Abstract
Shoot branching is fundamentally important in determining soybean yield. Here, through genome-wide association study, we identify one predominant association locus on chromosome 18 that confers soybean branch number in the natural population. Further analyses determine that Dt2 is the corresponding gene and the natural variations in Dt2 result in significant differential transcriptional levels between the two major haplotypes. Functional characterization reveals that Dt2 interacts with GmAgl22 and GmSoc1a to physically bind to the promoters of GmAp1a and GmAp1d and to activate their transcription. Population genetic investigation show that the genetic differentiation of Dt2 display significant geographic structure. Our study provides a predominant gene for soybean branch number and may facilitate the breeding of high-yield soybean varieties.
Similar content being viewed by others
Introduction
Shoot branching is fundamentally important to the growth and productivity of crops1. By unconscious or conscious selection of the superior alleles of key genes that confer branching architecture, humans have significantly increased crop yields. One typical example is the selection and utilization of the TEOSINTE BRANCHED1 (TB1) gene during the domestication of maize2. TB1 and its orthologs play profound roles in modulating branching architecture via determination of the bud activation potential in plants1,3,4,5. Cultivated maize (Zea mays) was domesticated from teosinte (Z. mays ssp. parviglumis). Largely benefitting from the selection of a higher-expression allele of TB1, cultivated maize was successfully domesticated as a high-yield crop with a single culm, in contrast to its highly branched ancestor teosinte6. Another promising advance is the application of IPA1 in rice. A point mutation in IPA1 leads to an ideal rice plant with fewer tillers, increased plant height, lodging resistance, and thus enhanced grain yield7,8,9. By introducing beneficial ipa1 alleles into widely cultivated cultivars, a series of new elite varieties with higher yields were developed7. An increasing number of regulatory genes that control shoot branching are being identified1,10, providing genetic resources for breeding high-yield crops via molecular design.
Soybean (Glycine max (L.) Merr.) is one of the most important crops that supplies more than half of global oilseed production and approximately one quarter of the world’s plant protein11. With the increasing population and continuous improvement in people’s living standards, it was estimated that the soybean yield has to be doubled by 2050 to meet the consumption demands12. In contrast to the dramatic increase in yield of major crops, such as rice and wheat, which greatly benefitted from the Green Revolution, soybean yield has not been improved significantly in the past six decades. Modulating branch number, one of the most profound traits that determines the final yield of soybean, is crucial for high-yield soybean breeding13. However, to date, the genes associated with soybean branching have seldom been reported14.
Here, we show that natural variation in Dt2 predominantly determine soybean branching. We also reveal that Dt2 interact with GmAgl22 and GmSoc1a to bind to the promoters of GmAp1a and GmAp1d to regulate their transcriptions. Interestingly, the selection of Dt2 is associated with geographic differentiation. Modulating Dt2 lines resulted in significantly increase of soybean adaptation and yield, which is also associated with its effect on maturity.
Results
Natural variation in Dt2 predominantly determines soybean branching
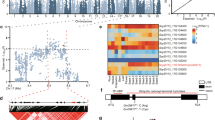
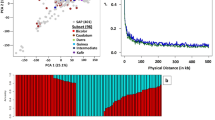
To identify the key genes that control branch number in soybean, we phenotyped 2409 accessions from our previous resequencing panel15 in 2017 and 2018. Phylogenetic and principal-component analysis of the landraces and cultivars did not show significant genetic differentiation (Supplementary Fig. 1a, b). Genome-wide association study (GWAS) performed using a mixed linear model revealed a stable association signal across the 2 years in a 40 kb interval block on chromosome 18 (Fig. 1a–d and Supplementary Fig. 1d–g). Within this 40 kb interval, a total of 5 protein-coding genes were annotated according to the reference genome ZH1316,17 (Fig. 1d), among which SoyZH13_18g242900 showed higher specific expression at the shoot apical meristem (Supplementary Fig. 2a), a tissue closely related to the final branching architecture1. Therefore, SoyZH13_18g242900 was considered to be the candidate gene controlling branch number in soybean.
a Manhattan plot of GWAS for branch number using 2 years of Blup data of the 2409 accessions. P values are calculated based on linear mixed model in GWAS and the dashed horizontal line indicates the genome-wide significance threshold (P = 1 × 10−7.9), which is determined by the Bonferroni test. −log10 P values are plotted against the position of SNPs on 20 chromosomes. b Quantile-quantile plot for the branch number. For quantile-quantile plot, −log10-transformed observed P values are plotted against −log10-transformed expected P values. c Genome-wide Manhattan plot in the 57–60 Mb region on chromosome 18. The red lead SNPs are shown above the threshold signals. d Linkage disequilibrium plot for SNPs in the 57.80–57.84 Mb region from a continuous association block. Navy blue box, genes. Pink box, candidate gene Dt2. Asterisk, position of the peak SNP. The color key (white to red) represents linkage disequilibrium value (r2) accessions. e Gene structure of Dt2. Two SNPs (−3259th T > C, −2580th T > A) in the promoter and a nonsynonymous SNP (+98th G > A) in the first exon are labeled on the gene sketch. f Haplotype analysis of Dt2HapI-1 (n = 1052 accessions), Dt2HapI-2 (n = 262 accessions) and Dt2HapII (n = 179 accessions). In each box plot (drawn by R 4.1.1 software), the center line indicates the median, the edges of the box represent the first and third quartiles, and the whiskers extend to span a 1.5 interquartile range from the edges. Different letters indicate statistically significant differences at P < 0.05 by one-way ANOVA test. g Promoter activity analysis of Dt2HapI and Dt2HapII using sequences 3,400-bp upstream from the translation initiation site (n = 5 biologically independent replicates), **P < 0.01, two-sided t-test, and P = 6.0 × 10−3. Data in (f, g) are the mean ± SEM. Source data are provided as a Source Data file.
