Abstract
The Tibetan Plateau’s Kobresia pastures store 2.5% of the world’s soil organic carbon (SOC). Climate change and overgrazing render their topsoils vulnerable to degradation, with SOC stocks declining by 42% and nitrogen (N) by 33% at severely degraded sites. We resolved these losses into erosion accounting for two-thirds, and decreased carbon (C) input and increased SOC mineralization accounting for the other third, and confirmed these results by comparison with a meta-analysis of 594 observations. The microbial community responded to the degradation through altered taxonomic composition and enzymatic activities. Hydrolytic enzyme activities were reduced, while degradation of the remaining recalcitrant soil organic matter by oxidative enzymes was accelerated, demonstrating a severe shift in microbial functioning. This may irreversibly alter the world´s largest alpine pastoral ecosystem by diminishing its C sink function and nutrient cycling dynamics, negatively impacting local food security, regional water quality and climate.
Similar content being viewed by others
Introduction
The Tibetan Plateau (TP) hosts the world’s largest high-altitude grasslands, contributing 2.5% to global SOC stocks1 but covering only 0.3% of the Earth’s total terrestrial area. It influences the Asian monsoon climate2, is the water source for one-fifth of the global population3 and provides grazing grounds for >8 million sheep, yaks, and goats4. Approximately one-fifth of the TP is covered by Kobresia grasslands2. Kobresia pygmaea forms a 2–4 cm high grazing lawn with low shoot biomass but very compact root mats (root-to-shoot ratio >20)5, induced by a long history of low-to-moderate grazing intensity, which increases belowground carbon allocation and root biomass6,7. Kobresia pygmaea’s dense root network protects it from trampling-induced soil erosion and enables fast regrowth after defoliation8.
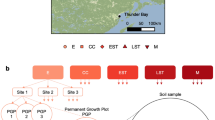
Pasture degradation has increased dramatically in recent decades2,3, and about 30% of the Tibetan grasslands are considered to be degraded9. This has indisputably severe consequences for ecosystem functions, most importantly the major decline in SOC and N storage. Three mechanisms contribute to this—erosion, decreased C and N input, and increased soil organic matter (SOM) mineralization—but their relative importance remains unclear10. We conducted a meta-analysis including 594 single observations from 49 literature studies published between 2002 and 2020 (Supplementary Table 1) to quantify the SOC and N losses for Kobresia pygmaea’s core area. In a detailed field study, we determined the relative contributions of erosion and net mineralization to SOC losses and identified the underlying changes in microbial community structure and functioning. For this, we categorized six successive stages of degradation, from intact Kobresia root mats (S0) to stages with increasing extents of surface cracks (S1–S4) to bare soil patches without root mats (S5) (Supplementary Fig. 1).
We hypothesize that substantial parts of the Kobresia pastures are close to a critical point of microbial functioning changes with substantial consequences for SOC and N storage. We further hypothesize that these abrupt shifts are characterized by changes in SOM quality and quantity that are coupled by feedback loops to microbial community structure and functions, which in turn control C and N mineralization.
Results
Root-mat cracking and soil erosion
The formation of polygonal surface cracks is widespread between the Qilian Mountains and the Himalayas (Supplementary Fig. 1). These surface cracks represent the early stages of a degradation process that, in its final stage (S5), is associated with a loss of 42% of SOC compared to non-degraded pastures (Fig. 1A, B; Supplementary Fig. 2). Nitrogen losses were comparably high (33%; Fig. 1C, D; Supplementary Fig. 2). Soil erosion induced preferential loss of the easily erodible fine particles, resulting in a relative accumulation of coarser soil material from S0 to S5 (Fig. 1G, Supplementary Fig. 3), with soil clay content being 60% lower at the most degraded sites compared to intact pastures (Fig. 1G; Supplementary Fig. 3). At the most extreme degradation stage (S5) of our own study site, 81 kg m−2 of the most fertile SOC- and N-rich topsoil had been lost to erosion. This corresponds to 5 kg C m−2 or 45% of the total soil C stock (Fig. 1A), in agreement with the mean soil C loss found in the literature study (42%). Degradation of the Kobresia turf was furthermore associated with a large decrease in penetration resistance (Fig. 2A) and root density (Fig. 2B) from S1 to S5. Intensified degradation decreased vegetation cover, leaving the soil prone to erosion with extended size and depth of cracks (Supplementary Table 2).
A soil organic carbon (SOC) core study site, B SOC literature study, C total nitrogen (N) core study site, D total N literature study, E bulk density (BD) core study site, F BD literature study, G clay content core study site. All parameters are presented for each soil horizon at the midpoint of the depth increment. Error bars display standard error. Exact data are provided in Supplementary Table 2.
A Penetration resistance, B root density, C δ13C of soil and roots, D δ13C of soil organic carbon (SOC), E content of lignin monomers vanillyl, syringyl, and cinnamyl (VSC). All parameters (means ± SE, n = 4) are presented for each soil horizon at depth midpoint, except for penetration resistance (A) and δ13C values of soil and roots (C), which are shown in 1 and 5 cm increments, respectively. Exact data are given in Supplementary Table 2. Progressive changes along the degradation sequence can be explained by the following processes: (a) root-mat cracking by desiccation and frost, (b) root death and decomposition, (c) kinetic 13C fractionation during root decomposition, (d) SOC loss due to reduced root carbon (C) input and greater SOC mineralization, (e) relative lignin accumulation and 13CSOC depletion (S0–S3), (f) relative lignin accumulation during stages S0–S3, (g) lignin degradation and 13CSOC enrichment (S4, S5).
