Abstract
Isogenic populations of cells exhibit phenotypic variability that has specific physiological consequences. Individual bacteria within a population can differ in antibiotic tolerance, but whether this variability can be regulated or is generally an unavoidable consequence of stochastic fluctuations is unclear. Here we report that a gene encoding a bacterial (p)ppGpp synthetase in Bacillus subtilis, sasA, exhibits high levels of extrinsic noise in expression. We find that sasA is regulated by multisite phosphorylation of the transcription factor WalR, mediated by a Ser/Thr kinase-phosphatase pair PrkC/PrpC, and a Histidine kinase WalK of a two-component system. This regulatory intersection is crucial for controlling the appearance of outliers; rare cells with unusually high levels of sasA expression, having increased antibiotic tolerance. We create a predictive model demonstrating that the probability of a given cell surviving antibiotic treatment increases with sasA expression. Therefore, multisite phosphorylation can be used to strongly regulate variability in antibiotic tolerance.
Similar content being viewed by others
Introduction
Many bacterial phenotypes, including antibiotic tolerance and virulence, often reflect the phenotype of a subset of the population rather than the average behavior1,2. Subpopulations of bacteria can arise through purely stochastic processes, as well as by regulatory and signaling pathways3. Theoretically, one way to create phenotypic diversity via a signaling pathway is multisite phosphorylation, in which each successive phosphorylation changes the activity of a protein4,5. However, it has not been experimentally shown in bacterial populations that multisite phosphorylation regulates variation in gene expression between cells, and subsequently, the emergence of phenotypic diversity. Recently, multisite phosphorylation of transcription factors have been observed in pathways involved in antibiotic tolerance and virulence6, suggesting that dynamics of multisite phosphorylation could have particular physiological relevance.
Bacterial signaling is often characterized in the context of two-component signal transduction systems (TCS) that generally consist of a histidine kinase that phosphorylates a response regulator on a single residue, which then acts as a transcription factor7. The stimulus-dependent response of this type of signaling system architecture has been analyzed theoretically8,9 and experimentally10,11, with little cell-to-cell variability observed (as quantified by CV), regardless of inducer level. This suggests that extensive cell-to-cell variability is not a general feature of bacterial TCS. However, some notable exceptions have been found for two-component systems with more complex architectures, such as the broad distribution of gene expression in the E. coli TorS/TorR regulon12 which has recently been shown to be an important factor for cell survival during oxygen depletion13. The network architecture of bacterial signal transduction systems may therefore play an underappreciated role in the dynamics and survival of bacterial populations.
In addition to TCS, bacteria also have eukaryotic-like (also called Hank’s type) Ser/Thr kinase – phosphatase pairs with homology to eukaryotic systems that perform reversible phosphorylation on Ser and Thr residues14. One particular subfamily of these systems appears to be universally conserved across Gram-positive bacteria and plays key roles in growth and virulence for many clinically important pathogens including the streptococci, S. aureus, M. tuberculosis, E. faecalis, and others6,15. Genetic and proteomic studies indicate that these Ser/Thr kinases can perform transcriptional regulation of key cellular processes involved in antibiotic tolerance and persistence through multisite phosphorylation of transcription factors. However, to date, the consequences of multisite phosphorylation for gene regulation at the single-cell-level has not been quantified. In this context the model gram-positive bacterium B. subtilis presents a comparatively straightforward system to quantify the contribution of the additional Ser/Thr phosphorylation in vivo: the homologous kinase-phosphatase pair is PrkC/PrpC, and it has been verified to regulate gene expression through additional phosphorylation of a response regulator.
It has been apparent for over 60 years that bacterial populations contain rare cells that display increased phenotypic resistance to antibiotics16. These cells, presumed to be quiescent, have been implicated in antibiotic treatment failure in genetically susceptible bacterial infections17. To date, it remains unclear to what extent the appearance of these rare cells is subject to regulation. Emerging evidence strongly implicates elevated levels of the nucleotide second messenger (p)ppGpp as a causative agent of quiescence in many bacterial species18,19. (p)ppGpp downregulates essential cellular processes such as transcription, translation, and DNA replication20. Although the precise mechanism of (p)ppGpp synthesis and its direct cellular targets vary between bacterial species, highly elevated levels of (p)ppGpp confer a quiescent state to the bacterial cell. As many antibiotics target active cellular processes, the resulting quiescent cells exhibit increased antibiotic tolerance21, suggesting that cell-to-cell variability in (p)ppGpp may be involved in phenotypic resistance to antibiotics.
The mechanistic origin of cell-to-cell variability in (p)ppGpp levels across bacterial populations remains a major open question. To date, this has been best studied in E. coli, in the context of the RelA (p)ppGpp synthetase and the SpoT hydrolase22. In contrast, other bacterial species often possess dedicated (p)ppGpp synthetases, termed small alarmone synthetases (SAS), in addition to bi-functional synthetase-hydrolases23. These SAS proteins can be activated transcriptionally20, suggesting that cell-to-cell variability in (p)ppGpp levels could originate in the transcriptional regulation of the synthetases themselves. In the Gram-positive bacterium B. subtilis, three distinct proteins, RelA, SasA, and SasB synthesize (p)ppGpp24. B. subtilis RelA is a bi-functional (p)ppGpp synthetase-hydrolase, and both SasA and SasB are dedicated synthetases. Although relA and sasB transcripts are both readily detectable during log phase growth, sasA (formerly ywaC) transcripts are found at considerably lower levels. However, sasA is inducible by certain classes of cell-wall-active antibiotics25,26, and its induction by alkaline shock increases the cellular levels of ppGpp24. Since sasA expression stops growth27, SasA-mediated (p)ppGpp synthesis provides a mechanism to induce cellular quiescence in response to environmental stresses. To date, SasA is only known to be regulated transcriptionally, so significant cell-to-cell variability in sasA expression could produce physiologically relevant cell-to-cell variability in (p)ppGpp levels. The pre-existing distribution of sasA expression may therefore be critical in predicting the relative survival of cells under conditions that do not specifically induce sasA.
In this work, we demonstrate that sasA expression displays physiologically relevant amounts of extrinsic noise, although the average level of sasA expression is very low during growth under non-inducing conditions. Furthermore, we find that both the distribution of sasA expression and the frequency of outliers are strongly regulated by the activity of a highly conserved eukaryotic-like Ser/Thr kinase system and its subsequent multisite phosphorylation of a transcription factor. Using quantitative analysis of the full distributions of sasA expression, we find that multisite phosphorylation is responsible for exponentially regulating the abundance of cells with a given level of SasA, and we generate a predictive model for sasA-expression-dependent antibiotic tolerance.