SoyZH13_18g242900, also known as Dt2, has been found to play important roles in regulating multiple agronomic traits, including stem growth habit, plant height, and flowering time18,19,20. Whether it could modulate branch number has not been investigated. Phylogenetic analysis found that Dt2 belongs to MADS-box transcription factor family and shares high homology with AGL79 in Arabidopsis, a member of the AP1/FUL subfamily (Supplementary Fig. 2c). Of the association polymorphisms with minor allele frequency (MAF) >0.05, two SNPs from the promoter regions (3259 bp and 2580 bp upstream of the translation start site, respectively) and six SNPs from the introns showed higher association values than the threshold, and a G/A SNP from the first exon that changed amino acid serine to asparagine showed a lower association value than the threshold (Supplementary Fig. 2b). Based on these higher-value association SNPs and the G/A nonsynonymous SNP, three major haplotypes of Dt2: Dt2HapI-1, Dt2HapI-2, and Dt2HapII were classified in the natural population (Fig. 1e, f). We found that the accessions harboring Dt2HapI-1 and Dt2HapI-2 did not exhibit a significant difference in branch number, whereas Dt2HapII showed a significantly lower branch number than Dt2HapI-1 and Dt2HapI-2 (Fig. 1f).
Transient transcription activity assays suggested that the promoter sequence of Dt2HapII had significantly higher transcriptional activity than that of Dt2HapI (Fig. 1g). We then randomly selected 20 representative natural accessions and investigated the gene expression level of Dt2 and branch number. A negative correlation between the Dt2 expression level and branch number was observed (Supplementary Fig. 3a). Investigation of a pair of Dt2 near-isogenic lines (NILs) also revealed that the Dt2HapII line exhibited a significantly decreased branch number than the Dt2HapI-1 line (Supplementary Fig. 3b, c). Consistently, quantitative real-time PCR (RT-qPCR) assays showed that Dt2 expression in the Dt2HapII line was significantly higher than that in Dt2HapI-1 (Supplementary Fig. 3d). The above results suggested that the variation at the promoter of Dt2 played an important role in determining branch number in soybean.
Functional validation of the role of Dt2 in controlling branch number
To validate the function of Dt2 in determining branch number, we knocked out Dt2 in Dong Nong 50 (DN50), a variety harboring Dt2HapII with a mean of four branches, by the CRISPR/Cas9 system and obtained two independent homozygous knockout lines (named Dt2CR-1 and Dt2CR-2, respectively) (Supplementary Fig. 4). Field characterization demonstrated that the Dt2CR lines exhibited increased branch number compared with the wild type DN50 (Fig. 2a, b, f). The Dt2CR lines also showed significantly delayed flowering and maturity, and increased plant height and stem node number, which were consistent with its reported functions18,19 (Fig. 2g and Supplementary Fig. 5a–c). Moreover, we found that the Dt2CR lines exhibited multiple yield related trait changes, including higher 100 seed weight, longer seed length and width, and higher grain weight per plant, resulting in significantly increased yield per plot (Fig. 2h and Supplementary Fig. 6a, c–f).
a Architecture of DN50 and Dt2CR plants at the harvest stage. b Branching ability of DN50 and Dt2CR after harvest. c Architecture of DN50 and Dt2OE plants at the harvest stage. d Branching ability of DN50 and Dt2OE after harvest. e Relative expression level of Dt2 in DN50 and different transgenic lines (n = 3 biologically independent replicates). f Branch number of DN50, Dt2CR and Dt2OE lines (n = 70 biologically independent plants). g Flowering time of DN50, Dt2CR and Dt2OE lines (n = 30 biologically independent plants). DAE indicates days after emergence. h Plot yield of DN50, Dt2CR and Dt2OE (n = 4 biologically independent replicates). The plot area is 10 m2. Scale bars in the (a–d) are 10 cm. Data in (e–h) are the mean ± SEM. Statistical significance is determined using a two-sided t-test. Source data are provided as a Source Data file.
We further overexpressed the coding sequence (CDS) of Dt2 driven by the 35 S promoter in DN50 and obtained two independent transgenic overexpression lines (named Dt2OE-1 and Dt2OE-2, respectively). RT-qPCR assays showed that the expression of Dt2 was increased approximately threefold in the Dt2OE lines compared to the wild type DN50 (Fig. 2e). In contrast to the results from the Dt2CR lines, the Dt2OE lines exhibited decreased branch number, promoted flowering time (Fig. 2c, d, f, h) and maturity, decreased plant height, decreased stem node number (Supplementary Fig. 5), decreased pod number per plant, and decreased seed length and width, thus exhibiting decreased yield per plot (Fig. 2h and Supplementary Fig. 6b, c–f). These experiments showed that Dt2 negatively regulated branch number in soybean. Growth state statistics of axillary buds in leaf axils between DN50 and Dt2CR lines showed that the effect of Dt2 on branch development may be related to inflorescence determinacy (Supplementary Fig. 7).
Dt2 interacts with GmAgl22 and GmSoc1a
Transcriptional profiling of different tissues showed that Dt2 was highly expressed in the lateral buds and shoot tip (Supplementary Fig. 8a). An in situ hybridization assay demonstrated that Dt2 was specifically expressed in the axillary meristem and shoot apical meristem (Supplementary Fig. 8b). To identify the protein interaction partners of Dt2, we carried out a yeast two-hybrid (Y2H) assay by screening the cDNA library constructed from the lateral buds and identified 136 unique cDNA clones that corresponded to 28 genes (Supplementary Table 1).
Of the interacting proteins, an ortholog of Arabidopsis SHORT VEGETATIVE PHASE (SVP)/AGL22, named GmAgl22 in this study, showed more repeats and was of interest to us. GmAgl22 encodes a MADS-box transcription factor in Arabidopsis and functions as a floral repressor in the thermosensory pathway21,22,23. The interaction between Dt2 and GmAgl22 was further confirmed by point-to-point Y2H, bimolecular florescence complementation (BiFC), split luciferase complementation (split-LUC) and coimmunoprecipitation (Co-IP) assays (Fig. 3a–c and Supplementary Fig. 9a, b). Detailed investigation demonstrated that the K-box domain of Dt2 was essential for the physical interaction between Dt2 and GmAgl22 (Supplementary Fig. 9d). In addition, we found that GmAgl22 could interact with itself in Y2H, split-LUC and Co-IP assays (Supplementary Fig. 10a–c), indicating that the protein develops dimers or polymers to exert its function. A previous study reported that Dt2 could interact with GmSoc1a to affect the soybean growth habit18,24. We suspected that GmAgl22 could also interact with GmSoc1a, which was then confirmed by Y2H, BiFC, split-LUC and Co-IP assays (Fig. 3d–f and Supplementary Fig. 9c). Expression pattern analysis found that GmAgl22 and GmSoc1a were highly expressed in the lateral bud and shoot tips (Supplementary Fig. 10d, e). In situ hybridization assays demonstrated that GmAgl22 and GmSoc1a, similar to Dt2, were specifically expressed in the axillary meristem in different developmental stages (Supplementary Fig. 11).
