Abstract
Signal amplification in biological systems is achieved by cooperatively recruiting multiple copies of regulatory biomolecules. Nevertheless, the multiplexing capability of artificial fluorescent amplifiers is limited due to the size limit and lack of modularity. Here, we develop Cayley tree-like fractal DNA frameworks to topologically encode the fluorescence states for multiplexed detection of low-abundance targets. Taking advantage of the self-similar topology of Cayley tree, we use only 16 DNA strands to construct n-node (n = 53) structures of up to 5 megadalton. The high level of degeneracy allows encoding 36 colours with 7 nodes by site-specifically anchoring of distinct fluorophores onto a structure. The fractal topology minimises fluorescence crosstalk and allows quantitative decoding of quantized fluorescence states. We demonstrate a spectrum of rigid-yet-flexible super-multiplex structures for encoded fluorescence detection of single-molecule recognition events and multiplexed discrimination of living cells. Thus, the topological engineering approach enriches the toolbox for high-throughput cell imaging.
Similar content being viewed by others
Introduction
Recruitment of multiple copies of effector molecules represents a natural strategy for amplifying weak signals from ligand-receptor interactions in various signal pathways in vivo1,2. This principle has been harnessed in synthetic systems for developing signal amplifiers for biosensors, cellular imaging, and genetic engineering3,4,5. For example, by recruiting multiple copies of fused fluorescent proteins to the repeating protein-binding motifs (e.g., MS2) localised on the target RNA of interest, Singer and coworkers established a method for directly visualising single RNA molecules in living cells6,7. Similar strategies have been employed for the CRISPR/Cas9 imaging of low-copy and non-repeating chromatin loci8,9. The more recently developed SunTag10,11,12,13 technology utilises peptide scaffolds that can recruit up to 24 copies of effector proteins to drastically amplify fluorescent signals for chromatin and RNA translation imaging14,15,16,17. However, due to the lack of precise spatial arrangments of the signal molecules on the peptide scaffolds, there is either incomplete labelling or background from excessive signal molecules that limit the multiplexity of SunTags18. The multiplexity of fluorescence imaging is also affected by the interference between FRET pairs with overlapping spectra that generally has a practical limit of 4–5 colours19,20,21.
Structural DNA nanotechnology provides a powerful platform to construct self-assembled DNA nanostructures with near-atomic resolution and to programme the spatial organisation of molecules and nanoparticles with sub-10-nm resolution22,23,24,25,26,27,28,29,30. Specifically, the use of DNA frameworks as biocompatible scaffolds for programmable biomolecular assembly has been actively explored for various diagnostic, bioimaging and drug delivery applications5,31,32,33,34,35,36,37,38,39,40,41. However, the regular, monomeric DNA frameworks do not satisfy the requirements for developing super-multiplex fluorescent amplifiers. For example, small-sized tetrahedral DNA nanostructures (TDNs) of 5–20 nm can only accommodate a few molecules in a single structure35,39,42; whereas the complexity to construct large-sized DNA origamis or single-stranded-tile-based structures of >100 nm with hundreds or even thousands of distinct DNA strands makes them practically difficult to use43,44,45,46.
In this work, we develop Cayley tree-like fractal DNA frameworks (FDFs) to break the size limit with minimal use of DNA strands, and to improve the modularity for multiplexing. Cayley trees are acyclic graph structures with each node linked to a consistent number of neighbour nodes (i.e., coordination number). We show that the use of TDNs as nodes with 4-coordination generates rigid-yet-flexible FDFs with molecular weights up to ~5 megadalton. By exploiting the self-similar topology (node connection arrangement), we construct FDF amplifiers with only a small set of 16 DNA strands which can encode up to 36 distinct colours with minimal crosstalk. These FDFs can topologically encode quantitative fluorescence states for single-molecule recognition and multiplexed cell discrimination.
Results
Construction of fractal DNA frameworks
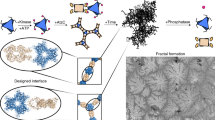
In our design, a Cayley tree-like FDF (denoted as Fn,i, where n is the coordination number of each node, and i is the iteration number of the structure, shown in Fig. 1a) is assembled with tetrahedral DNA nodes (TDNs). Each TDN (edge length 20 bp, or ~6.8 nm, adapted from previous studies35,42, see Methods and Supplementary Table 1) is assembled from four oligonucleotides, and carries n (n = 2, 3 or 4) single-stranded overhangs for inter-node recognition (denoted as TDN-II, TDN-III or TDN-IV, respectively, shown in Supplementary Fig. 1). The root TDN (shell-0) with n overhangs can recognise and assemble with n TDNs in the exterior shell (shell-1) through hybridisation of the overhangs to form an Fn,1 structure containing n + 1 nodes. The remaining overhangs of the shell-1 nodes in turn recruit additional nodes to form Fn,2 and so forth (Supplementary Table 2). Using this shell-by-shell assembly approach (in a sequence of shell-0 + shell-1 + shell-2 + shell-3), we synthesised FDFs with 1–3 shells (including F2,1, F2,2, F2,3, F3,1, F3,2, F3,3, F4,1, F4,2 and F4,3, respectively). Alternatively, fractal assembly was used for the synthesis of 3-shell trees (F2,3, F3,3 and F4,3, shown in Fig. 1a): shell-0 nodes were first assembled with shell-1 nodes (shell-0 + shell-1) to form F2,1, F3,1 and F4,1 (same to the shell-by-shell approach); in parallel, shell-2 nodes were coupled with shell-3 nodes elsewhere to form partial assemblies (shell-2 + shell-3), which were then assembled with the Fn,1 structures to form whole 3-shell trees ((shell-0 + shell-1) + (shell-2 + shell-3)).
