Abstract
Across the evolutionary history of insects, the shift from nitrogen-rich carnivore/omnivore diets to nitrogen-poor herbivorous diets was made possible through symbiosis with microbes. The herbivorous turtle ants Cephalotes possess a conserved gut microbiome which enriches the nutrient composition by recycling nitrogen-rich metabolic waste to increase the production of amino acids. This enrichment is assumed to benefit the host, but we do not know to what extent. To gain insights into nitrogen assimilation in the ant cuticle we use gut bacterial manipulation, 15N isotopic enrichment, isotope-ratio mass spectrometry, and 15N nuclear magnetic resonance spectroscopy to demonstrate that gut bacteria contribute to the formation of proteins, catecholamine cross-linkers, and chitin in the cuticle. This study identifies the cuticular components which are nitrogen-enriched by gut bacteria, highlighting the role of symbionts in insect evolution, and provides a framework for understanding the nitrogen flow from nutrients through bacteria into the insect cuticle.
Similar content being viewed by others
Introduction
Advanced genomic technologies and bioinformatics approaches have increased exponentially the diversity of known microbes in recent years. Nevertheless, understanding the role of microbial symbionts in hosts remains elusive. One approach for studying symbiosis is to manipulate microbes and monitor the consecutive changes within the host. This strategy has been applied successfully in the field of insect–microbe interactions demonstrating that beneficial bacteria increase the fitness1,2, tolerance to abiotic stressors3, nutrient assimilation3,4 chemical defense5,6, and cuticle formation7,8 of hosts. Although a limited number of biochemical processes by which microbes contribute to host’s metabolism are known, new molecular pathways are likely to be discovered.
The evolution of ants (∼150 million years)9 has been shaped by mutualistic interactions with microbes, which have permitted ants to radiate into new ecological niches10 becoming one of the most ecologically important insect groups11. The shift from nitrogen-rich carnivore/omnivore diets to nitrogen-poor herbivorous diets has evolved multiple times during ant evolution12. For example, the intracellular symbiont Blochmannia, associated with Camponotus ants produces several amino acids that help ants assimilate nitrogen—consequently improving larval development, fitness, and reproductive success13,14. Another example is the herbivorous turtle ants of the genus Cephalotes that that have low nitrogen diets and benefit from the nutritional contribution of gut microbes10. Genomic analysis of gut bacteria from turtle ant host species revealed the occurrence of bacterial genes related to nitrogen recycling and assimilation for the biosynthesis of essential and non-essential amino acids15. The microbially derived amino acids are taken up by the host, which presumably benefits them and supports a low nitrogen herbivorous diet. Interestingly, the conserved and phylogenetically distant gut symbionts of Cephalotes ants (Burkholderiales, Opitutales, Pseudomonadales, Rhizobiales, and Xanthomonadales) possess similar and complementary biosynthetic genes to recycle urea to produce multiple amino acids, suggesting, among other things, a high amino acid demand by the host.
It has been suggested that the amino acid tyrosine (Tyr) produced in significant quantities by bacteria during insect development is involved in cuticle formation7. This is supported by previous work in insect cuticle biochemistry showing the precursor role of aromatic amino acids in the metabolic pathways of melanization16 and sclerotization17,18. In beetles, several studies have shown that different host species have vertically transmitted endosymbionts, which contribute to the host’s cuticle formation, and in the absence of associated bacteria reported significantly thinner, softer, and pale/reddish cuticles instead of dark and thick cuticles7,8,19,20. The color, roughness, thickness, permeability, and mechanical properties of the cuticle vary across insects and these traits have played a role in insect adaptation into new ecological niches21. However, there are still gaps in our understanding of which cuticular constituents depend on the symbiont’s contribution as these past studies have mostly only noted physical changes in the absence of symbionts without identifying any molecular components of the cuticle impacted by symbionts. Symbiont contribution to cuticle formation emphasizes the potential important impact of symbiosis in insect evolution. Therefore, based on the fact that Cephalotes cuticles are very tough and often dark, we hypothesize that gut bacteria participate in the cuticle formation of arboreal herbivorous turtle ants. Although the combination of 15N isotopic enrichment experiment with 15N solid-state NMR has been a successful methodology to reveal the molecular structure of the chitin–catecholamide–protein matrix, specifically for characterizing the nitrogen–carbon bonds in the protein–catecholamide and inter-catecholamide cross-links22,23, this approach has not been used in recent studies of insect cuticle and never to investigate if host-associated bacteria contribute to these processes.
