Abstract
Serotonin receptors (5-HT3AR) directly regulate gut movement, and drugs that inhibit 5-HT3AR function are used to control emetic reflexes associated with gastrointestinal pathologies and cancer therapies. The 5-HT3AR function involves a finely tuned orchestration of three domain movements that include the ligand-binding domain, the pore domain, and the intracellular domain. Here, we present the structure from the full-length 5-HT3AR channel in the apo-state determined by single-particle cryo-electron microscopy at a nominal resolution of 4.3 Å. In this conformation, the ligand-binding domain adopts a conformation reminiscent of the unliganded state with the pore domain captured in a closed conformation. In comparison to the 5-HT3AR crystal structure, the full-length channel in the apo-conformation adopts a more expanded conformation of all the three domains with a characteristic twist that is implicated in gating.
Similar content being viewed by others
Introduction
The ion channel class of serotonin receptors (5-HT3AR) are cation selective and belong to the super family of pentameric ligand-gated ion channels (pLGICs)1,2. The 5-HT3AR pentameric complex is ~250 KDa in molecular weight and is made up of five homologous subunits A (5-HT3A) or a heterologous combination of subunit A with other subunits (B–E) arranged around a pseudo five-fold symmetric axis3. These channels, located in the dorsal vagal complex of the brainstem and in the gastrointestinal (GI) tract, form a circuit that controls gut motility, visceral perception, secretion, and the emesis reflex4,5. 5-HT3AR are implicated in a number of psychiatric and GI disorder conditions including anxiety, depression, bipolar disorder, and irritable bowel syndrome6,7. Currently, serotonin receptor (5-HT3AR) antagonists are in clinical use to alleviate nausea and vomiting caused by chemotherapy and radiotherapy, and for the management of post-infection diarrhea and irritable bowel syndrome8,9. However, in several cases, adverse side effects have led to restrictions in use of these drugs10. A better understanding of the structural correlates of 5-HT3AR function will therefore facilitate ongoing drug design efforts for safer therapeutics.
At the functional level, the 5-HT3AR gating cycle involves transitions between at least three distinct conformational states, namely: the resting state, a non-conductive conformation with low agonist-affinity; the open state, a conductive conformation with higher agonist-affinity; and the desensitized state, a non-conductive conformation, with the highest agonist-affinity among the three states. In the absence of the agonist (serotonin), the channel resides predominantly in the resting or closed conformation, while in the presence of the agonist, the channel transiently opens and eventually transitions to the desensitized conformation. Several therapeutically interesting compounds act as orthosteric or allosteric ligands and modulate 5-HT3AR channel activity by shifting the equilibrium between these pre-formed functional states. At the structural level, 5-HT3AR is a dynamic allosteric protein where binding of the neurotransmitter serotonin in the N-terminal extracellular domain (ECD) elicits a conformational change leading to pore opening within the transmembrane domain (TMD). In addition to these two domains, the channel has a large intracellular domain (ICD) formed by the region between the transmembrane M3 and M4 helices. Although high resolution structures of several prokaryotic and eukaryotic pLGICs are now available11,12,13,14,15,16,17,18,19, the key questions regarding the conformational coupling between the ECD, TMD, and the ICD still remain unclear. The ICD is implicated to play a role in receptor trafficking and clustering at the synapse plasma membrane, gating, post-translational modification, and intracellular regulation of channel function20. Additionally, in 5-HT3AR, the relatively large ICD also modulates single-channel channel conductance, rectification, and desensitization kinetics21,22. In the pLGIC structures solved so far, the ICD is intrinsically missing, or has been genetically removed or considerably digested by trypsin13,14,15,16,17. In the trypsin-digested 5-HT3AR crystal structure, the ICD was partially resolved as two split α-helices; a short horizontal MX helix extending from the post-M3 loop and the MA helix extending from the cytoplasmic side toward M416. Moreover, the 5-HT3AR was crystallized in the presence of single-chain antibodies (referred to as nanobodies) that bound to the channel in the vicinity of the serotonin binding site. Functional analysis showed that in the nanobody-bound form, the channel is not activated by serotonin, and hence the channel conformation likely corresponds to that of an inhibited, non-conducting channel16. Although the inhibited state (at least in the case of competitive antagonism) could be expected to resemble the resting (closed) conformation, experimental evidences from voltage-clamp fluorometry, X-ray crystallography, and theoretical predictions in pLGIC homologs suggest that the inhibitors elicit conformational changes of their own even though the channel is electrically silent23,24,25,26,27. Interestingly, the antagonist-induced structural changes at some positions are similar while others distinct to those evoked by the agonist suggesting that the antagonist stabilizes the receptor in a different non-conductive conformation23,25. Thus, to determine the basic tenets of the resting conformation, we sought to determine the structure of the full-length 5-HT3AR channel in the apo-conformation by single-particle cryo-electron microscopy (cryo-EM). The cryo-EM structure of apo-5-HT3AR at 4.3 Å resolution is similar to the 5-HT3AR crystal structure but is captured in a distinct state with differences in the ECD, TMD, and the ICD conformations.