Results
sasA expression exhibits high levels of extrinsic noise
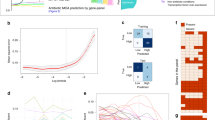
While the population average level of sasA expression during growth is extremely low25, the average behavior may mask important phenotypic variation between cells. We therefore generated a transcriptional reporter for sasA (PsasA-yfp) to study the population at the single-cell level. Surprisingly, there was considerable cell-to-cell variability in PsasA-yfp (coefficient of variation, CV ~ 4.95 ± 0.42, mean ± SEM), with most cells having very low expression, and rare cells showing significantly higher levels of expression (Fig. 1a). Quantification of YFP fluorescence revealed that a small fraction of the population had much higher (>~10×) levels of fluorescence than the mean, and rare cells had ~100×. Note that a typical bacterial gene has a CV in the range 0.1–128,29,30. Consistently, we measured the B. subtilis gene veg to have a CV of 0.4 ± 0.02 (mean ± SEM, Supplementary Fig. 1).
sasA exhibits cell-to-cell variability in expression. a PsasA-yfp expression in log phase culture. Left: YFP image. Color scale indicates the fold change relative to the population average. Right: Phase contrast. Scale bar indicates 10 µm. b Schematic of the dual-color transcriptional reporter strain used to characterize the noise in sasA expression. Two copies of the sasA promoter driving different reporters (PsasA-yfp, PsasA-mcherry) are inserted at separate ectopic loci. c Evidence for extrinsic noise in sasA expression. Images of single cells from the dual reporter strain with significant fluorescence shown in mCherry (Left), YFP (Center), and phase contrast (Right) channels. Scale bar indicates 5 µm. d Quantification of cellular fluorescence intensities for single cells in the experiment described in b, c. Values were normalized relative to the population mean value for each channel. Data shown was measured on ~580 cells with detectable fluorescence in both channels. The fluorescence of the two reporters have a Pearson’s correlation coefficient of r ~ 0.90 ± 0.08 (mean ± SEM, 4 experiments). For each reporter, the CV ~ 4.95 ± 0.42 (mean ± SEM, 4 experiments)
The high levels of cell-to-cell variability in sasA expression could be caused by intrinsic noise from the promoter itself, or by extrinsic noise originating in an upstream process31. To differentiate between these mechanisms, we used a strain with dual fluorescent reporters, PsasA-yfp and PsasA-mcherry (Fig. 1b). Expression of the dual reporters in individual cells was highly correlated (Pearson’s correlation coefficient, r ~ 0.90 ± 0.08, mean ± SEM; Spearman’s correlation coefficient ~ 0.78 ± 0.06, mean ± SEM), demonstrating that the noise was largely extrinsic to the promoter (Fig. 1c, d).
Some variability in PsasA is expected to originate in natural variability in protein expression and accumulation between cells in a population32. Therefore, we compared the variability observed in PsasA-yfp to a presumably unrelated promoter known to be constitutively active during log phase growth, Pveg-mcherry33 (Supplementary Fig. 1). We found that YFP and mCherry levels were not highly correlated, suggesting that the high levels of variability in sasA expression are largely caused by a sasA-specific pathway.
We then tested a previously characterized sasA regulator, the sigma factor σM (SigM)34, that is required for sasA expression26 (Supplementary Fig. 2A). sigM expression levels were generally detectable at low levels across the population (Supplementary Fig. 2B). We also found that sasA expression did not correlate strongly with sigM expression (Supplementary Fig. 2C, D) demonstrating that SigM levels alone do not predict variability in sasA expression. To our surprise, we also found that the relatively rare outliers in sigM expression were not predictive of the outliers we observed in sasA. Since the outliers in sasA expression would likely exhibit the strongest sasA-dependent physiological effects on the cell, we sought to determine the regulatory factors responsible for regulating their frequency in the population.
The Ser/Thr kinase PrkC represses sasA expression through WalR
Another potential regulator of sasA is the WalR transcription factor observed to bind the sasA promoter in a genome-wide screen35. WalR is the response regulator of the essential WalRK two-component system and is activated by phosphorylation of Asp-53 by WalK36. Once phosphorylated, WalR can activate and/or repress genes in its regulon. A reversible second phosphorylation on WalR Thr-101 by the eukaryotic-like Ser/Thr kinase-phosphatase pair PrkC/PrpC37 further increases WalR activity at both activating and repressing sites38. In rich media (LB), the multisite phosphorylation of WalR affects gene expression (e.g., enhanced activation of yocH) specifically in post-log phase38. However, in the commonly used defined minimal media S7-glucose, there is a consistent PrkC-dependent effect on the population average level of yocH expression throughout log phase (Supplementary Fig. 3).
sasA is known to be activated by antibiotics such as bacitracin25 through σM activation. We first tested whether the PrkC/PrpC–WalR system regulates sasA at the population level to determine if WalR activates or represses sasA. We found that PrkC activity represses sasA expression through WalR Thr101~P during bacitracin treatment (Supplementary Fig. 4). Based on these bulk measurements, we developed a model for sasA regulation (Fig. 2a) in which PrkC activity further potentiates WalR-repressing activity at sasA through a second phosphorylation of WalR at Thr-101. However, it remained unclear to what extent multisite phosphorylation of WalR affects pre-existing cell-to-cell variability and physiologically relevant outliers in sasA expression under non-inducing conditions.
The Ser/Thr kinase PrkC and phosphatase PrpC regulate cell-to-cell variability in sasA. a Model for PrkC-dependent regulation of sasA. WalR binding to the sasA promoter represses sasA expression. WalR activity is primarily controlled by phosphorylation on Asp-53 by its cognate histidine kinase WalK, and secondarily by phosphorylation on Thr-101 by the Ser/Thr kinase PrkC. WalR Thr-101~P can be dephosphorylated by the phosphatase PrpC. Thr-101~P further enhances the repressing activity of WalR Asp-53~P, resulting in lower expression of sasA under conditions with high Thr-101~P (e.g., ΔprpC). Under conditions lacking Thr-101~P (e.g., ΔprkC), the increased repression of sasA is relieved. b Histograms demonstrating PrkC-dependent cell-to-cell variability in sasA. Top: PsasA-yfp reporter activity was quantified by flow cytometry in wild type (WT, gray), ΔprpC (orange), ΔprkC (green), and Δ(prpC-prkC) (blue) backgrounds. Shaded region of each plot indicates the range of cellular autofluorescence observed. Each histogram was computed from data on ~3.0 × 104 events. Bottom: Smoothed, overlaid histograms of the data shown above for comparison. c Percentage of outliers in each genetic background. At least 4 independent experiments, similar to and including the representative one shown in b, were performed in walRWT (Left) and walRT101A (Right) backgrounds. Each experiment was normalized to a control and outliers were defined as cells above a fixed threshold level of normalized fluorescence (~1250 A.U.). Dots represent the percentage of each population that is above the threshold; bars and lines represent the mean and SEM, respectively. d Summary of each genotype, its effect on the maximal occupancy of the two known WalR phosphosites, and the expected effect on WalR activity relative to WT. For each genotype the mean percentage of outliers in sasA expression (shown in c) is also summarized. Note that in the ΔprpC background, the Thr-101 phosphorylation is stabilized (denoted as T101*)
PrkC regulates noise in sasA through WalR Thr-101 phosphorylation
Cell-to-cell variability in gene expression can be tuned by changing repressor-binding affinities39,40, suggesting that multisite phosphorylation of WalR may play a critical role in setting the observed distribution of sasA expression across the population. To test this, we measured the distribution of sasA expression in wild type (WT) cells and compared it to genetic backgrounds that alter the phosphorylation state of WalR: ΔprpC (no phosphatase, high levels of T101~P), and ΔprkC (no kinase, no detectable T101~P) (Fig. 2b, Supplementary Fig. 5). Qualitatively, in the ΔprpC background, the frequency of cells with high levels of sasA expression was strongly reduced, whereas it was strongly increased in the ΔprkC background. The PrpC-dependent effect on sasA expression requires PrkC, since the distribution of sasA expression in a strain lacking both the kinase and phosphatase (Δ(prpC-prkC)) is very similar to a strain lacking just the kinase.
