Abstract
Endoplasmic reticulum (ER)-associated degradation (ERAD) and ER-phagy are two principal degradative mechanisms for ER proteins and aggregates, respectively; however, the crosstalk between these two pathways under physiological settings remains unexplored. Using adipocytes as a model system, here we report that SEL1L-HRD1 protein complex of ERAD degrades misfolded ER proteins and limits ER-phagy and that, only when SEL1L-HRD1 ERAD is impaired, the ER becomes fragmented and cleared by ER-phagy. When both are compromised, ER fragments containing misfolded proteins spatially coalesce into a distinct architecture termed Coalescence of ER Fragments (CERFs), consisted of lipoprotein lipase (LPL, a key lipolytic enzyme and an endogenous SEL1L-HRD1 substrate) and certain ER chaperones. CERFs enlarge and become increasingly insoluble with age. Finally, we reconstitute the CERFs through LPL and BiP phase separation in vitro, a process influenced by both redox environment and C-terminal tryptophan loop of LPL. Hence, our findings demonstrate a sequence of events centered around SEL1L-HRD1 ERAD to dispose of misfolded proteins in the ER of adipocytes, highlighting the profound cellular adaptability to misfolded proteins in the ER in vivo.
Similar content being viewed by others
Introduction
Cells have evolved various quality control mechanisms to maintain proteostasis in various organelles to prevent the formation of potentially toxic protein aggregates1,2. The collapse of protein quality control system(s) may indeed contribute to disease pathogenesis such as in aging and diabetes1. In the endoplasmic reticulum (ER), misfolded proteins are cleared by two degradative mechanisms, namely ER-associated degradation (ERAD) and selective macroautophagy known as ER-phagy3. While ERAD degrades proteins in a substrate-specific manner4,5,6,7,8, ER-phagy represents a mechanism for lysosome-mediated bulk clearance of ER fragments or subdomains. Recent studies have demonstrated the importance of ER-phagy in vitro toward protein aggregates such as disease-relevant mutant proteins of proinsulin9, procollagen10,11, POMC12,13, and NPC114. However, while it has been reported that ER-phagy degrades ERAD-resistant model substrates in vitro15, the physiological relevance and significance of ER-phagy and its crosstalk with ERAD in vivo remain largely unclear16,17,18,19.
The ER-resident SEL1L-HRD1 protein complex (Hrd3p-Hrd1p in yeast) represents the most conserved branch of ERAD4,5,20,21,22,23, where SEL1L not only controls the stability of the E3 ligase HRD120,21,22,23,24, but defines substrate specificity and HRD1 function24,25. Loss of SEL1L or HRD1, either in germline or acutely in adult mice, cause lethality24,26,27,28, highlighting the importance of SEL1L-HRD1 ERAD in organismal development and function. Indeed, using cell type-specific mouse models, we and others have shown that SEL1L-HRD1 ERAD is indispensable for fundamental physiological processes such as immune cell development and function, gut homeostasis, water balance, food intake, systemic energy homeostasis, nutrient metabolism and organellar crosstalk in a substrate-dependent manner5,20,24,29,30,31,32,33,34,35,36,37,38,39,40,41. Peculiarly, in most cell types examined to date, SEL1L-HRD1 ERAD deficiency is coupled with only very mild ER stress or unfolded protein response (UPR), with no detectable cell death before, at or even after disease initiation in vivo4,5,20, pointing to the existence of compensatory mechanism(s) in response to impaired SEL1L-HRD1 ERAD function42.
Using adipocyte-specific genetic knockout (KO) mouse models powered by a combination of state-of-the-art ultrastructural imaging, proteomic screens, biophysical and biochemical assays, we uncover the sequence of events underlying the intimate crosstalk between SEL1L-HRD1 ERAD and ER-phagy in adipocytes in vivo, to dispose of a misfolded proteins in the ER, such as lipoprotein lipase (LPL). In the absence of SEL1L-HRD1 ERAD, autophagy becomes active to clear ER fragments containing misfolded proteins and/or aggregates. In adipocytes lacking both SEL1L-HRD1 ERAD and autophagy, these ER fragments are coalesced into a unique cellular compartment termed CERFs. LPL, a key enzyme involved in systemic lipid metabolism and partitioning43, and ER chaperone BiP are two main components of CERFs and undergo phase separation in vitro.
Results
Activation of autophagy in adipocytes lacking SEL1L
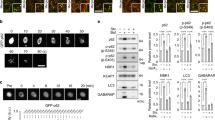
We previously showed that Sel1L deficiency in adipocytes (Sel1LAdipCre) is associated with very mild UPR activation31 as measured by Xbp1 mRNA splicing (Supplementary Fig. 1a), and without cell death as measured by cleaved caspase 3 and TUNEL staining (Supplementary Fig. 1b, c), pointing to the existence of compensatory mechanism(s) in response to impaired SEL1L-HRD1 ERAD function. Indeed, KEGG pathway enrichment analysis of cDNA profiling microarray analysis of white adipose tissues (WAT) (GSE56918)31 revealed autophagy among the top pathways upregulated in Sel1LAdipCre WAT compared to WT WAT (Supplementary Fig. 1d). Indeed, ATG7-dependent conversion of microtubule-associated protein 1 light chain (LC3) LC3-I to LC3-II (LC3-phosphatidylethanolamine conjugate) was doubled in WAT of Sel1LAdipCre mice compared to that of WT littermates (Fig. 1a). On the other hand, protein level of the ubiquitin-binding autophagy receptor and substrate P6244,45 was largely unchanged while highly accumulated in Atg7AdipCre WAT (Fig. 1a), excluding the possibility of defective lysosomal function in Sel1LAdipCre WAT. Moreover, transmission electron microscopy (TEM) analysis of WAT revealed that there were more double-membrane autophagosomes (arrows) containing ER-like structures (asterisks) in Sel1LAdipCre WAT compared to that in WT WAT under basal state (arrows, Fig. 1b, c and Supplementary Fig. 1e), pointing to ER-phagy. Hence, similar to our recent report in pancreatic β cells42, our data showed an activation of autophagy and ER-phagy in adipocytes lacking SEL1L.
a Immunoblot analysis of LC3B and P62 in gonadal WAT with quantitation of LC3B-II to I conversion ratio shown on the right (n = 7 for WT and 5 for Sel1LAdipCre mice). b, c Representative TEM images of gonadal WAT showing ER-like structures (*) in the autophagosomes (arrow), with quantitation of autophagic vacuoles (AV) shown in (c); n = 50 WT and 63 Sel1LAdipCre adipocytes from 3 mice each. LD, lipid droplet; M, mitochondria. d Immunoblot analysis of SEL1L-HRD1 ERAD and autophagy in gonadal WAT (n = 4 mice per genotype). e Body weight of female (F, n = 19 for WT, 12 for Sel1LAdipCre, 8 for Atg7AdipCre and 17 for DKO) and male (M, n = 12 for WT, 7 for Sel1LAdipCre, 8 for Atg7AdipCre and 11 for DKO) littermates at the age of 12 weeks. Weight (f) and representative pictures (g) of gonadal WAT of female (F, n = 18 for WT, 6 for Sel1LAdipCre, 8 for Atg7AdipCre and 15 for DKO) and male (M, n = 11 for WT, 6 for Sel1LAdipCre, 9 for Atg7AdipCre and 9 for DKO) littermates at 12 weeks of age. h Weight of gonadal WAT of female littermates at 5 and 12 weeks of age (n = 6 mice at 5 weeks of age per genotype, n = 14 mice at 12 weeks of age per genotype). i, j Representative H&E images of gonadal WAT of female littermates at 12 weeks of age (n = 3 mice per genotype), with quantitation of area of lipid droplets (LD) shown in (j) (n = 960 and 1042 cells from two mice per genotype). k Immunoblot analysis of (cleaved) caspase 3 in gonadal WAT of 12-week-old mice (n = 4 mice per genotype). Mouse thymus, a positive control. Histogram was plotted as mean with SD; each data points were derived from biologically independent mice. n.s., not significant; P values were derived by two-sided Student’s t-test (a), One-way ANOVA followed by Tukey’s test (e, f), Two-way ANOVA (h) and Chi-square test (j). Source data are provided as a Source Data file.
Generation of mice with adipocyte-specific deletion of Sel1L and Atg7
To examined the physiological significance of the crosstalk between ERAD and autophagy in adipocytes, we generated a mouse model lacking both Sel1L and Atg7 in adipocytes (Sel1LAdipCre;Atg7AdipCre, hereafter DKO) using the Adiponectin-Cre mice31. Wildtype (WT) and adipocyte-specific Sel1L or Atg7 deficient (Sel1LAdipCre or Atg7AdipCre) littermates were included as controls. Both genders of adult mice were used throughout this study. Western blot analyses confirmed the reduction of SEL1L, HRD1 or ATG7 protein levels in WAT (Fig. 1d). At 12 weeks of age, DKO mice, of both genders, exhibited comparable body weights as the other three cohorts (Fig. 1e). The size and weight of white fat pads were reduced in DKO WAT compared to the other cohorts (Fig. 1f, g), which was likely caused by the inability of WAT to expand from 5 to 12 weeks of age (Fig. 1h). Histologically, DKO adipocytes exhibited smaller lipid droplets and cell size compared to the other cohorts at 12 weeks of age (Fig. 1i, j). There was no detectable cell death as measured by caspase-3 cleavage and TUNEL staining (Fig. 1j and Supplementary Fig. 2a), while a modest increase of Xbp1 mRNA splicing (Supplementary Fig. 2b, c), a marker of UPR IRE1α activity46,47, in DKO WAT compared to other cohorts.