In this work, we perform feeding experiments with urea-15N2 in the presence or absence of antibiotics to reveal the contribution of bacteria in cuticle formation during insect development. Our approach consists of using NMR to document and characterize the changes in the gut and cuticle including the 15N-enriched products in the chitin–catecholamide–protein matrix. Overall, we report the biomolecular mechanism of cuticle formation assisted by gut bacteria in herbivorous ants.
Results and discussion
Study species and experimental manipulation
In herbivorous Cephalotes turtle ants their nitrogen-poor diet is supplemented by amino acids produced by gut bacteria from recycling nitrogen host waste such as urea15. These bacteria almost exclusively belong to families Burkholderiales, Opitutales, Pseudomonadales, Rhizobiales, and Xanthomonadales, are highly related across Cephalotes species, and possess genes involved in nitrogen recycling and assimilation. Labeled urea-15N2 was added to the sterile diet of Cephalotes varians of six different colonies for a sufficient period of time to ensure the full development of individuals from larvae to adults and also treated half of the colonies with antibiotics to compare to untreated colonies. Despite the difficulty of rearing Cephalotes species in laboratory, five larvae from each group fully developed to the adult stage and individuals from independent colonies were pooled for analysis.
Bacterial community, isotopic, and cuticle response across treatment
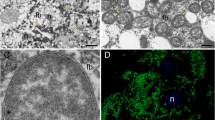
To assess if turtle ant gut bacteria were disrupted in our paired feeding experiments, we confirmed the bacterial communities had been knocked down through 16S rRNA qPCR and amplicon sequencing. Our antibiotic treatment reduced the bacterial quantity in our samples from 14,033 to 6607 16S rRNA copy numbers (Supplementary Table 1), and analysis of the bacterial community diversity in our antibiotic treatment samples found almost all transient and environmental contaminates compared to our intact bacterial communities, which were dominated by symbiotic bacterial amplicon sequence variants (Supplementary Fig. 1). To measure the contribution of gut bacteria to the cuticle, we have quantitative and qualitative evidence. The shift in δ15N values in the cuticle measured by isotope-ratio mass spectroscopy demonstrates an impressive contribution of bacteria with a fourfold increase in the untreated samples compare to sample from the antibiotics experiment (Supplementary Table 2). Scanning electronic microscopy reveals that the cuticle of untreated C. varians individual is twofold thicker compare to treated individual (Supplementary Fig. 2).
Quantification of aromatic amino acids in turtle ant guts
For the NMR experiments of pooled gut extracts, a proton 1H spectrum was recorded in order to track the aromatic amino acids, which are the precursors of melanin and the chitin–protein catecholamine cross-linkers in the cuticle (Supplementary Fig. 3). Resonances of interest appeared at low intensity in the aromatic region (6.5–8.5 ppm). The recorded 2D 1H–1H correlation spectroscopy (COSY) map (Supplementary Fig. 4) shows a number of correlations between metabolites, specifically two major coupled spin systems in the aromatic area (6.9/7.2 and 7.3/7.4 ppm), which were tentatively identified as Tyr and phenylalanine (Phe) resonances (Supplementary Fig. 4). To improve the identification of aromatic amino acids, a zero-quantum filtered total COSY experiment was carried out (Supplementary Figs. S5 and S6). The correlations between aliphatic resonances Hα and Hβ were detected for Phe and Tyr and the aromatic area of the 2D TOCSY spectra shows the Phe and Tyr intra-aromatic correlations as well as other less concentrated aromatic amino acids histidine (His) and tryptophan (Trp) (Supplementary Fig. 6). The diffusion order spectroscopy (DOSY) experiment unambiguously extracts the 1H NMR spectra of Phe and Tyr due to their close but different logD values (−9.44 for Tyr and −9.42 for Phe) (Supplementary Fig. 6), thus making it possible to quantify the respective concentration of Phe and Tyr in a single measurement of five pooled individuals of the gut extract samples from the antibiotic treated and untreated experiments (Supplementary Table 3). A similar concentration of Phe and Tyr at ~11 μM without antibiotic treatment was calculated and this value drops to twofold lower (~5 μM) in the gut extract of the antibiotic treated group.