SOC losses through mineralization and decreased root carbon input
The SOC and N contents were closely positively correlated with root density (Supplementary Fig. 4), and showed a consistent relationship in isotopic composition (δ13C) with the associated roots at each depth (Fig. 2C). This demonstrates the importance of roots as the dominant SOC source for intact Kobresia pastures. An association of increasing δ13C values with declining SOC content (Fig. 2D) due to kinetic isotope fractionation during decomposition was observed only at the final degradation stage. With degradation from S1 to S4, the SOC content in the upper 20 cm decreased (Fig. 1A, B), while δ13C values already decreased from S0 to S1 (Fig. 2D). We quantified the sum of lignin monomers vanillyl, syringyl, and cinnamyl as an indicator of complex organic compounds11. Vanillyl, syringyl, and cinnamyl contents increased from S0 to S4 (Fig. 2E), indicating selective lignin preservation during early degradation stages. The close negative correlation between δ13C and vanillyl, syringyl, and cinnamyl contents in the mineral horizon indicates the selective enrichment of the isotopically light lignin (Supplementary Fig. 5A). However, the decrease of vanillyl, syringyl, and cinnamyl contents from S4 to S5 in the mineral soil (Fig. 2E), accompanied by 13C enrichment (Fig. 2D) and a decrease in SOC content (Supplementary Fig. 5B), indicates a pronounced lignin decomposition at the final degradation stage (S5).
Soil microbial community structure and functions
The bacterial (Supplementary Figs. 6A, and 9) and fungal (Supplementary Figs. 6B, and 10) community composition changed significantly (p < 0.05, MANOVA) along the degradation sequence. Bacterial groups (Supplementary Table 3) associated with decomposition of low-molecular-weight organic compounds, e.g., Actinobacteria (Supplementary Figs. 7A, and 9), declined with pasture degradation, whereas lignin-degrading groups such as Rhizobiales increased (Supplementary Fig. 7D). Nitrifying (Nitrospirales and Nitrosomonadaceae; Supplementary Fig. 7B, C) and denitrifying (Pseudomonadales; Supplementary Fig. 7E) bacteria increased from S0 to S3 and then declined towards S5. These shifts indicate severe changes in N-cycling towards the accelerated microbial transformation of organic N to mineral N in the early stages of degradation. Similarly, fungal groups specialized in efficient lignin degradation increased in relative abundance at later degradation stages, as evidenced by Agaricomycetes12,13 (Supplementary Fig. 7F), which include the brown-rot and white-rot fungi. Correspondingly, groups generally incapable of efficient lignin degradation12, such as Ascomycota (Supplementary Fig. 7G), decreased along the degradation sequence (Supplementary Fig. 10a, b). After a peak at degradation stage S2, Kobresia pygmaea successively lost its arbuscular mycorrhizal fungal partners (Glomeromycota, Supplementary Fig. 7H), which were replaced by ectomycorrhizal ones (Thelephoraceae and Inocybaceae, Supplementary Fig. 7I). When Kobresia pygmaea largely disappeared (S5), the abundance of its ectomycorrhizal partners also declined and the first arbuscular mycorrhizal partners of the newly establishing pioneer plants appeared (Supplementary Fig. 7H).
Non-metric multidimensional scaling (NMDS), including C/N, δ13C, total N, δ15N, total phosphorus (P), pH, cation exchange capacity, calcium, potassium, iron and aluminum content, clay content and vanillyl, syringyl, and cinnamyl content as explanatory factors revealed a strong correlation between microbial community structure and SOC quality (C/N, δ13C, N, P and vanillyl, syringyl, and cinnamyl) during early degradation stages (Fig. 3). However, changes in microbial community structure during later degradation stages (from S3 to S5) were mainly driven by abiotic soil properties such as increasing pH and decreasing clay content, which had a stronger effect on fungi than on bacterial communities.
Generalized changes in microbial community composition and enzyme activities (A). Simplified non-metric multidimensional scaling (NMDS) plots derived from terminal restriction fragment length polymorphism (t-RFLP) data for the bacterial (B) and fungal (C) communities. Shaded areas mark each degradation stage, symbols indicate soil depth, and arrows show most important driving factors (canonical correspondence analysis, p < 0.05). Individual variables underlying the processes described by the blue and red arrows are shown in Supplementary Fig. 6.
Maximal activities of enzymes for utilizing easily hydrolyzable C sources (β-glucosidase and xylanase) significantly increased from S0 to S3 and declined when pasture degradation reached stage S4 (Supplementary Figs. 7J, and 12). Likewise, the activity of N- and P-mobilizing enzymes (urease and alkaline phosphatase) declined from S3 to S4 (Supplementary Figs. 7J, and 12). In contrast, the activity of lignin-degrading phenol oxidases increased steadily with the degradation from S2 to S4 along with the relative increase in vanillyl, syringyl, and cinnamyl monomers (Supplementary Figs. 7J, and 12).
Disentangling processes contributing to SOC losses
In the study area, SOC stocks at 0–30 cm depth declined strongly by 7.5 kg C m−2 from the stage of intact Kobresia root mats (S0) to that of bare soil patches (S5). This corresponds to a 45% reduction (Fig. 4) and matches well with the mean SOC loss observed for degraded areas across the whole TP (42%). SOC losses were attributed to topsoil erosion, which accounted for two-thirds of the decline (for S5: 5.0 kg C m−2, Fig. 4B), with the combined effects of decreasing root carbon input and accelerated mineralization of root litter and SOC accounting for the remaining one-third of the decline (2.5 kg C m−2; Fig. 4B).
Total SOC stocks along the degradation sequence (S0–S5) down to 30 cm for each horizon (A). SOC losses by erosion and by reduced C input and increased SOC mineralization (B) during degradation (S1–S5). Lowercase letters indicate significant differences in SOC stocks between degradation stages. The absence of significant differences (n.s.) between quantified total SOC losses (red outline) and the calculated sum of erosion- and mineralization-induced SOC losses confirms the validity of the estimations. Error bars display standard error.
Discussion
Above- and belowground linkages of organic matter and energy flux play a central role in the responses of the TP Kobresia ecosystem to anthropogenic forcing and climate warming, as reported for mountain ecosystems worldwide14. Severe soil degradation of the Kobresia ecosystem results from complex reinforcing interactions of biotic and abiotic processes, as well as anthropogenic pressure (Fig. 5). Frost events at the beginning of the cold season, when the soil is moist, cause the formation of polygonal surface cracks, a process severely enhanced by permafrost thaw15. Overgrazing-induced trampling, followed by root death and decomposition, expand these surface cracks (S1, S2). Exacerbated polygonal cracking then promotes soil erosion with preferential loss of fine-textured material, containing SOC and nutrients (S3–S4). This implies, among others, a strong reduction in the water holding capacity with as yet unforeseeable consequences for the water supply and quality of one-fifth of the world’s population. Moreover, erosion of soil material and the associated loss of SOC carry severe consequences for ecosystem energy, water, and C fluxes, e.g., regionally earlier initiation of convection and cloud generation2, affecting the strength and variability of the TP summer monsoon2.