ER fragments spatially coalesce into clusters (CERFs) in DKO adipocytes
The forementioned data prompted the question how DKO cells deal with misfolded proteins in the ER and cytosol. Loss of autophagy is known to cause the formation of protective P62-positive cytosolic inclusions, containing ubiquitinated proteins and protein aggregates44, which may include ER fragments and contents as suggested by recent studies48,49. Indeed, under transmission electron microscopy (TEM), we noted the presence of the typical P62 inclusions in DKO WAT and brown adipose tissues (BAT) (Fig. 2a). This was further confirmed using P62 immunogold labeling followed by TEM (Fig. 2b) and confocal microscopy (Supplementary Fig. 3a). P62-containing inclusions were also detected in Atg7AdipCre WAT/BAT, albeit of much smaller size and at much lower frequency than those in DKO WAT/BAT (Supplementary Fig. 3a). Upon co-staining with the ER marker KDEL (for two major ER chaperones BiP and GRP94), we noted that KDEL proteins were largely excluded from the P62 inclusions in DKO WAT (Fig. 2b). Hence, we concluded that misfolded proteins in the ER are not predominantly shunted to the P62 inclusions.
a Representative TEM images of P62 inclusions in DKO gonadal WAT (n = 4 mice) and BAT (n = 3 mice). #, P62 inclusion; LD, lipid droplet; M, mitochondria. b, c Representative EM images following immunogold co-labeling of P62 (pink dotted line) and the ER marker KDEL (green dotted line) in DKO gonadal WAT (n = 2 mice). Color-coded images, P26 (pink) and KDEL (blue). d–f Representative TEM images (d, e, n = 3-4 mice each) and TEM with immunogold labeling of the ER marker BiP (f, n = 3 mice) in DKO gonadal WAT or BAT. Arrows, membranes. g 3D reconstruction of a CERF in DKO gonadal WAT from 370 FIB-SEM images taken at every 10 nm (n = 2 independent repeats). h Diameters of the CERFs (n = 52) and P62 inclusions (n = 49) in DKO gonadal WAT based on TEM analysis of 4 mice with the mean of diameters indicated. i, j TEM images showing possible clustering (i) and fusion (j) of ER fragments in DKO BAT (n = 3 mice). Arrows, likely fused membranes inside CERFs. Source data are provided as a Source Data file.
Excitingly, under immunogold-TEM, we frequently observed clusters of the KDEL signals in addition to the P62 inclusions in the cytosol of DKO WAT (green dotted lines, Fig. 2c), which were not present in other cohorts (Supplementary Fig. 3b). These ER clusters had very few detectable P62 signals inside (Fig. 2c), hence representing a cellular compartment distinct from the P62 inclusions. TEM analysis indeed revealed a unique, membranous, cellular compartment in DKO WAT, but not in other cohorts (arrows, Fig. 2d and Supplementary Fig. 3c, d). The same structure of clusters of ER fragments was observed in DKO BAT as well (Fig. 2e, f). 3D FIB-SEM analysis showed that this architecture was a distinct oval-shaped compartment composed of ER fragments (Fig. 2g and Supplementary Video 1). The diameters of the clusters ranged between 2 and 7 μm, slightly smaller than those of the P62 inclusions in DKO WAT (Fig. 2h). Providing further support for the nature of ER fragments, we noted clusters of large ER fragments in DKO adipocytes (Fig. 2i), some of which might have fused at the core (arrows, Fig. 2j and Supplementary Video 1). Taken together, these data suggested that ER fragments may coalesce into a unique cellular architecture in adipocytes lacking SEL1L and ATG7, which we termed as the coalescence of ER fragments (CERFs).
LPL is a misfolded SEL1L-HRD1 ERAD substrate
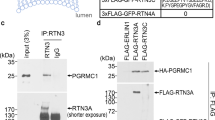
We next explored the key protein component(s) of CERFs. The observation that CERFs are clusters of ER fragments only in DKO cells prompted us to hypothesize that ER fragments contain misfolded ERAD substrates. To this end, we designed and performed an exploratory immunoprecipitation mass spectrometry (IP-MS) experiment (without replication) to identify potential SEL1L-HRD1 ERAD substrates in BAT. As HRD1 deficiency allows the capture of the interactions between SEL1L and substrates5,20, we performed SEL1L immunoprecipitation in BAT from mice with brown adipocyte-specific HRD1 deficiency (Hrd1 Ucp1Cre)39, with WT and Sel1L Ucp1Cre BAT as controls (Supplementary Fig. 4a). Only proteins showing 4- and 2-fold increases in peptide-spectrum matches (PSMs) in Hrd1Ucp1Cre samples over Sel1L Ucp1Cre and WT samples, respectively, were considered as potential substrates. A total of 40 candidates were identified (Supplementary Table 1), which included known SEL1L interactors such as HSP90B1 (GRP94), OS9 and ERLEC1 (XTP3B)50 (Supplementary Fig. 4b), hence validating our experimental system. Excitingly, lipoprotein lipase (LPL), a rate-limiting enzyme catalyzing the hydrolysis of triglycerides packed in plasma lipoproteins into fatty acids43, was identified among the top hits (Supplementary Fig. 4b).
We previously showed that LPL is largely trapped in the ER of Sel1L-deficient adipocytes31, but it was unclear whether LPL is an endogenous substrate of SEL1L-HRD1 ERAD. LPL interacted with SEL1L in Hrd1–/– but not WT adipocytes (Fig. 3a). Protein levels of LPL were stabilized and accumulated in Sel1L–/– or Hrd1–/– adipocytes versus that in WT adipocytes (Fig. 3b, c). Consistently, LPL accumulated by threefold to fourfold in Sel1L-deficient WAT (Fig. 3d, e), suggesting that LPL is indeed a misfolded SEL1L-HRD1 ERAD substrate. Unlike in Sel1LAdipCre WAT, Atg7 deficiency in WAT had a subtle effect on LPL protein level (Fig. 3d, e). Moreover, LPL protein levels were further increased by another ~5-folds in DKO WAT over that in Sel1LAdipCre WAT (Fig. 3d, e). The accumulation of LPL in Sel1LAdipCre or DKO WAT was not a general phenomenon for membrane or secretory proteins as others such as calnexin and glucose transporter 4 (GLUT4) were unchanged compared to those in WT WAT (Fig. 3d, f, g). These changes in LPL protein levels were uncoupled from gene transcription as mRNA level of Lpl was comparable among all four cohorts (Fig. 3h).
a Immunoblot analysis following immunoprecipitation of endogenous LPL in differentiated adipocytes (3 independent repeats). b Immunoblot analysis of LPL in differentiated adipocytes pre-treated with Brefeldin A for 30 min followed by cycloheximide (CHX) treatment for the indicated times (2 independent repeats). c Immunoblot analysis of LPL in differentiated adipocytes (3 independent repeats). d–g Immunoblot analysis of LPL and other proteins in gonadal WAT (d) with quantitation shown in e-g (n = 3 mice per genotype for LPL and GLUT4 and n = 2 mice per genotype for CALNEXIN). h qPCR analysis of Lpl mRNA levels in gonadal WAT (n = 14 per WT and DKO, n = 9 for Sel1LAdipCre and n = 11 for Atg7AdipCre). Histogram was plotted as mean with SD; each data points were derived from biologically independent mice/samples. P values were derived by one-way ANOVA followed by Tukey’s test; n.s. not significant. Source data are provided as a Source Data file.
LPL is a major component of CERFs
LPL is normally secreted from adipocytes to the capillary where it hydrolyzes triglyceride-rich lipoproteins43. Endoglycosidase H (EndoH) digestion showed that over 90% of LPL in Sel1LAdipCre and DKO WAT was retained in the ER (i.e. EndoH sensitive), unlike those in WT and Atg7AdipCre WAT with 60% being in extra-ER compartments (i.e. EndoH resistant) (Fig. 4a, b), suggesting that SEL1L is required for the maturation of nascent LPL in the ER. Immunofluorescent staining showed that LPL was trapped intracellularly and largely absent from the endothelium in Sel1LAdipCre and DKO WAT (white arrows, Fig. 4c, d and Supplementary Fig. 5a). By contrast, majority of LPL was co-localized with the capillary marker GPIHBP1 in WT and Atg7AdipCre WAT (yellow arrows, Fig. 4c, d and Supplementary Fig. 5a). In keeping with intracellular retention of LPL, DKO mice exhibited mild postprandial hyperlipidemia compared to other cohorts (Supplementary Fig. 5b).
a, b EndoH analysis of LPL in gonadal WAT using Western blot with quantitation of percentage of EndoH resistant LPL in (b) (n = 6 for WT, 3 for Sel1LAdipCre, 4 for Atg7AdipCre and DKO). s, i and r, EndoH sensitive, intermediate and resistant, respectively. c, d Representative immunofluorescent images of LPL without (c) and with co-labeling of the endothelial marker GPIHBP1 (d) in gonadal WAT (n = 3 mice per WT and DKO; n = 2 mice per Sel1LAdipCre and Atg7AdipCre). Capillary, intracellular and LPL CERF are marked by yellow, white, and green arrows, respectively. e Representative immunofluorescent images of co-labeling of LPL and cell surface marker caveolin 1 (CAV1) in gonadal WAT (n = 2 mice per genotype). Green arrows, LPL CERFs. f Representative EM images following immunogold labeling of LPL in gonadal WAT (n = 3 mice per genotype). Images with color coded gold particles shown on the right. Green dot line, LPL CERFs. g Representative immunofluorescent images of co-labeling of P62 and LPL in gonadal WAT (n = 3 mice per genotype). Red and green arrows, P62 inclusion and LPL CERFs, respectively. h Representative immunofluorescent images of triple labeling of LPL (gray), ubiquitin (Ub, red) and P62 (green) in gonadal WAT (n = 2 mice per genotype). Yellow and gray arrows, Ub-containing P62 inclusions and LPL CERFs, respectively. Histogram was plotted as mean with SD; each data points were derived from biologically independent mice/samples. P values were derived by one-way ANOVA followed by Tukey’s test; n.s. not significant. Source data are provided as a Source Data file.