Figure 1b shows representative atomic force microscopic (AFM) images of the synthesised FDFs in our study. For a perfect Cayley tree-like FDF Fn,i, the number of nodes in the kth shell (k > 1) is given by Nk = n(n−1)k−1, thus the total node number of the FDF is given by Eq. (1)
We found that the resulting structures under AFM (Fig. 1b and Supplementary Fig. 2) contained expected numbers of TDNs. F2,1, F2,2 and F2,3 contained 3, 5 or 7 TDNs; F3,1, F3,2 and F3,3 contained 4, 10 and 22 TDNs; F4,1, F4,2 and F4,3 contained 5, 17 and 53 TDNs, respectively. These results confirm the formation of FDFs with precise node number defined by the coordination number and iteration number of the TDNs. Given that the molecular weight (MW) of a TDN is ~90 kilodalton, the FDF with 53 TDNs (F4,3) possesses a MW of ~5 megadalton (Fig. 1c). It’s worth noting that this supersized structure is composed of only 16 distinct DNA strands, much less compared to a DNA origami structure of similar size (typically requires hundreds of DNA strands). The gel electrophoresis images and chromatograms (Supplementary Fig. 3) present discrete bands or peaks corresponding to FDFs with different shell numbers, which allows unambiguous discrimination of them via common gel electrophoresis or high-performance liquid chromatography (HPLC). Moreover, we found that for multi-shell FDFs, e.g., F3,3, the fractal assembly strategy led to a higher yield (84%) of end product in the final step, compared to the shell-by-shell strategy (45%, see Supplementary Fig. 4). The self-similarity of FDFs facilitates the fabrication of supersized DNA nanomaterials with minimal use of DNA strands.
Structural properties of FDFs
We next investigated the structural rigidity of FDF structures. We observed that among different FDFs, the TDNs under AFM exhibit a uniform size with apparent lateral length of ~12 nm and height of ~2.6 nm, which are consistent with that of monomer TDNs. Hence, TDNs remain intact and structurally rigid in FDFs. However, we found that the F2,3 structures generallly adopted curved conformations, resulting in shorter lateral lengths than the corresponding contour lengths (Fig. 2a). Histogram analysis shows the broad inter-node angle distribution of F2,n ranging from 50° to 180° (Fig. 2b), suggesting these chain-like F2,3 structures have a high degree of flexibility, albeit with rigid nodes. This feature allows the construction of rigid-yet-flexible DNA materials.
a Schematic structures of F2,i (i = 0-3) with representative atomic force microscopic (AFM) images and line profiles across both terminal tetrahedral DNA nanostructures (TDNs) (marked with blue cross) for F2,i. The full width of half maximum height (FWHM) are measured from the line profiles. b Histogram statistics of the inter-node angles in F2,i (N = 360). c Schematic structures of F4,3 and their stochastic optical reconstruction microscopic (STORM) images (large view and individual speckles magnified from the upper image). Red speckle, Gaussian speckle (surrounded by blue dashed circle) resulted from Alexa647 labelled on the outer-shell TDNs. Yellow cross, fluorescence location of the root TDN (Cy3). Blue dot, geometric centre of the circle. d Schematic and representative AFM images of F4,1 and F2,2. e Diameter distribution of F4,1 and F2,2 from theoretical simulations (N = 1000) and AFM measurements (N = 100). Curves are Gaussian fittings of the histograms.
We also characterised the flexibility of a dendrimer-like FDF (F4,3) in solution using stochastic optical reconstruction microscopy (STORM, detailed in Supplementary Methods). The outer-shell (shell-3) nodes of the F4,3 were labelled with Alexa647 (A647) while the root node was labelled with Cy3. As shown in Fig. 2c, we observed flower-like Guassian speckles of A647 (reconstructed from 30,000 frames of imaging), suggesting the quasi-spherical morphology of the F4,3 structures in solution. We also found that the fluorescence locations of the root nodes in some extent deviate (79 ± 36 nm, Supplementary Fig. 5) from the geometric centres of the A647 speckles, suggesting the flexible feature of the F4,3 structure.