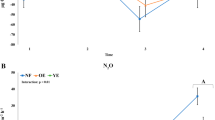
δ13C patterns in roots and SOC in undegraded pastures demonstrate that SOC accumulation depends heavily on root biomass input and rhizodeposition. A close correlation between SOC stocks and root density across all degradation stages demonstrates that decreasing C input with degradation is a major driver of decreasing SOC content. Decreasing SOC content along the degradation gradient indicates that root necromass was not stabilized in degraded soils, and consequently labile constituents of root necromass were rapidly mineralized. The large root litter input from dying Kobresia stimulated microorganisms to decompose the old SOC pool (root litter priming). This was confirmed by the increased activity of hydrolyzing enzymes during the early degradation stages. Such priming effects were triggered by the three to four times higher C/N ratio in roots compared to soil, driving microbes in this N-limited ecosystem5 to decompose SOM to obtain N (N mining)16. Similar effects of N limitation were observed along a thawing permafrost chronosequence in grassland soils of the TP, where N limitation in the late stages of permafrost collapse induced SOC decomposition through priming effects17. In the case of permafrost thawing, a decrease in metabolic efficiency was the major mechanism underlying priming, without a response in hydrolytic enzyme activity. The root litter decomposition observed along our degradation sequence was, however, clearly related to an increase in hydrolytic enzyme activity from S0 to S3. Thus, the successive degradation and subsequent mineralization of the organic N stocks increased the abundance of nitrifying and denitrifying bacteria during the early degradation stages (S0–S3). Using N-cycling gene abundances (amoA), Che et al.18 identified an up to five-fold increase in nitrifiers associated with a major shift in the N cycle during the degradation of Tibetan pasture meadows. Nitrification produces easily leachable nitrate, a limiting substrate for denitrification in alpine meadows19, which consequently increases in periods of high soil moisture or in moist soil microhabitats. We observed an increase in N-hydrolyzing enzymes, an increase in plant litter decomposers, and an increasing abundance of nitrifier- and denitrifier-hosting bacterial orders and families (Nitrospirales and Nitrosomonadaceae (Supplementary Fig. 7B, C) and Pseudomonadales (Supplementary Fig. 7E), respectively) from degradation state S0–S3. This suggests that all three processes - N mineralization, nitrification, and denitrification—increase along the degradation gradient, which leads to severe nitrogen losses from this ecosystem via leaching, emissions of nitrous oxide and nitrogen gas, and NH3 volatilization20. Quantification of the net and gross rates of these processes21 as well as corresponding gene abundances22 confirmed that these three key steps of the N cycle accelerate with increased grazing intensity in Tibetan meadows. Concomitant increases in mineral N availability21 induced a change in the mycorrhizal partners of Kobresia pygmaea. Arbuscular mycorrhizal fungi, capable of capturing nutrients released by other microorganisms’ hydrolytic enzymes, increased along the sequence of increasing hydrolytic organic matter decomposition (S0–S2) and optimized the access to otherwise rapidly leached or consumed ammonia and nitrate. However, toward stage S4, there was a shift in favor of ectomycorrhizal partners of Kobresia pygmaea, being specialized for the active mobilization of organic matter-bound nutrients using their own oxidative enzymes. The concurrent decrease in urease activity from S3 to S4, reflecting successive depletion of the easily accessible N in the soil, is evidence of a shift in ecosystem nutrition strategy: mineral nutrient release through AMF-mediated and prokaryote-driven hydrolysis was replaced by fungi-based nutrient scavenging for nutritional elements “locked up” in complex structural root-mat tissues using oxidative enzymes. Such fundamental shifts in nutritional strategies and biogeochemical cycles suggest that degradation will have large-scale feedback on multiple environmental processes.
As a result of organic matter decomposition, we expected kinetic isotope fractionation to induce a continuous δ13C increase along the degradation gradient23. However, the opposite was found for early degradation stages. δ13C values decreased from S0 to S4 within the upper 20 cm, while δ13C values increased from S4 to S5. We attribute the δ13C decrease (S0–S4) to a relative accumulation of more complex, 13C-depleted organic compounds such as lignin24. Assuming a 4.3‰ depletion in lignin δ13C compared to bulk plant material24 and a recovery of 38% of the lignin C by the vanillyl, syringyl, cinnamyl method25, the relative increase of lignin from 0.9 to 4.7% of SOC accounts for nearly 23% of the observed δ13C shift across the degradation sequence. However, lignin is only one compound class among others (cutins, suberins, etc.) with complex structures and depleted δ13C values. Lipids have even more depleted δ13C values than lignin24. The negative δ13C shifts observed with soil degradation from S0 to S4 likely reflect the relative accumulation of these rather persistent compounds in addition to lignin. The intact Kobresia root mats at the research site consist of about 6 kg root dry matter m−25, including many woody roots that contain large amounts of lignin and suberin. Root dieback during soil degradation, therefore, provides a substantial source of complex, 13C-depleted compounds. Relative lignin enrichment of the SOC pool was evident not only in the δ13C decrease but also in the increased abundance of fungal and bacterial specialists decomposing complex SOM (e.g., Rhizobiales and Agaricomycetes). The elevated abundance of these groups was accompanied by increased phenoloxidase activity, which catalyzes the nonspecific oxidation of complex organic substances, especially lignin. Consequently, after easily available C pools are depleted (S3), complex organic compounds are successively degraded as microbial communities adapt to these distinct carbon sources. While microbial communities changed gradually from S0 to S3, a profound shift in community structure and functions occurred from S3 to S4 (evidenced by a vertical shift in direction in the NMDS plots; Supplementary Figs. 6 and 8). This transition coincided with (a) the exhaustion of easily accessible, hydrolyzable N sources, (b) the complete loss of the topsoil through mineralization and erosion, and (c) the first establishment of pioneer plants on the bare subsoil between Kobresia root-mat remnants. The new fungal symbionts of these pioneer plants explain the particularly pronounced shift in the fungal community from S3 to S4, representing this new ecosystem state. Degradation exceeding S4, therefore, constitutes a severe alteration of microbial community composition and functioning, beyond which biotic and abiotic ecosystem properties have changed so severely that ecosystem recovery is unlikely.