We first sought to quantify the effect of WalR multisite phosphorylation on the frequency of “outliers”: cells with a level of sasA expression above a fixed threshold in each population. We therefore compared independent measurements of the distribution of sasA expression in WT, ΔprpC, and Δ(prpC-prkC) backgrounds (Fig. 2c, left) and found that PrkC significantly affects the mean frequency of outliers >8 fold by this measure (walRWT, ΔprpC vs. Δ(prpC-prkC): **p-value ~ 0.004, Kolmogorov-Smirnov test). We repeated the measurements in a walRT101A background (Fig. 2c, right) and found that PrkC no longer has a significant effect on the mean frequency of outliers in the phosphosite mutant background (walRT101A, ΔprpC vs. Δ(prpC-prkC): p-value ~ 0.56, ns, Kolmogorov-Smirnov test). These results are consistent with increased WalR activity by Thr-101 phosphorylation causing increased repression of sasA, and thereby regulating the frequency of sasA outliers (Fig. 2d). Furthermore, heterologous expression of PrkC was sufficient to reduce the frequency of outliers observed in the ΔprkC background (Supplementary Fig. 6A). Heterologous expression of PrkC was also able to further reduce the variability to below that observed in the ΔprpC background, approaching the level of cellular autofluorescence (Supplementary Fig. 6B). This suggests that at least some of the remaining variability in the ΔprpC background arises due to incomplete saturation of WalR T101~P.
This definition of outliers, however, relies on the definition of a cutoff threshold and therefore does not fully address how multisite phosphorylation affects the distribution of sasA expression across the entire population. To quantify the effect of PrkC on the distribution of cell-to-cell variability in sasA expression, we deconvolved the measured data from the cellular autofluorescence (see Methods section). This resulted in autofluorescence-free distributions of sasA expression, allowing better quantitative comparison of expression between genetic backgrounds (Fig. 3a). This deconvolution method uses only the first two moments (i.e., the mean and variance) of the observed distributions of fluorescence. As such, the auto-fluorescence free distributions are relatively insensitive to the observed “outliers” in each distribution, but makes a statistical prediction for those frequencies. To verify the accuracy of the predictions, these calculated distributions were re-convolved with the cellular autofluorescence and the reconstructed data set compared to the original data (Supplementary Fig. 7). Calculation of the relative enrichment of cells with a given level of sasA in each genetic background revealed that maximal T101~P (ΔprpC) results in exponential changes in the relative abundance of cells at a given level of sasA compared to the absence of T101~P (ΔprkC) (Fig. 3b).
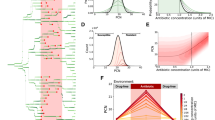
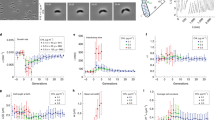
Multisite phosphorylation causes exponential changes in the abundance of cells with a given level of sasA expression. a Functional fits of autofluorescence-free distributions of sasA expression. A deconvolution algorithm was used to remove the contribution of autofluorescence from the measured distributions of sasA expression shown in Fig. 2b. Validation of the fits are shown in Supplementary Fig. 7. b Effect of PrkC on the distribution of sasA expression. For each genotype in a the frequency of cells with a given level of sasA expression was normalized against the equivalent frequency in the WT population. On this semi-logarithmic plot, straight lines indicate that the enrichment factor changes ~exponentially with fluorescence. c PrkC-dependent regulation of cell-to-cell variability in sasA is WalR Thr-101-dependent. PsasA-yfp reporter activity was quantified by flow cytometry in a ΔprpC walRT101A (yellow, solid) background and compared to ΔprpC (orange), Δ(prpC-prkC) (blue), and Δ(prpC-prkC) walRT101A (yellow, dashed) backgrounds in the same experiment. Plots were generated by applying the deconvolution method described in a to the measured data in Supplementary Fig. 8A. d Gradual inhibition of PrkC results in progressive enrichment of cells with elevated levels of sasA. PsasA-yfp reporter activity was quantified by flow cytometry during treatment with increasing concentrations of staurosporine: 0 (solvent only; black), 50 nM, 100 nM, and 1 µM (shades of magenta), in otherwise WT populations. For reference, a Δ(prpC-prkC) (blue) population treated with solvent only is shown; note that the effect of staurosporine saturates at this level. Plots were generated by applying the deconvolution method described in a to the measured data in Supplementary Fig. 8B, C. For each condition the frequency of cells with a given level of sasA expression was normalized against the equivalent frequency in the WT population. As in b, straight lines indicate that enrichment increases ~exponentially with fluorescence. Enrichment becomes more pronounced as the concentration of staurosporine increases and saturates at the level of complete inhibition ~Δ(prpC-prkC)
Together, the model and the outlier analysis in Fig. 2 suggest that PrkC-dependent regulation of the distribution of sasA expression requires the second WalR phosphosite at Thr-101. To test this, we repeated the deconvolution procedure for a strain expressing a WalR mutant that lacks the Thr-101 phosphosite, WalR T101A, and found that the PrkC-dependent effect on sasA expression is indeed WalR Thr-101-dependent (Fig. 3c, Supplementary Fig. 8A). Thus, multisite phosphorylation is responsible for the exponential depletion of cells with medium to high levels of sasA expression in the ΔprpC background (Fig. 3b). We note that exponential depletion has a strong effect on the frequency of outliers with high levels of sasA expression, and only modestly effects the first two moments of the distribution (mean and variance). Consistently, we found that although the effect on the frequency of outliers is large, the CV of the distributions measured by flow cytometry changes by only ~30% (For the data shown in Fig. 3b: WT ~ 2.87, ΔprpC ~ 2.2, ΔprkC ~ 2.61, Δ(prpC-prkC) ~ 2.65).