We then asked whether the structural rigidity of FDF structures can be tuned via topological engineering (i.e. engineering their internode connection arrangements). To answer this question, we compared the structure of a chain-like FDF (F2,2) and a 4-branched FDF (F4,1). They have identical node number (five nodes) but different topological arrangements (i.e., two-way connections vs. four-way connections, shown in Fig. 2d). In F2,2, the two terminal nodes are separated by four flexible linkages and three nodes; while in F4,1, any two terminal nodes are across two linkages and one node. Thus, they should have different mechanical properties. Indeed, we found that under AFM (Fig. 2d, e and Supplementary Figs. 6 and 7), the F2,2 structures present larger size and larger size deviation (83 ± 11 nm in diameter) compared to the F4,1 structures (59 ± 5 nm), in agreement with the molecular dynamic simulations (F2,2, 80 ± 8 nm, F4,1, 59 ± 7 nm, see Methods). These results indicate that the F4,1 structure is relatively compact and rigid while the F2,2 structure is loose and flexible. This difference can be attributed to their different topologies for organising TDNs.
FDFs encode quantised fluorescence states
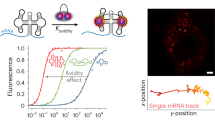
Taking advantage of the rigid-yet-flexible FDFs, we further test whether fluorophores can be precisely organised on these scaffolds to generate quantised fluorescence with minimised crosstalk (Fig. 3a). The location of each TDN node in the FDFs was measured. The lateral distance between two adjacent TDNs is 18.5 ± 3.8 nm under AFM (Fig. 3b), which is consistent with the result obtained by super-resolution stochastic optical reconstruction microscopy (STORM) at the single-molecule level (Supplementary Fig. 8), showing that the distance between two fluorophores individually anchored on two terminal TDNs of an F2,1 was ~35 nm, suggesting that two connected TDNs should be at least ~17 nm apart. Meanwhile, any two non-connected yet proximal TDNs in a folded FDF structure or in different structures were also apart for over 15 nm (Supplementary Fig. 9) Moreover, we synthesised an F2,3 structure carrying three Cy3 and four Cy5 (one fluorophore per node) for a Förster resonance energy transfer (FRET) analysis. Although the Cy3-Cy5 pair typically exhibit energy transfer when brought close to one another, we found that its fluorescence spectra were similar to that of a simple mixture of [TDN-Cy5]3 and [TDN-Cy3]4 (Supplementary Fig. 10), indicating that the distance between Cy3–Cy5 pair is beyond the effective FRET distance (>10 nm). In contrast, if the Cy3 and Cy5 were placed on the same TDN (with a distance of ~4 nm, Supplementary Fig. 10), a significant Cy3–Cy5 FRET effect was observed (quenching of Cy3 along with excitation of Cy5), in agreement with previous studies35,47.
a Schematic illustration of the FDF structure with spatial control of fluorophores. b Representative AFM image of a two-way FDF (F2,3) structure, and statistical distribution of the internode distance of the F2,n structures (N = 200). TDNs are marked with dashed circles. Measured distances between nodes (from centre to centre) are marked with numbers. c Correlation between fluorescence intensity and the copy number of A488 on the two-way FDF. Embedded images are fluorescence images of droplets containing corresponding FDFs in pseudocolour. d Correlation between fluorescence intensity and the copy number of ROX and Cy5 anchored on the F2,3 structures. Embedded images are fluorescence images of droplets containing corresponding FDFs in pseudocolour. Error bars indicate standard deviations from three independent tests. e 36 FDF barcodes encoded with different fluorophore combinations, which can be decoded by fluorescence spectroscopy/microscopy. The three-digit IDs refer to the numbers of Cy5, ROX and A488 in a single FDF structure (e.g., “124” refers to the presence of 1 Cy5, 2 ROX and 4 A488).
Next, we prescribed four anchoring sites on each TDN node in the F2,3 structure, which can accommodate up to 28 fluorophores. By anchoring 4–28 molecules of Alexa488 (A488, max emission wavelength = 517 nm) on F2,3, we synthesised a series of FDFs with amplified fluorescence. Remarkably, the measured fluorescence intensity of the tree increases nearly linearly with the copy number of A488 (R2 > 0.99) (Fig. 3c). In particular, F2,3 with a copy number of 28 exibits ~24-fold increase in brightness as compared to a single A448 anchored on a monomeric TDN (Fig. 3c and Supplementary Fig. 11), suggesting that the ~4-nm interspace in a TDN is adequate to avoid self-quenching of A488 fluorophores, making FDF a quantitative amplifier for high-brightness fluorescence imaging. We further evaluated the fluorescence crosstalk among differently-coloured fluorophores in an FDF structure. Three types of fluorophores with distinct emission wavelengths (Cy5, 668 nm; ROX, 606 nm; A488, 517 nm, see Supplementary Fig. 12) were anchored onto FDFs with up to seven nodes, one fluorophore per node. Regardless of the presence of other types of fluorophores, the fluorescence intensity in each colour channel is proportional to the number of the corresponding fluorophores (R2 > 0.99, Fig. 3d and Supplementary Fig. 13). Taken together, FDFs enable tailorable spatial separation of fluorophores. For probes carrying heterogenous fluorophores with a larger Förster radius (i.e. the inter-fluorophore distance with 50% efficiency of FRET)48, the sparse fluorophore arrangement (one fluorophore per node) maintains an inter-fluorophore distance of >10 nm, thus minimising FRET quenching to ensure the linearity of fluorescence. For probes carrying a single species of fluorophores with a smaller Förster radius (typically < 4 nm), the dense arrangement (up to four fluorophores per node) can be used to realise high brightness.