Taken together, overgrazing of Kobresia pastures on the TP induces a highly interrelated sequence of abiotic and biotic degradation processes. Polygonal crack extension and root-mat dieback as a result of freeze-thaw cycles and overgrazing induce topsoil erosion and progressive SOC decomposition and N loss. This is accompanied by a shift in microbial community structure and functions to adapt to the altered availability of SOC and N. Studies of the past decade predict an intensification of the degradation of this pastoral ecosystem because of the large population that is dependent on livestock products3, increasing stocking rates close to settlements4 and diminishing availability of high-quality grazing grounds following degradation9. Once the critical point between S3 and S4 has been passed, conventional strategies for mitigating degradation are unlikely to prevent complete loss of the entire C-rich topsoil, with cascading effects on microbial functioning and the related ecosystem services.
The challenges brought about by TP degradation can only be addressed through lowering livestock densities, and perhaps more importantly increasing livestock mobility to maintain important functions of the vulnerable Kobresia ecosystem. In 2011, China proposed the Ecological Conservation Redline strategy that protects ecologically fragile zones like the TP26 by prioritizing and prohibiting rangeland degradation. Tibetan pastoralists increasingly manage their rangelands through contracting, assessing limits to livestock holdings based on land availability27, e.g., by “multi-household management pattern”. This practice has been shown to reduce SOC losses considerably28, more closely resembling pre-settlement grassland management. Further, the reintroduction of seasonal livestock grazing in the framework of community-based villages can protect or even restore vegetation and soil29, provided that a certain degradation stage has not been surpassed. Once this critical point is crossed, reseeding has been proposed as a restoration strategy30. However, reseeding of heavily degraded areas generally requires N and P fertilization30, which carries the risk of nutrient leaching and headwater pollution and is nevertheless often unsuccessful31. Dong et al.31 studied the restoration potential of TP grasslands and their soil chemical and physical properties in three depth increments (0–4, 4–10, and 10–20 cm). The results show that neither the decreased water holding capacity, nor the SOC and N content could be restored in the reestablished artificial grasslands this was attributed to the loss of the topsoil, changed soil texture, and changes in vegetation composition.
This demonstrates the threshold behavior of TP’s pastures, where a complete recovery from highly degraded stages is unlikely, due to slow pedogenic processes and vegetation restoration, as well as continuously increasing global warming.
Methods
Literature study
Literature considering the effect of pasture degradation on SOC, N, and clay content, as well as bulk density (BD), was assembled by searching (i) Web of Science V.5.22.1, (ii) ScienceDirect (Elsevier B.V.) (iii) Google Scholar, and (iv) the China Knowledge Resource Integrated Database (CNKI). Search terms were “degradation gradient”, “degradation stages”, “alpine meadow”, “Tibetan Plateau”, “soil”, “soil organic carbon”, and “soil organic matter” in different combinations. The criteria for including a study in the analysis were: (i) a clear and comprehensible classification of degradation stages was presented, (ii) data on SOC, N, and/or BD were reported, (iii) a non-degraded pasture site was included as a reference to enable an effect size analysis and the calculation of SOC and N losses, (iv) sampling depths and study location were clearly presented. (v) Studies were only considered that took samples in 10 cm depth intervals, to maintain comparability to the analyses from our own study site. The degradation stages in the literature studies were regrouped into the six successive stages (S0–S5) according to the respective degradation descriptions. In total, we compiled the results of 49 publications published between 2002 and 2020.
When SOM content was presented, this was converted to SOC content using a conversion factor of 2.032. SOC and N stocks were calculated using the following equation:
where elemental stock is SOC or N stock [kg ha−1]; content is SOC or N content [g kg−1]; BD is soil bulk density [g cm−3] and depth is the soil sampling depth [cm].
The effect sizes of individual variables (i.e., SOC and N stocks as well as BD) were quantified as follows:
where ES is the effect size in %, D is the value of the corresponding variable in the relevant degradation stage and R is the value of each variable in the non-degraded stage (reference site). When ES is positive, zero, or negative, this indicates an increase, no change, or decrease, respectively, of the parameter compared to the non-degraded stage.
Experimental design of the field study
Large areas in the study region are impacted by grassland degradation. In total, 45% of the surface area of the Kobresia pasture ecosystem on the TP is already degraded2. The experiment was designed to differentiate and quantify SOC losses by erosion vs. net decomposition and identify underlying shifts in microbial community composition and link these to changes in key microbial functions in the soil C cycle. We categorized the range of Kobresia root-mat degradation from non-degraded to bare soils into six successive degradation stages (S0–S5). Stage S0 represented non-degraded root mats, while stages S1–S4 represented increasing degrees of surface cracks, and bare soil patches without root mats defined stage S5 (Supplementary Fig. 1). All six degradation stages were selected within an area of about 4 ha to ensure equal environmental conditions and each stage was sampled in four field replicates. However, the studied degradation patterns are common for the entire Kobresia ecosystem (Supplementary Fig. 1).
Site description
The field study was conducted near Nagqu (Tibet, China) in the late summer 2013 and 2015. The study site of about 4 ha (NW: 31.274748°N, 92.108963°E; NE: 31.274995°N, 92.111482°E; SW: 31.273488°N, 92.108906°E; SE: 31.273421°N, 92.112025°E) was located on gentle slopes (2–5%) at 4,484 m a.s.l. in the core area of the Kobresia pygmaea ecosystem according to Miehe et al.8. The vegetation consists mainly of K. pygmaea, which covers up to 61% of the surface. Other grasses, sedges, or dwarf rosette plants (Carex ivanoviae, Carex spp., Festuca spp., Kobresia pusilla, Poa spp., Stipa purpurea, Trisetum spp.) rarely cover more than 40%. The growing season is strongly restricted by temperature and water availability. At most, it lasts from mid-May to mid-September, but varies strongly depending on the onset and duration of the summer monsoon. Mean annual precipitation is 431 mm, with roughly 80% falling as summer rains. The mean annual temperature is −1.2 °C, while the mean maximum temperature of the warmest month (July) is +9.0 °C2.