a Interaction between Dt2 and GmAgl22 in the yeast two-hybrid assay. Transformed yeast cells are grown on DDO (Trp/Leu) or QDO (Trp/Leu/His/Ade) synthetic dropout medium. The number at the top indicates three serial dilutions. AD, GAL4 activation domain; BD, GAL4 DNA-binding domain. b BiFC analysis of the interaction between Dt2 and GmAgl22. Scale bars, 10 mm. cYFP C-terminal portion of YFP. nYFP N-terminal portion of YFP. c Co-IP analysis of the protein interactions between Dt2 and GmAgl22. Flag-tagged GmAgl22 is co-transformed with HA-tagged Dt2 into Arabidopsis protoplasts. d Interaction between GmSoc1a and GmAgl22 in the yeast two-hybrid assay. e BiFC analysis of the interaction between GmSoc1a and GmAgl22. Scale bars, 10 mm. f Co-IP analysis of the protein interactions between GmSoc1a and GmAgl22. g Phenotypic comparison between W82 and GmSoc1aCR lines. Scale bars, 20 cm. h Branch number statistic in W82 and GmSoc1aCR lines (n = 20 biologically independent plants). i Phenotypic comparison between DN50 and GmAgl22OE lines. Scale bar, 10 cm. j Branch number statistic in DN50 and GmAgl22OE lines (n = 10 biologically independent plants). All the data in the graphs represent the mean ± SEM. Statistical significance is determined using a two-sided t-test. For (b, e), at least 5 independent cells are observed and a representative result is shown. For (c, f), at least 3 independent replicates are performed and a representative result is shown. Source data are provided as a Source Data file.
Previously, we generated a GmSoc1a knockout mutant by the CRISPR/Cas9 system (named GmSoc1aCR) and determined that the mutation could significantly affect flowering time. We also obtained two GmAgl22 overexpression lines (named GmAgl22OE-1 and GmAgl22OE-2) (Supplementary Fig. 10f). We then compared the branch numbers of GmSoc1aCR and GmAgl22OE with those of non-transgenic parents, respectively. We found that the GmSoc1aCR line showed significantly increased branch number (Fig. 3g, h), and the GmAgl22OE lines showed significantly decreased branch number (Fig. 3i, j). These results indicated that Dt2, GmAgl22 and GmSoc1a may function by forming a complex to control branching in soybean.
Dt2 regulates the expression of the GmAp1 gene family
To mine the downstream targets and regulatory network of Dt2, transcriptome profiling by RNA-Seq was performed with the lateral buds from WT (DN50), Dt2CR and Dt2OE. Using a P value < 0.05 and the fold change larger than 2 as thresholds, 646 up-regulated and 296 down-regulated genes were identified as differentially expressed genes (DEGs) between the wild type and Dt2CR lines (WT/Dt2CR) and 1160 up-regulated and 464 down-regulated genes were identified between the wild type and Dt2OE lines (WT/Dt2OE) (Supplementary Fig. 12a–d). Gene Ontology (GO) term analysis demonstrated that the DEGs from the WT/Dt2CR and WT/Dt2OE lines were enriched among metabolic process terms. Notably, multiple terms were related to carbohydrate metabolism and photosynthesis processes (Supplementary Fig. 12e, f), which was consistent with previous findings that sugar metabolism or signaling played an important role in axillary bud outgrowth1,25,26.
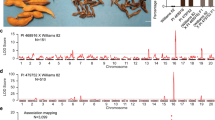
We then further narrowed down the DEGs by selecting the genes with an opposite pattern in the WT/Dt2OE and WT/Dt2CR panels: (1) up-regulated in the WT/Dt2OE panel but down-regulated in the WT/Dt2CR panel, and (2) down-regulated in the WT/Dt2OE panel but up-regulated in the WT/Dt2CR panel. In total, 30 genes meeting these criteria were identified (Fig. 4a). Interestingly, half of these 30 genes were annotated as agamous-liked genes, and most of these agamous-liked genes showed higher expression in the shoot meristem and flower (Supplementary Fig. 13a). Further comparison of the DEGs with previously reported Dt2 ChIP-seq data20 revealed that 4 genes might be the direct targets of Dt2: SoyZH13_16G083100 (GmAp1a), SoyZH13_01G060300 (GmAp1c), SoyZH13_02G112700 (GmAp1d) and SoyZH13_18G251400 (GmRPP13). The three GmAp1 gene family members were significantly up-regulated in the Dt2OE lines and down-regulated in the Dt2CR line, whereas GmPRR13 was down-regulated in the Dt2OE lines and up-regulated in the Dt2CR lines (Fig. 4a and Supplementary Fig. 13b). In this study, we focused on a functional assay of GmAp1 gene family using GmAp1a and GmAp1d as representatives. A previous in situ hybridization assay demonstrated that GmAp1a was specifically expressed in the shoot apices in the V2 stage27, and we also found that GmAp1a had a similar expression pattern as GmAp1d (Supplementary Fig. 13c), indicating that they may perform a similar function in branching development.
