crystal structure of AbPcob revealed that steric hindrance by this His residue interferes with BV binding, unlike in SasPcob. Molecular dynamics simulations and ligand docking (Supplementary Figs. 7–9, Supplementary Table 1 and 2, and Supplementary Data 1) demonstrate that rearrangement of surrounding residues could allow BV to bind with AbPcob in certain conformations, where hydrophobic interactions with residues in the binding pocket (e.g., Leu252, Trp289, Leu294, Arg297, Val299 and Val303) contribute to retain BV in the correct orientation (Fig. 6c−h). The lack of the main binding interaction with the Asp residue in AbPcob may explain the absence of any obvious spectral shift and increase in absorbance at 700 nm upon BV binding that is observed for SasPcob (Fig. 2). These findings could explain the lower BV occupancy levels for AbPcob whilst retaining the ability to respond to red light (Fig. 3 and Supplementary Fig. 4).
In SasPcob, the close interaction and relative position of the B12- and BV-binding domains brings the two chromophores in close proximity, with the edge of the BV C and D rings within van der Waal’s distance of the conserved Trp61 and the corrin ring B, respectively. Furthermore, a direct hydrogen bond is formed between one of the corrin ring B amide groups and the BV ring D (Fig. 2e, f). This likely explains the B12 allosteric effects on BV affinity observed in solution. A network of direct and water mediated polar interactions surround this central chromophore edge-to-edge contact. These include direct hydrogen bonds between the corrin ring and Trp296 from the globin domain, and a putative salt bridge formed between the propionic acid of ring C of BV and His137 of the opposite monomer (Fig. 2e). The juxtaposition of both chromophores likely underpins the optical coupling observed.
In order to investigate the effects of illumination on the photocobilins, we determined the light-exposed SasPcob crystal structure (Figs. 5, 7). Similar to CarH, the SasPcob light state corresponds to a monomeric species (Supplementary Fig. 1), with the B12-binding domain dimer interface disrupted by the light-triggered release of the adenosine group (Fig. 7 and Supplementary Figs. 10−21). However, the SasPcob light state structure lacks the drastic large-scale domain repositioning that accompanies formation of a bis-His ligated cobalamin in CarH. Instead, more modest changes occur at the adenosine binding pocket in SasPcob culminating with the formation of a water/hydroxide ligated Co(III)balamin. While the majority of the Rossmann and globin binding domains are relatively unperturbed, the N-terminal four helix bundle adapts to the removal of the adenosine through changes in the position of the Glu71/His72 containing helix. The C-alpha displacement of ~2.4 Å at the Glu71/His72 region is such that, when superposed on the corresponding dark SasPcob dimer, it would lead to a severe clash at the dimer interface (Fig. 7c). Therefore, we postulate that rearrangement of the adenosine binding pocket in response to illumination occurs concomitantly with monomer formation.
Light-induced functional changes in photocobilins
To investigate how the photocobilin light-response can affect functional change, we investigated the activity of the AbDPcob DGC domain. This domain catalyses the formation of cyclic-di-GMP, involved in regulating a number of cellular processes22,23,24, from GTP with the release of pyrophosphate. We successfully expressed the full-length AbDPcob protein and measured DGC activity through the detection of both cyclic-di-GMP and pyrophosphate (Fig. 8 and Supplementary Tables 4 and 5). Initially, the production of cyclic-di-GMP, confirmed by LC-MS, was found to increase upon illumination when compared to the apo (i.e., no chromophore bound) and AdoCbl-bound dark states of the protein (Fig. 8a). Steady-state kinetics were measured using a pyrophosphate assay for the apo and AdoCbl-bound dark/light states of the AbDPcob protein, as well as the truncated DGC domain only (Fig. 8b). Although the Michaelis constant, Km, for GTP is similar for all protein forms, the apparent turnover number, kcat, varies significantly and suggests major differences in DGC activity (Fig. 8b). All full-length AbDPcob forms are more active than the isolated DGC domain, indicating that the presence of the photocobilin domain plays a role in maintaining the DGC activity. The binding of AdoCbl to the full-length protein results in a decrease in DGC activity compared to the apo protein. However, upon illumination, there is a > 10-fold increase in apparent turnover number, implying that photoconversion of the AdoCbl-bound photocobilin domain to the light state activates the enzymatic activity of the neighbouring DGC domain. Simple replacement of the AdoCbl with the smaller upper ligand found in methylcobalamin, hydroxycobalamin or cyanocobalamin does not increase DGC activity (Fig. 8c). This implies that the presence of the adenosyl group or changes mediated through AdoCbl photochemistry that lead to the removal of the adenosyl group are important in this dark-inhibition and light-activation process. The oligomerisation state of the protein is not affected by GTP binding so this process is light-mediated (Supplementary Fig. 22).