A characteristic feature of Kobresia pastures is their very compact root mats, with an average thickness of 15 cm at the study site. These consist mainly of living and dead K. pygmaea roots and rhizomes, leaf bases, large amounts of plant residue, and mineral particles. Intact soil is a Stagnic Eutric Cambisol (Humic), developed on a loess layer overlying glacial sediments and containing 50% sand, 33% silt, and 17% clay in the topsoil (0–25 cm). The topsoil is free of carbonates and is of neutral pH (pH in H2O: 6.8)5. Total soil depth was on average 35 cm.
The site is used as a winter pasture for yaks, sheep, and goats from January to April. Besides livestock, large numbers of plateau pikas (Ochotona) are found on the sites. These animals have a considerable impact on the plant cover through their burrowing activity, in particular the soil thrown out of their burrows, which can cover and destroy the Kobresia turf.
Sampling design
The vertical and horizontal extent of the surface cracks was measured for each plot (Supplementary Table 2). Vegetation cover was measured and the aboveground biomass was collected in the cracks (Supplementary Table 2). In general, intact Kobresia turf (S0) provided high resistance to penetration as measured by a penetrologger (Eijkelkamp Soil and Water, Giesbeek, NL) in 1 cm increments and four replicates per plot.
Soil sampling was conducted using soil pits (30 cm length × 30 cm width × 40 cm depth). Horizons were classified and then soil and roots were sampled for each horizon directly below the cracks. Bulk density and root biomass were determined in undisturbed soil samples, using soil cores (10 cm height and 10 cm diameter). Living roots were separated from dead roots and root debris by their bright color and soft texture using tweezers under magnification, and the roots were subsequently washed with distilled water to remove the remaining soil. Because over 95% of the roots occurred in the upper 25 cm5, we did not sample for root biomass below this depth.
Additional soil samples were taken from each horizon for further analysis. Microbial community and functional characterization were performed on samples from the same pits but with a fixed depth classification (0–5 cm, 5–15 cm, 15–35 cm) to reduce the number of samples.
Plant and soil analyses
Soil and roots were separated by sieving (2 mm) and the roots subsequently washed with distilled water. Bulk density and root density were determined by dividing the dry soil mass (dried at 105 °C for 24 h) and the dry root biomass (60 °C) by the volume of the sampling core. To reflect the root biomass, root density was expressed per soil volume (mg cm−3). Soil and root samples were milled for subsequent analysis.
Elemental concentrations and SOC characteristics
Total SOC and total N contents and stable isotope signatures (δ13C and δ15N) were analyzed using an isotope ratio mass spectrometer (Delta plus, Conflo III, Thermo Electron Cooperation, Bremen, Germany) coupled to an elemental analyzer (NA 1500, Fisons Instruments, Milano, Italy). Measurements were conducted at the Centre for Stable Isotope Research and Analysis (KOSI) of the University of Göttingen. The δ13C and δ15N values were calculated by relating the isotope ratio of each sample (Rsample = 13C/12C or 15N/14N) to the international standards (Pee Dee Belemnite 13C/12C ratio for δ13C; the atmospheric 15N/14N composition for δ15N).
Soil pH of air-dried soil was measured potentiometrically at a ratio (v/v) of 1.0:2.5 in distilled water.
Lignin phenols were depolymerized using the CuO oxidation method25 and analyzed with a gas chromatography-mass spectrometry (GC–MS) system (GC 7820 A, MS 5977B, Agilent Technologies, Waldbronn, Germany). Vanillyl and syringyl units were calculated from the corresponding aldehydes, ketones, and carboxylic acids. Cinnamyl units were derived from the sum of p-coumaric acid and ferulic acid. The sum of the three structural units (VSC = V + S + C) was considered to reflect the lignin phenol content in a sample.
DNA extraction and PCR
Samples were directly frozen on site at −20 °C and transported to Germany for analysis of microbial community structure. Total DNA was extracted from the soil samples with the PowerSoil DNA isolation kit (MoBio Laboratories Inc., Carlsbad, CA, USA) according to the manufacturer’s instructions, and DNA concentration was determined using a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific, Wilmington, DE, USA). The extracted DNA was amplified with forward and reverse primer sets suitable for either t-RFLP (fluorescence marked, FAM) or Illumina MiSeq sequencing (Illumina Inc., San Diego, USA): V3 (5’-CCT ACG GGN GGC WGC AG-3’) and V4 (5’-GAC TAC HVG GGT ATC TAA TCC-3’) primers were used for bacterial 16 S rRNA genes whereas ITS1 (5’-CTT GGT CAT TTA GAG GAA GTA A-3’), ITS1-F_KYO1 (5’-CTH GGT CAT TTA GAG GAA STA A-3’), ITS2 (5’-GCT GCG TTC TTC ATC GAT GC-3’) and ITS4 (5’-TCC TCC GCT TAT TGA TAT GC-3’) were used for fungi33,34. Primers for Illumina MiSeq sequencing included adaptor sequences (forward: 5’-TCG TCG GCA GCG TCA GAT GTG TAT AAG AGA CAG-3’; reverse: 5’-GTC TCG TGG GCT CGG AGA TGT GTA TAA GAG ACA G-3’)33. PCR was performed with the Phusion High-Fidelity PCR kit (New England Biolabs Inc., Ipswich, MA, USA) creating a 50 µl master mix with 28.8 µl H2Omolec, 2.5 µl DMSO, 10 µl Phusion GC buffer, 1 µl of forward and reverse primer, 0.2 µl MgCl2, 1 µl dNTPs, 0.5 µl Phusion HF DNA Polymerase, and 5 µl template DNA. PCR temperatures started with initial denaturation at 98 °C for 1 min, followed by denaturation (98 °C, 45 s), annealing (48/60 °C, 45 s), and extension (72 °C, 30 s). These steps were repeated 25 times, finalized again with a final extension (72 °C, 5 min), and cooling to 10 °C. Agarose gel electrophoresis was used to assess the success of the PCR and the amount of amplified DNA (0.8% gel:1.0 g Rotigarose, 5 µl Roti-Safe Gelstain, Carl Roth GmbH & Co. KG, Karlsruhe, Germany; and 100 ml 1× TAE-buffer). PCR product was purified after initial PCR and restriction digestion (t-RFLP) with either NucleoMag 96 PCR (16 S rRNA gene amplicons, Macherey-Nagel GmbH & Co. KG, Düren, Germany) or a modified clean-up protocol after Moreau (t-RFLP)35: 3× the volume of the reaction solution as 100% ethanol and ¼x vol. 125 mM EDTA was added and mixed by inversion or vortex. After incubation at room temperature for 15 min, the product was centrifuged at 25,000 × g for 30 min at 4 °C. Afterwards the supernatant was removed, and the inverted 96-well plate was centrifuged shortly for 2 min. Seventy microliters ethanol (70%) were added and centrifuged at 25,000 × g for 30 min at 4 °C. Again, the supernatant was removed, and the pallet was dried at room temperature for 30 min. Finally, the ethanol-free pallet was resuspended in H2Omolec.