Results
Function and the cryo-EM structure of 5-HT3AR
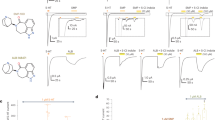
The full-length mouse 5-HT3AR was cloned into an oocyte expression vector for functional measurements. In response to application of serotonin, robust concentration-dependent macroscopic inward currents were elicited that desensitize in the continued presence of the activating ligand (Supplementary Fig. 1A). A plot of peak response as a function of serotonin concentration yielded an EC50 of 2.7 ± 0.09 μM which is in agreement with the previous studies28 (Supplementary Fig. 1A). For structural studies, the full-length 5-HT3AR gene was codon-optimized, cloned into pFastBac1 plasmid, and expressed in Spodoptera frugiperda (Sf9) cells. Milligram quantities of 5-HT3AR samples were purified to homogeneity. The purified protein was deglycosylated with PNGase treatment and gel-filtration fractions corresponding to pentamers (Supplementary Fig. 1B) were used for single-particle cryo-EM analysis (Supplementary Fig. 1C). Three dimensional reconstructions of the detergent-solubilized, full-length 5-HT3AR channel led to a density map with a nominal resolution of 4.3 Å with imposed C5 symmetry (Supplementary Figs. 1D, 2A, and 2B). The final reconstruction was built from 108,727 particles (Supplementary Fig. 3) and had a local resolution in most regions between 3.5 and 4.5 Å (Supplementary Fig. 1D). The map contained density for the entire ECD, TMD, and a large region of the ICD, and was used for model building and refinement (Supplementary Fig. 4). The final refined apo-5-HT3AR model contains residues Asp6-Gln338 and L397-Ser461 (Table 1).
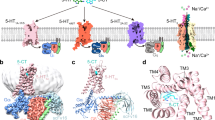
When viewed from the plane of the bilayer, perpendicular to the pore axis, the apo-5-HT3AR pentamer is 165 Å in length and has a diameter of 80 Å (Fig. 1a, b). The ECD reveals protruding non-protein densities corresponding to N-glycans. The mouse 5-HT3AR has three N-glycosylation sites at positions Asn82, Asn148, and Asn164, and strong densities for glycans were observed at each of the asparagine sidechains. Site-directed mutagenesis shows that prevention of glycosylation at sites Asn82, Asn148, and Asn164 abolishes the expression of functional receptors on the membranes29. Densities of the glycans were also resolved in several previous pLGIC structures including 5-HT3AR, GABA-β3, nAChR, GlyR, and GluCl13,16,17,30. The overall topology of apo-5-HT3AR is as previously observed in other members of the pLGIC family with each subunit comprised of a twisted β-sandwich ECD and a four-helical TMD. In addition, in apo-5-HT3AR, the density for the ICD is only partially resolved with post-M3 loop, MX, and MA helices visible.
Cryo-EM structure of apo-5-HT3AR. a The 3D reconstruction map from the full-length 5-HT3AR at 4.3 Å resolution. The views, going from left to right, are parallel to the membrane (side view), from the extracellular side (top view), and from the intracellular side (bottom view). Individual subunits are depicted in different colors, and the three domains are labeled. The solid lines denote putative membrane limits. b Cartoon representations of the 5-HT3AR structural model based on the EM reconstruction. The views correspond to the orientations shown in a. For each subunit, three sets of glycans (green) and one lipid (brown) molecule are shown as stick representation
We then compared the apo-5-HT3AR conformation with that of 5-HT3AR crystal structure16. An alignment of the two structures reveals an expansion of all the three domains in apo-5-HT3AR resulting from a radial outward twist of the ECD outer β-sheet and an iris-like movement of the TM helices and the ICD (Fig. 2a). The ECD undergoes a clockwise twist with respect to the crystal structure, which changes the relative positions of the interfacial loops at the ECD–ECD and ECD–TMD interfaces (Fig. 2b). Particularly, the movement in the outer β-strands is more pronounced than the inner β-strands. In the TMD, the M3 and M4 helices show an outward displacement with respect to an axis through the intrasubunit helical bundle composed of M1–M4 helices, while there is very little change in the position of the M2 helix (Fig. 2b). Changes in M3 and M4 conformations relative to M2 lead to a widening of the intrasubunit cavity lined by M1–M4 helices (indicated by dotted lines, Fig. 2b). These structural differences are also reflected as an increase in total surface area (protein-only region) of the apo-5-HT3AR pentamer (102,850 Å2) in comparison to the crystal structure (85,000 Å2). The increase in total surface area is accompanied by an increase in solvent accessible areas for apo-5-HT3AR pentamer (73,210 Å2) in comparison to the crystal structure (54,530 Å2). The analysis of inter-subunit interfaces reveals that the buried areas between adjacent principal and complementary subunits are slightly reduced in apo-5-HT3AR (2858 Å2) compared to the crystal structure (3045 Å2). A comparison of different pLGIC structures captured in various conformational states seems to suggest that activation involves a concerted counter-clockwise twist of the ECD around the pore axis15,19,30. It was previously noted that the nanobody-bound 5-HT3AR is twisted counter-clockwise with respect to the pLGIC putative closed conformations such that it lies further along the pathway that defines the global motion going from closed to open structures16. In apo-5-HT3AR, the ECD twist is clockwise with respect to the crystal structure, and in the direction toward the closed conformation.
Alignment of the apo-5-HT3AR with the crystal structure of nanobody-bound 5-HT3AR. a A view of the ECDs from the extracellular end when aligned with respect to the TMDs (left). The view of the TMDs from the intracellular end when aligned with respect to the ECDs (right). The apo-5-HT3AR structure and the 5-HT3AR crystal structure are shown in salmon red and pale green, respectively. The arrows show the putative direction of displacements between the two structures. b A comparison of the ECD of the apo-structure with the crystal structure when aligned with respect to the TMD of the (−) subunit (left). A comparison of the TMD between the two structures when aligned with the ECD of the (−) subunit. The relative tilt of the axis parallel to the TM helices between the two structures are indicated. The dotted lines highlight the differences in the intrasubunit cavity volume. The spheres indicate the position of residues M1 (Leu227), M2 (Leu266), M3 (Met291), and M4 (Trp456). The arrows show the putative direction of displacements between the two structures (right). The alignment in b highlights the relative changes in the two structures, both with respect to the neighboring subunit, as well as with respect to the other domain
The ion permeation pathway and the TMD
Viewing from the extracellular end (Fig. 3a, b), the ion permeation pathway in apo-5-HT3AR is wide open with a radius greater than 3.5 Å throughout the ECD followed by a narrower pathway in the TMD and the ICD with multiple constriction sites (Supplementary Fig. 5). A slight constriction is seen at the center of the ECD lined by Lys108 in the β4–β5 loop, although the extent of constriction is less than that seen in the crystal structure (Fig. 3b). A negative charge in the vicinity of Lys108 (at position Asp105) has been shown to regulate single-channel conductance in pLGICs31. The Asp105 position is conserved among most cation-selective pLGICs, whereas this position is a lysine in anion-selective members of the family32.