We then measured how intermediate levels of multisite phosphorylation regulate the distribution of sasA expression using the kinase inhibitor staurosporine to progressively inhibit PrkC activity41 (Supplementary Fig. 8B; 9). The distributions of sasA expression were again deconvolved (Supplementary Fig. 8C), and we calculated the relative enrichment of cells with a given level of sasA fluorescence at increasing concentrations of staurosporine (Fig. 3d). Titration of PrkC activity resulted in exponential enrichment of cells with a given level of sasA. Therefore, even small changes in PrkC activity result in large changes in the abundance of “outliers”, cells with unusually high levels of sasA.
sasA expression level continuously predicts antibiotic tolerance
Cell-to-cell variability in (p)ppGpp production has been proposed to result in cell-to-cell variability in antibiotic survival42,43,44,45. However, a direct and quantitative relationship between the expression of a transcriptionally regulated (p)ppGpp synthetase and the probability of survival for an individual cell has not been demonstrated. We therefore sought to determine if cells with pre-existing high levels of sasA preferentially survive antibiotic exposure, and if so, to provide a model for how the level of sasA expression influences the probability of survival for a given cell.
We used ciprofloxacin, a DNA gyrase inhibitor that does not significantly increase the population average level of sasA expression25. We measured (Supplementary Fig. 10) and deconvolved (Fig. 4a, b) the distributions of sasA expression (PsasA-yfp) both pre-ciprofloxacin and post-ciprofloxacin treatment that results in ~99% killing in both WT and ΔsasA backgrounds when measured at the bulk population level (Supplementary Fig. 11). We note that, importantly, the starting distributions of PsasA-yfp are very similar in both genetic backgrounds, allowing a direct comparison. Using these distributions, we calculated the relative enrichment of cells with a given level of sasA following antibiotic treatment (Fig. 4c), yielding a simple model for the effect of antibiotic treatment on the distribution of sasA expression (see Methods section). Because survival after antibiotic treatment can be affected by many processes, we separated out the size of the sasA-dependent effect by using a ΔsasA mutant as a control. WT populations exhibited a significant increase in the fraction of cells with elevated levels of sasA (Fig. 4c). This effect is strongly reduced in the ΔsasA background, demonstrating that sasA has a significant contribution to survival after ciprofloxacin treatment. We repeated this analysis on Δ(prpC-prkC) and Δ(prpC-prkC) ΔsasA strains to determine if a similar effect holds for the phosphorylation mutant that generates an increased frequency of cells with high levels of sasA expression (Supplementary Fig. 12). We found that Δ(prpC-prkC) populations have a ~50% increase in cells with high levels of sasA both pre-ciprofloxacin and post ciprofloxacin treatment. Consistent with the results on WT populations, the effect is partially sasA-mediated. This suggests that rare, pre-existing cells with increased probability of survival are more abundant in Δ(prpC-prkC) than in WT populations.
Cells with elevated sasA expression preferentially survive antibiotic treatment. a Autofluorescence-free distributions of sasA expression before (dark gray) and after (light gray) ciprofloxacin treatment in an otherwise WT background based on the data shown in Supplementary Fig. 10A. b Autofluorescence-free distributions of sasA expression before (dark red) and after (light red) ciprofloxacin treatment in a ΔsasA background based on the data shown in Supplementary Fig. 10B. c Enrichment plot following antibiotic treatment for WT and ΔsasA populations. For each genotype in a, b the frequency of cells with a given level of PsasA-yfp after antibiotic treatment was normalized against the corresponding frequency prior to antibiotic treatment. Dashed lines and dots indicate the average values used in d, e. d Histograms of PsasA-yfp fluorescence in WT (black) and ΔsasA (red) backgrounds measured by flow cytometry prior to sorting into “low” (4.1 ± 1.8) or “high” (779.5 ± 134.9) expression groups (relative mean ± SEM, 3 experiments). Each histogram is comprised of data obtained from ~3.0*104 events and is from a representative experiment used in e. e Relative survival of the “high” and “low” expression populations in d after treatment with ciprofloxacin for ~3.5 h as in a, b. Bars indicate the mean survival ratio of the “high” to “low” expressing populations, lines indicate the SEM. The data shown represents 3 independent experiments (paired t-test, p-value ~ 0.009, **). The red dashed line indicates a survival ratio of 1, indicating no advantage
We then sought to determine whether our enrichment model reflects the probability of survival for cells as a function of sasA expression. Since SasA is a (p)ppGpp synthetase, the sasA-dependent enrichment we observe post-ciprofloxacin treatment could be due to an expression-dependent probability of surviving antibiotic treatment. Alternately, since the measured increase in mean fluorescence was relatively small (~2 fold), ciprofloxacin could act in a complex, expression-dependent, manner to generate the observed post-treatment distribution of sasA expression without affecting survival. To differentiate between these hypotheses, we used FACS to sort bacteria prior to ciprofloxacin treatment from both WT and ΔsasA populations into “high” (upper ~ 1%) and “low” (~average) PsasA-yfp expression groups (Fig. 4d). Importantly, sorting cells by sasA expression level prior to ciprofloxacin treatment allows more sensitive measurements of the survival advantage conferred by SasA than traditional bulk population level CFU measurements. In addition, cells were measured and sorted prior to antibiotic treatment and assayed by CFUs to avoid confounding survival measurements with changes in cell physiology, fluorescent protein expression, or lysis in response to ciprofloxacin. Following ciprofloxacin treatment (as in Fig. 4a, b, Supplementary Fig. 10), the relative survival of “high” and “low” expression cells, or the survival ratio, was assayed by plating for CFUs (Fig. 4e). We observed that cells with high levels of sasA expression had a survival advantage of 9.5 ± 0.6 for WT, but only 1.8 ± 0.7 in ΔsasA background, demonstrating that cells with high levels of sasA expression prior to antibiotic treatment preferentially survive.
The average fluorescence cutoff values used in the FACS experiments, low: 4.1 ± 1.8 and high: 779.5 ± 134.9 (mean ± SEM, 3 experiments), were then used as inputs for a model where the enrichment of cells with increased levels of sasA (Fig. 4c) is caused by increased survival (see Methods section). The model yielded good agreement with the results of the FACS experiments: it predicted relative survival ratios of ~9 for wild-type, and ~2 for ΔsasA, respectively, compared to the measured values of ~9.5 ± 0.6 (WT) and 1.8 ± 0.7 (ΔsasA) (Fig. 4e). We found that relaxing the experimental cutoff for the “high” threshold to ~60% of the value in Fig. 4d resulted in a strong reduction of the survival advantage for WT: 2.3 ± 0.7 fold relative survival for WT, and 0.9 ± 0.4 fold for ΔsasA (mean ± range, 2 experiments) (Supplementary Fig. 13). This is in reasonably good agreement with a model prediction of 3.7 and 2.0-fold, respectively. Therefore, the enrichment of cells with elevated levels of sasA post-ciprofloxacin treatment can be largely attributed to the increased survival probability of pre-existing cells in the population with elevated sasA expression.