Unlike that in Sel1LAdipCre WAT, LPL protein formed large puncta in DKO WAT (green arrows, Fig. 4c, d). These LPL-containing puncta were formed inside of adipocytes as demonstrated by co-staining with a surface membrane protein caveolin1 (CAV1) (green arrows, Fig. 4e). Immunogold labeling of LPL followed by TEM imaging demonstrated that LPL was enriched in the CERFs (Fig. 4f). LPL-containing foci were distinct from the P62 inclusions in WAT as shown using confocal microscopy (Fig. 4g). Furthermore, unlike the P62 inclusions which were ubiquitin positive, LPL-containing CERFs were devoid of ubiquitin (Fig. 4h). Taken together, these data demonstrated that the LPL-containing CERFs and P62 inclusion are two distinct cellular quality-control compartments.
LPL interacts extensively with ER chaperones in the CERFs
The identification of LPL as a protein composition of the CERFs made it possible to further explore the biophysical and biochemical nature of the CERFs. First, we performed an exploratory LPL IP-MS (without replication) in WAT to identify LPL interactomes, where two clusters of LPL-interacting proteins were found to be enriched in either WT or DKO WAT. Cluster 1 contained 47 protein hits with PSMs at least 2-fold higher in WT than DKO WAT, while Cluster 2 contained 101 hits with PSMs at least 4-fold higher in DKO than WT WAT (Supplementary Fig. 6a). Cluster 1 included cytoskeleton proteins (e.g. Myosin-11 MYH11), secretory/scaffold proteins (e.g. Caveolin 1 CAV1, Clusterin CLU and Gephyrin GPHN), and ribosomal subunits (RPL3, 4, 6, 7a, 18) (Supplementary Fig. 6b and Supplementary Table 2). These Cluster 1 hits were similarly enriched in Atg7AdipCre, but not Sel1LAdipCre WAT (Supplementary Fig. 6b and Supplementary Table 2), pointing to reduced biosynthesis and trafficking of LPL in ERAD-deficient WAT. On the other hand, Cluster 2 included ER chaperones involved in post-translational modifications such as glycosylation and hydroxylation (e.g. ribophorin 1 RPN1, Malectin MLEC and procollagen-Lysine,2-oxoglutarate 5-dioxygenase 3 PLOD3), folding and degradation (e.g. HSPA5/BiP, HSP90B1/GRP94, OS9 and GPX3)51 (Supplementary Fig. 6c and Supplementary Table 3). Most proteins in Cluster 2 were also upregulated in Sel1LAdipCre, not Atg7AdipCre WAT (Supplementary Fig. 6c and Supplementary Table 3), indicating that LPL is accumulated in the ER in the absence of ERAD. Consistently, pathway analyses of the LPL interactomes showed that, while ribosome and exosome pathways were enriched in WT WAT, glycan-related pathways and protein processing in the ER were highly represented in DKO WAT (Supplementary Fig. 6d).
Indeed, IP of endogenous LPL pulled down endogenous CAV1 in WT WAT, but much less so in DKO WAT (Fig. 5a). By contrast, endogenous LPL interacted strongly with endogenous BiP, OS9, MLEC and PLOD3 in DKO WAT, but not in WT WAT (Fig. 5b). Immunofluorescent staining and TEM following immunogold labeling confirmed colocalization of chaperone proteins such as OS9, MLEC and PLOD3 with LPL in the CERFs in DKO WAT (Fig. 5c–f and Supplementary Fig. 7a–f). Taken together, our data suggested that LPL is present, either as folding intermediates or terminally misfolded, in the CERFs of DKO adipocytes.
a, b Immunoblot analysis following immunoprecipitation of endogenous LPL in gonadal WAT (n = 3 mice each). c–e Representative immunofluorescent images of co-labeling of LPL and PLOD3 (c, n = 3 mice per genotype), MLEC (d, n = 2 mice per genotype), and OS9 (e, n = 2 mice per genotype), the top hits identified in cluster 2 (Supplementary Fig. 6c), in gonadal WAT. f Representative TEM images following immunogold labeling of LPL and PLOD3 in gonadal WAT (n = 3 mice). Green dotted line outlines the CERFs. Images with color coded gold particles shown on the right. Source data are provided as a Source Data file.
CERFs progressively enlarge and become insoluble with age
Next, we examined the biochemical properties of CERFs using LPL as a readout. Western blot analysis following non-denaturing SDS-PAGE showed that LPL formed significantly more HMW aggregates in detergent-soluble fractions of DKO WAT than those in the other cohorts (Fig. 6a). These aggregates were formed via aberrant disulfide bonds as they were sensitive to the treatment of β-ME (Fig. 6a). In addition, significantly more LPL was detected in the detergent-insoluble fractions of DKO WAT compared to those of the other cohorts (Fig. 6b). Indeed, about half of LPL proteins were insoluble at 12 weeks of age vs. 30% at 5 weeks of age (Fig. 6c), suggesting that LPL gradually becomes insoluble with age. Similar observation was obtained for BiP (Fig. 6c). Indeed, some CERFs from 12-week-old WAT appeared hardened under TEM (Fig. 6d). In keeping with these findings, the size and abundance of LPL CERFs were markedly elevated in DKO WAT from 5 to 12 weeks of age (arrows, Fig. 6e and Supplementary Fig. 8a). These CERFs were not amyloidogenic as revealed by the negative Thioflavin S staining, unlike the amyloidogenic β-amyloid plaques in the brains of Alzheimer’s disease 5xFAD mouse model (arrows, Supplementary Fig. 8b, c). Hence, these data demonstrated that CERFs grow and become insoluble with age.
a Immunoblot analysis of LPL under reducing (left) and non-reducing (right) conditions in gonadal WAT lysates of 12-week-old mice (n = 4 mice per WT and DKO, and 2 mice per Sel1LAdipCre and Atg7AdipCre). Stats shown comparing DKO vs. Sel1LAdipCre (P = 0.01) by one-way ANOVA followed by Tukey’s test. b Immunoblot analysis of LPL in gonadal WAT lysates of 12-week-old mice fractionated in 1% NP40 (S, soluble; P, pellet). HSP90 and H2A, loading controls for S and P fractions, respectively (n = 3 mice per genotypes). c Immunoblot analysis of LPL in gonadal WAT lysates of 5- and 12-week-old WT and DKO mice fractionated in 1% NP40 as in b (n = 3 mice per group). LPL*: insoluble LPL that is resistant to heat and β-Mercaptoethanol (β-ME). d TEM images of a CERF (*) in gonadal WAT of 12-week-old mice with the diameter indicated (n = 4 mice). e Representative immunofluorescent images of co-labeling of LPL and BiP in gonadal WAT of mice at the age of 5 and 12 weeks, respectively (n = 3 mice per group). Arrows, LPL CERFs. Data are mean ± SD. Source data are provided as a Source Data file.
In vitro reconstitution of LPL condensates, coated by BiP
We noticed that, while BiP colocalized with LPL (Fig. 6e and Supplementary Fig. 8a), BiP seemed to wrap around LPL in the CERFs of DKO WAT in a 3D reconstruction of the confocal images (Fig. 7a, b) and immunogold co-labeling of BiP and LPL followed by TEM (Fig. 7c). This distribution pattern was not seen in other cohorts (Supplementary Fig. 9). This unique spatial organization prompted us to explore how LPL proteins assemble in CERFs by asking whether LPL protein can self-assemble or condense spontaneously in vitro using purified human recombinant LPL (Supplementary Fig. 10a, b).
a, b Representative immunofluorescent images of co-labeling of LPL and BiP (a) and its 3D reconstruction (b) in gonadal WAT of DKO mice (n = 3 mice). Arrows, CERFs. c Two representative EM images following immunogold co-labeling of LPL and BiP in gonadal WAT (n = 3 mice). Images with color coded gold particles against LPL (red) and BiP (blue) are shown on the right. d Representative images of mCherry-LPL phase-separated condensates at the indicated protein concentrations (n = 3 independent repeats). e Representative images of mCherry-LPL (10 μM) phase-separated condensates in the presence or absence of EGFP-BiP (n = 3 independent repeats). f Fluorescent images and intensity profile across the focal plane (arrow) of an LPL and BiP co-condensed droplet. g Negative stain TEM images showing LPL-BiP co-condensates (n = 2 independent repeats). h Turbidity at 395 nm (OD395) was measured for mCherry-LPL phase-separated condensates, in the presence or absence of EGFP-BiP or BSA (n = 3 independent repeats). Graph was plotted as mean with SD. P values were derived by Two-way ANOVA followed by Tukey’s multiple comparisons test; n.s., not significant. i Fluorescence intensity recovery curves of mCherry-LPL condensates in vitro in the presence or absence of EGFP-BiP post photo-bleaching. Source data are provided as a Source Data file.