Profile of ion permeation pathway. a The pore profile generated by the HOLE program60 depicts an ion permeation pathway of ~165 Å encompassing the ECD, TMD, and the ICD. Only two subunits are shown for clarity. Sidechains of residues that line the constricted areas are shown as sticks. b A comparison of the pore radii along the pore axis for the 5-HT3AR cryo-EM structure (salmon red) with that of the crystal structure (pale green). The dashed line indicates an approximate radius of a hydrated Na+ ion35. The pore is constricted below 3 Å radius at three sites: L260 and E250 along M2 and R416 in the ICD. c Non-protein densities along the pore axis were modeled as water (red), Na+ (magenta), and Cl− (green). The map around the ions is shown as a mesh representation calculated at various σ values (outer water ring: 4 σ; inner water ring:5 σ; Na+ ion: 6 σ; Cl− ion: 7 σ)
The pore of apo-5-HT3AR is lined by M2 which appear as straight helices lying parallel to the pore axis (Fig. 3c). At the mouth of the extracellular end, the pore is lined by Asp20′, followed by three rings of hydrophobic residues at Ile17′, Val13′, and Leu9′. Below Leu9′, the pore is polar lined by Thr6′, Ser2′, and negatively charged at the intracellular end with Glu-1′. The Asp-4′ is positioned further away from the pore axis. The radius of the pore at Asp20′ is 3.8 Å and decreases to 2.1 Å at the level of Leu9′ position. Strong density (7–6 σ) and weaker density (5–4 σ) are present within the permeation pathway in the TMD and the ICD (Fig. 3c). The weaker densities are arranged as two sets of five-membered rings: one as a loose arrangement, in-plane with Asp20′ sidechain with a mean distance of 3.94 Å and the other as a tight pentagon ring beneath it with a mean distance of 2.4 Å. These values are close to those estimated for water polygons in high-resolution protein crystals33. Based on these distances, the pore radii at this location (>hydrated Na+ ion radius), and the B-factors for the modeled ions and waters (Table 1), we suggest that these densities likely correspond to ions and water. A similar arrangement of ions and water pentagons was also observed in the intracellular end of M2 in the GLIC open structure34. Further down, the pore at Leu9′ is constricted to ~2.3 Å which is too small to allow a hydrated Na+35 to pass through and in addition the hydrophobicity of this region prevents ion dehydration. There are two additional constrictions below the Leu9′ position, at Thr6′ and Ser2′ with a radius of 2.7 Å and 2.1 Å, respectively. Although the pore radii at these locations are smaller than a hydrated Na+, the hydroxyl sidechain at these positions may allow coordination of partially dehydrated ions. The intracellular end is lined by two rings of negative charges at positions Glu-1′ and Asp-4′ which are involved in governing charge selectivity and constitute the selectivity filter36. This suggests that the channel in this conformation is non-conducting and the Leu9′ position acts as a gate impeding ion permeation. This is not surprising considering that the channel was imaged in the absence of any activating ligands. The finding that the activation gate is located at the center of M2 is also in agreement with SCAM studies in 5-HT3AR which showed that extracellularly positioned Cys in M2, particularly the 17′ position, were accessible even in the closed state. In contrast, residues below the 13′ are accessible in the open but not in the closed state37. In the strychnine-bound GlyR structure, the channel is constricted primarily at the extracellular end with constrictions to below 3 Å radii at 20′, 13′, and 9′ positions15. The “closed” GLIC conformation is also much narrower in the extracellular end with positions 16′ and 17′ constricted to 2 Å19. On the other hand, the 5-HT3AR structures are similar to the apo-GluCl structure with a relatively wider pore at the external end30 (Supplementary Fig. 6).
At the extracellular end, M2 extends into the M2–M3 linker, which in apo-5-HT3AR is bent and tilted upward (above the level of the cys loop), such that it is in close proximity with the pre-M1 region as well as the β8–β9 loop of the neighboring subunit (Supplementary Fig. 7A). Residue Ala277 in the M2–M3 linker makes a H-bond contact with Arg219 in the pre-M1 region, while Thr276 interacts with the backbone carbonyl of Gly185 in the β8–β9 loop. The M2–M3 linker was only partially resolved in the 5-HT3AR crystal structure16. The M2–M3 linker is seen in multiple orientations in different pLGIC structures with this region adopting a distinct extended conformation in the apo-5-HT3AR (Supplementary Fig. 7B). The M2–M3 linker in the agonist-bound GlyR structure is retracted from the inter-subunit interface in comparison to the antagonist-bound structure15. The M2–M3 linker and the pre-M1 region are implicated to be a part of the signal transduction machinery that communicates ligand-binding events in the ECD to the channel gate in the M2 bundle38. Consequently, mutations in these regions are shown to impact channel function39.
The M4 segment has a well-defined density and the tip of M4 is α-helical extending above the putative membrane limits (Supplementary Fig. 8). The tip of M4 is in close proximity to the β6–β7 (Cys) loop with Trp459 in M4 oriented toward the conserved Phe144 in the Cys loop in a potential stacking interaction. The tip of M4 was not well-resolved in the 5-HT3AR crystal structure and Trp459 was built only in two subunits16. Of the two, one of the Trp sidechains was in a similar stacking orientation. In nAChR, the interaction between the post-M4 and cys loop is predicted to modulate gating40.