a LC-MS chromatograms showing the production of cyclic-di-GMP for the apo, dark and light states of the AbDPcob protein (cps, counts per second). Cyclic-di-GMP standard was used for identifying the product. b Michaelis-Menten plots showing the rate of cyclic-di-GMP formation for AbDPcob protein under different conditions and the truncated DGC domain only. The rates were normalised for enzyme concentration and fitted to the Michaelis-Menten equation to determine the kinetic parameters. c DGC activity of AbDPcob upon addition of different cobalamin cofactors under dark and light condition. Ordinary one-way ANOVA F-test with multiple comparisons was performed, F (7, 16) = 804.7, P < 0.0001, ***P < 0.001, ****P < 0.0001, n = 3 independent experiments. d The rates of cyclic-di-GMP formation by AbDGC upon addition of the dark or light state of different photocobilin domains (upper panel, including AdoCbl as a control) or different concentrations of the AdoCbl-bound AbPcob domain (lower panel). The hatch patterns indicate samples for light-adapted states. For upper panel, ordinary one-way ANOVA F-test with multiple comparisons was performed, F (8, 18) = 85.38, P < 0.0001, **P < 0.01, ****P < 0.0001, n = 3 independent experiments. For lower panel, ordinary one-way ANOVA F-test with multiple comparisons was performed, F (6, 14) = 4024, P < 0.0001, **P < 0.01, ****P < 0.0001, n = 3 independent experiments. All data are presented as mean values ± SDs, n = 3 independent experiments. Source data are provided in Source Data file.
Our attempts to crystallise the full-length AbDPcob protein were unsuccessful, most likely because of the long flexible linker between the photocobilin and DGC domains. To understand the photo-activation process in more detail, we performed small-angle X-ray scattering (SAXS) measurements using the ‘dark’ and ‘light’ states of the full-length protein (SASBDB SASDUS3 and SASDUT3 respectively). The results show a significant difference in the scattering signal between the dark and light state (Fig. 9a−c). From the SAXS signals the radius of gyration Rg and the extrapolated intensity at q → 0 I(0) can be extracted. Upon illumination Rg increases from 42.1 Å (dark) to 52.2 Å (light), thus revealing an expansion of the protein, while I(0) remains almost unchanged (1.03 × 10³ and 1.08 × 10³ for dark and light state, respectively), indicating that the protein remains in its dimeric form. Fourier transform of the SAXS signals produces the signal distribution function p(r), which reveals a transition from a compact single peaked distribution for the dark state to a larger multidomain conformation for the light state (Fig. 9b). Low-resolution ab initio modelling based on the p(r) distributions shows that the AlphaFold225 prediction for AbDPcob is compatible with the dark conformation in solution, while the light state is characterised by an extended space arrangement largely different with respect to the dark state (Fig. 9c and Supplementary Fig. 23). Molecular dynamics simulations were conducted to compare dynamics of AbDPcob in the light and dark states (Supplementary Figs. 7–9). AbDPcob showed larger conformational change in the light state compared to that in the dark state indicated by structural deviation (RMSD) from the same starting structure (Supplementary Fig. 8). Specifically, the Pcob domain stays more stable than the DGC domain in both light and dark state. The DGC region was observed to show higher flexibility in the light state particularly at the interface with the Pcob region (Supplementary Fig. 7). Further analysis revealed an obvious opposite movement of the Pcob and DGC domains in the light state but not in the dark state (Fig. 9d, Supplementary Fig. 23 and Supplementary Movie 1 and 2). This indicates that photoactivation of AbDPcob is modulated by local protein conformation changes.
a SAXS signals of AbDPcob in the dark state and after illumination at 530 nm for 10 min. The inset shows the difference between the two signals. b The distance distribution p(r) reveals an expansion of the protein upon illumination. c Low-resolution ab-initio modelling based on the SAXS data provides the envelope of the protein electron density distribution for both dark and light state. Results in (c) show that the AlphaFold2 prediction (orange structure) is compatible with the dark structure in solution, while the light state is characterised by an extended conformation. d Superposition of AbPcob in dark and light state form in molecular dynamics simulation results of AbDPcob. AlphaFold2 predicted dimer was used as the starting model. The final dark and light state structures are aligned in Pymol. Protein surface is shown according to domain arrangement. In dark state, B12 domain is shown as lemon-green, BV as light-pink and DGC as light-blue. In light state, B12 domain is shown as green, BV as hot-pink and DGC as marine-blue. The linker regions in both dark and light state are shown as grey. The arrows in the figure indicate the protein movement upon conversion from the dark to the light state. The SAXS data collection parameters are shown in Supplementary Table 3.
Although a clear photoactivation role has been established for the photocobilin domain fused to the DGC enzyme domain, the function of the standalone photocobilin photoreceptors is less clear. To study this aspect, we measured the activity of the isolated AbDGC domain upon mixing with an equal molar ratio of either of the two photocobilin proteins, isolated AdoCbl-bound forms of AbPcob or SasPcob (with and without bound BV) (Fig. 8d). Similar to the previous findings, the light-adapted states of the B12-bound AbPcob and the holo SasPcob (i.e., both AdoCbl and BV bound) activate AbDGC catalysis. The increase in enzyme activity is dependent on the concentration of the photocobilin protein (Fig. 8d), suggesting a potential role for protein-protein interactions in the activation process. Surprisingly, in the absence of BV, the AdoCbl-bound SasPcob dark state activates AbDGC activity, whereas the light state appears to inhibit activity (Fig. 8d). Further evidence that the two proteins interact is provided by size-exclusion chromatography multi-angle light scattering measurements, which show dissociation of the AbDGC dimer into monomers upon addition of the photocobilin domain (AbPcob). (Supplementary Fig. 2). Higher concentrations of the photocobilin domain give rise to the appearance of a higher molecular weight species, which is likely to represent a heterodimer between the photocobilin and the AbDGC protein. Taken together, these results suggest that the standalone photocobilin photoreceptors can also regulate protein function through protein:protein interactions (PPIs), a mechanism that could potentially influence a wide range of metabolic pathways.