Identification of cuticular components impacted by gut bacteria
The 13C cross-polarization magic-angle spinning (CP-MAS) NMR spectra of samples (antibiotics treated or untreated C. varians individuals) were recorded (Supplementary Fig. 7). Although resonances from chitin, proteins, and catecholamide were detected in agreement with previous work22, no significant difference was noticed between 13C resonances of antibiotic treated and untreated samples as expected as we did not enrich 13C in their diet. In contrast, the CP-MAS 15N NMR spectrum appeared greatly dissimilar (Fig. 1). The 15N resonances in antibiotic treated samples were qualitatively (number of peaks) and quantitatively (peak intensity) less compared to 15N resonances in the spectra of untreated samples, and different from the 15N NMR spectra at natural isotopic abundance of unlabeled untreated C. varians ants which is characterized by a single large resonance at 120 ppm (Fig. 1). Based on previous work on solid-state 15N NMR spectroscopy of insect cuticle22,23, we attempted to annotate all 15N resonances (Fig. 1 and Supplementary Table 4). The main 15N-enriched resonances in untreated samples, a broad peak at 100–145 ppm with a maximum at 119 ppm, represent the overlapping peaks of different 15N amide resonances in chitin (Supplementary Figs. S8 and S9), cuticular proteins, and catecholamide cross-linkers. We confirm the presence of 15N-enriched chitin in untreated C. varians individuals with a double cross-polarization {1H}–{15N}–13C experiment using a cryogenically cooled CP-MAS probe, which shows the resonances of the C1 and C2 carbons in close proximity to the 15N amide to allow polarization exchange of the 15N amide function in chitin (Supplementary Fig. 10). This suggests that the aminotransferase adds a glutamine amino acid 15N-enriched by gut bacteria to fructose-6-phosphate to form 15N-labeled chitin in Cephalotes cuticle (Fig. 2b)24. The broad peak (100–145 ppm) reflects the amide functional group in chitin overlapping with additional amide resonances of cuticular proteins composed of 15N-enriched amino acids and catecholamide cross-linkers N-acetyldopamine (NADA) and N-β-alanyldopamine (NBDA) from presumably microbially derived 15N-enriched Phe and Tyr precursors (Fig. 2c).
Untreated (red line) and antibiotic treated (blue line) C. varians samples. Both spectra are normalized with respect to sample mass in the rotor and the number of accumulations. Enriched nitrogen is represented in pink. Inset depicts the 15N NMR spectrum of cuticle at natural isotopic abundance of unlabeled untreated C. varians ants.
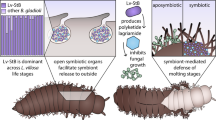
a Biosynthetic pathways impacted by gut bacteria in the production of amino acids from previous work15 and molecular components of the cuticle inferred in this study (b–d). e Overall representation of gut bacteria contribution to ant cuticle (proteins, chitin, and cross-linkers are shown in different colors). Enriched nitrogen is represented in pink. Ant photograph used with permission from the Field Museum of Natural History (FMNHINS 0000 062 654).