According to Cayley’s formula in graph theory49, the FDFs have high data capacity resulting from the diversity of node combinations. To demonstrate this capacity, a set of 36 FDFs (F2,3) with all possible combination (C31+C32‧C61+C31‧C51) of Cy5/ROX/A488 fluorophores were constructed (Fig. 3e). Each FDF is given a three-digit colour ID corresponding to the copy numbers of Cy5, ROX and A488, respectively (see characteristic fluorescence spectra in Supplementary Fig. 14). The confocal fluorescence images (Fig. 3e and Supplementary Fig. 15) show a palette comprised of 36 pseudocolours. Likewise, we constructed a series of F3,1 structures encoding 15 colours, and F4,1 structures encoding 21 colours, respectively (Supplementary Figs. 14 and 15). These colours are beyond the conventional multiplexing limit of fluorescence imaging (typically 4–5 colours).
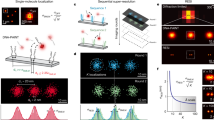
We next applied the FDF-encoded barcodes as probes to single-molecule recognition and imaging. The key to this goal is to decode the fluorescence signal of a single barcode molecule, i.e. determine the number and type of fluorophores on a barcode of a specific colour. Given the fact that the fluorescence states of a barcode are quantised, the fluorescence under high-power excitation should present a stepwise photobleaching trace, as illustrated in Fig. 4a. A barcode can be decoded by the number of steps in the photobleaching trace, which correspond to the copy number of fluorophores50,51,52.
a Schematic illustration of the quantised photobleaching of different FDF barcodes (ID: 140 and 410). b Fluorescent microscopic images of the FDF barcodes. Scale bar, 5 µm. c Single-molecule fluorescence quenching snapshots, kinetics of the FDF barcodes, and distributions of quenching step counts (N = 100 for each barcode).
To validate this decoding strategy, we studied the photobleaching traces of two FDF barcodes (barcode ID: 140 and 410, both with Cy5 and ROX but in different ratios, as shown in Fig. 4b) with total internal reflection fluorescence microscopy (TIRF). In the initial state, spots with different fluorescence ratios from both Cy5 channel and ROX channel can be monitored (Fig. 4b). These fluorescent spots confirmed the location of the barcode molecules. Then, selected spots were illuminated by the 647-nm laser and the fluorescence kinetics was monitored within 70 s in the Cy5 channel (Fig. 4c, N = 100 for each type of barcodes). For barcode 140, descending fluorescence traces (78%) with one step were observed corresponding to one Cy5 fluorophore. While for barcode 410, we observed in majority (59%) four-step photobleaching traces, indicating that there were four Cy5 fluorophores present. Next, we used the 561-nm laser for excitation and photobleaching of ROX (Fig. 4c). Barcode 140 (54%) presented four-step fluorescence quenching traces while barcode 410 (84%) presented one-step traces, in accordance with their prescribed ROX numbers. These results confirmed that the quantised FDF barcodes can be decoded at the single-molecule level via photobleaching and single-particle fluorescence tracking. Although errors may occur in fluorophore counting, we estimated that over 80% barcodes can be correctly decoded (Supplementary Fig. 16), and the decoding errors might be further suppressed by choosing subsets of barcodes with high fluorescence intensity ratios between different colours.
For a single-molecule recognition experiment, an FDF probe (barcode ID: 131) was constructed to respond to specific DNA targets (Supplementary Table 1). As illustrated in Fig. 5a, two probe sequences (complementary to two DNA targets) were separately placed in two linker regions (one between ROX-TDN and Cy5-TDNs, the other between A488-TDN and ROX-TDN) of the FDF structure. The presence of these specific targets could thus trigger toehold-mediated strand-displacement reactions and break these linkers, resulting in “quantised” release of certain TDN nodes (Supplementary Fig. 17). We tracked the single-molecule fluorescence kinetics in response to the target inputs (Fig. 5b–d). Without any input, the counts of fluorescence photobleaching steps in all three channels matched the corresponding fluorophore numbers of the probe (88%, 73% and 92% of the FDFs presented one-, three- and one-step quenching traces for Cy5, ROX and A488, respectively. N = 100 each). After the input of target 1, we observed that the fluorescence of Cy5 disappeared, while the stepped photobleaching kinetics of A488 and ROX remained, indicating that the target 1 solely induced the specific release of Cy5-nodes. After the input of target 2, we observed the disappearance of both Cy5 and ROX fluorescence, leaving only the A488 showing a one-step photobleaching kinetics, indicating that the presence of target 2 triggered the release of ROX-nodes. In contrast, when we challenged the probe with target 2 first, the fluorescence signals of Cy5 and ROX were lost (showing no quenching steps), leaving mostly a one-step quenching trace of A488, indicating the breaking of the linkage between A488-TDN and ROX-TDN. As expected, this result remained unchanged with the subsequent input of target 1 (Supplementary Figs. 18 and 19). Collectively, using the quantised FDFs, we demonstrate a multi-state molecule recognition strategy that can translate target-induced probe cleavage into readable quantised fluorescence state changes at the single-molecule level in a sequence-sensitive manner. This strategy provides a means to develop molecular probes that can identify targets or multiple target states.