A wider role for photoglobins in cellular processes
Previous bioinformatics approaches have shown that photoglobin domains can be fused to several other types of functional domain and have proposed that coupled photoglobin and B12-binding domains (i.e., photocobilins) may be involved in the regulation of different enzymes and transcription factors13. We have now shown that the photocobilins have the ability to regulate enzyme activity through PPIs. As such, we have investigated potential PPI networks to explore possible biological functions in cell metabolism (Supplementary Fig. 24). In Saccharothrix species, SasPcob potentially interacts with STAS (Sulphate Transporter and AntiSigma factor antagonist) domain, Stage II sporulation protein E (SpoIIE) and Sigma factor PP2C-like phosphatases (Supplementary Figs. 24 and 25a), suggesting a putative role in regulation of the cell stress response. Other photocobilins were frequently involved in one carbon pool by folate, amino acid metabolism and purine biosynthesis, suggesting potentially broad participation in cell metabolism (Supplementary Figs. 24 and 25a). Sequence similarity networks shows there are a wide range of photocobilin photoreceptor proteins with a high similarity of overall sequence, structure, motifs and domain architecture, either fused to other enzyme domains or as standalone proteins (Supplementary Figs. 26−30). A comparison of our SasPcob structure with other photocobilin proteins (predicted structures obtained from the recently published Alphafold2 database26), shows they can be divided into three main groups depending on the location of the AdoCbl-binding domain at either the N- or C-terminus, and whether they contain the additional output domain in the fused proteins (Supplementary Figs. 25 and 31). The majority of the predicted structures align well with SasPcob (average RMSD = 1.27 Å). Some of these putative photocobilin proteins have variable lengths of the linker region between the AdoCbl- and BV-binding domains, which may affect the arrangement of the overall scaffold, while a small number of the fusion proteins lack a complete photocobilin domain. As the Alphafold2 structural models only show the monomeric form of the protein it is likely that these structures may differ in the higher order oligomeric form of the protein.
Conclusions
In conclusion, we have identified and characterised a new sub-family of AdoCbl-dependent photoreceptors that uniquely use two different chromophores to expand the wavelength range of B12 for sensing and responding to light. The close proximity of the B12 and BV cofactors in the photocobilins allows interaction and ‘cross-talk’ between them, meaning excitation of BV triggers B12 photochemistry. This leads to structural change in the photocobilin protein that can promote enzyme activity either through direct fusion to the enzyme domain or by protein-protein interactions. The photocobilins are likely to have multiple functions in regulating cell metabolism and provides further light-sensing roles for B12 in biology. The unique coupling of B12 and BV in these novel photoreceptor proteins now opens a window to their exploitation as new optogenetic components.
Methods
Protein expression and purification
The proteins were expressed and purified as previously described with slightly modifications27. The potential photoreceptor DNA sequences were selected, codon optimised and synthesised by GeneArt® (ThermoFisher) company. Synthesised genes were sub-cloned into a pET21a (Novagen) vector with a C-terminal 6·His tag. The recombinant plasmid was transformed into E. coli BL21(DE3) for protein expression. Different IPTG concentrations and auto-induction LB medium were used to optimise soluble expression. The large-scale protein expression was carried out with auto-induction LB media (FormediumTM, glucose/lactose ratio 1:4) containing 50 μg/mL ampicillin. After 24 h incubation at 25 °C, cells were harvested by 10 min centrifugation at 6000 g, 4 °C. Harvested cells were resuspended in 20 mM HEPES pH 7.0, 500 mM NaCl, 25 mM imidazole (lysis buffer) supplemented with protease inhibitor cocktail and then lysed by a cell disruptor at 25 kpsi (Constant Systems). The cell lysate was centrifuged at 20,000 rpm (48,000 × g) for 1 h at 4 °C to remove cell debris. The supernatant was collected and loaded onto a HisTrap column (Cytiva). After washing with cell lysis buffer, bound protein was eluted with 20 mM HEPES pH 7.0, 500 mM NaCl, and 250 mM imidazole. The peak fractions were collected and incubated with ligands (either AdoCbl, MeCbl or BV) for at least 2 h at 4 °C under dark conditions. The sample was then loaded on to a size exclusion column (HiLoad 16/600 Superdex 200) to remove free ligands and further purification. Absorbance spectra were used to confirm ligand binding and protein fractions with ligand bound were collected for further experiments. The CarH protein was expressed and purified for photoproduct determination (Supplementary Figs. 10−21) as described27.