T-RFLP fingerprinting
The purified fluorescence-labeled PCR products were digested with three different restriction enzymes (MspI and BstUI, HaeIII) according to the manufacturer’s guidelines (New England Biolabs Inc., Ipswich, MA, USA) with a 20 µl master mix: 16.75 µl H2Omolec, 2 µl CutSmart buffer, 0.25 or 0.5 µl restriction enzyme, and 1 µl PCR product for 15 min at 37 °C (MspI) and 60 °C (BstUI, HaeIII), respectively. The digested PCR product was purified a second time35, dissolved in Super-DI Formamide (MCLAB, San Francisco, CA, USA) and, along with Red DNA size standard (MCLAB, San Francisco, USA), analyzed in an ABI Prism 3130 Genetic Analyzer (Applied Biosystems, Carlsbad, CA, USA). Terminal restriction fragments shorter than 50 bp and longer than 800 bp were removed from the t-RFLP fingerprints.
16 S rRNA gene and internal transcribed spacer (ITS) sequencing and sequence processing
The 16 S rRNA gene and ITS paired-end raw reads for the bacterial and fungal community analyses were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) and can be found under the BioProject accession number PRJNA626504. This BioProject contains 70 samples and 139 SRA experiments (SRR11570615–SRR11570753) which were processed using CASAVA software (Illumina, San Diego, CA, USA) for demultiplexing of MiSeq raw sequences (2 × 300 bp, MiSeq Reagent Kit v3).
Paired-end sequences were quality-filtered with fastp (version 0.19.4)36 using default settings with the addition of an increased per base phred score of 20, base-pair corrections by overlap (-c), as well as 5′- and 3′-end read trimming with a sliding window of 4, a mean quality of 20 and minimum sequence size of 50 bp. Paired-end sequences were merged using PEAR v0.9.1137 with default parameters. Subsequently, unclipped reverse and forward primer sequences were removed with cutadapt v1.1838 with default settings. Sequences were then processed using VSEARCH (v2.9.1)39. This included sorting and size-filtering (—sortbylength,—minseqlength) of the paired reads to ≥300 bp for bacteria and ≥140 bp for ITS1, dereplication (—derep_fulllength). Dereplicated sequences were denoised with UNOISE340 using default settings (—cluster_unoise—minsize 8) and chimeras were removed (—uchime3_denovo). An additional reference-based chimera removal was performed (—uchime_ref) against the SILVA41 SSU NR database (v132) and UNITE42 database (v7.2) resulting in the final set of amplicon sequence variants (ASVs)43. Quality-filtered and merged reads were mapped to ASVs (—usearch_global–id 0.97). Classification of ASVs was performed with BLAST 2.7.1+ against the SILVA SSU NR (v132) and UNITE (v7.2) database with an identity of at least 90%. The ITS sequences contained unidentified fungal ASVs after UNITE classification, these sequences were checked (blastn)44 against the “nt” database (Nov 2018) to remove non-fungal ASVs and only as fungi classified reads were kept. Sample comparisons were performed at the same surveying effort, utilizing the lowest number of sequences by random selection (total 15,800 bacteria, 20,500 fungi). Species richness, alpha and beta diversity estimates, and rarefaction curves were determined using the QIIME 1.9.145 script alpha_rarefaction.py.
The final ASV tables were used to compute heatmaps showing the effect of degradation on the community using R (Version 3.6.1, R Foundation for Statistical Computing, Vienna, Austria) and R packages “gplots”, “vegan”, “permute” and “RColorBrewer”. Fungal community functions were obtained from the FunGuild database46. Plant mycorrhizal association types were compiled from the literature38,39,40,41,47,48,49,50. If no direct species match was available, the mycorrhizal association was assumed to remain constant within the same genus.
Enzyme activity
Enzyme activity was measured to characterize the functional activity of the soil microorganisms. The following extracellular enzymes, involved in C, N, and P transformations, were considered: two hydrolases (β-glucosidase and xylanase), phenoloxidase, urease, and alkaline phosphatase. Enzyme activities were measured directly at the sampling site according to protocols after Schinner et al.51. Beta-glucosidase was incubated with saligenin for 3 h at 37 °C, xylanase with glucose for 24 h at 50 °C, phenoloxidase with L-3,4-dihydroxy phenylalanine (DOPA) for 1 h at 25 °C, urease with urea for 2 h at 37 °C and alkaline phosphatase on P-nitrophenyl phosphate for 1 h at 37 °C. Reaction products were measured photometrically at recommended wavelengths (578, 690, 475, 660, and 400 nm, respectively).
SOC stocks and SOC loss
The SOC stocks (in kg C m−2) for the upper 30 cm were determined by multiplying the SOC content (g C kg−1) by the BD (g cm−3) and the thickness of the soil horizons (m). SOC losses (%) were calculated for each degradation stage and horizon and were related to the mean C stock of the reference stage (S0). The erosion-induced SOC loss of the upper horizon was estimated by considering the topsoil removal (extent of vertical soil cracks) of all degraded soil profiles (S1–S5) and the SOC content and BD of the reference (S0). To calculate the mineralization-derived SOC loss, we accounted for the effects of SOC and root mineralization on both SOC content and BD. Thus, we used the SOC content and BD from each degradation stage (S1–S5) and multiplied it by the mean thickness of each horizon (down to 30 cm) from the reference site (S0). The disentanglement of erosion-derived SOC loss from mineralization-derived SOC loss was based on explicit assumptions that (i) erosion-derived SOC losses are mainly associated with losses from the topsoil, and (ii) the decreasing SOC contents in the erosion-unaffected horizons were mainly driven by mineralization and decreasing root C input.