The second resonances of interest which was found in the untreated sample only is the peak at 153 ppm identified as the N-substituted nitrogen (Nε2) in the protonated imidazole ring of His (Fig. 1). The complexity of isotropic chemical shifts of the Nδ1 and Nε2 sites in His has been thoroughly described for NMR spectroscopy of enzymes25,26,27. The chemical shift of 153 ppm was assigned to the N-substituted Nε2 site of protonated His covalently bonded to chitin, whereas the non-protonated Nε2 site of neutral His was putatively attributed to the resonance at 139 ppm. Neither of the resonances of the protonated and non-protonated Nδ1 site in His were observed, respectively, at 144–147 and 225–230 ppm. The attributed Nε2 resonances of His only detected in untreated C. varians ants highlight an unexpected contribution of the gut bacteria to the nitrogen connecting cuticular proteins and chitin in the cuticle. Such enrichment has to occur early in His biosynthesis with a 15N-enriched aspartame as the source of 15N (Fig. 2d and Supplementary Fig. 11)28.
The 15N resonances between 70 and 100 ppm are attributed to the guanidine nitrogen (Nη1, Nη2, Nε) of arginine and the Nη resonance at 77 ppm has a quantitative higher enrichment in antibiotic treated samples. Enhanced resonance in the 15N spectra of untreated samples, between 40 and 50 ppm, represent the secondary amine resonance previously identified in the inter-catecholamide cross-links23. In insect cuticle, catecholamide cross-linkers NADA and NBDA connect chitin and proteins during sclerotization. It has been shown that another type of cross-link can occur from the reaction between NBDA and N-β-alanylnorepinephrine to form catecholamide–catecholamide cross-links23. These results underline that 15N-recycling by gut bacteria contribute to the formation of different types of cross-links in the cuticular chitin–protein matrix (Fig. 2e).
Originally, we expected 15N-enrichment to occur in the melanization and sclerotization pathways since the gut bacteria increase the concentration of Tyr and Phe, which are the biosynthetic precursors of melanin and catecholamide cross-linkers (Supplementary Fig. 12). The broad 15N NMR resonances of melanin, which is characterized by multiple resonance peaks (100–190 ppm) from a complex mixture of catechol- and dihydroxyindole-based oligomers29,30, were not identified in our samples, as confirmed by comparison with spectra of melanin standards (Supplementary Figs. 13 and 14). This is likely because the quantity of melanin in a ∼10 mg sample was too small to be detected. Even though the cuticle of antibiotic treated C. varians individuals appears to be as dark as untreated individuals it was not possible to assess the contribution of gut bacteria to melanin production.
Implications for insect evolution
From an evolutionary standpoint, the functional redundancy related to the nitrogen flux in the genomes of the conversed symbionts (Burkholderiales, Opitutales, Pseudomonadales, Rhizobiales, and Xanthomonadales) in the gut of herbivorous turtle ants is intriguing. Maybe having different bacterial families, which can feed on various nutritional sources, allow for the constant production of amino acids for the host. Such multipartite mutualism is contrasting with the single strain Nardonella bacterial endosymbiont of several weevil beetle species, which contributes to cuticle formation7. Nardonella’s small genome (0.2 Mb) retains only the Tyr metabolic pathway and no other amino acid biosynthetic pathway was identified. Similar results were reported for the Sitophilus cereal weevil associated with the Sodalis pierantonius endosymbiont which provides Tyr and Phe during metamorphosis to contribute to cuticle formation19. Tyr and Phe produced by the symbiont are assumed to be involved in sclerotization and melanization processes as reddish and soft cuticles are formed in the absence of the symbiont. In comparison, we have demonstrated that turtle ant-associated bacteria not only contribute to the production of Tyr and Phe but to the formation of chitin, cuticular proteins, and catecholamine cross-linkers (Fig. 2e). Therefore the macromolecular structure, thickness, permeability, and mechanical properties of ant cuticle are likely impacted by gut bacteria suggesting that the conserved symbionts have indirectly contributed to the cuticle-associated traits, which have played a role in turtle ant adaptation into new ecological niches.
As we discover the roles played by mutualistic bacteria in hosts, the study of nitrogen flux is revealing which host physiological functions benefit from the microbial enrichment of amino acids. We report multiple lines of evidence that the nitrogen-recycling activity of gut bacteria is involved in several biosynthetic pathways contributing to host cuticle formation. We identify the contribution of gut bacteria at the molecular level to produce cuticular components, i.e., chitin, proteins, and cross-linkers. Overall these results demonstrate the important role of symbiotic bacteria in insect ecology and evolution.