Conformation of the ECD
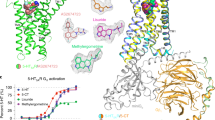
The neurotransmitter binding site has been extensively studied and a wealth of structural data is available from the acetylcholine binding protein (AChBP) and from the 5-HT3A-AChBP chimera bound to a battery of pLGIC agonists and antagonists24,41. The serotonin binding site is located at the subunit interface lined by residues from Loops A, B, and C on the (+) subunit and Loops D, E, and F from the (−) subunits42. There are clear densities for aromatic residues (Fig. 4a) from each of these loops lining the binding pocket, particularly Trp156-Loop B, Phe199 and Tyr207-Loop C, Trp63-Loop D, Tyr126-Loop E. Densities for Trp168, Arg169, and D177 in Loop F are also shown and are implicated in ligand binding. In comparison to the crystal structure, the β-sheets forming the ECD undergo a clockwise twist (viewed from the extracellular end) (Fig. 4b) leading to a widening of the ligand-binding pocket with rearrangement of the binding site residues (Fig. 4c). In addition, Loop C, which is at the periphery of the neurotransmitter binding pocket, in the apo-5-HT3AR structure is placed outward in an open or “extended” conformation. The Loop C conformation has been implicated to correlate both with the occupancy of the ligand at the neurotransmitter binding site and the functional state of the channel24. Particularly in AChBP, in comparison to the apo-state, in the antagonist-bound state, Loop C adopts an “open” conformation, while in the agonist-bound state it takes a “closed” conformation. A partly “closed” conformation of Loop C in the crystal structure is likely from its interaction with the nanobody bound in the vicinity. The orientation of Trp156 in Loop B, lining the back wall of the neurotransmitter binding pocket, in apo-5-HT3AR is flipped relative to its position in the 5-HT3AR crystal structure. This position is highly conserved as a Trp in 5-HT3AR, α-subunits of nAChRs, and AChBP, while it is a Tyr in the anionic members of pLGIC family. The indole ring of Trp is involved in a cation-π interaction with the quaternary ammonium ligands binding to the neurotransmitter binding pocket43. An alignment with AChBP and nAChR shows that the Trp156 orientation in apo-5-HT3AR is similar to the conformation seen in these structures (Supplementary Fig. 9). Further studies are necessary to understand the potential role of Trp156 re-orientation in channel 5-HT3AR inhibition.
The neurotransmitter binding site. a The map around the aromatic residues at the subunit interface that constitutes the neurotransmitter binding site (top). The map for the residues in Loop F that are involved in ligand binding (bottom). b Alignment of 5-HT3AR apo (salmon red) and crystal (pale green) structures reveals a twist and an expansion at the region lined Loop C, Loop B, and Loop F. The arrows indicate the direction of movement. c A comparison of the orientations of the residues that are involved in neurotransmitter binding
Conformation of the ICD
The ICD sequence is the least conserved region among the pLGIC family, and in human isoforms the length is always greater than 70 residues and varies up to 150 residues. In apo-5-HT3AR, the post-M3 region forms a loop that extends away from the pore axis and terminates in an α-helix referred to as MX helix. MX is a short helix which forms a belt around the MA helical bundle, lying parallel to the plane of the membrane, and is seen to bend downward toward the cytoplasmic region (Fig. 5a). Although we see densities for the entire MX and MA helices and for a short region of the unstructured portion from MX, the density for a stretch of 59 residues is missing until beginning of the MA helix, presumably due to the unstructured and flexible nature of this loop. Not surprisingly, this is also the region that is chipped away during trypsin digestion16. The apo-5-HT3AR structure suggests the presence of multiple intrasubunit interactions between MX, unstructured loop, and MA. The post-M3 loop is held in its position through potential polar interactions with M3 (Arg306) and M4 helix (Trp426) within the same subunit (Fig. 5b). Although in comparison to the 5-HT3AR crystal structure the post-M3 loop is positioned away from the pore axis (with minimal interaction with the neighboring subunit) (Fig. 5a), it still partially blocks the inter-subunit cavity or lateral portals which have been implicated to serve as conduits for ion permeation12.
The Intracellular domain. a The ICD is comprised of the post-M3 loop, the MX, helix, a stretch of unstructured region, followed by the MA helix. The density from the MX helix is bent downward to the intracellular end of the MA helix, but the unstructured region is not resolved. Superposition of the 5-HT3AR apo and crystal structures reveals differences in the conformation of ICD in the two structures. The (−) subunit of the two structures are aligned. The expansion of the ICD resulting from an outward displacement of MA and MX helices are indicated by arrows. b The residues within the ICD involved in several potential intra and inter-subunit interactions. c The solvent-accessible electrostatic potential map generated using the APBS tool. The inset shows a zoomed-in view of the lipid-binding pocket within the dotted green box. The lipid (partially built phospholipid) and the interacting residues (R306 and R435) are shown in stick. The map around the lipid is shown as blue mesh
The ICD reveals a notable constriction to 2.1 Å at the level of Arg416 with a strong density (6 σ) at the center of the pore axis, which we modeled as a Cl− (Fig. 3c and Supplementary Fig. 5). Interestingly, previous studies have suggested that intracellular phosphates interact with MA arginine residues near the cytoplasmic mouth of the channel facilitating pore block and inhibition of ion conduction44. Besides the constriction at Arg416, the MA helix offers a continuous conduit of radius over 3 Å. Serotonin-mediated activation, therefore, likely evokes a conformational change within the ICD to allow permeation. 5-HT3AR has an unusually low conductance which is well below the resolution of single-channel recordings. The Arg416 residue is a part of a cluster of arginine residues in the MA helix that are shown to be a critical determinant of 5-HT3AR conductance and mutations at these positions lead to significant increases in single-channel conductance21. In contrast to the tightly packed arrangement of MA helices in the crystal structure with an obstruction at the hydrophobic patch at the C-terminal end of the helix lined by Leu402, Leu406, Ile409, and Leu413, the apo-5-HT3AR structure reveals a more loose packing of the MA helices with the hydrophobic residues positioned farther away from the pore axis. This finding is in agreement with mutagenesis studies that suggest that this lower region has a lesser impact on channel conductance in comparison to the charged cluster further up45.