a Venn diagram among overlapping DEGs from DN50, Dt2CR and Dt2OE lines. b A probe sequence that could be bound by the MADS-box protein in the promoter of the GmAp1a is predicted in Plantpan (http://plantpan.itps.ncku.edu.tw/). c MBP-Dt2HapI-1 or MBP-Dt2HapI-2 directly bind to the promoter of GmAp1a in EMSA. MBP, maltose-binding protein. At least 3 independent replicates are performed and a representative result is shown. d, e Transient dual luciferase (dual-LUC) assays of Dt2HapI-1 (d) and Dt2HapI-2 (e) binding to the promoter of GmAp1a in tobacco leaves. f Transient dual luciferase (dual-LUC) assay of Dt2 on the promoter of GmAp1a in Arabidopsis protoplast (n = 3 biologically independent replicates). Different letters indicate statistically significant differences at P < 0.05 by one-way ANOVA test. g, h Transient dual-LUC assay in tobacco leaves of Dt2HapI-1 (g) or Dt2HapI-2 (h) and its interacting proteins GmAgl22 and GmSoc1a on the promoter of GmAp1a. i Transient dual-LUC assay of Dt2 and its interacting proteins GmAgl22 and GmSoc1a on the promoter of GmAp1a in Arabidopsis protoplasts (n = 3 biologically independent replicates). Different letters indicate statistically significant differences at P < 0.05 by one-way ANOVA test. j, k Phenotypic comparison between W82 and GmAp1aOE lines. Scale bars, 20 cm. l Branch number statistics of W82 and GmAp1aOE lines (n = 20 biologically independent plants). Statistical significance is determined using a two-sided t-test. m, n Phenotypic comparison between the HX3 and GmAp14m mutation lines in branch number. Scale bars, 20 cm. o Branch number statistics of the HX3 and GmAp14m mutation lines (n = 20 biologically independent plants). Data in (f, i, l and o) are mean ± SEM. Source data are provided as a Source Data file.
A previous study showed that Dt2, a MADS-box protein, could bind to CArG elements20. Putative binding motif prediction using PlantPAN 3.0 (http://plantpan.itps.ncku.edu.tw/) indicated that a sequence located −1274 bp from the GmAp1a translation initiation site (named Probe1) (Fig. 4b) and a sequence located −678 bp from the GmAp1d translation initiation site (named Probe2) (Supplementary Fig. 14a) might be the target sites of Dt2. Subsequently, we performed an electrophoretic mobility shift assay (EMSA) and found that the cold probe concentration could abolish the binding activities (Fig. 4c and Supplementary Fig. 14b), confirming that Dt2 could bind to the promoters of GmAp1a and GmAp1d.
Dt2-GmAgl22-GmSoc1a could activate the transcription of GmAp1a and GmAp1d
To further explore how Dt2 affects the transcription of GmAp1, we performed transient dual luciferase (Dural-LUC) assay in tobacco leaf system and Arabidopsis protoplasts. The results showed that Dt2 functioned as a transcriptional activator to promote the transcription of GmAp1a and GmAp1d (Fig. 4d–f and Supplementary Fig. 14c–e), which was consistent with the changes in the expression of GmAp1 in Dt2OE and Dt2CR lines (Supplementary Fig. 13b). Similarly, we found that Dt2-GmAgl22-GmSoc1a together showed stronger activity than Dt2 alone (Fig. 4g–i and Supplementary Fig. 14f–h).
To check whether GmAp1 indeed affects the branching of soybean, we overexpressed GmAp1a (named GmAp1aOE) and also knocked out the four homologous genes of GmAp1 (named GmAp14m) to eliminate their functional redundancy27. We found that the GmAp1aOE line exhibited significantly decreased branch number (Fig. 4j–l and Supplementary Fig. 13d), and the quadruple mutant GmAp14m exhibited a significantly increased branch number (Fig. 4m–o and Supplementary Fig. 13e), confirming that GmAp1 was indeed involved in branching development in soybean.
Selection of Dt2 natural variations under adaptation
Soybean is a vital crop in China and is planted nationwide from the high-latitude northeast to the low-latitude south. Since branching has a great influence on soybean yield, it has been considered and strongly selected in soybean breeding. Interestingly, we found that the branch number of the soybean accessions from different ecoregions exhibited significant differences: the average branch number of the accessions from higher latitudes was lower than that of accessions from lower latitudes (Fig. 5a).
a Geographic distribution of Dt2 haplotypes in three different ecoregions. The column diagram is the percentage ratio between Dt2HapI and Dt2HapII. The bottom left box plot represents the branch number comparison of accessions in I (n = 695 accessions), II (n = 821 accessions) and III (n = 222 accessions) ecoregions. Data are the mean ± SEM. In box plot (drawn by R 4.1.1 software), the center line indicates the median, the edges of the box represent the first and third quartiles, and the whiskers extend to span a 1.5 interquartile range from the edges. Different letters indicate statistically significant differences at P < 0.05 by one-way ANOVA test. b Proposed working model of the role of Dt2 in controlling branch number in soybean. IM inflorescence meristem. AM axillary meristem. SAM shoot apical meristem. The purple triangles represent the transitions from AM to IM.
An investigation of the haplotypes of Dt2 using our 2898 previously re-sequenced accessions15 revealed that Dt2HapII did not exist in wild soybean and exhibited an increased ratio from landraces to cultivars (Supplementary Fig. 15a). As Dt2 is a dominant locus controlling branching in soybean natural population (Fig. 1e, f), we speculated that the natural variation in Dt2 may be related to the branch number variation in different planting ecoregions. We then investigated the haplotypes of Dt2 in the cultivated accessions (including landraces and cultivars) and found that the two Dt2 haplotypes exhibited different geographical distributions: an increased ratio of Dt2HapI/Dt2HapII from higher latitudes to the lower latitudes, which was consistent with the branch number change pattern (Fig. 5a, Supplementary Table 2 and Supplementary Data 1). It has been suggested that the domestication of soybean may have originated in China in the Huanghuai region (ecoregion II in Fig. 5a) and then radiated to the northern and southern regions28. FST analysis showed that the Dt2 locus exhibited a genetic differentiation tendency between ecoregions II/I, but not between ecoregions II/III (Supplementary Fig. 15b), indicating that the distinct geographic distribution of Dt2 haplotypes may be related to soybean adaptation to different latitudes.
When a soybean accession from higher latitudes is planted at lower latitudes, it usually exhibits a significant yield decrease due to the early flowering and maturity29. The geographic and genetic differentiations of Dt2 inspired us that modification of Dt2 may improve the adaptation of soybean. DN50, the parent used for genetic modification of Dt2 in this study, is an accession from Heilongjiang Province (northeastern of China; 45°77′ N and 126°68′ E), a region located at high latitudes. When DN50 was planted in Beijing (in the middle of China; 40°22′ N and 116°23′ E) and Hainan (in the southern China, close to the equator; 18°09′ N and 108°48′E), the yields were significantly decreased (Supplementary Table 3). However, the Dt2CR lines showed significantly higher yields than DN50, either at a lower planting density or a higher planting density, which was also associated with its effect on maturity (Supplementary Fig. 16).