Crystallisation, data collection, and structure determination
Purified proteins were exchanged into 20 mM HEPES pH 8.0, 150 mM NaCl and concentrated to 20 mg/ml. Crystallisation was performed in the dark using the sitting drop vapour diffusion technique (200 nL crystallisation reagent mixed with 200 nL protein). The preparation of the dark sample was conducted in the dark with dimmed red light. In order to achieve full conversion to the final light-adapted state, the protein samples were exposed to a 530 nm LED light for a duration of 5 min. Absorbance spectral analysis was performed to ensure complete and effective illumination. The SasPcob dark crystals were obtained from LMB crystallisation screen B7 with 38% v/v 1,4-dioxane. The SasPcob light crystals were obtained from 0.2 M Potassium citrate tribasic monohydrate with 20% w/v PEG 3350. The AbPcob crystals were obtained from 8% w/v PEG 20000, 8% v/v PEG 550 MME, 0.1 M sodium acetate, pH 5.5, 0.2 M potassium thiocyanate. Crystals were cryo- protected by the addition of 20% PEG 200 to the reservoir solution and flash frozen in liquid nitrogen. Individual datasets were collected from single crystals at beamlines i03 (Diamond Light Source). All data were indexed, scaled and subsequently integrated with Xia2 Dials. Structure determination was initially performed by molecular replacement in Phaser28 using a search model generated by Alphafold225. A combination of automated and manual rebuilding and refinement in Refmac29 and COOT30 were used to produce the refined models. Chromophore ligands were obtained from Refmac monomer library by 3 letter code and refined using COOT30 and Refmac29. Validation with both Molprobity31 and PDB_REDO32 were integrated into the iterative rebuild process. Complete data collection and refinement statistics are available in Table 1. The atomic coordinates and experimental data have been deposited in the Protein Data Bank (www.pdb.org). All figures were made using open-source PyMOL 2.5 software.
LED illumination and absorbance spectroscopy
Absorbance spectra were collected using a Cary 60 spectrophotometer (Agilent Technologies). All measurements were carried out under dimmed red light. The dark spectrum was collected prior to any illumination. A TDS3032C 300 MHz Digital Phosphor Oscilloscope (Tektronix) and TGP110 10 MHz Pulse Generator with Delay (Thurlby Thandar Instruments) were used to generate a 100 ms LED (Thorlabs Inc.) pulse at either 530 nm (green light) or 660 nm (red light). After each LED pulse, the spectrum was collected until there were no further significant changes. The difference spectra were obtained by subtracting the dark spectrum from the illuminated spectrum. Data were normalised and plotted using Origin 9.0 software (OriginLab, Northampton, MA).
B12 and BV binding measurements
The binding ratio of B12 and BV ligands was calculated based on their extinction coefficient. AdoCbl, 8.0 mM−1•cm−1 at 540 nm33. MeCbl, 7.7 mM−1•cm−1 at 540 nm34. BV, 90 mM−1•cm−1 at 700 nm35. The protein concentration was determined by utilising the ProtParam tool36 to calculate its theoretical extinction coefficient derived from its protein sequence. SasPcob, 55.46 mM−1•cm−1 at 280 nm. AbDPcob, 63.91 mM−1•cm−1 at 280 nm. The binding constant was measured by monitoring the spectral changes at increasing protein concentrations. The absorbance change at 540 nm was plotted for B12 binding and the absorbance change at 700 nm was plotted for BV binding. With tight binding ligand, the binding constant (Kd) is much lower than ligand concentration used in the titration assay, tight binding equation (Eq. 1) should be used to get a more accurate Kd value. BV binding data were fitted to the tight binding equation (Eq. 1) and AdoCbl binding data to apparent binding equation (Eq. 2) to obtain the Kd value using Origin 9.0 software (OriginLab, Northampton, MA).
where [EL] is the concentration of the enzyme and ligand complex formed, [E] is the enzyme concentration and [L] is the concentration of the ligand, c is the constant.
Diguanylate cyclase activity assay
The diguanylate cyclase activity was determined by using both liquid chromatography–mass spectrometry (LC-MS) and a pyrophosphate assay. For LC-MS, the reaction was prepared with 10 µM enzyme, 200 µM GTP in 20 mM HEPES buffer, pH 6.8, 150 mM NaCl, 10 mM MgCl2.The reaction was carried out at 30 °C for 5 min, then deactivated at 95 °C for 5 min. The dark state reaction was carried out in a black tube and the light state reaction in a transparent tube after illumination with white light. All samples were then cooled down on ice and centrifuged for 15 min at 15,000 rpm. Samples were diluted 50 times with HPLC grade water and transferred to LC-MS glass vials for quantification on a Waters ACQUITY Xevo TQ-S UPLC–MS/MS (Waters Corporation) under negative mode (ESI-). The analysis was carried out using an Acquity UPLC HSS T3 column (1.7 µm, 50 × 2.1 mm) with an optimised gradient programme. The mobile phases were as follows; A: 10 mM ammonium acetate in water contains 0.1% acetate acid (v/v); B: methanol; flow rate: 0.6 mL/min; column temperature: 30 °C. The gradient starts from 97% A and hold for 0.5 min then decreased to 2% in 0.3 min, hold the 98% B for 0.4 min then returned to 97% A in 0.1 min and equilibrated for 0.7 min for waiting another injection. The detection of cyclic-di-GMP was optimised by employing the multiple reaction monitoring transition of 690.9 > 152.0 to quantify the yield of the reaction. The mass spectrometer parameters were as follows: source desolvation temperature of 600 °C and source ion block temperature of 150 °C were used; Nitrogen (≥95%) desolvation gas was set at 1000 L/h, and nebuliser gas at 7.00 Bar; Argon (zero grade) collision gas flow was set at 0.15 mL/min. The system was trained with a series of standard solutions of cyclic-di-GMP and the limit of detection limit was determined. The quantification standard curve was obtained ranging from 6 nM to 20 µM.