The electrostatic potential map along the ion permeation pathway shows that in contrast to the ECD and the TMD, where the surface charge is electronegative (lined by Asp52, Glu53 (β1-β2 loop), and Glu186 (β8–β9 loop) in the ECD and Glu-2′ and Asp-4′ (M2) in the TMD), the ICD has a cluster of positively charged residues (highlighted by the blue region in Fig. 5c). This is a unique feature of 5-HT3AR among the cation-selective pLGICs. At the edge of the TMD–ICD interface, there is a density present in the cavity lined by M3 of one subunit and M1 and M4 helices of the neighboring subunit. This density was partially modeled as a phospholipid based on putative polar contacts with neighboring Arg sidechains (Arg306 in M3 and Arg435 in M4). Interestingly, a recently solved structure of the GABA (β3–α5) chimeric channel reveals a neurosteroid binding site that aligns with the lipid site suggesting that this lipid pocket may be involved in allosteric modulation of pLGICs46.
Discussion
The cryo-EM structure of apo-5-HT3AR determined in the solution state, in the absence of inhibitors, Fab fragments, truncation, or constraints from crystal lattice, bears the hallmarks of the resting conformation. In comparison to the 5-HT3AR crystal structure, the full-length channel adopts an expanded conformation with a clockwise twist in the β-strands of the ECD and the Loop C in an extended position. In addition, the locations of constriction that occlude ion permeation in the resting conformation can be inferred. The pore-lining region reveals non-protein densities, the arrangement of which suggests that they correspond to ions and water molecules. Further, the structure suggests that the ICD is involved in several inter-subunit interactions that may underscore its role in signal transduction across the channel. If the crystal structure and the apo-5-HT3AR structure were to truly represent physiologically inhibited and resting conformations, respectively, the conformational differences in the ECD, TMD, and ICD provide insights into mechanisms underlying channel modulation. The apo-5-HT3AR structure serves as a starting conformation to determine ligand-induced conformational changes leading to open and desensitized states.
Methods
5-HT3AR cloning and electrophysiological measurements in oocytes
The gene encoding 5-HT3AR (purchased from GenScript USA Inc.) was inserted into a Xenopus laevis oocyte expression vector (pTLN) and confirmed by DNA sequencing. This construct was first linearized with Mlu1 restriction enzyme overnight at 37 °C and then used for mRNA synthesis using the mMessage mMachine kit (Ambion). The mRNA was purified with RNAeasy kit (Qiagen) and stored at −20 °C. For measurement of 5-HT3AR macroscopic currents, Xenopus laevis oocytes (stages V–VI) were injected with 3–10 ng of mRNA. As a control, oocytes were injected with the corresponding volume of water to verify endogenous currents were not present. The oocytes used in this study were kindly provided by Dr. Walter F. Boron. Female X. laevis were purchased from Nasco. All animal experimental procedures were approved by Institutional Animal Care and Use Committee (IACUC) of Case Western Reserve University. Both sets of injected oocytes were maintained at 18 °C in OR3 media (GIBCO BRL Leibovitz media containing glutamate and supplemented with 500 units each of penicillin and streptomycin, pH 7.5. The osmolarity was adjusted to 197 mOsm). After 2–5 days of injection, two electrode voltage-clamp (TEVC) experiments were performed at room temperature on a Warner Instruments Oocyte clamp OC-725. Oocytes were clamped at −60 mV holding potential, and macroscopic current traces were recorded in response to the application of serotonin hydrochloride (at indicated concentrations). The solution-exchange was performed using a syringe-pump perfusion system operating at a flow-rate of 6 ml/min. The current was sampled and digitized at 500 Hz with a Digidata 1332A. The traces were analyzed by Clampfit 10.2 (Molecular Devices). The electrophysiological solutions had the following buffer composition: 96 mM NaCl, 2 mM KCl, 1.8 mM CaCl2, 1 mM MgCl2, and 5 mM HEPES (osmolarity adjusted to 195 mOsm, pH 7.4). All chemical reagents used in these experiments were purchased from Sigma-Aldrich.
Cloning and transfection
Codon-optimized mouse 5-HT3AR gene (NCBI Reference Sequence: NM_001099644.1) was purchased from GenScript USA Inc. Subsequently, it was subcloned into pFastBac1 vector containing four Strep-tags (WSHPQFEK) at the N-terminus, followed by a linker sequence (GGGSGGGSGGGS) and a TEV-cleavage sequence (ENLYFQG). The construct also includes a C-terminal 1D4-tag47. S. frugiperda (Sf9) cells (purchased from Expression System) were cultured in ESF921 medium (Expression System) and incubated at 28 °C without CO2 exchange. The culture media did not contain antibiotics. Sub-confluent cells were then transfected with recombinant 5-HT3AR bacmid DNA using Cellfectin II transfection reagent (Invitrogen) using manufacturer recommended instructions. At 72 h post-transfection, the cell-culture supernatant was collected and centrifuged at 1000×g for 5 min to remove cell debris and harvest progeny 1 (P1) recombinant baculovirus. The P2 virus was obtained through a consecutive round of Sf9 cells infection with the P1 virus stock. The supernatant containing P2 virus was then used to infect Sf9 cells, thus generating P3 virus. The level of protein expression was checked by western blot using both P2 and P3 viruses. The P3 virus was used for recombinant protein production and analysis.