Statistical analyses
Statistical analyses were performed using PASW Statistics (IBM SPSS Statistics) and R software (Version 3.6.1). Soil and plant characteristics are presented as means and standard errors (means ± SE). The significance of treatment effects (S0–S5) and depth was tested by one-way ANOVA at p < 0.05. Prior to this, we checked for normality and homogeneity of variance using the Shapiro–Wilk test and Levene’s test, respectively.
Post-hoc multiple comparisons were carried out using the LSD or Tukey HSD ANOVA, if normality was indicated. In cases of non-normal distribution, the nonparametric Kruskal–Wallis test was implemented coupled with a Bonferroni correction. To detect relationships between various plant and soil characteristics, we used linear and nonlinear regressions. Correlations were deemed significant for single regressions at p < 0.05.
Before testing for significant differences, three outliers were detected by Grubbs outlier test (p < 0.05) in the bacterial t-RFLP and MiSeq datasets: MspI: S0_B_0-5, S1_E_0-5; BstuI: S5_A_15-35; MiSeq: S3_E_5-15 and were excluded from the analysis. T-RFLP and MiSeq data of all degradation stages were compared for significant differences with MANOVA, based on the Bray–Curtis index for dissimilarity. For pairwise multilevel comparisons, “pairwiseAdonis” was used52. Differences in microbial community data from t-RFLP and MiSeq were displayed in non-metric multidimensional scaling (NMDS) plots and environmental factors were correlated by canonical correspondence analysis (CCA). Statistics on community data were carried out with R statistical software (Version 3.6.1).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data generated in this study have been deposited in the PANGAEA Open Access library under accession code https://doi.org/10.1594/PANGAEA.918249. The 16 S rRNA and ITS gene paired-end raw reads for the bacterial and fungal community analyses have been deposited in the National Center for Biotechnology In-formation (NCBI) Sequence Read Archive (SRA) under the accession code PRJNA626504; BioProject: Tibetan plateau microbiome.
References
Genxu, W., Qian, J., Cheng, G. & Lai, Y. Soil organic carbon pool of grassland soils on the Qinghai-Tibetan Plateau and its global implication. Sci. Total Environ. 291, 207–217 (2002).
Babel, W. T. et al. Pasture degradation modifies the water and carbon cycles of the Tibetan highlands. Biogeosciences 11, 6633–6656 (2014).
Qiu, J. Trouble in Tibet. Nature 529, 142–145 (2016).
Zhou, H., Zhao, X., Tang, Y., Gu, S. & Zhou, L. Alpine grassland degradation and its control in the source region of the Yangtze and Yellow Rivers. China Grassl. Sci. 51, 191–203 (2005).
Schleuss, P.-M. et al. Nitrogen uptake in an alpine Kobresia pasture on the Tibetan Plateau: localization by 15N labeling and implications for a vulnerable ecosystem. Ecosystems 18, 946–957 (2015).
Ingrisch, J. et al. Carbon pools and fluxes in a Tibetan alpine Kobresia pygmaea pasture partitioned by coupled eddy-covariance measurements and 13CO2 pulse labeling. Sci. Total Environ. 505, 1213–1224 (2015).
Wang, Y. & Wesche, K. Vegetation and soil responses to livestock grazing in Central Asian grasslands: a review of Chinese literature. Biodivers. Conserv. 25, 2401–2420 (2016).
Miehe, G. et al. The Kobresia pygmaea ecosystem of the Tibetan highlands—origin, functioning and degradation of the world’s largest pastoral alpine ecosystem. Sci. Total Environ. 648, 754–771 (2019).
Harris, R. B. Rangeland degradation on the Qinghai-Tibetan plateau: a review of the evidence of its magnitude and causes. J. Arid Environ. 74, 1–12 (2010).
Wang, C. T., Long, R. J., Wang, Q. L., Jing, Z. C. & Shi, J. J. Changes in plant diversity, biomass and soil C, in alpine meadows at different degradation stages in the headwater region of three rivers, China. Land Degrad. Dev. 20, 187–198 (2009).
Kögel-Knabner, I. The macromolecular organic composition of plant and microbial residues as inputs to soil organic matter. Soil Biol. Biochem. 34, 139–162 (2002).
Cragg, S. M. et al. Lignocellulose degradation mechanisms across the Tree of Life. Curr. Opin. Chem. Biol. 29, 108–119 (2015).
Andlar, M. et al. Lignocellulose degradation: an overview of fungi and fungal enzymes involved in lignocellulose degradation. Eng. Life Sci. 18, 768–778 (2018).
Hagedorn, F., Gavazov, K. & Alexander, J. M. Above- and belowground linkages shape responses of mountain vegetation to climate change. Science 365, 1119–1123 (2019).
Wang, T. et al. Permafrost thawing puts the frozen carbon at risk over the Tibetan Plateau. Sci. Adv. 6, eaaz3513 (2020).
Craine, J. M., Morrow, C. & Fierer, N. Microbial nitrogen limitation increases decomposition. Ecology 88, 2105–2113 (2007).
Chen, L. et al. Nitrogen availability regulates topsoil carbon dynamics after permafrost thaw by altering microbial metabolic efficiency. Nat. Commun. 9, 3951 (2018).
Che, R. et al. Increase in ammonia-oxidizing microbe abundance during degradation of alpine meadows may lead to greater soil nitrogen loss. Biogeochemistry 136, 341–352 (2017).
Xie, Z. et al. Identifying response groups of soil nitrifiers and denitrifiers to grazing and associated soil environmental drivers in Tibetan alpine meadows. Soil Biol. Biochem. 77, 89–99 (2014).
Liu, S., Schleuss, P.-M. & Kuzyakov, Y. Carbon and nitrogen losses from soil depend on degradation of Tibetan Kobresia pastures. Land Degrad. Devel. 28, 1253–1262 (2017).
Gao, Y. H., Luo, P., Wu, N., Chen, H. & Wang, G. X. Impacts of grazing intensity on nitrogen pools and nitrogen cycle in an alpine meadow on the Eastern Tibetan Plateau. Appl. Ecol. Environ. Res. 6, 69–79 (2008).