The EnzCheck pyrophosphate assay kit (ThermoFisher) was used to follow the kinetics of diguanylate cyclase activity. The reaction was carried out with 1−10 µM enzyme, 0−100 µM GTP in 20 mM HEPES buffer, pH 6.8, 150 mM NaCl, 10 mM MgCl2. The reaction mixture was incubated in the dark or by illumination with a 530 nm LED for 3 min at 30 °C, prior to the addition of GTP to initiate the reaction. Pyrophosphate standard (0−60 μM Na4P2O7) in the same buffer condition was used to generate the standard curve for accurately quantifying pyrophosphate concentration. The pyrophosphate produced was measured and quantified and then converted to cyclic-di-GMP production rate (two molecules of pyrophosphate equals to one molecule of cyclic-di-GMP). All data were collected in triplicate and plotted against GTP concentration. The Km was obtained by fitting data into the Michaelis-Menten equation (Eq. 3).
SAXS data collection and analysis
Static X-ray scattering data were collected at 20 °C at the BM29 BIOSAXS beamline of the ESRF Synchrotron (Grenoble, France). X-ray solution scattering signals in the SAXS region (q = 0.0025−0.5 Å−1) were collected on AbDPcob at 5 mg/mL concentration using a monochromatic X-ray beam (double multilayer monochromator) centred at 12.5 keV and a Pilatus3 2 M (Dectris) detector. Two datasets were collected, before and after irradiation of the AbDPcob solution with a LED light source at 530 nm for 10 min. For each dataset, 10 X-ray scattering signals (each registered with 1 s of X-ray exposure) were collected with 10 analogous signals of the buffer before and after each protein measurement. Each two-dimensional pattern was converted to a one-dimensional scattering profile by azimuthal integration. Corresponding scattering profiles were averaged and protein signals were obtained by subtraction of the buffer signal. The distance distribution functions p(r) were computed using GNOM37, and the Rg and I(0) parameters were determined from the reduced data using routines from the ATSAS suite38. Low-resolution molecular envelopes were computed using DAMMIF37.
Molecular dynamics simulations of AbDPcob and AbPcob in light and dark state
The full-length AbDPcob and AbPcob was modelled using AlphaFold225 predicted and solved crystal structure respectively. The AdoCbl and cobalt was placed in AbDPcob by taking the coordinate in the crystal structure of AbPcob after structure alignment to Pcob domain. OHCbl was created by a simple mutation of the upper ligand in PyMol 2.5. We simulate the dynamics of AbDPcob and AbPcob in light and dark state by including different upper ligand into the protein: AdoCbl for dark state and OHCbl for light state. The input files for the calculation of parameters for AdoCbl and OHCbl were generated by MCPB.py protocol39 in AmberTools 2240,41 package. The starting structure of AdoCbl from the crystal structure was used as the starting geometry and the starting structure of OHCbl was created by replacing the adenosyl group with OH group in AdoCbl. Both ligands were first optimised by B3LYP/6-31 G*42 in Gaussian 0943 and the optimised structure was used for calculating point charges and force constants. The protonation state of protein residues were calculated by PROPKA44 in the presence of AdoCbl/OHCbl. AMBER99SB force field45 was applied to model protein and ions. The complex structure was solvated by TIP3P modelled water molecules46 in cubic box, keeping at least 14 Å between any atoms of protein and the boundary of the box. Counterions (19 Na+ ions for truncated AbPcob and 58 Na+ ions for full-length AbDPcob) were added to neutralise the system. MD simulations were carried out by Gromacs version 5.047. We use the same protocol for the simulation for the protein in light and dark state.
The system was first minimised to remove possible clashes after adding hydrogens to the protein, followed by a heating step to 300 K under NVT ensemble. The initial velocities were generated to maintain a Maxwell−Boltzmann distribution at the desired temperature, 300 K. The stepwise equilibration steps are conducted in order to make sure the structure gradually relaxed in the solvent environment: firstly, protein-complex were restrained to minimise/equilibrate (100 ps) water and ions; secondly, heavy atoms of protein and AdoCbl/OHCbl-Cobalt were restrained to minimise/equilibrate (100 ps) protein hydrogens; next main chain of protein and heavy atoms of AdoCbl/OHCbl and Cobalt were restrained to minimise/equilibrate (2 ns) protein side chains; then, protein Cα atoms and heavy atoms of AdoCbl/OHCbl and Cobalt were restrained to minimise/equilibrate (2 ns) main chain of protein; in last step, all restraints were remove to equilibrate for 2 ns. Harmonic restraint was applied before full relaxation of structures with a force constant of 1000 kJ/mol/nm2. Production was run at 300 K by NPT ensemble for 1500 ns based on the equilibrated structure (500 ns for AbPcob). V-rescale thermostat algorithm48 was used to control the temperature with a response time 1.0 ps. The pressure was kept at 1.0 bar using the Parrinello-Rahman pressure coupling scheme49 with a response time 1.0 ps. Hydrogen atoms were constrained during the production by LINCS50. The cut-off was set to 12 Å for electrostatic interactions calculation. Additional sampling was carried out on AbDPcob using three parallel simulations as follows: 50 × 1 ns cycles of simulated annealing (700 ps at 300 K, 100 ps heating to 400 ps, 100 ps at 400 K, 100 cooling to 300 K) followed by an additional 350 ns of MD for each run, which was then used for analysis. The increased sampling is clear from the Cα RMSDs for these simulations (more details see supplementary information).