FDF barcodes for cell imaging and discrimination
Having established the FDF barcodes, we further sought to implement multiplexed cell imaging and discrimination. We found that the F4,1 structure could maintain structural stability within 12 h in serum culture and showed minimal cytotoxicity (Supplementary Figs. 20 and 21), which supports the use of FDF barcodes in live-cell imaging. We built four F4,1-based FDF barcodes (colour IDs: 500, 401, 104 and 005) carrying overhangs of distinct sequences (Fig. 6a). Four groups of HeLa cells were separately tagged with cholesterol-modified ssDNAs (chol-DNAs, which can be embedded in cell membranes and allow DNA hybridisation) orthogonally complementary to the overhangs of the FDF barcodes. Then the cells were mixed up and incubated with the mixture of the four FDF barcodes. The flow cytometric result (Fig. 6b) shows that the cell mixture could be sorted into four separated populations with different A647/A488 fluorescence intensity ratios, which characteristically correspond to the four FDF barcodes, and are in agreement with the results from separately tagged cells (Supplementary Fig. 22). In the wide-view fluorescence image (Fig. 6c), the cells can also be classified into four groups by their fluorescence intensity profiles (Fig. 6d), matching the four FDF barcodes. Taken together, these results suggest that the multiplexity of FDF barcodes allows discrimination of multiple cell populations in a cell mixture.
a Schematic illustration of FDF barcodes for discrimination of cell populations. b Cytometric scatter plot and representative snapshots of FDF-tagged cells during cell sorting. c Wide-view fluorescence image of FDF-tagged cells and d representative fluorescence intensity profiles (along the dashed lines in the cell images). Scale bar, 25 μm.
Discussion
In this study, we report a topological engineering approach to construct multiplexed FDF amplifiers. The Cayley tree-like fractal assembly features simplicity, modularity and high capacity for fluorescence encoding and decoding, which compares favourably with previously reported DNA assembly methods that either require high number of DNA strands or lack well-defined structures23,44,53,54,55,56. Luo’s group have developed a series of dendrimer-like DNA structures based on Y-DNA, and demonstrated their use in fluorescence encoding34,54,57. But their structural compactness, topological engineerability and cargo/information capacity are still limited. Here, we employed tetrahedral DNA nanostructures as rigid nodes for the Cayley tree framework, which have defined coordination number (2–4), higher rigidity and molecular weights, leading to FDF structures with fixed numbers of nodes and guest molecules. By using only a small set of oligonucleotides (as few as 16 distinct sequences), we can construct FDFs with up to 53 TDN nodes and ~5-megadalton in molecular weight.
We show that the structural flexibility of FDFs can be topologically engineered, which allows modular design of DNA nanomaterials with tuneable mechanical properties. Moreover, these nodes have large information capacity with minimal cross-talk. By individually anchoring fluorophores with different emitting wavelength on each TDN node, we synthesised colour-encoded multiplexed probes for identification of different molecules and live cells. Hence, FDFs provide a highly generic amplification toolset for multiplexed and quantitative detection of low-abundance biological targets.
Methods
Preparation and purification of TDN nodes and the FDFs
To synthesise TDN nodes, four component oligonucleotides (S1, S2, S3 and S4, synthesised by Invitrogen, USA, sequences were listed in Supplementary Table 1) of the TDN structure were mixed in equimolar concentration (1 µM each) in TM buffer (10 mM Tris-HCl, 5 mM MgCl2, pH 8.0). The mixture was heated to 95 °C for 10 min, then rapidly cooled to 4 °C in a thermal cycler (Applied Biosystems Veriti 96 well Thermal Cycler). HPLC purification and characterisation were carried out using an Agilent 1260 system. An SEC column (Phenomenex BioSec-SEC-S4000, 300 × 7.8 mm) was used to purify the TDN nodes, and the chromatograms were recorded at 260 nm. The mobile phase was 25 mM Tris-HCl, pH 7.2, 450 mM NaCl with a flow rate of 1 ml min−1. To fabricate FDFs, the purified TDN nodes with complementary linking strands were mixed in equimolar concentration (0.5 µM each) and incubated at 25 °C for 2 h. The products were again purified with the SEC column.