Indeed, LPL spontaneously phase separated into hollow droplets in a protein concentration-dependent manner (Fig. 7d). Surprisingly, while the addition of purified BiP had no noticeable effect on the morphology of LPL droplets, BiP formed an outer layer surrounding the LPL (Fig. 7e), bearing striking resemblance to the situation in vivo (Fig. 7a–c). Indeed, BiP coated outside of the LPL droplets, as revealed by the quantified fluorescence intensities of red and green channels (Fig. 7f), suggesting that this in vitro reconstituted structure resembles the pattern seen in CERFs. These LPL-BiP droplets were solid and spheric shaped as visualized by TEM (Fig. 7g). The turbidity of LPL droplets decreased in the presence of BiP, but not BSA (Fig. 7h). Fluorescence recovery after photobleaching (FRAP) analysis showed that LPL droplets exhibited relatively low mobility (Fig. 7i and Supplementary Fig. 10c), indicating the LPL droplets were more gel-like in nature. The addition of EGFP-BiP slightly increased LPL mobility (Fig. 7i and Supplementary Fig. 10c).
Redox environment and C-terminal W loop influence LPL self-assembly or aggregation
We next explored the molecular mechanisms underlying LPL phase separation. LPL has five disulfide bonds (Supplementary Fig. 8a)43. Indeed, in the absence of a reducing agent β-mercaptoethanol (β-ME), LPL protein readily aggregates in addition to form phase-separated droplets (arrows, Fig. 8a). These aggregates could not be reversed by the addition of BiP (arrows, Fig. 8b). To further explore how LPL self-assembly occurs, we assessed the importance of LPL structural domains in this process. Truncating the C-terminal domain (CTD) of LPL significantly hampered its phase separation (Fig. 8c, d). Taking a cue from a recent study showing that the tryptophan (W) loop at the CTD of LPL is critical for the formation of helical LPL oligomers52, we mutated three tryptophan residues (W417, W420 and W421 in human LPL) in this loop to asparagine (W3N) (Fig. 8e). Indeed, W3N mutation significantly reduced the efficiency and size of LPL droplets (Fig. 8f, h), and moreover, promoted LPL aggregation under non-reducing conditions (arrows, Fig. 8g). These data indicated that LPL self-assembly in vitro is sensitive to the redox environment and, at least partially, mediated by its C-terminal tryptophan loop.
Representative images of phase-separation of mCherry-LPL (a) and LPL + EGFP-BiP (b) in the absence of β-mercaptoethanol (β-ME) (n = 3 independent repeats). Arrows, aggregates. c Predicted structure of LPL monomer (PDB ID: 6U7M) showing the N- (NTD, in cyan) and C-terminal domain (CTD, in pink). The tryptophan (W) loop is highlighted in magenta. d Representative images of phase-separated condensates of mCherry-LPL WT and the mutant lacking the CTD (amino acids 340-475) (n = 3 independent repeats). e Predicted structure of LPL oligomers with the inset on the right showing the tetrameric interface with two W loops colored in gray. The side chains of three tryptophan residues W417, W420, and W421 in magenta are shown as sticks. Representative images of phase-separated condensates of mCherry-LPL WT and the tryptophan mutant (W3N) in the presence (f) and absence (g) of β-ME. Arrows, aggregates (n = 3 independent repeats). h Quantitation of the total area and the number of droplets per unit focal plane shown in (f) (n = 3 independent repeats). Histogram was plotted as mean with SD; each data points were derived from biologically independent samples. P values were derived by Two-way ANOVA followed by Sidak’s multiple comparison test; n.s., not significant. Source data are provided as a Source Data file.
Discussion
Our data demonstrate a sequence of events leading to the clearance of misfolded proteins in the ER by SEL1L-HRD1 ERAD and autophagy in adipocytes in vivo (Fig. 9). The observations that SEL1L-HRD1 ERAD limits basal autophagy activity and that ER fragments containing misfolded proteins and aggregates form in Sel1L-deficient adipocytes establish this ERAD branch as a central player in the maintenance of ER homeostasis, at least under physiological condition. SEL1L-HRD1 ERAD not only degrades misfolded proteins, but also may affect ER-phagy activity by limiting its substrate (ER fragments) availability and the autophagy activity under basal conditions. Our data provide an in vivo visualization of ER fragments in mammalian cells, and their subsequent coalescence into CERFs when ER-phagy is absent. These findings highlight the importance of the crosstalk among the branches of ER quality control pathways in the maintenance of ER homeostasis under physiological settings, which underscores the profound ability of cells to clear misfolded proteins from the ER (Fig. 9).
SEL1L-HRD1 ERAD plays a major role in the degradation of misfolded LPL in the ER under basal state. Without ERAD, ER fragments containing misfolded protein and aggregates form, which are cleared by autophagy, i.e. ER-phagy. In adipocytes lacking both SEL1L-HRD1 ERAD and autophagy, ER fragments accumulate, cluster and fuse to form CERFs.
Our data reveal a profound cellular adaptation capability in dealing with misfolded proteins or protein aggregates in the ER. Adipocytes were able to survive the lack of SEL1L-HRD1 ERAD, autophagy or both – two principal degradative mechanism in the cells – without instigating an overt UPR. We speculate that the formation of CERFs as a strategy to sequester potentially toxic protein aggerates in the ER, without triggering deleterious cellular consequences including overt UPR and cell death in adipocytes. CERFs are storage compartments for misfolded proteins and aggregates, which progressively enlarge and become insoluble with time. Intriguingly, this unique structure was not seen in β cells lacking SEL1L-HRD1 ERAD and autophagy as we recently reported42. The dynamic and intramembranous features of the CERFs make it unique and distinct from the homogenous ER inclusions previously reported in the ER storage diseases, such as those containing fibrinogen or α1 anti-trypsin in the liver, proAVP in the AVP neurons, and CFTR disease mutant53,54. Using IP-MS, we identified LPL as a protein substrate of SEL1L-HRD1 ERAD and a principal component of CERFs – a significant finding as it made possible subsequent biochemical and biophysical studies of the CERFs. Indeed, our data further show that these CREFs are also composed of many ER chaperones, which are involved in the folding and refolding of misfolded proteins or aggregates such as BiP, OS9 and MLEC.
We in vitro reconstituted the CERFs through LPL and BiP phase separation. Phase separation has been recently reported as an important mechanism for the formation of intracellular biological condensates55, including the Z-α1-antitrypsin ER inclusion56. Consistent with the notion that LPL protein is misfolding-prone31,43, and tends to polymerize into helical filaments52, our data reveal that purified LPL readily forms not only precipitates, but also phase separation in vitro. Unlike many other proteins57,58, phase separation of LPL is unlikely mediated by the low-complexity sequences as LPL is predicted to be an ordered protein by IUPred259. Instead, LPL oligomerization and phase separation may require protein-protein interactions among structured domains60. Indeed, our data showed that both redox environment and the tryptophan loop of LPL may play important roles in its phase separation. In addition, our data showed that BiP surrounds LPL in the CERFs and droplets in vivo and in vitro, respectively, and that the interaction with BiP may alter LPL physical properties, including solubility and mobility in the CERFs. Further studies are required to delineate the importance of LPL phase separation in its basal physiological function and trafficking61.
Our data demonstrate that SEL1L-HRD1 ERAD is a critical regulator of LPL biology by degrading misfolding LPL in the ER. If not cleared efficiently, these LPL forms aggregates in the ER, which are cleared by ER-phagy. We acknowledge that the use of Atg7 KO mouse model is not ideal to study the ER-phagy process per se. However, despite of our limited understanding of ER-phagy in vivo, we were able to visualize the ER-phagy process specifically with the use of powerful imaging tools including confocal microscopy, immunogold labeling and TEM. One notable hit in the IP-MS of LPL is a non-canonical ER-phagy receptor CCPG116 – an active investigation in our laboratory.
Under basal state with functional SEL1L-HRD1 ERAD, our data show that autophagy plays a minor role in the maturation of LPL in the ER. In both WT and Atg7AdipCre WAT, it should be noted that LPL does form small extra-ER puncta, likely representing those in storage vesicles as previously reported43,52,62. This finding is consistent with previous studies showing that inhibiting macroautophagy increased the amount of LPL at the cell surface63. By contrast, when SEL1L-HRD1 ERAD is impaired, autophagy/ER-phagy plays a major role in clearing misfolded LPL and its aggregates in fragmented ER. While molecular events underlying the formation of ER fragments and subsequent coalescence remain vague, this study uncovers the biochemical chain of events involved in disposing of misfolded protein in the ER of adipocytes. Further characterization of the pathways and mechanisms underlying the formation of ER fragments and cellular inclusions as well as their interaction with various quality control pathways in vivo is key to the understanding of the disease pathogenesis associated with ER protein misfolding, and basis for future therapy.