Discussion
Shoot branching is both an agronomically important and a complex developmental trait that can be affected by many factors, of which the transition from the vegetative to reproductive stage is particularly important30,31. Functional variation in the genes related to vegetative-reproductive transition tends to cause a coupled phenotypic change in shoot branching morphology and flowering time, as observed, for the Hd3a gene in rice32 and the VEG1 gene in pea33. In this study, we revealed that the dominant gene controlling branch number in soybean natural population, Dt2, is a gene related to vegetative-reproductive transition. In addition, the interaction genes and downstream targets of Dt2 are also involved in flowering regulation21,22,23,27,34. Therefore, taking into account the abovementioned factors, we proposed a possible working model for the role of Dt2 in modulating soybean branching (Fig. 5b): GmAp1 functions as a positive regulator in the transition from vegetative growth into reproductive growth by promoting the transition of axillary meristems (AM) into the inflorescence meristem (IM); Dt2 interacts with GmAgl22 and GmSoc1 to activate the transcription of GmAp1; the Dt2HapII promoter has higher transcriptional activity than the Dt2HapI promoter, which in turn enhances the expression of GmAp1 and promotes earlier transition from vegetative growth to reproductive growth, resulting in earlier flowering and reduced branching. Nevertheless, of the multiple polymorphisms between Dt2HapI and Dt2HapII, we have not disclosed the causal genetic variations responsible for the functional divergence of the two haplotypes. A further investigation of upstream regulatory genes may help us to determine which polymorphisms are essential for the transcription of Dt2, which will make the regulatory network more complete.
MADS-box transcription factor genes have been extensively studied in plants and were reported to play important roles in plant development, especially in vegetative-reproductive transition and plant architecture. For instance, SUPPRESSOR OF OVEREXPRESSION OF CO1 and FRUITFULL were found to not only promote flowering, but also affect the determinacy of meristems in Arabidopsis35. Previous studies in soybean have illuminated that the members involved in the working model of Dt2 from this study, including Dt2, GmAgl22, GmSoc1 and GmAp1, all regulate flowering time by being involved in inflorescence transition18,19,20,24,27. Similarly, MtFUL1-c and VEG1/PsFUL1-c, orthologs of Dt2 from Medicago and pea respectively, were also found to play important roles in the reproductive phase transition33,36. Here, we found that these genes not only affect flowering time, but also control branch number in soybean. Consistently, the Arabidopsis homologs of Dt2-GmSoc1a, FUL-SOC1, were also found to regulate branch number35,37,38. These results suggested that Dt2 and other MADS box genes are highly pleiotropic in regulating vegetative-reproductive transition and plant architecture. In addition, these results also indicated that the network of MADS box genes in regulating plant architecture may be functionally conserved among plant species. Taken together previous studies18,19,20,24,27, our results revealed that Dt2, GmAgl22 and GmSoc1 function as a complex to regulate GmAp1. Interestingly, the genes involved in the working model of Dt2 all belong to the MADS-box gene family, indicating a complicated regulatory network among different MADS-box transcription factor genes although they belong to the same transcription factor family. Further investigation of more MADS-box members is needed to reveal more detailed regulatory networks of this important transcription factor gene family.
Soybean was originally domesticated in China and was then introduced into different countries39. Because soybean is highly sensitive to photoperiod, selection of the beneficial alleles of the genes that confer adaptation is critical during the spreading process13,40. For instance, selection of natural variation in the J gene improved the soybean adaptation to the tropical regions and significantly enhanced yield29, making Brazil one of the largest soybean producing countries today. Here, we determined that the natural variation in Dt2 is also related to the adaptation of soybean (Fig. 5a, Supplementary Fig. 15), which provides a genetic candidate for adaptation breeding in soybean. In the future, a trial involving a combinational selection/modification of J and Dt2 may enhance soybean adaptation ability.
In addition to the vegetative-reproductive transition, other factors/pathways also determine shoot branching development1. Here, we found that the dominant gene associated with branch number in soybean is related to vegetative-reproductive transition. Next, fixing the effect of vegetative-reproductive transition, particularly the effect of the dominant gene Dt2 may help to identify additional genes functioning in sobyean shoot branching, and will provide more insight for soybean improvement.
Methods
Plant materials and growth conditions
The 2409 soybean accessions used for the GWAS were planted at the experimental station of the Tianjin Academy of Agricultural Science, Wuqing (39° 38′ N and 117° 04′ E) during the summer season in 2017 and the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing (40° 22′ N and 116° 23′ E) in 2018. At the full-pod stage (R4), 5 representative plants were selected from each accession and the primary effective branch number was identified. The Dt2HapI-1 material was PI 548533 and Dt2HapII was PI 547501.
For the BiFC and dual Luciferase reporter assays, Arabidopsis ecotype Col-0 was planted in the greenhouse at 22 °C under an 8 h light and 16 h dark photoperiod. Tobacco (Nicotiana benthamiana) was planted in the greenhouse at 22 °C under a 16 h light and 8 h dark photoperiod.
Vector construction and transformation
To construct the Dt2 overexpression plasmid, the CDS of Dt2 was amplified from DN50 and ligated into pTF101, a binary vector containing the CaMV 35 S promoter and a terminator, with the restriction sites XbaI and SacI. To construct the GmAgl22 overexpression plasmid, the CDS of GmAgl22 was ligated into pTF101 with the restriction sites XbaI and SacI. These constructs were introduced into Agrobacterium tumefaciens strain EHA101 and then transformed into DN50.
For the CRISPR/Cas9 system experiments, two sgRNAs were designed using Primer Design software. Two U6 promoters were used for the guide RNA oligonucleotide pair. The U6 promoter driving a single guide RNA cassette was cloned into the PMDC123 vector41. These constructs were introduced into Agrobacterium tumefaciens strain EHA105 and then transformed into DN50. The relevant primers used are listed in Supplementary Data 2.
RNA extraction and expression analysis
Total RNA was extracted using an RNA isolation kit (Tiangen, DP432) according to the manufacturer’s protocol and three biological replicates were performed in each experiment. Reverse transcription was performed using a cDNA synthesis kit (Transgen, AE311). Then the cDNA sequence was used as the template for the quantitative real-time PCR. qPCR was performed using LightCycler 480 SYBR Green I Master (Transgen, AQ101-01) on a LightCycler 480 instrument (Roche). Gene expression was normalized to the expression of the soybean gene ACTIN11. Fold changes were calculated from the 2−ΔΔCt values. The relevant primers used are listed in Supplementary Data 2.