AFM imaging
AFM images were taken with a Bruker multimode 8 atomic force microscope (Veeco Inc., USA). Freshly cleaved mica was treated with a 0.5% (v/v) APTES (Sigma-Aldrich) for 2 min, washed with Milli-Q water (18 MΩ cm), and dried with compressed air prior to nanostructures deposition. In all, 10 nM of the the sample in 30 µl TM buffer was deposited on the mica surface and let sit 5 min for adsorption. The samples were imaged in tapping mode in solution using SNL-10 tips.
Fluorescence spectroscopic measurements
All fluorescence spectroscopic measurements were carried out using an Edinburgh FS920 fluorescence spectrometer operating in photon counting mode. The fluorescence emission spectra of Alexa488 labelled structures were collected from 495 to 590 nm with excitation at 488 nm. The fluorescence emission of ROX was collected from 595 to 640 nm with excitation at 588 nm. The fluorescence of Cy5 was collected from 655 to 720 nm by with excitation at 650 nm. All spectra were collected at 25 °C.
Confocal microscopic imaging
For multi-colour FDF fluorescence imaging, solutions of FDFs were dropped onto the slides to form 6 × 6 Dot Matrix shapes using a SpotBot 2 microarrayer programmed by SpotApp software (ArrayIt SpotBot 3). Then, confocal microscopic imaging of these spots was carried out on a laser confocal microscope (Leica TCS SP8) at 10× magnification. For Cy5, exitation (ex) 647 nm, emission (em) 650–720 nm; For ROX, ex 561 nm and em 575–620 nm. For A488, ex 488 nm, em 495-550 nm.
Single molecule imaging
FDF barcodes (ID:140 or 410, 20 pM) were adsorbed on clean glass coverslips respectively for 10 min. Total internal reflection fluorescence (TIRF) imaging was performed on a commercial super-resolution microscope (N-STORM, Nikon) with a 100× objective lens (NA 1.49) and an electron multiplying charge-coupled device (EMCCD) camera (Andor, iXon 3). The FDF barcodes were first excited with 647-nm solid state laser for 70 s. The second round of excitation was carried out by using 561-nm laser for 70 s. The fluorescence intensity kinetics were analysed from the fluorescence spots across the entire movie through Nikon N-STORM analysis software.
For single-molecule recognition, a clean glass coverslip was modified with the mixture of biotin-PEG and PEG as described previously58. Streptavidin was then adsorbed on the coverslip via biotin-avidin interaction. Next, the biotinylated FDF single-molecule probe (ID: 131, 20 pM) was immobilised on the coverslip. Then, a target strand was added into the solution (final concentration, 100 pM). After an incubation at 25 °C for 30 min, the sample was washed with 1× PBS three times. The target-triggered strand-displacement reaction was verified using gel electrophoresis (Supplementary Fig. 16). 647-nm, 561-nm and 488-nm solid state laser (Coherent) were used to excite Cy5, ROX and A488 (70 s each), successively. The fluorescence intensity kinetics of the fluorescent spots were recorded.
Cell imaging and discrimination
For cell discrimination, four F4,1-based barcodes loaded with different ratios of A647 and A488 (colour ID: 500, 401, 104 and 005) were built. Each of them carried a distinct overhang sequence (sequences listed in Supplementary Table 1). For cell labelling, four groups of HeLa cells (~6 × 104 cells each group) were separately incubated (for 0.5 h at 37 °C) with cholesterol-modified ssDNAs (chol-DNAs, sequences listed in Supplementary Table 1) orthogonally complementary to the barcodes. The cells were washed with PBS solution for three times to remove the free chol-DNAs. Next, the four groups of cells were trypsinized and mixed up in serum-free culture medium and incubated with the mixture of the FDF barcodes (chol-DNAs) for 1 h at 37 °C. These cells were washed three times with PBS and centrifugation at 1000 rpm, and resuspended in 100 μL of PBS. The cell mixture was underwent flow cytometry analysis using a Merk Millipore amins Image streamX Mark II system (A647 channel, Ex 642 nm and Em 642–745 nm. A488 channel, Ex 488 nm and Em 505–560 nm). Confocal imaging of these cells was carried out on a Leica TCS SP8 confocal laser scanning microscope. The cells nuclei were stained with Hoechst33258 for 10 min prior to imaging. For A647 channel, Ex 633 nm and Em 650–720 nm; A488 channel, Ex 488 nm and Em 495-550 nm; Hoechst channel, Ex 405 nm and Em 420–470 nm. The fluorescence profiles were extracted and analysed using ImageJ.
Data availability
The data that support the findings of this study are available within the paper and its Supplementary Information and are available from the corresponding authors upon reasonable request.
References
Agalioti, T. et al. Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 103, 667–678 (2000).
Croce, L. D. et al. Methyltransferase recruitment and DNA hypermethylation of target promoters by an oncogenic transcription factor. Science 295, 1079–1082 (2002).