Methods
Mice
All animal procedures were approved by the Institutional Animal Care and Use Committee of the University of Michigan Medical School (PRO00010658) and in accordance with the National Institutes of Health (NIH) guidelines. All the mouse models used in this study were on the C57BL/6 J background. Sel1Lflox/flox 24, Atg7flox/flox64 and Sel1LAdipCre (Sel1Lflox/flox;Adipoq-Cre+/–)31 mice have been described previously. Atg7flox/flox mice were provided by Rajat Singh (University of California, Los Angeles, USA) with the permission of Masaaki Komatsu and Keiji Tanaka (Tokyo Metropolitan Institute of Medical Science, Tokyo, Japan). Atg7AdipCre (Atg7flox/flox;Adipoq-Cre+/–) mice were generated by breeding Atg7flox/flox mice with Adipoq-Cre mice65. The single knockouts were then intercrossed to generate adipocyte-specific (Sel1Lflox/flox;Atg7flox/flox;Adipoq-Cre+/–). Sel1LAdipCre;Atg7AdipCre/+ and Sel1LAdipCre/+;Atg7AdipCre mice were generated as Sel1LAdipCre and Atg7AdipCre littermate controls, respectively. Sel1Lflox/flox;Atg7flox/flox, Sel1Lflox/+;Atg7flox/flox, Sel1Lflox/flox;Atg7flox/+ and Sel1Lflox/+;Atg7flox/+ littermates were used as WT controls. Brown adipocyte-specific Sel1LUcp1Cre (Sel1Lflox/flox;Ucp1-Cre+/–) and Hrd1Ucp1Cre (Hrd1flox/flox;Ucp1-Cre+/–) were generated by breeding Sel1Lflox/flox and Hrd1flox/flox66 with Ucp1-Cre mice (The Jackson Laboratory, #024670) as previously described in ref. 39. Experiments were performed using age- and sex-matched littermates at the ages of 5 and 10–12 weeks. The Alzheimer’s disease 5xFAD female mice67 at the age of 6 months and their WT littermates were gifts form Dr. Geoffrey Murphy. All mice were housed in a pathogen-free animal facility at 22 ± 1 °C on a 12-h light/dark cycle with 40–60% humidity and fed a low-fat diet (13% fat, 57% carbohydrate, and 30% protein, LabDiet 5LOD), unless otherwise indicated.
Animal experiments
The fasting-refeeding and oil-gavage experiments were performed as described in ref. 31. Briefly, mice were fasted for 16 h with refeeding with HFD (60% fat, 20% carbohydrate and 20% protein, Research Diet D12492) for 3 h or olive oil gavage (1 μl/1 g body weight) for 2.5 h. For measurement of blood TG, 12-week-old mice were fasted for 16 h followed by oral gavage of olive oil. Blood was withdrawn through cardiac puncture after 2.5 h. TG levels in the serum were measured using LabAssay™ Triglyceride kit (Wako Chemicals, 290-63701) per manufacturer’s instruction.
Cell lines
Wildtype (WT) and inducible Sel1L-deficient (Sel1LERCre) pre-adipocytes were isolated from brown adipose tissue of mice and immortalized using SV40 large T antigen and differentiated into adipocytes as previously described in refs. 39,68. Cells were cultured at 37 °C with 5% CO2 in DMEM with 10% fetal bovine serum (Fisher Scientific). Cultures were passaged every 2–3 days, when 80% confluence was reached. Cells were treated with 400 nM 4-hydroxytamoxifen (4-OHT, Sigma H7904) at day three of differentiation for 2 days to generate Sel1L–/– adipocytes. To induce ER stress, differentiated adipocytes were incubated in fresh DMEM medium in the presence and absence of 5 µg/ml tunicamycin (Calbiochem) for 4 h as described in refs. 31,69. To measure the stability of LPL, fully differentiated adipocytes were incubated with 2 μg/ml brefeldin A (MilliporeSigma) for 30 min followed by the addition of 50 μg/ml cycloheximide (Calbiochem), as described in ref. 31.
To generate CRISPR/Cas9-based adipocytes, CRISPR-targeting oligonucleotides were designed for mouse SEL1L (5’-GCCAGCAACTACTTTGCCCG-3’) and HRD1 (5’-ATCCATGCGGCATGTCGGGC-3’). Target sequences were inserted into lentiCRISPR v2. The third-generation lentiviral vector system (RRE [gag/pol], VSV-G and REV) were used to generate lentiviral particles in HEK293T cells. Media containing lentivirus were harvested at 24 and 48 h post-transfection for cell infection. One day after infection, pre-adipocytes positive for CRISPR/Cas9 were selected in the medium containing 2 μg/ml puromycin for 72 h and then maintained in regular medium without selection drugs.
Western blot
Mouse tissues and cells were harvested, snap-frozen in liquid nitrogen, and processed for (non)-denaturing SDS-PAGE and Western blot as described in ref. 30. Primary antibodies used are: anti-HSP90 (1:10,000, Santa Cruz, sc-7947), anti-GAPDH (1:10,000, ProteinTech, 60004-1-Ig), anti-LPL (1:500, gift from Dr. Andre Bensadoun, Cornell University and R&D Systems AF7197-SP)31, anti-SEL1L (1:1000, Abcam, ab78298; 1:10,000, homemade)39, anti-HRD1 (1:200, gift from Richard Wojcikiewicz, Syracuse University; 1:1000, ProteinTech, 13473-1-AP)31, anti-ATG7 (1:1000, Cell Signaling, 8558 S), anti-P62 (1:5000, MBL, PM066; 1:5000, Enzo, BML-PW9860), anti-LC3B (1:2000, Cell Signaling, 2775 S), anti-BiP (1:2000, Abcam ab21685), anti-OS9 (1:2000, Abcam, ab109510), anti-GLUT4 (1:2000, ProteinTech, 66846-I-Ig), anti-CALNEXIN (1:5000, ProteinTech, 10427-2-AP), anti-MLEC (1:2000, ProteinTech, 26655-1-AP), anti-PLOD3 antibody (1:2000, 11027-1-AP), anti-CAV 1 (1:2000, ABclonal, A1555), anti-Caspase 3 (1:1000, Cell Signaling, 9662 S); anti-cleaved Caspase 3 (1:1000, Cell Signaling, 9661 S), anti-H2A (1:1000, Cell Signaling, 2578), anti-mCherry (1:1000, ABclonal, AE002), and anti-His (1:1000, ABclonal, AE003). Membranes were washed with TBST and incubated at room temperature for 1 h with secondary antibodies: donkey anti-goat IgG-HRP (1:5000, ThermoFisher, #PA1-28664), goat anti-rabbit IgG-HRP (1:5000, Bio-Rad, #1706515), goat anti-mouse IgG-HRP (1:5000, Bio-Rad, #1706516) and goat anti-guinea pig IgG-HRP (1:5000, Bio-Rad, #AHP863P). The uncropped blots were provided in the Source Data file.
Detergent-solubility assay
The NP40 solubility assay was performed as described in ref. 30. Briefly, frozen WAT was lysed in cold NP40 lysis buffer (1% NP40, 50 mM Tris-HCL, PH 7.5, 150 mM NaCl, 1 mM EDTA) with complete protease inhibitors (MilliporeSigma) and PhosSTOP (MilliporeSigma) by gentle sonication. After centrifugation at 13,400 g for 20 min at 4 °C, the supernatant (NP40S fraction) was collected and protein concentration in soluble fraction was determined using Bradford assay followed by normalization. The supernatant was mixed with 5x SDS protein sample buffer and heated at 95 °C for 5 min. The pellet (NP40P fraction) was washed with lysis buffer, centrifuged to remove fat and mixed with 1x reducing SDS-sample buffer with ¼ volume of total volume of the soluble fraction. The pellet was then gently sonicated and heated at 95 °C for 30 min. Samples of soluble and pellet fractions (10 μl each) were loaded onto the gel. Bands intensity of soluble (S) and pellet (P) fractions were determined using Image lab (Bio-Rad). To quantitate the relative amount of protein in pellet fraction (P%), the band intensity of pellet fraction was divided by 4 (P/4) and P% was calculated as (P/4) / [(P/4) + S] x 100%.
Immunoprecipitation (IP) and IP-based LC-Tandem Mass spectrometry (IP-MS)
For SEL1L IP-MS, BAT from 12-week-old female mice (one per genotype) was used. LPL IP-MS was performed in two separate experiments using frozen WAT pooled from 12-week-old mice (5 WT, 6 Sel1LAdipCre and 7 DKO mice in experiment #1; and 4 WT and 3 Atg7AdipCre mice in experiment #2). Tissues were lysed in cold IP lysis buffer (1% Triton X-100, 150 mM NaCl, 25 mM Tris-HCl pH 7.5), supplemented with protease and phosphatase inhibitors. Overall 5–8 mg of protein lysates was used for IP, pre-cleared with Protein G/A agarose beads for 4 h at 4 °C. Samples were then incubated with IgG (MBL, PM094 or Cell Signaling, #2729), LPL antibody (2 mg/ml, gift from Dr. Andre Bensadoun)31 or SEL1L antibody (2 mg/ml, homemade)39 overnight at 4 °C. Immunocomplexes were precipitated by mixing samples with 150 µl Protein G/A agarose beads under gentle rocking for 2 h at 4°C. The beads were washed with cold IP buffer followed by cold PBS for three times at 4°C. The beads containing immunocomplexes were stored at −80 °C until further analysis. The samples were submitted to the Proteomics Resource Facility at the University of Michigan Medical School on a fee-for-service basis. Proteins were identified by searching the MS/MS data using Proteome Discoverer (v2.4, Thermo Scientific). False discovery rate (FDR) was determined using Percolator and proteins/peptides with a FDR of ≤1% were retained for further analysis. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE70 partner repository with the dataset identifier PXD038310 and PXD040899. For Western blot, proteins on the beads were eluted using 2X protein sample buffer and heated at 95 °C for 5 min. For LPL Western blot, Trueblot anti-goat IgG HRP (1:1000, Rockland, mouse monoclonal eB270, #18-8814-31) was used as secondary antibody.