Methods
Collection
C. varians colonies (N = 6) were collected in the Florida Keys, USA (Florida Department of Environmental Protection scientific research permit 04251635; US Fish and Wildlife research permit FFO4RFKD-2015-0). Voucher specimens are deposited in the Cornell University Insect Collection.
Feeding experiment
For the feeding experiment with urea-15N2 (Sigma-Aldrich, ref: 316930), six colonies were split into two treatment groups. In the first treatment, the colony were subjected to antibiotic feeding to remove their gut bacteria through rearing on 30% (weight/volume) sucrose water containing 0.01% (w/v) of each Tetracycline, Rifampicin, and Kanamycin. Untreated colonies from the second treatment group consumed only 30% sucrose water. After 3 weeks of antibiotics treatment both treated and untreated colonies were reared upon the same 30% sucrose water diet also containing 1% (w/v) urea-15N2.
Bacterial 16S rRNA qPCR and amplicon sequencing
The abdomen of one adult from each treatment group was extracted with the Qiagen Powersoil Kit (Qiagen, Germantown, MD) following the manufactures protocol. We amplified the bacterial region of 16S rRNA with primers described in Caporaso et al.31, following the Earth Microbiome Project protocol [515f primer (5′-GTGCCAGCMGCCGCGGTAA) and 806r primer (5′-GGACTACHVHHHTWTCTAAT); for details see: http://www.earthmicrobiome.org/emp-standard-protocols/16s/]. An Illumina MiSeq run using MiSeq V2 Reagent Kit 300 Cycles (150 × 150) was performed using the primers and procedures described previously31. We measured the quantity of bacterial DNA present with quantitative PCR of the bacterial 16S rRNA gene using the same primers as above. All qPCRs were performed in triplicate on a CFX Connect Real-Time System (Bio-Rad, Hercules, CA). Standard curves were created from serial dilutions of linearized plasmid containing inserts of the E. coli 16S rRNA gene and melt curves were used to confirm the absence of qPCR primer dimers. The resulting triplicate quantities were averaged before calculating the number of bacterial 16S rRNA gene copies per microliter of DNA extraction.
Preparation of ant samples for chemical analysis
After 8 weeks of rearing the larvae that became adults over the course of feeding experiments were collected and individuals were pooled for each treatment group (treated N = 5, and untreated N = 5). The gut of these individuals (N = 5 per treatment) was dissected and metabolites were extracted by grinding pooled guts with a pestle in an Eppendorf tube (1.5 mL) containing a solvent mix (0.5 mL, ethyl acetate/methanol/water: 3:2:1). The mixture was centrifuged (2 min, 1677 × g rpm) and the supernatant was transferred to a vial prior to drying under vacuum. After dissecting guts and removing internal tissue, ant cuticles from the whole organisms were crushed in a mortar and washed two times successively with dichloromethane and methanol. The washed cuticle parts were dried at 50 °C for 3 h before further solid-state NMR experiments. For solution-state NMR, samples were composed of extracts dissolved in D2O and buffered at pH = 7.42 with 0.8 mM TSP-d4 as a chemical shift and concentration reference.
Solution-state NMR spectroscopy
1H 1D and 2D NMR spectra were acquired on a Bruker Avance III HD spectrometer at 700.13 MHz, equipped with a 1H/13C/15N/2H cryoprobe at a low temperature: 277 K to prevent any degradation of the metabolites. For all 1D and 2D experiments, a continuous wave presaturation on the residual H2O was applied during the recycling time. Quantitative 1H 1D NMR spectra were recorded with a recycling time d1 of 25 s (as five times the highest T1 1H), in quadriplicate with TSP-d4 as an internal reference. For the 2D DOSY experiment a stimulated echo with bipolar gradients including a longitudinal eddy current delay was chosen. A 6k × 56 matrix was acquired, zero-filled to 16k × 56 and apodized in F2 by a decreasing exponential with 2 HZ line broadening. A recycling time d1 = 5 s was used as well as a diffusion time d20 = 200 ms, with a gradient length of 900 and 200 ms of resting delay, a linear gradient ramp was used from 2 to 95% of the maximum gradient amplitude (50 G cm−1) and a number of accumulations NS = 200. An inverse Laplace transform was applied to obtain the 2D DOSY map displaying the diffusion coefficients along the indirect dimension. 2D COSY and ZQF-TOCSY spectra were also recorded. Further experimental parameters can be found in the figure captions. Tyr and Phe resonances were identified relying on the Biological Magnetic Resonance Bank database.