Taken together, our results demonstrate that an important consequence of PrkC-dependent multisite phosphorylation of WalR is the regulation of cell-to-cell variability, or noise, in the WalR regulon gene sasA. By comparing the full distributions of gene expression, we demonstrate that this effect is not just confined to the regulation of outliers in gene expression above an arbitrary threshold, but has an exponential effect on the relative abundance of cells with a given level of expression in the population. By analyzing the full distributions of expression, we are also able to demonstrate that sasA expression continuously affects the antibiotic tolerance of individual cells: specifically, the survival probability during a fixed course antibiotic treatment. This model (see Methods section) is consistent with cell sorting experiments that explicitly demonstrate that the observed distributions are a consequence of differential survival probabilities.
Discussion
Antibiotic tolerance is believed to be an important factor in the failure of antibiotic treatments and a key step toward the development of antibiotic resistance46. Noise in expression of genes that regulate cellular quiescence are hypothesized to play an important role in cell-to-cell variability in tolerance. We therefore sought to trace the origin of the cell-to-cell variability in expression of the (p)ppGpp synthetase sasA and determine if it can be regulated by genetic or chemical means. Noise in gene expression can be conceptually separated into intrinsic and extrinsic noise. Although it is difficult to design strategies to specifically target events generated by intrinsic noise, extrinsic noise may have upstream regulatory pathways that can be modulated. Therefore, it is significant that the cell-to-cell variability in sasA was dominated by extrinsic noise at high levels of expression (Fig. 1) that have the strongest effect on antibiotic tolerance (Fig. 4). Furthermore, since multisite phosphorylation is responsible for setting the observed distribution of cell-to-cell variability (Figs. 2, 3), this regulatory pathway could be a novel antibiotic target.
Multisite phosphorylation can expand the range of a protein’s function, generating both switch-like47,48, and graded49,50 changes in average activity. In contrast, here we observed only minimal changes in the average levels of sasA expression as a function of PrkC activity, but measured up to a ~100-fold effect on the frequency of “outliers”, cells with particularly high levels of expression (Fig. 3b). This response was shown to be graded, rather than switch-like, likely arising as a consequence of the integration of signals from two distinct signaling systems. A single phosphorylation at WalR Asp-53 strongly, but imperfectly, represses the sasA promoter. The addition of the second phosphorylation at Thr-101 by a distinct signaling system then acts as a second input to further regulate WalR. Interestingly, even small changes in activity of the second system result in marked changes in the frequency of outliers. Heterologous expression of PrkC is capable of reducing the variability observed to nearly cellular autofluorescence, but does not eliminate it completely. This remaining variability in sasA may be due to PrkC overexpression still being unable to completely saturate WalR phosphorylation, intrinsic noise at the promoter, or as yet unidentified sources. This demonstrates that PrkC activity is likely heterogeneous in our culture condition, but does not pinpoint the cause or origin of the variability. For example, from this study we cannot rule out that cells with high levels of sasA expression originate in a PrkC-dependent manner from the colony itself.
Transcriptional regulation of outliers in eukaryotes has been shown to be predictive of which cancer cells survive drug treatment51. Here, we found that transcriptional regulation by multisite phosphorylation is also critical for setting the pre-existing distribution of survival probabilities for cells within a bacterial population. Distinct from bacterial persistence, which is characterized by bi-phasic killing, these survival probabilities reflect antibiotic tolerance or the killing kinetics during a relatively short, fixed time-course, antibiotic treatment52. In the ΔsasA background, we observed a much weaker dependence of antibiotic tolerance on sasA expression. This residual dependence is consistent with previous results that have implicated many global processes in antibiotic tolerance including heterogeneity in growth state53,54,55 and enhanced expression of drug efflux pumps56,57,58. This is also consistent with the relatively weak correlation in expression between sasA and the constitutive promoter veg (Supplementary Fig. 2). Indeed, it remains to be seen precisely how cellular physiology changes in a sasA-expression dependent manner. SasA has been shown to be important for ribosome dimerization in B. subtilis27 and for survival during envelope stress in S. aureus59. More generally, various cellular processes are known to be directly and indirectly affected by rising (p)ppGpp levels including inhibition of DNA primase activity60, and reduction in intracellular GTP pools61 thereby downregulating rRNA transcription62. As our results show that antibiotic survival increases continuously with sasA expression, they suggest that SasA exerts a continuous effect proportional to its level on physiological processes that effect ciprofloxacin killing. Therefore, multisite phosphorylation may provide a “bet-hedging” strategy to regulate the phenotypic diversity of a bacterial population, serving as a broadly useful mechanism to tune the frequency of rare phenotypes that facilitate survival under adverse conditions.
Methods
Strain construction
For a listing of strains used in figures, see Supplementary Table 1. All strains are derivatives of B. subtilis 168 trpC2 unless otherwise noted. For additional details of strain and plasmid construction, see Supplementary Data 1 and 2, respectively. For a table of oligos used in this study, see Supplementary Table 2.
Media and culture conditions
B. subtilis cultures were grown in the chemically defined minimal medium S760, modified from63, supplemented with trace elements64 and L-tryptophan to early log phase. In each experiment, strains were streaked out from frozen stocks on LB Lennox and grown overnight (~15 h) at 37 °C. Single colonies were used to inoculate 3 ml liquid cultures in S7 (supplemented with inducers as indicated) and cultures were grown to early log phase (OD600 ~ 0.1–0.2). Dilutions for OD600 matching, if required, were no more than 1:3.
Microscopy and image analysis
Microscopy was performed on live cells immobilized on 1% agarose pads prepared with S7 media. Imaging was performed using a Nikon 90i or a TE2000 microscope with a Phase contrast objective (CFI Plan Apo Lambda DM ×100 Oil, NA 1.45), an X-Cite light source, a Hamamatsu Orca ER-AG, and the following filter cubes: YFP (ET Sputter 500/20×, Dm515, 535/30 m), and mCherry (ET Sputter Ex560/40 Dm585 Em630/75). To generate representative fields of log phase cultures, cultures were concentrated ~20–100× immediately prior to imaging. Images were processed using Fiji65.
Quantitative measurement of gene expression was performed similarly to ref. 66. Briefly, phase contrast and fluorescence images (e.g., YFP, mCherry) were acquired of well separated bacterial cells immobilized on agarose pads. The resulting image stacks were segmented based on the phase contrast image, and the corresponding average florescence per pixel within each cell was calculated for each fluorescence channel using Matlab. A non-fluorescent control strain, treated with antibiotics as needed, was used as a control to subtract background and autofluorescence in each channel.