Expression and purification of recombinant protein
The recombinant 5-HT3AR protein production was carried out by infection of approximately 2.5 × 106/ml Sf9 cells with P3 virus. The cells were harvested at 72 h post-infection and centrifuged at 8000×g for 20 min at 4 °C to separate the supernatant from the cell pellet. The cells were then re-suspended in a buffer containing 20 mM Tris-HCl and 36.5 mM sucrose at pH 7.5 and supplemented with 1% protease inhibitor cocktail (Sigma-Aldrich). The cells were disrupted by sonication on ice and the non-lysed cells were removed by centrifugation (3000×g for 15 min). The membrane fraction was separated by ultracentrifugation (167,000×g for 1 h) and solubilized with 1% C12E9 (Anatrace) in a buffer containing 500 mM NaCl, 50 mM Tris pH 7.4, 10% glycerol, and 0.5% protease inhibitor by rotating for 2 h at 4 °C. Non-solubilized material was removed by ultracentrifugation (167,000×g and 15 min). The supernatant was collected and bound with 1D4 beads equilibrated with 150 mM NaCl, 20 mM HEPES pH 8.0, and 0.01% C12E9 for 2 h at 4 °C. The beads were then washed with 100 column volumes of 150 mM NaCl, 20 mM Hepes pH 8.0, and 0.01% C12E9 (Buffer A). The protein was then eluted with Buffer A supplemented with 3 mg/ml 1D4 peptide (NH2-TETSQVAPA-CO2H). Eluted protein was then concentrated and deglycosylated with PNGase F (NEB) by incubating 5 units of the enzyme per 1 μg of the protein for 2 h at 37 °C under gentle agitation. Deglycosylated protein was then applied to a Superose 6 column (GE healthcare) equilibrated with Buffer A. The peak fractions around 13.9 ml were pooled and concentrated to 2–3 mg/ml using 50 kDa MWCO Millipore filters (Amicon) and used subsequently for cryo-EM studies.
Sample preparation and cryo-EM data acquisition
The 5-HT3AR protein (~2.5 mg/ml) was filtered and incubated with 3 mM Fluorinated Fos-choline 8 (Anatrace) to improve particle distribution48. The sample was double blotted (3.5 µl per blot) onto Cu 300 mesh Quantifoil 1.2/1.3 grids (Quantifoil Micro Tools), and immediately after the second blot, the grid was plunge frozen using a Vitrobot (FEI). The grids were imaged on a Titan Krios microscope (FEI), operating at 300 kV, and equipped with a K2-Summit direct detector camera (Gatan). 40-frame movies were collected at 130,000× magnification (set on microscope) in super-resolution mode with a physical pixel size of 0.532 Å/pixel. The dose rate was 4 electrons/pixel/second, with a total exposure time of 12 s. The defocus values ranged from −0.75 µm to −2.5 µm (input range setting for data collection) as per the automated imaging software49.
Image processing
Movies were motion-corrected to compensate for the beam-induced motion using MotionCor250 with a B-factor of 150 pixels2. Super-resolution counting images were 2 × 2 binned in Fourier space with a pixel size of 1.064 Å. All subsequent data processing was performed using RELION 2.0351. The defocus values of the motion-corrected micrographs were estimated using Gctf software52. Approximately, 2000 particles were manually picked from the 3550 micrographs and sorted into two-dimensional (2D) classes. The best of these classes were then used as templates for auto-picking. A loose auto-picking threshold was selected to ensure no good particles were missed. This resulted in ~327,000 auto-picked particles that were subjected to multiple rounds of 2D classification to remove suboptimal particles. An initial three-dimensional (3D) model was generated from 5-HT3AR crystal structure (PDB code: 4PIR) and low-pass filtered to 60 Å using EMAN253.The best ~117,000 particles were then subjected to 3D auto-refinement, followed by 3D classification into three classes. The best 3D class, containing ~108,000 particles, was subjected to a final round of 3D auto-refinement and post-processing to yield a 5-HT3AR map at an overall resolution of 4.3 Å (calculated based on the gold-standard Fourier shell coefficient (FSC) = 0.143 criterion). In the RELION post-processing step, a soft mask was applied to the two half-maps before the FSC was calculated. The post-processing step also included B-factor estimation and map sharpening. ResMap software54 was used for estimation of local resolutions.
5-HT3AR model building
The 5-HT3AR crystal structure (PDB-ID: 4PIR)16 was used as an initial model and aligned to the 5-HT3AR cryo-EM map calculated with RELION 2.03. The cryo-EM map was converted to an .mtz format using mapmask and sfall tools in CCP4i software55. The mtz map was then used for model building in COOT56. In comparison to the 5-HT3AR crystal structure, for each monomer, 15 additional residues were built in our model and sidechains were built for seven additional residues. After initial model building, the model was refined against the EM-derived maps using the phenix.real_space_refinement tool from the PHENIX software package57, employing rigid body, local grid, NCS, and gradient minimization. This model was then subjected to additional rounds of manual model-fitting and refinement which resulted in a final model-to-map cross-correlation coefficient of 0.774. Stereochemical properties of the model were evaluated by Molprobity58.
Protein surface area and interfaces were analyzed by using PDBePISA server (http://www.ebi.ac.uk/pdbe/pisa/). To compare apo-5-HT3AR and the 5-HT3AR crystal structure, all ligands, ions, water molecules, and nanobodies (in the crystal structure) were removed from the PDB files. Additional residues in the apo-5-HT3AR structure were also removed before analysis so that surface area comparisons were made between identical construct lengths. Electrostatic surface potential calculations were carried out using the APBS tools plug-in PyMOL59. The pore profile was calculated using the HOLE program60.
Data availability
The coordinates of the 5-HT3AR structure and the cryo-EM map have been deposited under PDB ID 6BE1 and EMD-7088 with the wwPDB and EMDB. Data supporting the findings of this manuscript are available from the corresponding author upon reasonable request.