In SEL1L IP-MS, top protein hits enriched in Hrd1Ucp1Cre BAT were identified based on the criteria: peptide spectrum matches (PSM) in Hrd1Ucp1Cre ≥ 2, PSMs in Hrd1Ucp1Cre / WT ≥ 2, and PSMs in Hrd1Ucp1Cre / Sel1LUcp1Cre ≥ 4 and PSMs in Hrd1Ucp1Cre / IgG ≥ 4. The full list of identified hits with raw PSM values was provided in Supplementary Table 1.
In LPL IP-MS: protein hits enriched in WT versus DKO WAT (Cluster 1) were identified based on the criteria: PSMs in WT/ DKO ≥ 2, PSMs in WT ≥ 2 and PSMs in IgG =0. Protein hits enriched in DKO versus WT (Cluster 2) were identified based on the criteria: PSMs in DKO / WT ≥ 4, PSMs in DKO ≥ 4, and PSMs in IgG =0. The full list of identified hits with raw PSM values was provided in Supplementary Tables 2 and 3. Heatmaps shown in Supplementary Fig. 6b and c were presented as fold change to WT in each experiment.
Histology and immunofluorescent staining
WAT was dissected and fixed in 10% neutral buffered formalin (VWR 95042-908) and further processed for embedding, sectioning and H&E staining by the Histology Core at the University of Michigan Medical School. The H&E slides were imaged using Aperio Scanscope (Leica). TUNEL labeling was performed per manufacturer’s protocol using the In-Situ Cell Death detection kit (Roche, 11684795910). For positive control, paraffin section of WAT was incubated with 1500 U/ml recombinant DNase I (Omega, E1091) at 37 °C for 30 min per manufacture’s instruction.
For immunofluorescence labeling, paraffin-embedded WAT was sectioned at 8 µm and deparaffinized in xylene and re-hydrated using graded ethanol series and distilled water. The antigen retrieval was performed by boiling the slides in 1 mM EDTA (pH 8.0) or citric acid buffer (pH 6.0) using microwave oven for 25 min, followed by blocking (10% normal donkey serum, 0.4% tritonX-100 in PBS) at room temperature for 1 h. The slides were incubated at 4 °C overnight with primary antibody: anti-LPL (1:200, gift from Dr. Andre Bensadoun;31 1:100, R&D Systems AF7197-SP), anti-GPIHBP1 (1:250, gift from Dr. Stephen Young);71 anti-KDEL (1:500, Novus NBP1-97469), anti-BiP (1:500, Abcam ab21685), anti-P62 (1:500, MBL, PM066), anti-P62 (1:500, Enzo, BML-PW9860), anti-OS9 (1:250, Abcam, ab109510), anti-PLOD3 antibody (1:500, ProteinTech,11027-1-AP), anti-CAV1 (1:250, ABclonal, A1555), anti-MLEC (1:200, ProteinTech, 26655-1-AP). Next day, the slides were incubated at room temperature for 1 h with secondary antibodies: 1:500 Alexa fluor 488 affinipure donkey anti-goat IgG (H + L) (Jackson ImmunoResearch, 705-546-147); 1:500 Alexa fluor plus 555 donkey anti-goat IgG (H + L) (Invitrogen, A32816); 1:500 Alexa fluor plus 647 donkey anti-goat IgG (H + L) (Invitrogen, A32849); 1:500 Alexa fluor 647 affinipure donkey anti-rat IgG (H + L) (Jackson ImmunoResearch, 712-606-150); 1:500 Alexa fluor plus 555 donkey anti-mouse IgG (H + L) (Invitrogen, A32773); 1:500 Alexa fluor 488 affinipure donkey anti-rabbit IgG (H + L) (Jackson ImmunoResearch, 711-546-152); 1:500 Alexa fluor plus 555 donkey anti-rabbit IgG (H + L) (Invitrogen, A32794); 1:500 Alexa fluor 647 affinipure donkey anti-rabbit IgG (H + L) (Jackson ImmunoResearch, 711-606-152); and 1:500 Alexa fluor 647 affinipure donkey anti-guinea pig IgG (H + L) (Jackson ImmunoResearch, 706-606-148). Both primary and secondary antibodies were diluted in PBS containing 10% normal donkey serum and 0.2% triton X-100. The nuclei and slides were stained and mounted by prolong gold antifade reagent with DAPI (ThermoFisher) followed by imaging using the Nikon A1 confocal microscope in Microscopy and Imaging Analysis Core (Michigan Diabetes Research Center, University of Michigan Medical School). The 3D re-construction of the immunofluorescent image was performed by Imaris (Oxford Instruments) at the Microscopy and Imaging Analysis Core.
Thioflavin S staining
For thioflavin staining, the paraffin-embedded WAT and brain sections were deparaffinized as mentioned above. The slides were then incubated with 1% thioflavin for 1 h at room temperature in dark followed by a series of washing steps in 80% ethanol, 70% ethanol, 50% ethanol and distilled water. Then, the slides were mounted and dried overnight followed by imaging using the Leica SP5 confocal microscope at the Microscopy and Imaging Analysis Core.
Transmission electron microscopy (TEM)
The TEM of adipose tissue was performed as described in ref. 39. Briefly, mice were anesthetized and perfused with 2.5% glutaraldehyde, 4% formaldehyde in 0.1 M sodium cacodylate buffer (Electron Microscopy Sciences). WAT and BAT were immediately dissected and fixed overnight at 4 °C in the perfusion buffer. In some experiments, WAT was quickly removed without perfusion and fully immersed in the fixation buffer (2.5% glutaraldehyde and 4% paraformaldehyde in 0.1 M sodium cacodylate) for 1-h incubation at room temperature and overnight incubation at 4 °C. The fixed tissue was cut into small pieces (about 1-2 mm cubes) and then submitted to the University of Michigan Microscopy Core for washing, embedding and sectioning at 70 nm. The grid containing tissue sections was imaged by a JEOL 1400-plus electron microscope (JEOL) at the University of Michigan Microscopy Core.
Focused ion beam-SEM (FIB-SEM)
The FIB-SEM was performed as described previously in ref. 39. In brief, the fixed white adipose tissue cubes were sent to the University of Michigan Microscopy Core for embedding. The sample block was then submitted at the Michigan Center for Materials Characterization (University of Michigan) for FIB-SEM analysis using a FEI Helios Nanolab 650 DualBeam. After determining the location of adipocyte with CERFs, the block was sequentially cut at 10 nm and imaged to obtain an image stack for tomographic reconstruction. The field of view of the SEM and the slice thickness were set to produce voxel size 5 × 6 × 10. The resulting stack of images was aligned and reconstructed using Avizo v.9.3 (Thermo Fisher Scientific) and Imaris x64 9.5.1 (Oxford Instruments). Segmentation was performed using AIVIA 10 (Leica): manual segmentation was performed to determine the boundaries of the CERF in 30 images of the tomographic dataset, which was then used to train the software to segment remaining images.
Immunogold labeling for TEM
WAT was fixed in buffer containing 3% formaldehyde, 0.2 M sucrose in 0.1 M Sorensen’s phosphate buffer (PH = 7.2) and cut into small pieces and processed by the University of Michigan Microscopy Core. The ultra-thin sections (80 nm) were cut, placed bare nickel grid and stored at 4 °C. For immunogold labeling, the grids were quenched in 80 mM glycine and incubated in blocking solution (EMS 25599) containing 0.2% Tween-20 for 1 h at room temperature followed by overnight incubation with primary antibodies: anti-LPL (1:70, gift from Dr. Andre Bensadoun; 1:50, R&D Systems AF7197-SP), anti-KDEL (1:50, Novus NBP1-97469), anti-BiP (1:50, Abcam ab21685), PLOD3 (1:10, ProteinTech,11027-1-AP), and anti-P62 (1:50, Enzo, BML-PW9860) at 4 °C in a humidifying chamber. The grids were next incubated with colloidal gold antibodies (1:25, Jackson ImmunoResearch) for 1 h at room temperature followed by post-fixation in 1% glutaraldehyde and contrast staining using 0.5% uranyl acetate. After carbon evaporation, the grids were imaged by JEOL electron microscope at the University of Michigan Microscopy Core.