Solid-state NMR spectroscopy
15N solid-state NMR experiments were carried out on a Bruker Avance III spectrometer operating at 9.4 T (Larmor frequency of 40.5 MHz) using a 4 mm magic-angle spinning (MAS) double resonance probe head. Around 10 mg of cuticle per treatment (five individuals) was packed in a 50 µL HRMAS rotor to ensure a proper centering of the sample. 1H–15N cross-polarization (CP-MAS) experiments were performed at a MAS frequency of 14,286 Hz with a contact time of 1 ms (1H and 15N radiofrequency fields during CP of ca. 55 and 40 kHz, respectively), and with a recycling delay of 1 s. The experimental time was about 2–2.5 days for each spectrum. The 15N spectra were referenced using L-tyrosine (δ = 40.4 ppm). The 1H–15N–13C double CP-MAS experiment was carried out on a Bruker Advance Neo spectrometer operating at 14.1 T (13C Larmor frequency of 150.9 MHz) using a 3.2 mm Bruker CP-MAS CryoProbeTM32. The MAS frequency was 11 kHz, and the 1H–15N and 15N–13C contact times were 1.5 and 7 ms, respectively. The recycle delay was 3.5 s, and the 65,536 scans were accumulated.
Isotope-ratio mass spectrometry
Analysis was performed on a Thermo Delta V isotope-ratio mass spectrometer interfaced to a NC2500 elemental analyzer. In-house standards were calibrated against international reference materials provided by the International Atomic Energy Association. For the analytical sample run the overall standard deviation for the internal BSSL-100 standard (plant tissue) was 0.53‰ for δ15N and 0.05‰ for δ13C. Delta values obtained between the amplitudes of 100 and 10,000 mV for δ15N have an error associated with linearity of 0.65‰ and between 200 and 10,000 mV for δ13C error is 0.19‰. Isotope corrections are performed using a two-point normalization (linear regression) for all data using two additional internal standards (USGS40 and USGS41).
Electronic microscopy
The gaster of one individual for each treatment group (treated and untreated) was removed from the body and cut along the dorsal midpoint of the second abdominal segment. Each gaster was mounted on double-sided carbon sticker attached to aluminum stubs. Secondary electron images were recorded on a FEI Quanta 250 ESEM operating between 10 and 15 kV and with a working distance of 10 mm. Images for each individual were taken in the same area of the cuticle.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Sequence data and NMR spectroscopic data are available, respectively, in the NCBI SRA database under the Accession Number PRJNA683914 and in the Dryad Digital Repository (https://datad ryad.org/) under https://doi.org/10.5061/dryad.d7wm37q0h.
References
Hussa, E. & Goodrich-Blair, H. It takes a village: ecological and fitness impacts of multipartite mutualism. Annu. Rev. Microbiol. 67, 161–178 (2013).
Berasategui, A. et al. Gut microbiota of the pine weevil degrades conifer diterpenes and increases insect fitness. Mol. Ecol. 26, 4099–4110 (2017).
Engel, P. & Moran, N. A. The gut microbiota of insects—diversity in structure and function. FEMS Microbiol. Rev. 37, 699–735 (2013).
Henry, S. M. The significance of microorganisms in nutrition of insects. Trans. N. Y. Acad. Sci. 24, 676–673 (1962).