Wang, J. Self-propelled affinity biosensors: moving the receptor around the sample. Biosens. Bioelectron. 76, 234–242 (2016).
Rosenbloom, A. B. et al. Optimized two-color super resolution imaging of Drp1 during mitochondrial fission with a slow-switching Dronpa variant. Proc. Natl Acad. Sci. USA 111, 13093–13098 (2014).
He, R. et al. Facile synthesis of pentacle gold-copper alloy nanocrystals and their plasmonic and catalytic properties. Nat. Commun. 5, 4327 (2014).
Hocine, S., Raymond, P., Zenklusen, D., Chao, J. A. & Singer, R. H. Single-molecule analysis of gene expression using two-color RNA labeling in live yeast. Nat. Methods 10, 119–121 (2013).
Halstead, J. M. et al. An RNA biosensor for imaging the first round of translation from single cells to living animals. Science 347, 1367–1671 (2015).
Qin, P. et al. Live cell imaging of low- and non-repetitive chromosome loci using CRISPR-Cas9. Nat. Commun. 8, 14725 (2017).
Ma, H. et al. Multiplexed labeling of genomic loci with dCas9 and engineered sgRNAs using CRISPRainbow. Nat. Biotechnol. 34, 528–530 (2016).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Pichon, X. et al. Visualization of single endogenous polysomes reveals the dynamics of translation in live human cells. J. Cell Biol. 214, 769–781 (2016).
Thakore, P. I., Black, J. B., Hilton, I. B. & Gersbach, C. A. Editing the epigenome: technologies for programmable transcription and epigenetic modulation. Nat. Methods 13, 127–137 (2016).
Morita, S. et al. Targeted DNA demethylation in vivo using dCas9-peptide repeat and scFv-TET1 catalytic domain fusions. Nat. Biotechnol. 34, 1060–1065 (2016).
Wu, B., Eliscovich, C., Yoon, Y. J. & Singer, R. H. Translation dynamics of single mRNAs in live cells and neurons. Science 352, 1430–1435 (2016).
Wang, C., Han, B., Zhou, R. & Zhuang, X. Real-time imaging of translation on single mRNA transcripts in live cells. Cell 165, 990–1001 (2016).
Morisaki, T. et al. Real-time quantification of single RNA translation dynamics in living cells. Science 352, 1425–1429 (2016).
Yan, X., Hoek, T. A., Vale, R. D. & Tanenbaum, M. E. Dynamics of translation of single mRNA molecules in vivo. Cell 165, 976–989 (2016).
Kamiyama, D. et al. Versatile protein tagging in cells with split fluorescent protein. Nat. Commun. 7, 11046 (2016).
Hu, F. et al. Supermultiplexed optical imaging and barcoding with engineered polyynes. Nat. Methods 15, 194–200 (2018).
Wei, L. et al. Super-multiplex vibrational imaging. Nature 544, 465–470 (2017).
Hong, S. L. et al. Live-cell stimulated raman scattering imaging of alkyne-tagged biomolecules. Angew. Chem. Int. Ed. 53, 5827–5831 (2014).
Funke, J. J. & Dietz, H. Placing molecules with Bohr radius resolution using DNA origami. Nat. Nanotechnol. 11, 47–52 (2015).
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068 (2017).
Jones, M. R., Seeman, N. C. & Mirkin, C. A. Programmable materials and the nature of the DNA bond. Science 347, 1260901 (2015).
Hao, Y., Labean, T. H., Feng, L. & Reif, J. H. Directed nucleation assembly of DNA tile complexes for barcode-patterned lattices. Proc. Natl Acad. Sci. USA 100, 8103–8108 (2003).
Lin, C. et al. Submicrometre geometrically encoded fluorescent barcodes self-assembled from DNA. Nat. Chem. 4, 832–839 (2012).
Li, J., Green, A. A., Yan, H. & Fan, C. Engineering nucleic acid structures for programmable molecular circuitry and intracellular biocomputation. Nat. Chem. 9, 1056–1067 (2017).
Li, Y., Liu, Z., Yu, G., Jiang, W. & Mao, C. Self-assembly of molecule-like nanoparticle clusters directed by DNA nanocages. J. Am. Chem. Soc. 137, 4320–4323 (2015).
Tan, L. H., Xing, H. & Lu, Y. DNA as a powerful tool for morphology control, spatial positioning, and dynamic assembly of nanoparticles. Acc. Chem. Res. 47, 1881–1890 (2014).
Nickels, P. C. et al. Molecular force spectroscopy with a DNA origami-based nanoscopic force clamp. Science 354, 305–307 (2016).
Lubbe, A. S., Szymanski, W. & Feringa, B. L. Recent developments in reversible photoregulation of oligonucleotide structure and function. Chem. Soc. Rev. 46, 1052–1079 (2017).
Wang, J. S., Yan, Y. H. & Zhang, D. Y. Modular probes for enriching and detecting complex nucleic acid sequences. Nat. Chem. 9, 1222–1228 (2017).