RT-PCR, qPCR and Microarray
Total RNA was extracted from tissues and cells using Trizol and BCP phase separation reagent (Molecular Research Center). RT–PCR for Xbp1 splicing (F:ACGAGGTTCCAGAGGTGGAG; R: AAGAGGCAACAGTGTCAGAG) and l32 (F: GAGCAACAAGAAAACCAAGCA; R: TGCACACAAGCCATCTACTCA) was performed as previously described in ref. 69. The ratio of Xbp1s level to total Xbp1 level was quantified by Image lab software (BioRad). All qPCR data for mouse tissues were normalized to the expression level of ribosomal 18 s. qPCR primer sequences are: Lpl F: CGAGAGGATCCGAGTGAAAG, R: TTTGTCCAGTGTCAGCCAGA; and 18 s F: GCAATTATTCCCCATGAACG, R: GGGACTTAATCAACGCAAGC. Microarray data31 (accession number GSE56918) were processed using the limma package72 (version 3.52.0) in the R environment (version 4.2.0). Genes with higher mean intensities in Sel1LAdipCre than in WT samples and q values smaller than 0.05 from 2-tailed paired intensity-based moderated t-statistics (IBMT) were selected as input for pathway enrichment analysis. The pathways enrichment analysis was performed using the enrich KEGG function implemented in the clusterProfiler package73 (version 4.4.1) in R, with the background genes adjusted based on the genes detected in the microarray data.
Protein expression and purification
Human LPL (amino acids 1–475) or BiP (amino acids 1-645) was inserted into pETL7a vector, with mCherry or EGFP tag at the N-terminus and 8×His-tag at the C-terminus. LPL truncations or mutations were constructed using standard mutagenesis procedures. EGFP-BiP, mCherry-LPL, and their mutants were over-expressed in E. coli strain BL21 (DE3). The bacteria were cultured in LB medium at 37 °C in a shaker incubator to OD600 0.6-0.8, then induced with 0.5 mM IPTG for 16 h at 16 °C. The bacteria were collected, resuspended in lysis buffer (20 mM Tris-HCl, 150 mM NaCl, 20 mM β-ME, 4 M Urea, 5% glycerol, 0.1 mM PMSF, 1× protease inhibitor cocktail, pH 7.5), and sonicated for 30 min on ice (180 W, 10 s on and 10 s off). The lysates were collected, and the supernatant was loaded onto Ni2+-NTA resin. The column was washed with wash buffer (lysis buffer with 30 mM imidazole) and eluted with elution buffer (lysis buffer with 300 mM imidazole). Protein purity was confirmed by SDS-PAGE.
In vitro phase separation
To obtain phase-separated mCherry-LPL and EGFP-BiP condensates, purified recombinant proteins were dialyzed at 4 °C overnight in phase separation buffer (20 mM Tris-HCl, 150 mM NaCl, 20 mM β-ME, 1 M Urea, pH 7.5). Images of the condensates were taken by Olympus SpinSR spinning disk confocal super-resolution microscope. For negative staining, the phase separated droplets were stained with 1 % uranyl acetate and imaged by Hitachi HT7800 transmission electron microscope at Tsinghua University.
Fluorescence recovery after photobleaching (FRAP)
FRAP experiments were carried out using Nikon A1 confocal microscope. The mCherry-LPL with or without EGFP-BiP were bleached using 488-nm or 561-nm laser beam. Recovery from photobleaching was recorded at the indicated times.
Turbidity assays
Proteins were diluted in a buffer containing 20 mM Tris-HCl, 150 mM NaCl, and pH 7.5, to desired concentrations. The absorbance at 395 nm (turbidity) was monitored by the Bio-Tek Cytation 5 imaging reader.
Statistical analyses
All results are presented as mean ± standard deviation (SD) using Graphpad Prism8 (Graph Pad Software Inc.). Most experiments have been independently repeated for at least twice with multiple independent biological samples from which representative data are shown. IP-MS were performed once as the experiments were intended to identify hits and generate hypothesis. Statistical differences were performed using unpaired two-tailed Student’s t-test for two groups and One-way or Two-way ANOVA analysis with a post hoc Tukey’s multiple comparisons test for more than 2 groups. P values less than 0.05 were considered statistically significant. The frequency distribution of lipid droplet size between WT and DKO WAT was compared using Chi-square test. In Supplementary Fig.6d, the P values are calculated by two-tailed rate ratio tests and adjusted for multiple testing by False Discovery Rate (FDR). The details for adjustment were described previously in ref. 74.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Proteomics datasets have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifiers PXD038310 and PXD040899. The predicted structure of LPL is available at PDB ID: 6U7B. Other data supporting the findings of this study are available within the article and in the Supplementary Information and Source Data files. Source data are provided with the paper. Source data are provided with this paper.
References
Balch, W. E., Morimoto, R. I., Dillin, A. & Kelly, J. W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Kopito, R. R. Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol 10, 524–530 (2000).
Ferro-Novick, S., Reggiori, F. & Brodsky, J. L. ER-Phagy, ER homeostasis, and ER quality control: Implications for disease. Trends Biochem Sci 46, 630–639 (2021).
Hwang, J. & Qi, L. Quality control in the endoplasmic reticulum: Crosstalk between ERAD and UPR pathways. Trends Biochem Sci 43, 593–605 (2018).
Bhattacharya, A. & Qi, L. ER-associated degradation in health and disease - from substrate to organism. J Cell Sci 132, jcs232850 (2019).
Hamasaki, M., Noda, T., Baba, M. & Ohsumi, Y. Starvation triggers the delivery of the endoplasmic reticulum to the vacuole via autophagy in yeast. Traffic 6, 56–65 (2005).
Khaminets, A. et al. Regulation of endoplasmic reticulum turnover by selective autophagy. Nature 522, 354–358 (2015).
Mochida, K. et al. Receptor-mediated selective autophagy degrades the endoplasmic reticulum and the nucleus. Nature 522, 359–362 (2015).
Cunningham, C. N. et al. Cells deploy a two-pronged strategy to rectify misfolded proinsulin aggregates. Mol Cell 75, 442–456. e444 (2019).
Forrester, A. et al. A selective ER‐phagy exerts procollagen quality control via a Calnexin‐FAM 134B complex. EMBO J 38, e99847 (2019).
Omari, S. et al. Noncanonical autophagy at ER exit sites regulates procollagen turnover. Proc Nat Acad Sci 115, E10099–E10108 (2018).
Parashar, S. et al. Endoplasmic reticulum tubules limit the size of misfolded protein condensates. eLife 10, e71642 (2021).
Chen, Y. J. et al. PGRMC1 acts as a size-selective cargo receptor to drive ER-phagic clearance of mutant prohormones. Nat Commun 12, 5991 (2021).
Schultz, M. L. et al. Coordinate regulation of mutant NPC1 degradation by selective ER autophagy and MARCH6-dependent ERAD. Nat Commun 9, 3671 (2018).
Houck, S. A. et al. Quality control autophagy degrades soluble ERAD-resistant conformers of the misfolded membrane protein GnRHR. Mol Cell 54, 166–179 (2014).
Smith, M. D. et al. CCPG1 is a non-canonical autophagy cargo receptor essential for ER-phagy and pancreatic ER proteostasis. Dev Cell 44, 217–232.e211 (2018).
Shi, Q. et al. Impact of RTN3 deficiency on expression of BACE1 and amyloid deposition. J Neurosci 34, 13954–13962 (2014).
Grumati, P., Dikic, I. & Stolz, A. ER-phagy at a glance. J Cell Sci 131, jcs17364 (2018).
Chino, H. & Mizushima, N. ER-phagy: Quality control and turnover of endoplasmic reticulum. Trends Cell Biol 30, 384–398 (2020).
Qi, L., Tsai, B. & Arvan, P. New insights into the physiological role of endoplasmic reticulum-associated degradation. Trends Cell Biol 27, 430–440 (2017).
Carvalho, P., Goder, V. & Rapoport, T. A. Distinct ubiquitin-ligase complexes define convergent pathways for the degradation of ER proteins. Cell 126, 361–373 (2006).
Gardner, R. G. et al. Endoplasmic reticulum degradation requires lumen to cytosol signaling. Transmembrane control of Hrd1p by Hrd3p. J Cell Biol 151, 69–82 (2000).
Hampton, R. Y., Gardner, R. G. & Rine, J. Role of 26S proteasome and HRD genes in the degradation of 3-hydroxy-3-methylglutaryl-CoA reductase, an integral endoplasmic reticulum membrane protein. Mol Biol Cell 7, 2029–2044 (1996).
Sun, S. et al. Sel1L is indispensable for mammalian endoplasmic reticulum-associated degradation, endoplasmic reticulum homeostasis, and survival. Proc Natl Acad Sci USA 111, E582–E591 (2014).
Vashistha, N., Neal, S. E., Singh, A., Carroll, S. M. & Hampton, R. Y. Direct and essential function for Hrd3 in ER-associated degradation. Proc Natl Acad Sci USA 113, 5934–5939 (2016).
Francisco, A. B. et al. Deficiency of suppressor enhancer lin12 1 like (SEL1L) in mice leads to systemic endoplasmic reticulum stress and embryonic lethality. J Biol Chem 285, 13694–13703 (2010).
Yagishita, N. et al. Essential role of synoviolin in embryogenesis. J Biol Chem 280, 7909–7916 (2005).
Fujita, H. et al. The E3 ligase synoviolin controls body weight and mitochondrial biogenesis through negative regulation of PGC-1beta. EMBO J 34, 1042–1055 (2015).