Brownlie, J. & Johnson, K. Symbiont-mediated protection in insect hosts. Trends Microbiol. 17, 348–354 (2009).
Florez, L., Biederman, P., Engl, T. & Kaltenpoth, M. Defensive symbioses of animals with prokaryotic and eukaryotic microorganisms. Nat. Prod. Rep. 32, 904–936 (2015).
Anbutsua, H. et al. Small genome symbiont underlies cuticle hardness in beetles. Proc. Natl Ac. Sci. USA 114, E8382–E8391 (2017).
Hirota, B. et al. A novel, extremely elongated, and endocellular bacterial symbiont supports cuticle formation of a grain pest beetle. mBio 8, e01482-17 (2017).
Moreau, C. S., Bell, C. D., Vila, R., Archibald, S. B. & Pierce, N. E. Phylogeny of the ants: diversification in the age of angiosperms. Science 312, 101–104 (2006).
Russell, J. A. et al. Bacterial gut symbionts are tightly linked with the evolution of herbivory in ants. Proc. Natl Acad. Sci. USA 106, 21236–21241 (2009).
Hölldobler, B. & Wilson, E. O. (eds) The Ants (Harvard University Press, 1990).
Nelsen, M. P., Ree, R. H. & Moreau, C. S. Ant-plant interactions evolved through increasing interdependence. Proc. Natl Acad. Sci. USA 115, 12253–12258 (2018).
Zientz, E., Beyaert, I., Gross, R. & Feldhaar, H. Relevance of the endosymbiosis of blochmannia floridanus and carpenter ants at different stages of the life cycle of the host. Appl. Environ. Microbiol. 72, 6027–6033 (2006).
Feldhaar, H. et al. Nutritional upgrading for omnivorous carpenter ants by the endosymbiont Blochmannia. BMC Biol. 5, 48 (2007).
Hu, Y. et al. Herbivorous turtle ants obtain essential nutrients from a highly conserved nitrogen-recycling gut microbiome. Nat. Commun. 9, 964 (2018).
Noh, M. Y., Muthukrishnan, S., Kramer, K. J. & Arakane, Y. Cuticle formation and pigmentation in beetles. Curr. Opin. Insect Sci. 17, 1–9 (2016).
Hopkins, T. L. & Kramer, K. J. Insect cuticle sclerotization. Annu. Rev. Entomol. 37, 273–302 (1992).
Andersen, S. O. Insect cuticular sclerotization: a review. Insect Biochem. Mol. Biol. 40, 166–178 (2010).
Vigneron, A. et al. Insects recycle endosymbionts when the benefit is over. Curr. Biol. 24, 2267–2273 (2014).
Engl, T. et al. Ancient symbiosis confers desiccation resistance to stored grain pest beetles. Mol. Ecol. 27, 2095–2108 (2018).
Moussian, B. “The arthropod cuticle”. in Arthropod Biology and Evolution (eds Minelli, A., Boxshall, G. & Fusco, G.) 171–196 (Springer, 2013).
Schaefer, J. et al. Aromatic cross-Links in Insect cuticle: detection by solid-state 13C and 15N NMR. Science 235, 1200–1204 (1987).
Merritt, M. E., Christensen, A. M., Kramer, J., Hopkins, T. L. & Schaefer, J. Detection of intercatechol cross-links in insect cuticle by solid-state carbon-13 and nitrogen-15 NMR. J. Am. Chem. Soc. 118, 11278–11282 (1996).
Zhu, K. Y., Merzendorfer, H., Zhang, W., Zhang, J. & Muthukrishnan, S. Biosynthesis, turnover, and functions of chitin in insetcs. Annu. Rev. Entomol. 61, 177–196 (2016).
Munowitz, M., Bachovchin, W. W., Herzfeld, J., Dobson, C. M. & Griffin, R. G. Acid-base and tautomeric equilibria in the solid state: 15N NMR spectroscopy of histidine and imidazole. J. Am. Chem. Soc. 104, 1192–1196 (1982).