Ensemble docking of BV to AbPcob
The AbPcob conformations in dark state after MD simulation was used for BV docking analysis. AbPcob dark conformations were clustered based on RSMD values of BV binding region. Gromacs gromos51 method was used for clustering simulated structure at 1.5 Å cut-off. 38 clusters were obtained with RMSD cut-off of 1.5 Å for two structures to be neighbour. Autodockvina52 was used to dock BV into clustered AbPcob structure with a grid box size of 18 Å × 18 Å × 18 Å. All docked binding poses were ranked by binding affinity. Structures with a value below −7.0 kcal/mol were selected for further docking analysis. To get more detailed binding possibilities, the search exhaustiveness was set to 72 to rerun molecular docking with selected structures. The docking results were used to analyse the possible binding pose of BV to AbPcob. A similar procedure was used for the MD simulations of AbDPcob following simulated annealing, with and RMSD cut-off of 1.0 Å, resulting in 100 clusters (15 for one binding pocket, 85 for the other) to ensure an exhaustive conformational sampling during docking, and an exhaustiveness of 25.
Size exclusion chromatography-multi-angle light scattering (SEC-MALS)
SEC-MALS was used to determine the oligomeric state of the target proteins and to probe any protein-protein interactions, under different conditions. An Agilent G7110B HPLC pump, degasser and autoinjector (Agilient, Santa Clara, USA) was used to auto load the samples (50 µL each run, 1 to 10 mg/mL). Superdex 200 10/300 GL column was used for chromatographic separations. MiniDAWN TREOS MALS detector and Optilab rEX refractive index metre (Wyatt, Santa Barbara, USA) were used to collect light scattering signals. The flow rate was set at 1 mL min−1 with a mobile phase of 20 mM HEPES, pH 6.8, 150 mM NaCl buffer. Samples were preincubated in dark for 30 min or with 530 nm LED for 5 min. All results were processed according to referenced protocol53. Peak alignment, band-broadening correction and normalisation procedures were performed by selecting the central 50% region of the peaks. Raw data were exported and plotted using Origin 9.0 software (OriginLab, Northampton, MA).
Genomic contextual and protein-protein interaction-based enrichment analysis
A computational gene neighbourhood analysis was performed using Genomic enzymology tools54. Briefly, a sequence similarity network (SSN) was constructed to facilitate the assignment of function for different proteins. Then, Genome Neighbourhood Networks (GNNs) were constructed based on SSN to provide statistical analysis of neighbouring Pfam families. The frequency of common domains identified in the GNN was analysed and visualised by R and R package ggplot2. As 506 sequences belong to divergent species, four species (Saccharothrix syringae, Streptomyces sp. M41, Micromonospora yangpuensis and Magnetospirillum magnetotacticum MS-1) that possess more than one hit among 506 were selected for further analysis. In each species, the sequence was submitted to the STRING database55 for the construction of protein-protein interaction (PPI) network using two shells (20 interactors and 5 interactors) with medium confidence (0.400). Enrichment analyses, including Gene ontology, Local network cluster (STRING), reactome pathways, protein domains, and features (Pfam and InterPro) were performed based on gene lists from the PPI network. Enrichment results were visualised using R and R package ggplot2 and ggpubr.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates and experimental data generated in this study have been deposited in the Protein Data Bank (www.pdb.org) under accession code: 8J2W, 8J2X, 8J2Y. Atomic coordinates referenced in the main text can be found through 1B33, 3MWN. SAXS data has been deposited in the small Angle Scattering Biological Data Bank (www.sasbdb.org) under accession codes SASDUS3 and SASDUT3. LCMS data has been deposited on Figshare (Jeffreys, Laura [2024]). Photoproduct determination - LCMS. Figshare. Journal contribution. https://doi.org/10.6084/m9.figshare.25219721. LCMS data has been deposited on Figshare (Jeffreys, Laura [2024]). Photoproduct determination - NMR. Figshare. Journal contribution. https://doi.org/10.6084/m9.figshare.25219832). MD parameters for AdoCbl and OHCbl have been deposited on Figshare (https://doi.org/10.6084/m9.figshare.25226480). Initial and final structures from MD simulations have been deposited on Figshare (https://doi.org/10.6084/m9.figshare.25226549). All other source data are provided as supplementary data files. Source data are provided in this paper. Source data are provided with this paper.