GWAS analysis
For GWAS, we used the previously reported SNP dataset15. A total of 4,072,231 SNPs were used for association analysis with a minor allele frequency (MAF) of >5% and a missing rate of <10%. GWAS was performed based on a mixed model using the EMMAX software package42. EIGENSOFT software43 was used to perform principal-component analysis of the population, and the first five principal components were included as fixed effects. The matrix of pairwise genetic distances derived from the simple matching coefficients was used as the variance-covariance matrix of the random effects. For the threshold, we defined the whole-genome significance cutoff as the Bonferroni test threshold44,45, the threshold was set as −log(0.05/total SNPs), and the genome-wide significance level for branch number was determined as 1 × 10−7.9.
Haplotype analysis of Dt2 in the soybean population
The SNPs in the 3.5 kb promoter region and full-length genomic region of Dt2 of 2409 varieties were obtained from the previously reported SNP and INDEL dataset15. Then the SNPs were filtered by applying a MAF > 5% cutoff, missing rate <10%, nonfunctional SNP mutation and low association signals SNPs, retaining 35 high-quality SNPs. The association polymorphisms classified the accessions into three major haplotypes.
Phylogenetic tree analysis
The homology of Dt2 proteins was searched in Phytozome 13 (https://phytozome-next.jgi.doe.gov/blast-search), focusing on genes with homology >50% in soybean, Arabidopsis and rice. MEGA 6.0 was used for sequence comparison and phylogenetic tree construction analysis and the bootstrap repetition value was 1000. The phylogenetic tree was further modified by the online tool evolview (https://www.evolgenius.info//evolview/)46.
Selection analysis
The genetic differentiation fixation index (FST) was calculated by using VCFtools (0.1.13) with a 20 kb slide window and 2 kb slide step47. The first 5% value was used as the threshold in the whole genome.
In situ hybridization
In situ hybridization treatments were performed as previously described48,49. Briefly, the soybean shoot apexes of 10, 16 and 22 day-old seedlings were fixed in 50% formol-acetic-alcohol. Subsequently, the 8 μm-thick SAM samples slices were observed in a conventional light microscope after sample fixation, embedding, sectioning and hybridization. The size of the Dt2 probe was 228 bp, that of the GmAgl22 probe was 137 bp, and that of the GmSoc1a probe was 140 bp. The primers are listed in Supplementary Data 2.
RNA-seq sample preparation and sequencing
Lateral buds from the same node of WT and transgenic soybean plants were collected for RNA-seq analysis. Three biological replicates were performed for each sample. The Illumina HiSeq 2000 platform was used to generate 150 bp paired-end reads. And the detailed bioinformatic analyses were performed as previously described50. Briefly, the high-quality sequencing reads were mapped to the reference genome with Hisat (v. 2.2.1). And the gene expression counts were calculated using StringTie (v.1.3.4d). The different expression genes analysis were analyzed by the R-edgeR library (https://bioconductor.org/packages/release/bioc/html/edgeR.html).
Yeast two-hybrid assays
Yeast two-hybrid assays were performed as described in the Yeast Protocols Handbook (Clontech). The coding region sequence of Dt2HapI-2 was introduced into the prey vector (pGBKT7). To construct the prey vectors, we ligated the full-length CDSs of GmAgl22 and GmSoc1a into the pGADT7 vector. Then pGBKT7-Dt2HapI-2 was transformed into the Y2HGold strain with pGADT7-GmAgl22 or pGADT7-GmSoc1a and selected on DDO (Synthetic Dropout Medium/-Tryptophan-Leucine) and QDO (Synthetic Dropout Medium/-Tryptophan-Histone-Leucine-Adenine) media (Clontech). The empty AD or empty BD served as a negative control. The primers are listed in Supplementary Data 2.
BiFC
For the construction of BiFC vectors, the Gateway-compatible vectors pUGW2-nYFP and pUGW2-cYFP were used to generate vectors in BIFC assays by using Gateway cloning technology. pUGW2-nYFP is the vector for N-terminal fusion to yellow fluorescent protein (nYFP), and pUGW2-cYFP is the vector for C-terminal fusion to YFP (cYFP). The full-length CDS of Dt2HapI-2 was cloned into pUGW2-nYFP. The full-length CDSs of GmAgl22 and GmSoc1a were cloned into pUGW2-cYFP. Arabidopsis protoplasts were prepared for the expression assays. Vectors were co-transformed into Arabidopsis protoplasts and incubated at 22 °C in the dark for 12–16 h. YFP fluorescence was visualized using confocal laser scanning microscope (Zeiss LSM 985 NLO).
Co-IP analysis
Co-IP analysis was performed using Arabidopsis protoplasts47. To construct the vector, full-length CDSs of Dt2HapI-1 and Dt2HapI-2 were cloned into pUC19-35S-HA vectors, and the CDSs of GmAgl22 and GmSoc1a were introduced into the pUC19-35S-Flag vector. A. thaliana protoplasts were transfected with 50 μg of plasmid and incubated overnight under low-light-intensity environment. Total protein was extracted from protoplasts after incubation for 12–16 h using extraction buffer (50 mM Tris-HCl (pH 7.5), 0.5 mM EDTA, 150 mM NaCl, 0.5% np-40, 1 mM PMSF, and 1× complete protease inhibitor cocktail (Roche, 04693132001). The protein lysis product were incubated with Flag magnetic beads (MBL) for 30 min to 1 h and wash the beads four times with a wash buffer that consisted of 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 20% glycerol, 0.1% Triton X-100, 1 mM EDTA (pH 8.0) and 1× complete protease inhibitor cocktail. The immunoprecipitates were separated using SDS-PAGE and transferred to a nitrocellulose membrane (GE Healthcare). Proteins were detected by treating the membranes with anti-HA (1:5,000, MBL, M180-7) or anti-DDDDK-tag mAb-HRP-DirectT antibodies (1:10,000, MBL, M185-7).