Ash, E. L., Sudmeier, J. L., De Fabo, E. C. & Bachovchin, W. W. A low-barrier hydrogen bond in the catalytic triad of serine proteases? Theory versus experiment. Science 278, 1128–1132 (1997).
Hu, J. et al. Histidines, heart of the hydrogen ion channel from influenza A virus: toward an understanding of conductance and proton selectivity. Proc. Natl Acad. Sci. USA 103, 6865–6870 (2006).
Vásquez-Salazar, A., Becerra, A. & Lazcano, A. Evolutionary convergence in the biosyntheses of the imidazole moieties of histidine and purines. PLoS ONE 13, e0196349 (2018).
Chatterjee, S. et al. Demonstration of a common indole-based aromatic core in natural and synthetic eumelanins by solid-state NMR. Org. Biomol. Chem. 12, 6730–6736 (2014).
Della Vecchia, N. F. et al. Building-block diversity in polydopamine underpins a multifunctional eumelanin-type platform tunable through a quinone control point. Adv. Funct. Mater. 23, 1331–1340 (2013).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Hassan, A. et al. Sensitivity boosts by the CPMAS CryoProbe for challenging biological assemblies. J. Magn. Reson. 311, 106680 (2020).
Acknowledgements
We thank Grant Doering, Hanna Klaeser, Dreyvon McCray, Steven Medina, Shauna Price, Destiny Reeves, Andrea Thompson, and Mitchell Trychta for help with the feeding experiments. We thank Manuela Ramalho for laboratory and analysis assistance of the 16S rRNA qPCR and amplicon gene sequencing. We thank Jed Sparks director and Kim Sparks manager of the Cornell Stable Isotope Laboratory for running the isotope-ratio mass spectroscopy analysis. We acknowledge the Corsaire Metabolomics Core Facility (Biogenouest network) and warmly thank Sandrine Bouchet for her unfailing assistance. Ant photograph used with permission from the Field Museum of Natural History (FMNHINS 0000 062 654). We are grateful to Sebastien Vidal for reviewing chemical structures. This work received financial support from an “Investissement d’Avenir” grant managed by Agence Nationale de la Recherche (CEBA, ref. ANR-10-LABX-25-01), and C.D. acknowledges financial support from the CNRS Mission pour les initiatives transverses et interdisciplinaire (“Osez l’interdisciplinarité!” grant). C.S.M. thanks the US National Science Foundation for supporting this work (NSF DEB 1900357), US National Science Foundation REU program (NSF DBI 1559779).
Author information
Authors and Affiliations
Contributions
C.D. and C.S.M. conceived the study. C.S.M. and J.T.W. collected ant colonies, reared them in the lab, conducted the antibiotic and isotopic enrichment feeding experiments, performed gut and cuticle dissections, quantified gut microbial communities, and interpreted 16S rRNA sequence data. C.D. prepared the gut extracts and cuticle samples. E.M. prepared the NMR liquid samples. J.F. and E.M. performed liquid NMR analysis, and V.S-K., D.M., B.P. and A.H. performed the solid-state NMR analysis. Y.E. generated microscopic images. C.D., J.F., P.G. and V.S.-K. interpreted NMR data. C.D. wrote the first draft and all authors contributed to the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Tobias Engl, and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Duplais, C., Sarou-Kanian, V., Massiot, D. et al. Gut bacteria are essential for normal cuticle development in herbivorous turtle ants. Nat Commun 12, 676 (2021). https://doi.org/10.1038/s41467-021-21065-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-021-21065-y
This article is cited by
-
Untangling the complex interactions between turtle ants and their microbial partners
Animal Microbiome (2023)
-
Cuticle supplementation and nitrogen recycling by a dual bacterial symbiosis in a family of xylophagous beetles
The ISME Journal (2023)
-
Cuticular profiling of insecticide resistant Aedes aegypti
Scientific Reports (2023)
-
Physiological and evolutionary contexts of a new symbiotic species from the nitrogen-recycling gut community of turtle ants
The ISME Journal (2023)
-
Convergent evolution of a labile nutritional symbiosis in ants
The ISME Journal (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.