References
Reshetnikov, V. V., Smolskaya, S. V., Feoktistova, S. G. & Verkhusha, V. V. Optogenetic approaches in biotechnology and biomaterials. Trends Biotechnol. 40, 858–874 (2022).
Ortiz-Guerrero, J. M., Polanco, M. C., Murillo, F. J., Padmanabhan, S. & Elias-Arnanz, M. Light-dependent gene regulation by a coenzyme B12-based photoreceptor. Proc. Natl. Acad. Sci. USA 108, 7565–7570 (2011).
Gruber, K., Puffer, B. & Krautler, B. Vitamin B12-derivatives-enzyme cofactors and ligands of proteins and nucleic acids. Chem. Soc. Rev. 40, 4346–4363 (2011).
Jost, M. et al. Structural basis for gene regulation by a B12-dependent photoreceptor. Nature 526, 536–541 (2015).
Jost, M., Simpson, J. H. & Drennan, C. L. The transcription factor CarH safeguards use of sdenosylcobalamin as a light sensor by altering the photolysis products. Biochemistry 54, 3231–3234 (2015).
Jiang, B. et al. Injectable, photoresponsive hydrogels for delivering neuroprotective proteins enabled by metal-directed protein assembly. Sci. Adv. 6, eabc4824 (2020).
Narayan, O. P., Mu, X., Hasturk, O. & Kaplan, D. L. Dynamically tunable light responsive silk-elastin-like proteins. Acta Biomater. 121, 214–223 (2021).
Wang, R., Yang, Z., Luo, J., Hsing, I. M. & Sun, F. B(12)-dependent photoresponsive protein hydrogels for controlled stem cell/protein release. Proc. Natl. Acad. Sci. USA 114, 5912–5917 (2017).
Yang, Z. et al. B(12)-induced reassembly of split photoreceptor protein enables photoresponsive hydrogels with tunable mechanics. Sci. Adv. 8, eabm5482 (2022).
Mansouri, M. et al. Smart-watch-programmed green-light-operated percutaneous control of therapeutic transgenes. Nat. Commun. 12, 3388 (2021).
Chatelle, C. et al. A green-light-responsive system for the control of transgene expression in mammalian and plant cells. ACS Synth. Biol. 7, 1349–1358 (2018).
Cheng, Z., Yamamoto, H. & Bauer, C. E. Cobalamin’s (Vitamin B12) surprising function as a photoreceptor. Trends Biochem. Sci. 41, 647–650 (2016).
Schneider, T., Tan, Y., Li, H., Fisher, J. S. & Zhang, D. Photoglobin, a distinct family of non-heme binding globins, defines a potential photosensor in prokaryotic signal transduction systems. Comput. Struct. Biotechnol. J. 20, 261–273 (2022).
Kutta, R. J. et al. The photochemical mechanism of a B12-dependent photoreceptor protein. Nat. Commun. 6, 7907 (2015).
Padmanabhan, S., Jost, M., Drennan, C. L. & Elias-Arnanz, M. A new facet of vitamin B(12): gene regulation by cobalamin-based photoreceptors. Annu. Rev. Biochem. 86, 485–514 (2017).
Gourinchas, G., Etzl, S. & Winkler, A. Bacteriophytochromes - from informative model systems of phytochrome function to powerful tools in cell biology. Curr. Opin. Struct. Biol. 57, 72–83 (2019).
Takala, H., Edlund, P., Ihalainen, J. A. & Westenhoff, S. Tips and turns of bacteriophytochrome photoactivation. Photochem. Photobiol. Sci. 19, 1488–1510 (2020).
Fushimi, K. et al. Rational conversion of chromophore selectivity of cyanobacteriochromes to accept mammalian intrinsic biliverdin. Proc. Natl. Acad. Sci. USA 116, 8301–8309 (2019).
Yang, Y. et al. Ultrafast proton-coupled isomerization in the phototransformation of phytochrome. Nat. Chem. 14, 823–830 (2022).
Reuter, W., Wiegand, G., Huber, R. & Than, M. E. Structural analysis at 2.2 A of orthorhombic crystals presents the asymmetry of the allophycocyanin-linker complex, AP.LC7.8, from phycobilisomes of Mastigocladus laminosus. Proc. Natl. Acad. Sci. USA 96, 1363–1368 (1999).
Soni, B. R. et al. Structure of the novel 14kDa fragment of alpha-subunit of phycoerythrin from the starving cyanobacterium Phormidium tenue. J. Struct. Biol. 171, 247–255 (2010).
Romling, U., Galperin, M. Y. & Gomelsky, M. Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol. Mol. Biol. Rev. 77, 1–52 (2013).
Hengge, R. Principles of c-di-GMP signalling in bacteria. Nat. Rev. Microbiol. 7, 263–273 (2009).
Schirmer, T. C-di-GMP synthesis: structural aspects of evolution, catalysis and regulation. J. Mol. Biol. 428, 3683–3701 (2016).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–d444 (2022).
Poddar, H. et al. An unusual light-sensing function for coenzyme B12 in bacterial transcription regulator CarH. in Methods in Enzymology, (Academic Press, 2022).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Vagin, A. A. et al. REFMAC5 dictionary: organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. D. Biol. Crystallogr. 60, 2184–2195 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Joosten, R. P., Long, F., Murshudov, G. N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. Iucrj 1, 213–220 (2014).