Sun, W. J. et al. Cocoon-like self-degradable DNA nanoclew for anticancer drug delivery. J. Am. Chem. Soc. 136, 14722–14725 (2014).
Li, Y., Cu, Y. T. H. & Luo, D. Multiplexed detection of pathogen DNA with DNA-based fluorescence nanobarcodes. Nat. Biotechnol. 23, 885–889 (2005).
Li, J. et al. Self-assembled multivalent DNA nanostructures for noninvasive intracellular delivery of immunostimulatory CpG oligonucleotides. ACS Nano 5, 8783–8789 (2011).
Chen, Y. J., Groves, B., Muscat, R. A. & Seelig, G. DNA nanotechnology from the test tube to the cell. Nat. Nanotechnol. 10, 748–760 (2015).
Veetil, A. T. et al. Cell-targetable DNA nanocapsules for spatiotemporal release of caged bioactive small molecules. Nat. Nanotechnol. 12, 1183–1189 (2017).
He, L. et al. Fluorescence resonance energy transfer-based DNA tetrahedron nanotweezer for highly reliable detection of tumor-related mRNA in living cells. ACS Nano 11, 4060–4066 (2017).
Liu, X. et al. A DNA nanostructure platform for directed assembly of synthetic vaccines. Nano Lett. 12, 4254–4259 (2012).
Lee, H. et al. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nat. Nanotechnol. 7, 389–393 (2012).
Woehrstein, J. B. et al. Sub-100-nm metafluorophores with digitally tunable optical properties self-assembled from DNA. Sci. Adv. 3, e1602128 (2017).
Goodman, R. P. et al. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science 310, 1661–1665 (2005).
Tikhomirov, G., Petersen, P. & Qian, L. Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns. Nature 552, 67–71 (2017).
Ong, L. L. et al. Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components. Nature 552, 72–77 (2017).
Wagenbauer, K. F., Sigl, C. & Dietz, H. Gigadalton-scale shape-programmable DNA assemblies. Nature 552, 78–83 (2017).
Fox, M. E., Szoka, F. C. & Frechet, J. M. J. Soluble polymer carriers for the treatment of cancer: the importance of molecular architecture. Acc. Chem. Res. 42, 1141–1151 (2009).
Walsh, A. S., Yin, H., Erben, C. M., Wood, M. J. & Turberfield, A. J. DNA cage delivery to mammalian cells. ACS Nano 5, 5427–5432 (2011).
Roy, R., Hohng, S. & Ha, T. A practical guide to single-molecule FRET. Nat. Methods 5, 507–516 (2008).
Bondy, J. A. & Murty, U. S. R. Graph Theory With Applications (Macmillan, London, 1976).
Zhang, H. L. et al. Programming chain-growth copolymerization of DNA hairpin tiles for in-vitro hierarchical supramolecular organization. Nat. Commun. 10, 1006 (2019).
Hariri, A. A., Hamblin, G. D., Gidi, Y., Sleiman, H. F. & Cosa, G. Stepwise growth of surface-grafted DNA nanotubes visualized at the single-molecule level. Nat. Chem. 7, 295–300 (2015).
Ulbrich, M. H. & Isacoff, E. Y. Subunit counting in membrane-bound proteins. Nat. Methods 4, 319–321 (2007).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Um, S. H. et al. Enzyme-catalysed assembly of DNA hydrogel. Nat. Mater. 5, 797–801 (2006).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–626 (2012).
Jochum, C. et al. Structure and stimuli-responsiveness of all-DNA dendrimers: theory and experiment. Nanoscale 11, 1604–1617 (2019).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and notch signaling. Science 340, 991–994 (2013).
Acknowledgements
This work was financially supported by the the National Key R&D Program of China (2016YFA0400900, 2016YFA0902600), the National Natural Science Foundation of China (21991134, 21834007, 21675167, 31571014, 21775157, 11575278 and 21505148), the National Science Foundation (1607832), Shanghai Municipal Science and Technology Commission (19JC1410302), the Open Large Infrastructure Research of Chinese Academy of Sciences, and the LU JIAXI International team program supported. C.F. acknowledges the K.C. Wong Education Foundation (SJTU). H.Y. acknowledges the Presidential Strategic Initiative Fund from Arizona
Author information
Authors and Affiliations
Contributions
C.F., L.W. and H.Y. directed the research. J.L., S.J., L.W., H.Y. and C.F. conceived and designed the experiments. J.D., M.X., T.Z., S.X., Y.Z, L.L., F.W., S.C. and Y.Y. carried out experiments and analysed data. Z.Q. performed numeric calculations. J.L., S.J., H.Y. and C.F. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Gaetan Bellot, Nico Stuurman and Yonggang Ke for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, J., Dai, J., Jiang, S. et al. Encoding quantized fluorescence states with fractal DNA frameworks. Nat Commun 11, 2185 (2020). https://doi.org/10.1038/s41467-020-16112-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-16112-z
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.