Luminescence assays
Luminescence assays were performed similarly to as described38. Briefly, the cultures were initially grown to early log phase in a roller drum at 37 °C. 150 μl of each culture was then loaded into 96 well plates and 100 µg/ml bacitracin added as indicated. Measurements were performed in a Tecan Infinite 200 plate reader. Luminescence and OD600 were measured at 5 min intervals with continuous shaking and the values of all samples at a defined time point after the luminescence reached ~steady state, about 1 h post-treatment, are reported.
Flow cytometry and cell sorting
Cultures were grown to early log phase and diluted 2–4 fold with additional S7 media to obtain the optimal density for flow cytometry. The resulting samples were vortexed vigorously prior to measurement to disrupt aggregates. Flow cytometry and sorting were performed on a BD Biosciences FACS Aria II-SORP or a Miltenyi MACSquant VYB (Supplementary Figures 6 and 12 only). YFP was detected using a blue laser (488 nm) with a 525/50 dichroic, and a 505 long pass filter for both flow cytometers. mCherry was detected using a yellow/green laser (561 nm) with a 582/15 dichroic, and a 570 long pass filter (BD Aria only). Fluorescence values were quantitatively compared between experiments by rescaling each experiment by the mean fluorescence of a control sample. Sorting was performed with a 70 nm nozzle at 70 PSI. Detection voltages were set such that the non-fluorescent control had a median value of ~100. Unless otherwise noted, flow cytometry data shown is representative of at least 3 biological replicates.
The sorting thresholds were chosen such that the “high” expression gate corresponds to approximately 1% of the starting population and can be sorted in ~10 min. Higher thresholds than this result in very long sort times due to decreases in overall sorting efficiency, and also appeared to introduce unwanted variability in cell physiology. Lower thresholds (e.g., as in Supplementary Fig. 13), resulted in a strong reduction of the size of the effect.
Survival assays
~1.5 × 104 cells were sorted by fluorescence and dispensed into equal volumes of chemically defined growth media (S7). Cultures were incubated in a roller drum at 37 °C for 10 min, then treated with 500 ng/ml ciprofloxacin for 3.5 h. Serial dilutions of each population were plated for colony forming units (CFUs) on LB and grown overnight at 30 °C and survival ratios were calculated by comparing the CFU values between the high and low fluorescence groups. In each experiment, wild-type and mutant strains were tested in parallel.
Autofluorescence deconvolution method and validation
For each flow cytometry experiment, data from both sasA reporter strains and a nonfluorescent control were measured. As nonfluorescent controls for each genotype were verified to have similar autofluorescence, typically a single control measurement was performed for each experiment. Raw measurements of fluorescence intensities from transcriptional reporter strains have two contributions: (i) the transcriptional reporter for sasA expression (e.g., PsasA-yfp), the signal of interest; and (ii) background fluorescence originating from other sources. Measuring the statistics of the background fluorescence using a non-fluorescent control allowed extraction (deconvolution) of the sasA transcriptional reporter signal from the total raw measurement consisting of the sum of this signal and background fluorescence67.
We note that on our data sets, brute force deconvolution in absence of any prior knowledge of the signal statistics is quite noisy and error prone. However, in bacteria, the statistics of protein copy numbers is known to be well described by the Gamma distribution, which is a great advantage as this distribution is fully specified in terms of its mean and variance. Given the measured mean and variance of the raw fluorescence intensity and of the autofluorescence, the mean and variance of the signal can be estimated by assuming that the signal from the sasA transcriptional reporter is not correlated with background fluorescence. (This assumption is supported by fluorescence microscopy experiments showing that cell size was not well correlated with sasA expression, and by computationally verifying that simple forms of dependency are inconsistent with the observed data.) The mean and variance of fluorescence from the sasA transcriptional reporter could then be uniquely determined, as described in more detail in the mathematical details of the deconvolution procedure.
As mentioned above, we used a Gamma distribution to model the distribution of the fluorescence signal for a given mean and variance (we separately verified that other distribution choices, such as lognormal, did not capture the data as well). The Gamma distribution is routinely applied to fit gene expression data,30 but here we additionally verified that it closely mimics the underlying signal coming from the reporter by an in-silico re-convolution of the Gamma-distribution-fitted autofluorescence-free signal with measured background fluorescence. This was done by drawing two random fluorescence intensities: one from the estimated Gamma distribution for the clean (autofluorescence-free) signal and one from the measured autofluorescence distribution. Adding these two random contributions gave a single, re-convolved, data point. Repeating this procedure for a number of times equaling the size of the measured data set yielded the full re-convolved data set. The empirical distributions of the measured and re-convolved data sets were then compared (Supplementary Fig. 7) and found to be almost indistinguishable from those that were originally measured (Supplementary Fig. 3). No smoothing was applied.
Mathematical details of deconvolution procedure
The raw fluorescence intensity (R) measured from transcriptional reporter strains were assumed to have two statistically independent contributions. One coming from the signal of interest (S), and one coming from other sources, collectively treated here as noise (N). One then has
for every measurement taken, and averaging over all measurements immediately gives the following relation
between the mean fluorescence values of the raw fluorescence intensity, the signal of interest, and the noise. Statistical independence then asserts that the variance in the fluorescence intensity could also be decomposed in a similar way
These relations, and the fact <R>, <N>, Var (R), Var (N) could all be directly computed from measured data allowed us to estimate the mean, <S>, and variance, Var (S), of the signal of interest. The full distribution of the signal of interest was then assumed to be well described by a Gamma distribution, since this has previously been shown to correctly describe gene expression data in bacteria30 and we have moreover verified that this assumption is internally consistent with the data (see Autofluorescence deconvolution method and validation). The probability density function describing S was therefore modeled as
where \(\Gamma \left( \cdot \right)\) denotes the gamma function, and the parameters \(k\) and θ are uniquely set by their relations to the mean
and variance
of the signal.
Mathematical details of enrichment model
Deconvolved fluorescence intensities of cells before, fb(s), and after, fa(s), ciprofloxacin treatment were modeled by the Gamma distribution to test the hypothesis that they are related via
where p(s) stands for the probability of a cell with fluorescence intensity s to survive the prescribed treatment and C is a normalization constant assuring that
Following treatment, we define the enrichment factor (in the probability density) of cells with fluorescence level s as
One could then write the ratio between the survival probability of cells with fluorescence s1 and s2 (Survival ratio) in the following way
This relation was then used to predict survival ratios and compare with data as described in the main text.