Jacobsen, D. W., Digirolamo, P. M. & Huennekens, F. M. Adenosylcobalamin analogues as inhibitors of ribonucleotide reductase and vitamin B12 transport. Mol. Pharmacol. 11, 174–184 (1975).
Kreft, J. U. & Schink, B. O-demethylation by the homoacetogenic anaerobe Holophaga foetida studied by a new photometric methylation assay using electrochemically produced cob(I)alamin. Eur. J. Biochem. 226, 945–951 (1994).
Lamparter, T., Michael, N., Mittmann, F. & Esteban, B. Phytochrome from Agrobacterium tumefaciens has unusual spectral properties and reveals an N-terminal chromophore attachment site. Proc. Natl. Acad. Sci. USA 99, 11628–11633 (2002).
Gasteiger E. et al. Protein Identification and Analysis Tools on the ExPASy Server. in The Proteomics Protocols Handbook (ed Walker J. M.) (Humana Press, 2005).
Franke, D. & Svergun, D. I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J. Appl. Crystallogr. 42, 342–346 (2009).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Li, P. & Merz, K. M. Jr. MCPB.py: a Python based metal center parameter builder. J. Chem. Inf. Modeling 56, 599–604 (2016).
Salomon-Ferrer, R., Case, D. A. & Walker, R. C. An overview of the Amber biomolecular simulation package. WIREs Comput. Mol. Sci. 3, 198–210 (2013).
Case, D. A. et al. The Amber biomolecular simulation programs. J. Comput. Chem. 26, 1668–1688 (2005).
Lee, C., Yang, W. & Parr, R. G. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37, 785–789 (1988).
Frisch, M.J. et al. Gaussian 09 Revision A.1 (Gaussian Inc., 2009).
Olsson, M. H. M., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Showalter, S. A. & Brüschweiler, R. Validation of molecular dynamics simulations of biomolecules using NMR Spin relaxation as benchmarks: application to the AMBER99SB force field. J. Chem. Theory Comput. 3, 961–975 (2007).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Daura, X. et al. Peptide folding: when simulation meets experiment. Angew. Chem. Int. Ed. 38, 236–240 (1999).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: new docking methods, expanded force field, and Python bindings. J. Chem. Inf. Model 61, 3891–3898 (2021).
Some, D., Amartely, H., Tsadok, A. & Lebendiker, M. Characterization of proteins by size-exclusion chromatography coupled to multi-angle light scattering (SEC-MALS). J. Vis. Exp. https://doi.org/10.3791/59615 (2019).
Zallot, R., Oberg, N. & Gerlt, J. A. The EFI web resource for genomic enzymology tools: leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry 58, 4169–4182 (2019).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49, D605–d612 (2021).
Brown, N. P., Leroy, C. & Sander, C. MView: a web-compatible database search or multiple alignment viewer. Bioinformatics 14, 380–381 (1998).
Acknowledgements
The work was funded by the Engineering and Physical Sciences Research Council International Centre-to-Centre grant no EP/S030336/1. We thank Diamond Light Source beamlines I03 & I04 (proposal numbers mx2447-65, 87, 89). This work was also supported by National Natural Science Foundation of China (No. 31870855) and NUDT Research Program (22-ZZCX-039, 22-02-15). The authors also thank the ESRF (proposal MX-2398) for beamtime and the staff of beamline BM29 for assistance with data collection. An ANR grant (PhotoGene, ANR-21-CE11-0036-01) awarded to GS supported the research reported in this paper. The IBS acknowledges integration into the Interdisciplinary Research Institute of Grenoble (IRIG, CEA). The authors would like to gratefully acknowledge the training and use of mass spectrometry instrumentation provided by Professor Perdita Barran’s group at the Michael Barber centre for collaborative mass spectrometry (MBCCMS) based at the University of Manchester.
Author information
Authors and Affiliations
Contributions
N.S.S., D.J.H., D.L. and S.Z. initiated and coordinated the project. S.Z., L.N.J., D.J.H., N.S.S. and D.L. designed experiments, analysed data and wrote the manuscript with contributions from other authors. S.Z. produced and crystallised the proteins. H.P., M.S. and C.W.L. helped with protein crystallisation and data collection. S.Z. processed diffraction data and solved the structures. Y.Y., L.O.J. and S.Z. performed the docking and molecular dynamics simulations. C.L. and L.Z. carried out bioinformatics analysis of Pcob proteins. K.P., L.N.J. and M.J.C. assisted with SasPcob characterisation. L.N.J. and C.Y. performed mass spectrometry analysis. L.N.J. and M.J.C. performed all NMR experiments. G.S. and M.W. performed SAXS measurements. N.S.S., D.J.H. and D.L. advised on all aspects. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. Correspondence and request for materials should be addressed to N.S.S., D.J.H. and S.Z.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, S., Jeffreys, L.N., Poddar, H. et al. Photocobilins integrate B12 and bilin photochemistry for enzyme control. Nat Commun 15, 2740 (2024). https://doi.org/10.1038/s41467-024-46995-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-46995-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.