Dong, S. et al. Grazing promoted soil microbial functional genes for regulating C and N cycling in alpine meadow of the Qinghai-Tibetan Plateau. Agric. Ecosyst. Environ. 303, 107111 (2020).
Schweizer, M., Fear, J. & Cadisch, G. Isotopic (13C) fractionation during plant residue decomposition and its implications for soil organic matter studies. Rapid Commun. Mass Spectrom. 13, 1284–1290 (1999).
Hobbie, E. A. & Werner, R. A. Intramolecular, compound-specific, and bulk carbon isotope patterns in C3 and C4 plants: A review and synthesis. N. Phytol. 161, 371–385 (2004).
Pepper, J. M., Casselman, B. W. & Karapally, J. C. Lignin oxidation. Preferential use of cupric oxide. Can. J. Chem. 45, 3009–3012 (1967).
Gao, J., Zou, C., Zhang, K., Xu, M. & Wang, Y. The establishment of Chinese ecological conservation redline and insights into improving international protected areas. J. Environ. Manag. 264, 110505 (2020).
Yeh, E. T., Samberg, L. H., Gaerrang, Volkmar, E. & Harris, R. B. Pastoralist decision-making on the Tibetan plateau. Hum. Ecol. 45, 333–343 (2017).
Cao, J. et al. Impact of grassland contract policy on soil organic carbon losses from alpine grassland on the Qinghai–Tibetan Plateau. Soil Use Manag. 33, 663–671 (2017).
Zhuang, M., Gongbuzeren, Zhang, J. & Wenjun, L. Community-based seasonal movement grazing maintains lower greenhouse gas emission intensity on Quinghai-Tibet Plateau of China. Land Unse Policy 85, 155–160 (2019).
Liu, S., Zamanian, K., Schleuss, P.-M., Zarebanadkouki, M. & Kuzyakov, Y. Degradation of Tibetan grasslands: Consequences for carbon and nutrient cycles. Agric. Ecosyst. Environ. 252, 93–104 (2018).
Dong, S. K. et al. Soil-quality effects of grassland degradation and restoration on the Qinghai-Tibetan plateau. Soil Sci. Soc. Am. J. 76, 2256–2264 (2012).
Pribyl, D. W. A critical review of the conventional SOC to SOM conversion factor. Geoderma 156, 75–83 (2010).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, 1–11 (2013).
Toju, H., Tanabe, A. S., Yamamoto, S. & Sato, H. High-coverage ITS primers for the DNA-based identification of ascomycetes and basidiomycetes in environmental samples. PLoS ONE 7, e40863 (2012).
Moreau, C. S. A practical guide to DNA extraction, PCR, and gene-based DNA sequencing in insects. HALTERES 5, 32–42 (2014).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv https://www.biorxiv.org/content/10.1101/081257v1.full.pdf+html (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Abarenkov, K. et al. The UNITE database for molecular identification of fungi—recent updates and future perspectives. N. Phytol. 186, 281–285 (2010).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 11, 2639–2643 (2017).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Nguyen, N. H. et al. FUNGuild: An open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 20, 241–248 (2016).
Shi, Z. Y., Feng, G., Christie, P. & Li, X. L. Arbuscular mycorrhizal status of spring ephemerals in the desert ecosystem of Junggar Basin, China. Mycorrhiza 16, 269–275 (2006).
Tian, H., Gai, J. P., Zhang, J. L., Christie, P. & Li, X. L. Arbuscular mycorrhizal fungi associated with wild forage plants in typical steppe of eastern Inner Mongolia. Eur. J. Soil Biol. 45, 321–327 (2009).
Wang, B. & Qiu, Y. L. Phylogenetic distribution and evolution of mycorrhizas in land plants. Mycorrhiza 16, 299–363 (2006).
Gai, J. P., Feng, G., Cai, X. B., Christie, P. & Li, X. L. A preliminary survey of the arbuscular mycorrhizal status of grassland plants in southern Tibet. Mycorrhiza 16, 191–196 (2006).
Schinner, F., Öhlinger, R., Kandeler, E. & Margesin, R. Methods in Soil Biology. (Springer-Verlag, 1996).
Martinez Arbizu, P. pairwiseAdonis: Pairwise multilevel comparison using adonis. R package version 0.3. https://github.com/pmartinezarbizu/pairwiseAdonis (2020).
Acknowledgements
We thank our colleagues T. Biermann, W. Babel, U. Bange, L. Becker, C. Blaser, H. Coners, I. Hoeft, P. Jannack, K. Krüger, K. Schützenmeister, L. Steingräber, Y. Sun, and S. Willinghöfer for helping in the field and the laboratory. The German Research Council (Priority Program 1372) and the Centre for Stable Isotope Research and Analysis (KOSI) of Göttingen supported this project. This research was funded by the German Research Foundation (DFG) in the frame of the Priority Program SPP 1372 “Tibetan Plateau: Formation—Climate—Ecosystems”, DFG. 2009–2015; KU 1184/14-2, GU 406/22-2. We acknowledge financial support by DFG within the funding programme Open Access-Publikationskosten.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
P.-M.S., G.G., Y.K., and S.S. designed the study. P.-M.S. and X.X. performed SOC, N, P CEC, and stable isotope signature (δ13C and δ15N) measurements. S.S. performed measurements of enzyme activity and lignin phenol content. S.L. conducted the literature study and meta-analysis. J.X., D.S., A.B., C.B., M.G., H.N., Y.H., and J.M. contributed to molecular biological analyses and biostatistics. P.-M.S., G.M., E.S., X.X., and S.S. performed soil and vegetation sampling, field measurements, and vegetation description. P.-M.S., F.H., A.B., T.d.l.H., S.L., T.D., and K.M.-J. performed statistical data analyses and interpretation. G.G., Y.K., M.A.D., and S.S. conceptualized and supervised the research. All authors were involved in the data interpretation, writing, and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Breidenbach, A., Schleuss, PM., Liu, S. et al. Microbial functional changes mark irreversible course of Tibetan grassland degradation. Nat Commun 13, 2681 (2022). https://doi.org/10.1038/s41467-022-30047-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-30047-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.