Transient dual luciferase (dual-LUC) assay
To generate the pGmAp1a:LUC and pGmAp1d:LUC constructs, we amplified 3-Kb promoter fragments upstream of each gene from Williams 82 and ligated them with the pGreen0800-LUC as the reporter vector. The p35S-Dt2HapI-1-Flag, p35S-Dt2HapI-2-Flag, p35S-GmAgl22-Flag and p35S-GmSoc1a-Flag constructs were used as effectors. Transient transactivation assays were performed using Arabidopsis protoplasts51. In the tobacco leaf system, the promoters of the GmAp1a and GmAp1d (3000 bp) were cloned into the transient expression vector CP461, which was constructed as the reporter vector. In addition, the p35S-Dt2HapI-1-Flag, p35S-Dt2HapI-2-Flag, p35S-GmAgl22-Flag and p35S-GmSoc1a-Flag constructs were used as effectors and these plasmids were transformed into A. tumefaciens strain GV3101. Then these strains were injected into tobacco leaves in different combinations with p19, which was used to suppress RNA silencing. Dual luciferase assay reagents (Promega, VPE1910) with the Renilla luciferase gene as an internal control were used for luciferase imaging. The relevant primers are listed in Supplementary Data 2.
Split luciferase (split-LUC) complementation assay
Luciferase complementation imaging assays were performed as described previously52. Briefly, to generate a luciferase complementation vector, pCAMBIA1300-35S-NLuc was fused with the C- termini of Dt2HapI-1, Dt2HapI-2 and GmAgl22, and pCAMBIA1300-35S-CLuc was fused with the C-termini of GmAgl22 and GmSoc1a. Transient expression in tobacco (Nicotiana benthamiana) leaves was conducted by GV3101 Agrobacterium infiltration. Plants were then incubated at 22 °C for 2 days before the LUC activity was measured. Images were captured using the low-light cooled charge-coupled device imaging apparatus NightOWL IILB 983.
Electrophoretic mobility shift assay
The full length of Dt2HapI-1 and Dt2HapI-2 CDSs were amplified and cloned into the Pmal-C5x vector. MBP-Dt2HapI-1 and MBP-Dt2HapI-2 recombinant proteins with MBP tags were introduced into Escherichia coli BL21 (DE3). The recombinant proteins were purified by using maltose resin (NEB, E8021S). DNA probes (Probe1 in GmAp1a and Probe2 in GmAp1d) were artificially synthesized and labeled with biotin at the 5′ end (Thermo Fisher Scientific). DNA gel shift assays were performed as the protocol described of the LightShift Chemiluminescent EMSA kit (Thermo Fisher Scientific, 20148). The probe sequences are listed in Supplementary Data 2.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The SNP data of 2409 natural population accessions were reported previously and have been deposited in the Genome Sequence Archive (GSA) database in the BIG Data Center under accession number PRJNA257011, PRJNA394629 and CRA002269 [https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA002030]. The RNA-seq data generated in this study have been deposited into the Genome Sequence Archive (GSA) database in the National Genomics Data Center under accession number SAMC797049-SAMC797057 of PRJCA009434. Requests for materials should be addressed to Z.T. Source data are provided with this paper.
References
Wang, B., Steven, M. S. & Li, J. Genetic regulation of shoot architecture. Annu. Rev. Plant Biol. 69, 437–468 (2018).
Doebley, J., Stec, A. & Gustus, C. teosinte branched1 and the origin of maize: evidence for epistasis and the evolution of dominance. Genetics 141, 333–346 (1995).
Seale, M., Bennett, T. & Leyser, O. BRC1 expression regulates bud activation potential but is not necessary or sufficient for bud growth inhibition in Arabidopsis. Development 144, 1661–1673 (2017).
González-Grandío, E., Poza-Carrión, C., Sorzano, C. & Cubas, P. BRANCHED1 promotes axillary bud dormancy in response to shade in Arabidopsis. Plant Cell 25, 834–850 (2013).
Martín-Trillo, M. et al. Role of tomato BRANCHED1-like genes in the control of shoot branching. Plant J. 67, 701–714 (2011).
Studer, A., Zhao, Q., Ross-Ibarra, J. & Doebley, J. Identification of a functional transposon insertion in the maize domestication gene tb1. Nat. Genet. 43, 1160–1163 (2011).
Zhang, L. et al. A natural tandem array alleviates epigenetic repression of IPA1 and leads to superior yielding rice. Nat. Commun. 8, 14789 (2017).
Miura, K. et al. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 42, 545–549 (2010).
Jiao, Y. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544 (2010).
Domagalska, M. A. & Leyser, O. Signal integration in the control of shoot branching. Nat. Rev. Mol. Cell Biol. 12, 211–221 (2011).
Graham, P. H. & Vance, C. P. Legumes: importance and constraints to greater use. Plant Physiol. 131, 872–877 (2003).
Ray, D. K., Mueller, N. D., West, P. C. & Foley, J. A. Yield trends are insufficient to double global crop production by 2050. PLoS ONE 8, e66428 (2013).
Liu, S., Zhang, M., Feng, F. & Tian, Z. Toward a “Green Revolution” for soybean. Mol. Plant 13, 688–697 (2020).
Zhang, M. et al. Progress in soybean functional genomics over the past decade. Plant Biotechnol. J. 20, 256–282 (2021).
Liu, Y. et al. Pan-genome of wild and cultivated soybeans. Cell 182, 162–176 (2020).
Shen, Y. et al. De novo assembly of a Chinese soybean genome. Sci. China Life Sci. 61, 871–884 (2018).
Shen, Y. et al. Update soybean Zhonghuang 13 genome to a golden reference. Sci. China Life Sci. 62, 1257–1260 (2019).
Liu, Y. et al. Innovation of a regulatory mechanism modulating semi-determinate stem growth through artificial selection in soybean. PLoS Genet. 12, e1005818 (2016).
Ping, J. et al. Dt2 is a gain-of-function MADS-domain factor gene that specifies semideterminacy in soybean. Plant Cell 26, 2831–2842 (2014).
Zhang, D. et al. A post-domestication mutation, Dt2, triggers systemic modification of divergent and convergent pathways modulating multiple agronomic traits in soybean. Mol. Plant 12, 1366–1382 (2019).
Richter, R. et al. Floral regulators FLC and SOC1 directly regulate expression of the B3-type transcription factor TARGET OF FLC AND SVP 1 at the Arabidopsis shoot apex via antagonistic chromatin modifications. PLoS Genet. 15, e1008065 (2019).