References
Maricq, A. V., Peterson, A. S., Brake, A. J., Myers, R. M. & Julius, D. Primary structure and functional expression of the 5HT3 receptor, a serotonin-gated ion channel. Science 254, 432–437 (1991).
Lummis, S. C. 5-HT(3) receptors. J. Biol. Chem. 287, 40239–40245 (2012).
Niesler, B., Frank, B., Kapeller, J. & Rappold, G. A. Cloning, physical mapping and expression analysis of the human 5-HT3 serotonin receptor-like genes HTR3C, HTR3D and HTR3E. Gene 310, 101–111 (2003).
Engel, M., Smidt, M. P. & van Hooft, J. A. The serotonin 5-HT3 receptor: a novel neurodevelopmental target. Front. Cell Neurosci. 7, 76 (2013).
Gehlert, D. R., Gackenheimer, S. L., Wong, D. T. & Robertson, D. W. Localization of 5-HT3 receptors in the rat brain using [3H]LY278584. Brain Res. 553, 149–154 (1991).
Gershon, M. D. Review article: serotonin receptors and transporters -- roles in normal and abnormal gastrointestinal motility. Aliment. Pharmacol. Ther. 20(Suppl 7), 3–14 (2004).
Costall, B. & Naylor, R. J. 5-HT3 receptors. Curr. Drug Targets CNS Neurol. Disord. 3, 27–37 (2004).
Perwitasari, D. A. et al. Anti-emetic drugs in oncology: pharmacology and individualization by pharmacogenetics. Int. J. Clin. Pharm. 33, 33–43 (2011).
Kato, S. Role of serotonin 5-HT(3) receptors in intestinal inflammation. Biol. Pharm. Bull. 36, 1406–1409 (2013).
Spiller, R. C. Targeting the 5-HT(3) receptor in the treatment of irritable bowel syndrome. Curr. Opin. Pharmacol. 11, 68–74 (2011).
Hilf, R. J. & Dutzler, R. X-ray structure of a prokaryotic pentameric ligand-gated ion channel. Nature 452, 375–379 (2008).
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4A resolution. J. Mol. Biol. 346, 967–989 (2005).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. X-ray structure of the human alpha4beta2 nicotinic receptor. Nature 538, 411–415 (2016).
Hibbs, R. E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011).
Du, J., Lu, W., Wu, S., Cheng, Y. & Gouaux, E. Glycine receptor mechanism elucidated by electron cryo-microscopy. Nature 526, 224–229 (2015).
Hassaine, G. et al. X-ray structure of the mouse serotonin 5-HT3 receptor. Nature 512, 276–281 (2014).
Miller, P. S. & Aricescu, A. R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Basak, S., Schmandt, N., Gicheru, Y. & Chakrapani, S. Crystal structure and dynamics of a lipid-induced potential desensitized-state of a pentameric ligand-gated channel. Elife 6, e23886 (2017).
Sauguet, L. et al. Crystal structures of a pentameric ligand-gated ion channel provide a mechanism for activation. Proc. Natl Acad. Sci. USA 111, 966–971 (2014).
Connolly, C. N. Trafficking of 5-HT(3) and GABA(A) receptors (review). Mol. Membr. Biol. 25, 293–301 (2008).
Peters, J. A. et al. The 5-hydroxytryptamine type 3 (5-HT3) receptor reveals a novel determinant of single-channel conductance. Biochem. Soc. Trans. 32, 547–552 (2004).
Baptista-Hon, D. T., Deeb, T. Z., Lambert, J. J., Peters, J. A. & Hales, T. G. The minimum M3-M4 loop length of neurotransmitter-activated pentameric receptors is critical for the structural integrity of cytoplasmic portals. J. Biol. Chem. 288, 21558–21568 (2013).
Pless, S. A. & Lynch, J. W. Ligand-specific conformational changes in the alpha1 glycine receptor ligand-binding domain. J. Biol. Chem. 284, 15847–15856 (2009).
Hansen, S. B. Structures of Aplysia AChBP complexes with nicotinic agonists and antagonists reveal distinctive binding interfaces and conformations. EMBO J. 24, 3635–3646 (2005).
Chang, Y. & Weiss, D. S. Site-specific fluorescence reveals distinct structural changes with GABA receptor activation and antagonism. Nat. Neurosci. 5, 1163–1168 (2002).
Brams, M. et al. A structural and mutagenic blueprint for molecular recognition of strychnine and d-tubocurarine by different cys-loop receptors. PLoS Biol. 9, e1001034 (2011).
Yu, R. et al. Agonist and antagonist binding in human glycine receptors. Biochemistry 53, 6041–6051 (2014).
Davies, P. A. et al. The 5-HT3B subunit is a major determinant of serotonin-receptor function. Nature 397, 359–363 (1999).
Monk, S. A., Williams, J. M., Hope, A. G. & Barnes, N. M. Identification and importance of N-glycosylation of the human 5-hydroxytryptamine3A receptor subunit. Biochem. Pharmacol. 68, 1787–1796 (2004).
Althoff, T., Hibbs, R. E., Banerjee, S. & Gouaux, E. X-ray structures of GluCl in apo states reveal a gating mechanism of Cys-loop receptors. Nature 512, 333–337 (2014).
Livesey, M. R., Cooper, M. A., Lambert, J. J. & Peters, J. A. Rings of charge within the extracellular vestibule influence ion permeation of the 5-HT3A receptor. J. Biol. Chem. 286, 16008–16017 (2011).
Sine, S. M., Wang, H. L., Hansen, S. & Taylor, P. On the origin of ion selectivity in the Cys-loop receptor family. J. Mol. Neurosci. 40, 70–76 (2010).
Lee, J. & Kim, S. H. Water polygons in high-resolution protein crystal structures. Protein Sci. 18, 1370–1376 (2009).
Sauguet, L. et al. Structural basis for ion permeation mechanism in pentameric ligand-gated ion channels. EMBO J. 32, 728–741 (2013).