Kim, G. H. et al. Hypothalamic ER-associated degradation regulates POMC maturation, feeding and age-associated obesity. J Clin Invest 128, 1125–1140 (2018).
Shi, G. et al. ER-associated degradation is required for vasopressin prohormone processing and systemic water homeostasis. J Clin Invest 127, 3897–3912 (2017).
Sha, H. et al. The ER-associated degradation adaptor protein Sel1L regulates LPL secretion and lipid metabolism. Cell Metab 20, 458–470 (2014).
Bhattacharya, A. et al. Hepatic Sel1L-Hrd1 ER-associated degradation (ERAD) manages FGF21 levels and systemic metabolism via CREBH. EMBO J 37, e99277 (2018).
Ji, Y. et al. The Sel1L-Hrd1 endoplasmic reticulum-associated degradation complex manages a key checkpoint in B cell development. Cell Rep 16, 2630–2640 (2016).
Yang, Y. et al. The endoplasmic reticulum-resident E3 ubiquitin ligase Hrd1 controls a critical checkpoint in B cell development in mice. J Biol Chem 293, 12934–12944 (2018).
Wei, J. et al. HRD1-ERAD controls production of the hepatokine FGF21 through CREBH polyubiquitination. EMBO J 37, e98942 (2018).
Sun, S. et al. Epithelial Sel1L is required for the maintenance of intestinal homeostasis. Mol Biol Cell 27, 483–490 (2016).
Yoshida, S. et al. Endoplasmic reticulum-associated degradation is required for nephrin maturation and kidney glomerular filtration function. J Clin Invest 131, e143988 (2021).
Shrestha, N. et al. Sel1L-Hrd1 ER-associated degradation maintains β cell identity via TGFβ signaling. J Clin Invest 130, 3499–3510 (2020).
Zhou, Z. et al. Endoplasmic reticulum-associated degradation regulates mitochondrial dynamics in brown adipocytes. Science 368, 54–60 (2020).
Thepsuwan, P. et al. Hepatic SEL1L-HRD1 ER-associated degradation regulates systemic iron homeostasis via ceruloplasmin. Proc Natl Acad Sci USA 120, e2212644120 (2023).
Bhattacharya, A. et al. SEL1L-HRD1 ER-associated degradation suppresses hepatocyte hyperproliferation and liver cancer. iScience 25, 105183 (2022).
Shrestha, N. et al. Integration of ER protein quality control mechanisms defines beta cell function and ER architecture. J Clin Invest 133, e163584 (2023).
Wu, S. A., Kersten, S. & Qi, L. Lipoprotein lipase and its regulators: An unfolding story. Trends Endocrinol Metab 32, 48–61 (2021).
Bjorkoy, G. et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 171, 603–614 (2005).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem 282, 24131–24145 (2007).
Sidrauski, C. & Walter, P. The transmembrane kinase Ire1p is a site-specific endonuclease that initiates mRNA splicing in the unfolded protein response. Cell 90, 1031–1039 (1997).
Qi, L., Yang, L. & Chen, H. Detecting and quantitating physiological endoplasmic reticulum stress. Meth Enzymol 490, 137–146 (2011).
Cha-Molstad, H. et al. Amino-terminal arginylation targets endoplasmic reticulum chaperone BiP for autophagy through p62 binding. Nat Cell Biol 17, 917–929 (2015).
Yang, H. et al. Sequestosome 1/p62 protein is associated with autophagic removal of excess hepatic endoplasmic reticulum in mice. J Biol Chem 291, 18663–18674 (2016).
Christianson, J. C. et al. Defining human ERAD networks through an integrative mapping strategy. Nat Cell Biol 14, 93–105 (2011).
Lederkremer, G. Z. Glycoprotein folding, quality control and ER-associated degradation. Current Opn Struct Biol 19, 515–523 (2009).
Gunn, K. H. et al. The structure of helical lipoprotein lipase reveals an unexpected twist in lipase storage. Proc Natl Acad Sci USA 117, 10254–10264 (2020).
Li, H. & Sun, S. Protein Aggregation in the ER: Calm behind the Storm. Cells 10, 3337 (2021).
Johnston, J. A., Illing, M. E. & Kopito, R. R. Cytoplasmic dynein/dynactin mediates the assembly of aggresomes. Cell Motility Cytoskel 53, 26–38 (2002).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18, 285–298 (2017).
Chambers, J. E. et al. Z-α1-antitrypsin polymers impose molecular filtration in the endoplasmic reticulum after undergoing phase transition to a solid state. Sci Adv 8, eabm2094 (2022).
Frey, S., Richter, R. P. & Gorlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science 314, 815–817 (2006).
Kato, M. & McKnight, S. L. A solid-state conceptualization of information transfer from gene to message to protein. Ann Rev Biochem 87, 351–390 (2018).
Dosztányi, Z. Prediction of protein disorder based on IUPred. Prot Sci 27, 331–340 (2018).
Li, P. et al. Phase transitions in the assembly of multivalent signalling proteins. Nature 483, 336–340 (2012).
Birrane, G. et al. Structure of the lipoprotein lipase–GPIHBP1 complex that mediates plasma triglyceride hydrolysis. Proc Nat Acad Sci 116, 1723–1732 (2019).
Roberts, B. S., Yang, C. Q. & Neher, S. B. Characterization of lipoprotein lipase storage vesicles in 3T3-L1 adipocytes. J Cell Sci 135, jcs258734 (2022).
An, M. et al. ULK1 prevents cardiac dysfunction in obesity through autophagy-meditated regulation of lipid metabolism. Cardiovasc Res 113, 1137–1147 (2017).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol 169, 425–434 (2005).
Eguchi, J. et al. Transcriptional control of adipose lipid handling by IRF4. Cell Metabol 13, 249–259 (2011).
Wu, T. et al. Hrd1 suppresses Nrf2-mediated cellular protection during liver cirrhosis. Genes Dev 28, 708–722 (2014).
Ghoweri, A. O. et al. Electrophysiological and imaging calcium biomarkers of aging in male and female 5× FAD mice. J Alzheimer’s Dis 78, 1419–1438 (2020).
Klein, J., Fasshauer, M., Klein, H. H., Benito, M. & Kahn, C. R. Novel adipocyte lines from brown fat: a model system for the study of differentiation, energy metabolism, and insulin action. Bioessays 24, 382–388 (2002).
Sha, H. et al. The IRE1α-XBP1 pathway of the unfolded protein response is required for adipogenesis. Cell Metabol 9, 556–564 (2009).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res 50, D543–D552 (2022).
Beigneux, A. P. et al. Highly conserved cysteines within the Ly6 domain of GPIHBP1 are crucial for the binding of lipoprotein lipase. J Biolog Chem 284, 30240–30247 (2009).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucl Acids Res 43, e47–e47 (2015).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Wei, X., Zhang, C., Freddolino, P. L. & Zhang, Y. Detecting Gene Ontology misannotations using taxon-specific rate ratio comparisons. Bioinformatics 36, 4383–4388 (2020).
Acknowledgements
We thank Drs. Andre Bensadoun, Geoffrey Murphy, and Stephen Young for providing reagents; Dr. Haibo Sha at Novartis Institutes for BioMedical Research for some pilot experiments; Dr Saskia Neher and members in the Arvan/Qi laboratories for technical assistance and insightful discussions. We also acknowledge the use of the instruments and the staff assistance from the Proteomics Resource Facility, the Microscopy Core, Microscopy and Imaging Analysis Core at the University of Michigan. This work is supported by grants from the Ministry of Science and Technology (2022ZD0213900 and 2022ZD0204900), National Natural Science Foundation of China (32170684 to Y.L), R01DK128077, R01DK132068 (S.S.), 1R01DK120330 (L.Q. and D.F.), 1R01DK120047 (L.Q.), and Protein Folding Disease Initiative (L.Q. and M.I.). S.A.W. is supported by American Heart Association Pre-doctoral Fellowship (828841), and Barbour Scholarship and Rackham International Student Fellowship at University of Michigan. X.W. is supported in part by Pandemic Research Recovery Grant (U078128). L.L.L. is supported in part by National Ataxia Foundation Post-Doctoral Fellowship (918037). M.T. is a Pew Latin American Postdoctoral Fellow.
Author information
Authors and Affiliations
Contributions
S.A.W. designed and performed most experiments; Y.L. and C.S. designed and performed the in vitro experiments; X.W. and Y.L. performed analysis of proteomics and microarray data; X.Z., M.T., S.W., X.C., L.L.L., A.H.H., and H.W. assisted with some experiments; S.S., D.F., and M.I. provided reagents and insightful discussion; L.Q. designed and directed the study; L.Q. and S.A.W. wrote the manuscript; all authors commented on and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Simon Wilkinson, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, S.A., Shen, C., Wei, X. et al. The mechanisms to dispose of misfolded proteins in the endoplasmic reticulum of adipocytes. Nat Commun 14, 3132 (2023). https://doi.org/10.1038/s41467-023-38690-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-38690-4
This article is cited by
-
SEL1L-HRD1 interaction is required to form a functional HRD1 ERAD complex
Nature Communications (2024)
-
Proteomic screens of SEL1L-HRD1 ER-associated degradation substrates reveal its role in glycosylphosphatidylinositol-anchored protein biogenesis
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.