Significance testing for % outlier data shown in Fig. 2
In Fig. 2c at least 4 independent experiments measuring sasA expression were performed in WT, ΔprpC, and Δ(prpC-prkC) in otherwise walRWT and walRT101A backgrounds. For each genotype, the % of cells above a fixed fluorescence threshold was measured and is shown as black dots. To test if the % values obtained for each genotype are statistically significant, we treated the independent measurements of %s as members of a distribution and performed Kolmogorov-Smirnov testing to determine the likelihood that the % outliers observed for a comparison genotype pair could be drawn from the same distribution.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are provided in the paper itself and associated source data files.
References
Arnoldini, M. et al. Bistable expression of virulence genes in salmonella leads to the formation of an antibiotic-tolerant subpopulation. PLoS Biol. 12, e1001928 (2014).
Norman, T. M., Lord, N. D., Paulsson, J. & Losick, R. Stochastic switching of cell fate in microbes. Annu. Rev. Microbiol 69, 381–403 (2015).
Maamar, H., Raj, A. & Dubnau, D. Noise in gene expression determines cell fate in Bacillus subtilis. Science 317, 526–529 (2007).
Gunawardena, J. Multisite protein phosphorylation makes a good threshold but can be a poor switch. Proc. Natl Acad. Sci. USA 102, 14617–14622 (2005).
Ortega, F., Garces, J. L., Mas, F., Kholodenko, B. N. & Cascante, M. Bistability from double phosphorylation in signal transduction. Kinet. Struct. Requir. FEBS J. 273, 3915–3926 (2006).
Wright, D. P. & Ulijasz, A. T. Regulation of transcription by eukaryotic-like serine-threonine kinases and phosphatases in Gram-positive bacterial pathogens. Virulence 5, 863–885 (2014).
Laub, M. T. & Goulian, M. Specificity in two-component signal transduction pathways. Annu. Rev. Genet 41, 121–145 (2007).
Batchelor, E. & Goulian, M. Robustness and the cycle of phosphorylation and dephosphorylation in a two-component regulatory system. Proc. Natl Acad. Sci. USA 100, 691–696 (2003).
Shinar, G., Milo, R., Martinez, M. R. & Alon, U. Input output robustness in simple bacterial signaling systems. Proc. Natl Acad. Sci. USA 104, 19931–19935 (2007).
Batchelor, E., Silhavy, T. J. & Goulian, M. Continuous control in bacterial regulatory circuits. J. Bacteriol. 186, 7618–7625 (2004).
Miyashiro, T. & Goulian, M. High stimulus unmasks positive feedback in an autoregulated bacterial signaling circuit. Proc. Natl Acad. Sci. USA 105, 17457–17462 (2008).
Roggiani, M. & Goulian, M. Oxygen-dependent cell-to-cell variability in the output of the Escherichia coli Tor phosphorelay. J. Bacteriol. 197, 1976–1987 (2015).
Carey, J. N. et al. Regulated stochasticity in a bacterial signaling network permits tolerance to a rapid environmental change. Cell. https://doi.org/10.1016/j.cell.2018.02.005 (2018).
Pereira, S. F., Goss, L. & Dworkin, J. Eukaryote-like serine/threonine kinases and phosphatases in bacteria. Microbiol. Mol. Biol. Rev. 75, 192–212 (2011).
Manuse, S., Fleurie, A., Zucchini, L., Lesterlin, C. & Grangeasse, C. Role of eukaryotic-like serine/threonine kinases in bacterial cell division and morphogenesis. FEMS Microbiol. Rev. 40, 41–56 (2016).
Bigger, J. W. Treatment of staphylococcal infections with penicillin by intermittent sterilisation. Lancet 244, 497–500 (1944).
Fauvart, M., De Groote, V. N. & Michiels, J. Role of persister cells in chronic infections: clinical relevance and perspectives on anti-persister therapies. J. Med. Microbiol. 60, 699–709 (2011).
Abranches, J. et al. The molecular alarmone (p)ppGpp mediates stress responses, vancomycin tolerance, and virulence in Enterococcus faecalis. J. Bacteriol. 191, 2248–2256 (2009).
Gaca, A. O. et al. Basal levels of (p)ppGpp in Enterococcus faecalis: the magic beyond the stringent response. MBio 4, e00646–00613 (2013).
Liu, K., Bittner, A. N. & Wang, J. D. Diversity in (p)ppGpp metabolism and effectors. Curr. Opin. Microbiol. 24, 72–79 (2015).
Nguyen, D. et al. Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 334, 982–986 (2011).
Cashel, M., Gentry, D., Hernandez, V. & Vinella, D. in Esherichia coli and Salmonella; Cellular and Molecular Biology Vol. 2, 1458–1496 (ASM Press, Washington DC, 1996).
Atkinson, G. C., Tenson, T. & Hauryliuk, V. The RelA/SpoT homolog (RSH) superfamily: distribution and functional evolution of ppGpp synthetases and hydrolases across the tree of life. PLoS ONE 6, e23479 (2011).
Nanamiya, H. et al. Identification and functional analysis of novel (p)ppGpp synthetase genes in Bacillus subtilis. Mol. Microbiol. 67, 291–304 (2008).
D’Elia, M. A. et al. Probing teichoic acid genetics with bioactive molecules reveals new interactions among diverse processes in bacterial cell wall biogenesis. Chem. Biol. 16, 548–556 (2009).
Eiamphungporn, W. & Helmann, J. D. The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses. Mol. Microbiol. 67, 830–848 (2008).
Tagami, K. et al. Expression of a small (p)ppGpp synthetase, YwaC, in the (p)ppGpp(0) mutant of Bacillus subtilis triggers YvyD-dependent dimerization of ribosome. MicrobiologyOpen 1, 115–134 (2012).
Pedraza, J. M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005).
Milo, R., Jorgensen, P., Moran, U., Weber, G. & Springer, M. BioNumbers-the database of key numbers in molecular and cell biology. Nucleic Acids Res. 38, D750–D753 (2010).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Leveau, J. H. & Lindow, S. E. Predictive and interpretive simulation of green fluorescent protein expression in reporter bacteria. J. Bacteriol. 183, 6752–6762 (2001).
Radeck, J. et al. The Bacillus BioBrick Box: generation and evaluation of essential genetic building blocks for standardized work with Bacillus subtilis. J. Biol. Eng. 7, 29 (2013).
Thackray, P. D. & Moir, A. SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress. J. Bacteriol. 185, 3491–3498 (2003).
Salzberg, L. I. et al. The WalRK (YycFG) and sigma(I) RsgI regulators cooperate to control CwlO and LytE expression in exponentially growing and stressed Bacillus subtilis cells. Mol. Microbiol. 87, 180–195 (2013).
Dubrac, S., Bisicchia, P., Devine, K. M. & Msadek, T. A matter of life and death: cell wall homeostasis and the WalKR (YycGF) essential signal transduction pathway. Mol. Microbiol. 70, 1307–1322 (2008).