Marcus, Y. Ionic radii in aqueous solution. Chem. Rev. 88, 1475–1498 (1988).
Thompson, A. J. & Lummis, S. C. A single ring of charged amino acids at one end of the pore can control ion selectivity in the 5-HT3 receptor. Br. J. Pharmacol. 140, 359–365 (2003).
Panicker, S., Cruz, H., Arrabit, C. & Slesinger, P. A. Evidence for a centrally located gate in the pore of a serotonin-gated ion channel. J. Neurosci. 22, 1629–1639 (2002).
Lee, W. Y., Free, C. R. & Sine, S. M. Nicotinic receptor interloop proline anchors beta1-beta2 and Cys loops in coupling agonist binding to channel gating. J. Gen. Physiol. 132, 265–278 (2008).
Grosman, C., Salamone, F. N., Sine, S. M. & Auerbach, A. The extracellular linker of muscle acetylcholine receptor channels is a gating control element. J. Gen. Physiol. 116, 327–340 (2000).
daCosta, C. J., Dey, L., Therien, J. P. & Baenziger, J. E. A distinct mechanism for activating uncoupled nicotinic acetylcholine receptors. Nat. Chem. Biol. 9, 701–707 (2013).
Price, K. L., Lillestol, R. K., Ulens, C. & Lummis, S. C. Varenicline interactions at the 5-HT3 receptor ligand binding site are revealed by 5-HTBP. ACS Chem. Neurosci. 6, 1151–1157 (2015).
Brejc, K. et al. Crystal structure of an ACh-binding protein reveals the ligand-binding domain of nicotinic receptors. Nature 411, 269–276 (2001).
Zhong, W. et al. From ab initio quantum mechanics to molecular neurobiology: a cation-pi binding site in the nicotinic receptor. Proc. Natl Acad. Sci. USA 95, 12088–12093 (1998).
Noam, Y., Wadman, W. J. & van Hooft, J. A. On the voltage-dependent Ca2+block of serotonin 5-HT3 receptors: a critical role of intracellular phosphates. J. Physiol. 586, 3629–3638 (2008).
Carland, J. E. et al. Mutagenic analysis of the intracellular portals of the human 5-HT3A receptor. J. Biol. Chem. 288, 31592–31601 (2013).
Miller, P. S. et al. Structural basis for GABAA receptor potentiation by neurosteroids. Nat. Struct. Mol. Biol. 24, 986–992 (2017).
MacKenzie, D., Arendt, A., Hargrave, P., McDowell, J. H. & Molday, R. S. Localization of binding sites for carboxyl terminal specific anti-rhodopsin monoclonal antibodies using synthetic peptides. Biochemistry 23, 6544–6549 (1984).
Zhang, Z. & Chen, J. Atomic structure of the cystic fibrosis transmembrane conductance regulator. Cell 167, 1586–1597 e1589 (2016).
Potter, C. S. et al. Leginon: a system for fully automated acquisition of 1000 electron micrographs a day. Ultramicroscopy 77, 153–161 (1999).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Scheres, S. H. Processing of structurally heterogeneous cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016).
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Collaborative Computational Project, N The CCP4 suite: programs for protein crystallography. Acta crystallographica. Section D, Biological crystallography 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Chen, V. B. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 376 (1996).
Acknowledgements
We acknowledge the imaging time and the use of instruments at the National Cryo-Electron Microscopy Facility at the NCI. We are grateful to the Cleveland Center for Membrane and Structural Biology for providing the access to the Cryo-EM instrumentation. We thank Denice Major for assistance with hybridoma and cell culture at the Department of Ophthalmology and Visual Sciences (supported by the National Institutes of Health Core Grant P30EY11373). We thank Dr. Walter F. Boron for kindly providing us with Xenopus oocytes and for unrestricted access of the oocyte rig. We are very grateful to the members of the Chakrapani lab for critical reading and comments on the manuscript. This research was, in part, supported by the National Cancer Institute’s National Cryo-EM Facility at the Frederick National Laboratory for Cancer Research. This work was supported by the National Institutes of Health grant (1R01GM108921), the Cryo-EM supplement (3R01GM108921-03S1) to S.C., the AHA postdoctoral Fellowship to S.B. (17POST33671152), and the National Institutes of Health grant (1R01GM103899) to V.M.-B.
Author information
Authors and Affiliations
Contributions
S.B. and S.C. conceived the project and designed the experimental procedures. S.B. purified the protein and carried out all aspects of cryo-EM sample preparation, screening, data analysis, model building, and refinement. Y.G. performed the two-electrode voltage-clamp recordings. A.S. and V.Y.M.-B. trained S.B. in freezing samples and provided guidance to optimize grid preparation. S.M. trained S.B. and S.C. in the use of Tecnai TF20 microscope for screening grids. M.F. and M.N. trained S.B. in insect cell expression. W.H., T.H., D.J.T., and V.Y.M.-B. provided guidance at various stages of cryo-EM analysis. S.C. supervised the execution of the experiments, data analysis, and interpretation. S.B. and S.C. drafted the manuscript. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Basak, S., Gicheru, Y., Samanta, A. et al. Cryo-EM structure of 5-HT3A receptor in its resting conformation. Nat Commun 9, 514 (2018). https://doi.org/10.1038/s41467-018-02997-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-02997-4
This article is cited by
-
Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment
Nature Communications (2024)
-
Structural switch in acetylcholine receptors in developing muscle
Nature (2024)
-
Structural basis for partial agonism in 5-HT3A receptors
Nature Structural & Molecular Biology (2024)
-
Structure of tetrameric forms of the serotonin-gated 5-HT3A receptor ion channel
The EMBO Journal (2024)
-
Structural determinants for activity of the antidepressant vortioxetine at human and rodent 5-HT3 receptors
Nature Structural & Molecular Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.