Méndez-Vigo, B., Martínez-Zapater, J. M. & Alonso-Blanco, C. The flowering repressor SVP underlies a novel Arabidopsis thaliana QTL interacting with the genetic background. PLoS Genet. 9, e1003289 (2013).
Tao, Z. et al. Genome-wide identification of SOC1 and SVP targets during the floral transition in Arabidopsis. Plant J. 70, 549–561 (2012).
Kou, K. et al. A functionally divergent SOC1 homolog improves soybean yield and latitudinal adaptation. Curr. Biol. https://doi.org/10.1016/j.cub.2022.02.046 (2022).
Rameau, C. et al. Multiple pathways regulate shoot branching. Front. Plant Sci. 5, https://doi.org/10.3389/fpls.2014.00741 (2015).
Luo, Z., Janssen, B. J. & Snowden, K. C. The molecular and genetic regulation of shoot branching. Plant Physiol. 187, 1033–1044 (2021).
Yue, L., Li, X., Fang, C. & Kong, F. FT5a interferes with the Dt1-AP1 feedback loop to control flowering time and shoot determinacy in soybean. J. Integr. Plant Biol. 63, 1004–1020 (2021).
Li, Y. et al. Genetic structure and diversity of cultivated soybean (Glycine max (L.) Merr.) landraces in China. Theor. Appl. Genet. 117, 857–871 (2008).
Lu, S. et al. Natural variation at the soybean J locus improves adaptation to the tropics and enhances yield. Nat. Genet. 49, 773–779 (2017).
Huang, X., Ding, J., Effgen, S., Turck, F. & Koornneef, M. Multiple loci and genetic interactions involving flowering time genes regulate stem branching among natural variants of Arabidopsis. N. Phytol. 199, 843–857 (2013).
Andrés, F. & Coupland, G. The genetic basis of flowering responses to seasonal cues. Nat. Rev. Genet. 13, 627–639 (2012).
Tsuji, H. et al. Hd3a promotes lateral branching in rice. Plant J. 82, 256–266 (2015).
Benlloch, R. et al. Genetic control of inflorescence architecture in legumes. Front. Plant Sci. 6, https://doi.org/10.3389/fpls.2015.00543 (2015).
Chen, L. et al. Soybean AP1 homologs control flowering time and plant height. J. Integr. Plant Biol. 62, 1868–1879 (2020).
Melzer, S. et al. Flowering-time genes modulate meristem determinacy and growth form in Arabidopsis thaliana. Nat. Genet. 40, 1489–1492 (2008).
Cheng, X., Li, G., Tang, Y. & Wen, J. Dissection of genetic regulation of compound inflorescence development in Medicago truncatula. Development 145, dev158766 (2018).
Karami, O. et al. A suppressor of axillary meristem maturation promotes longevity in flowering plants. Nat. Plants 6, 368–376 (2020).
Bemer, M. et al. FRUITFULL controls SAUR10 expression and regulates Arabidopsis growth and architecture. J. Exp. Bot. 68, 3391–3403 (2017).
Dashiell, K. Soybeans: improvement, production, and uses. Agr. Syst. 83, 110–111 (2005).
Lin, X., Liu, B., Weller, J. L., Abe, J. & Kong, F. Molecular mechanisms for the photoperiodic regulation of flowering in soybean. J. Integr. Plant Biol. 63, 981–994 (2021).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Churchill, G. A. & Doerge, R. W. Empirical threshold values for quantitative trait mapping. Genetics 138, 963–71 (1994).
Wei, X. et al. Genetic discovery for oil production and quality in sesame. Nat. Commun. 6, 8609 (2015).
Zhang, H., Gao, S., Lercher, M. J., Hu, S. & Chen, W. H. EvolView, an online tool for visualizing, annotating and managing phylogenetic trees. Nucleic Acids Res. 40, 569–572 (2012).
Wang, M. et al. Parallel selection on a dormancy gene during domestication of crops from multiple families. Nat. Genet. 50, 1435–1441 (2018).
Javelle, M., Marco, C. F. & Timmermans M. In situ hybridization for the precise localization of transcripts in plants. J. Vis. Exp. 57, e3328 (2011).
Traas, J. Whole-mount in situ hybridization of RNA probes to plant tissues. CSH Protoc. 1, https://doi.org/10.1101/pdb.prot4944 (2008).
Shen, Y. et al. Global dissection of alternative splicing in paleopolyploid soybean. Plant Cell 26, 996–1008 (2014).
Wang, J. et al. Arabidopsis CSN5B interacts with VTC1 and modulates ascorbic acid synthesis. Plant Cell 25, 625–636 (2013).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol. 146, 368–376 (2008).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (grant no. 2021YFF1001201), National Natural Science Foundation of China (grant no. 31788103, 32090064), Hainan Yazhou Bay Seed Laboratory Project (grant no. B21HJ0002), and “Strategic Priority Research Program” of the Chinese Academy of Sciences (grant no. XDA24030501).
Author information
Authors and Affiliations
Contributions
Z.T. designed and supervised the project; Q.L., L.C., X.Y., H.Y., K.K., L.F., Z.D., Y.Y., S. Liang, X.L., G.Z., and M.Z. performed the experiments; Q.L., S. Liu, Z.Z., Y.L., F.K., and Z.T. analyzed the data; Q.L. and Z.T. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Carlos Alonso-Blanco, Thorsten Schnurbusch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, Q., Chen, L., Yang, X. et al. Natural variation of Dt2 determines branching in soybean. Nat Commun 13, 6429 (2022). https://doi.org/10.1038/s41467-022-34153-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-34153-4
This article is cited by
-
The type III effector NopL interacts with GmREM1a and GmNFR5 to promote symbiosis in soybean
Nature Communications (2024)
-
Natural variation in GmSW17 controls seed size in soybean
Nature Communications (2024)
-
Innovations in functional genomics and molecular breeding of pea: exploring advances and opportunities
aBIOTECH (2024)
-
Toward cis-regulation in soybean: a 3D genome scope
Molecular Breeding (2023)
-
Promoter variations in a homeobox gene, BrLMI1, contribute to leaf lobe formation in Brassica rapa ssp. chinensis Makino
Theoretical and Applied Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.