Gaidenko, T. A., Kim, T. J. & Price, C. W. The PrpC serine-threonine phosphatase and PrkC kinase have opposing physiological roles in stationary-phase Bacillus subtilis cells. J. Bacteriol. 184, 6109–6114 (2002).
Libby, E. A., Goss, L. A. & Dworkin, J. The Eukaryotic-Like Ser/Thr Kinase PrkC Regulates the Essential WalRK Two-Component System in Bacillus subtilis. PLoS Genet. 11, e1005275 (2015).
Jones, D. L., Brewster, R. C. & Phillips, R. Promoter architecture dictates cell-to-cell variability in gene expression. Science 346, 1533–1536 (2014).
Sanchez, A., Garcia, H. G., Jones, D., Phillips, R. & Kondev, J. Effect of promoter architecture on the cell-to-cell variability in gene expression. PLoS Comput. Biol. 7, e1001100 (2011).
Shah, I. M., Laaberki, M. H., Popham, D. L. & Dworkin, J. A eukaryotic-like Ser/Thr kinase signals bacteria to exit dormancy in response to peptidoglycan fragments. Cell 135, 486–496 (2008).
Fisher, R. A., Gollan, B. & Helaine, S. Persistent bacterial infections and persister cells. Nat. Rev. Microbiol. 15, 453–464 (2017).
Hahn, J., Tanner, A. W., Carabetta, V. J., Cristea, I. M. & Dubnau, D. ComGA-RelA interaction and persistence in the Bacillus subtilis K-state. Mol. Microbiol. 97, 454–471 (2015).
Germain, E., Castro-Roa, D., Zenkin, N. & Gerdes, K. Molecular mechanism of bacterial persistence by HipA. Mol. Cell 52, 248–254 (2013).
Kaspy, I. et al. HipA-mediated antibiotic persistence via phosphorylation of the glutamyl-tRNA-synthetase. Nat. Commun. 4, 3001 (2013).
Levin-Reisman, I. et al. Antibiotic tolerance facilitates the evolution of resistance. Science 355, 826–830 (2017).
Ferrell, J. E. Jr. & Ha, S. H. Ultrasensitivity part II: multisite phosphorylation, stoichiometric inhibitors, and positive feedback. Trends Biochem. Sci. 39, 556–569 (2014).
Nash, P. et al. Multisite phosphorylation of a CDK inhibitor sets a threshold for the onset of DNA replication. Nature 414, 514–521 (2001).
Zaytsev, A. V. et al. Multisite phosphorylation of the NDC80 complex gradually tunes its microtubule-binding affinity. Mol. Biol. Cell 26, 1829–1844 (2015).
Pufall, M. A. et al. Variable control of Ets-1 DNA binding by multiple phosphates in an unstructured region. Science 309, 142–145 (2005).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Kester, J. C. & Fortune, S. M. Persisters and beyond: mechanisms of phenotypic drug resistance and drug tolerance in bacteria. Crit. Rev. Biochem. Mol. Biol. 49, 91–101 (2014).
Balaban, N. Q., Merrin, J., Chait, R., Kowalik, L. & Leibler, S. Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004).
Conlon, B. P. et al. Persister formation in Staphylococcus aureus is associated with ATP depletion. Nat. Microbiol. 1, 16051 (2016).
Gutierrez, A. et al. Understanding and sensitizing density-dependent persistence to quinolone antibiotics. Mol. Cell 68, 1147–1154 (2017). e1143.
Adams, K. N. et al. Drug tolerance in replicating mycobacteria mediated by a macrophage-induced efflux mechanism. Cell 145, 39–53 (2011).
Pu, Y. et al. Enhanced efflux activity facilitates drug tolerance in dormant bacterial cells. Mol. Cell 62, 284–294 (2016).
El Meouche, I. & Dunlop, M. J. Heterogeneity in efflux pump expression predisposes antibiotic-resistant cells to mutation. Science 362, 686–690 (2018).
Geiger, T., Kastle, B., Gratani, F. L., Goerke, C. & Wolz, C. Two small (p)ppGpp synthases in Staphylococcus aureus mediate tolerance against cell envelope stress conditions. J. Bacteriol. 196, 894–902 (2014).
Wang, J. D., Sanders, G. M. & Grossman, A. D. Nutritional control of elongation of DNA replication by (p)ppGpp. Cell 128, 865–875 (2007).
Kriel, A. et al. Direct regulation of GTP homeostasis by (p)ppGpp: a critical component of viability and stress resistance. Mol. Cell 48, 231–241 (2012).
Krasny, L. & Gourse, R. L. An alternative strategy for bacterial ribosome synthesis: Bacillus subtilis rRNA transcription regulation. EMBO J. 23, 4473–4483 (2004).
Vasantha, N. & Freese, E. Enzyme changes during Bacillus subtilis sporulation caused by deprivation of guanine nucleotides. J. Bacteriol. 144, 1119–1125 (1980).
Harwood, C. R. & Cutting, S. M. in Modern Microbiological Methods (Wiley, New York, 1990).
Schindelin, J. et al. Fiji: an open-source least 4 independent experimentsplatform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Miyashiro, T. & Goulian, M. Single-cell analysis of gene expression by fluorescence microscopy. Methods Enzymol. 423, 458–475 (2007).
Corsetti, J. P. et al. Correction of cellular autofluorescence in flow cytometry by mathematical modeling of cellular fluorescence. Cytometry 9, 539–547 (1988).
Acknowledgements
We thank Eric Brown for the strain EB1385, Isaac Plant (Silver lab) for pIP384 and IP563, and Amir Figueroa at the Microbiology and Immunology Core Facility at Columbia University for assistance with flow cytometry and cell sorting. This work was supported by NIH GM114213 and a BWF Investigators in the Pathogenesis of Infectious Disease award to J.D., a grant from the Department of Systems Biology at Harvard Medical School to E.A.L., and S.R. was supported by NIH GM095784 and gratefully acknowledges support from the Azrieli Foundation.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments E.A.L. and J.D. E.A.L. performed the experiments. E.A.L. and S.R. analyzed data. Contributed reagents/materials/analysis tools: E.A.L. and S.R. Wrote the paper: E.A.L., S.R., and J.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Libby, E.A., Reuveni, S. & Dworkin, J. Multisite phosphorylation drives phenotypic variation in (p)ppGpp synthetase-dependent antibiotic tolerance. Nat Commun 10, 5133 (2019). https://doi.org/10.1038/s41467-019-13127-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-13127-z
This article is cited by
-
Growth rate is modulated by monitoring cell wall precursors in Bacillus subtilis
Nature Microbiology (2023)
-
Many birds with one stone: targeting the (p)ppGpp signaling pathway of bacteria to improve antimicrobial therapy
Biophysical Reviews (2021)
-
Association of mprF mutations with cross-resistance to daptomycin and vancomycin in methicillin-resistant Staphylococcus aureus (MRSA)
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.