Abstract
Circulating CCR2+ monocytes are crucial for maintaining the adult tissue-resident F4/80hiMHCIIhi macrophage pool in the intestinal lamina propria. Here we show that a subpopulation of CCR2-independent F4/80hiMHCIIlow macrophages, which are the most abundant F4/80hi cells in neonates, gradually decline in number in adulthood; these macrophages likely represent the fetal contribution to F4/80hi cells. In colon adenomas of ApcMin/+ mice, F4/80hiMHCIIlow macrophages are not only preserved, but become the dominant subpopulation among tumour-resident macrophages during tumour progression. Furthermore, these pro-tumoural F4/80hiMHCIIlow and F4/80hiMHCIIhi macrophages can self-renew in the tumour and maintain their numbers mostly independent from bone marrow contribution. Analyses of colon adenomas indicate that CSF1 may be a key facilitator of macrophage self-renewal. In summary, the tumour microenvironment creates an isolated niche for tissue-resident macrophages that favours macrophage survival and self-renewal.
Similar content being viewed by others
Introduction
Macrophages comprise a heterogeneous population of tissue-resident immune cells that contribute to tissue homeostasis, support the host defence system and can impact on the initiation and propagation of several diseases, including cancer (reviewed in refs.1,2). The majority of tissue-resident macrophages are established prenatally3,4,5 and are mainly independent from any further haematopoietic input due to their capacity to self-renew in situ to maintain population size6,7,8,9,10. For example, microglia, the resident macrophage population of the central nervous system, develop early during embryonic development from yolk sac precursors and effectively self-maintain in situ throughout adulthood5,6,11. Although the majority of the tissue-resident macrophages are maintained independently from bone marrow (BM)-derived monocytes, numerous studies have demonstrated that macrophages of certain tissues, including the dermis5,12, mammary gland13, heart14, pancreas15 and intestine5,16,17, require the input of monocytes to retain their pool during adulthood. In particular, the intestinal macrophages depend on the constant and fast replenishment of circulating blood Ly6Chi monocytes, not only during inflammation but also under normal healthy conditions16,17,18. In a healthy unperturbed colon, monocytes attracted to the intestine gradually differentiate into tissue-resident macrophages by losing Ly6C expression, up-regulating the expression of macrophage markers, such as CX3CR1, F4/80, CD64 and CD11c16,19, and secreting or responding to the anti-inflammatory cytokine interleukin-10 20,21. Mice lacking either the chemokine receptor CCR2 or its ligand CCL2 have reduced numbers of intestinal macrophages17, thus implying that the homeostatic recruitment of macrophage precursors to the gut lamina propria (LP) may be dependent on a CCR2–CCL2 axis. Upon intestinal inflammation, tissue-resident macrophages are still derived from circulating monocytes, but convert from being anti-inflammatory macrophages to highly Toll-like receptor-responsive inflammatory cells16.
Tissue-resident macrophages are also naturally present in various tumours, including lung and mammary tumours where macrophages are monocyte-derived22,23 or even self-renewing24. The developmental origins and the maintenance kinetics of resident macrophages in intestinal tumours, however, have not been studied thus far.
Here we show that the colon LP of normal adult mice contains a major population of F4/80hiMHCIIhi macrophages and a population of gradually disappearing F4/80hiMHCIIlow cells. Both F4/80hi macrophage populations are relatively slowly replenished by BM-derived cells, particularly MHCIIlow cells, compared to F4/80int monocyte/macrophage cells that have a fast turnover. The tumour microenvironment, however, is enriched with F4/80hiMHCIIlow macrophages. Furthermore, while F4/80hiMHCIIlow macrophages do not require CCR2+ monocytes for their maintenance in the healthy gut LP or in tumours, we found that F4/80hiMHCIIhi macrophages become independent from CCR2+ monocytes in tumours only, potentially due to tumour-derived colony stimulating factor-1 (CSF1) that supports macrophage self-renewal. Moreover, we demonstrate that CCR2-independent intratumoural F4/80hi macrophages, and not the CCR2-dependent monocytes or monocyte-derived macrophages, support cancer progression, suggesting this particular F4/80hi cell fraction as an attractive therapeutic target.
Results
Skewed representation of myeloid cells in colon adenomas
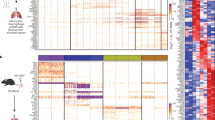
We first profiled colon LP myeloid subpopulations from wild-type (WT) C57BL/6J mice, previously treated with dextran sodium sulphate (DSS), and compared them with their tumoural counterparts obtained from the polyps collected from DSS-treated ApcMin/+ mice (Fig. 1). Similar as previously reported5, colon LP CD45+ cells comprised three major myeloid cell fractions (I–III) after excluding CD11chiMHCIIhi dendritic cells: F4/80hiCD11b+ tissue-resident (tumour-resident in polyps) macrophages (fraction I), F4/80intCD11b+ (fraction II) and F4/80lowCD11b+ neutrophils (fraction III) (Fig. 1a). The majority (>85%) of fraction I cells expressed high levels of major histocompatibility complex (MHC) class II and only a small proportion (~5%) of cells were low for MHC class II expression. Fraction II could be further subdivided into four distinct subpopulations in a waterfall-shaped distribution on a Ly6C vs. MHCII dot plot consisting of three distinct monocyte-differentiation stages (P1: Ly6ChiMHCII−; P2: Ly6ChiMHCII+; P3: Ly6C−MHCII+) and one eosinophil fraction (Ly6C−MHCII−) (Fig. 1a). We then analysed the presence of these subpopulations in DSS-accelerated colon tumours from ApcMin/+ mice. The same LP populations were also identified in colon polyps but with markedly different frequencies (Fig. 1b). In these tumours, neutrophils (fraction III) were the most abundant myeloid cell type (>40%) followed by monocyte-derived cells (fraction II) and tumour-resident macrophages (fraction I). Interestingly, eosinophils in tumours represented only a minor fraction (~3%) of the total CD11b+ cells. Furthermore, tumour-resident macrophages (fraction I) were clearly enriched in MHCIIlow cells (Fig. 1a,b), which were almost neglectable in normal adult colon LP.
Myeloid cell heterogeneity in colon lamina propria and colon tumours. WT C57BL/6J and ApcMin/+ mice aged 6 weeks old were administered drinking water supplemented with dextran sodium sulphate for 7 days, and analysed 4 weeks later. a Flow cytometry representative dot plots of colon lamina propria (LP) (upper panel) and tumour cell subpopulations (lower panel). Three different myeloid fractions (I–III) are defined by the differential expression of F4/80. Fraction I represents F4/80hi tissue-resident macrophages, which can be further subdivided into MHCIIhi and MHCIIlow. Fraction II contains monocytes (P1), two monocyte-derived macrophage subpopulations (P2 and P3) and eosinophils (eos), based on differential expression of MHCII and Ly6C. Fr III consists of neutrophils. Gating strategy is shown in Supplementary Fig. 4. b Bar charts of the distinct myeloid cell subpopulations obtained from the colon LP (n = 8) (upper bar chart) and 2–3 mm colon tumours (n = 20) (lower bar chart). White bars: fraction I; light blue bars: fraction II and black bars: fraction III. Error bars represent the s.e.m. c Pie charts show the proportions of F4/80hi tissue-resident macrophages (MHCIIhi and MHCIIlow), monocytes (P1), monocyte-derived macrophages (P2–P3), neutrophils and eosinophils across colon LP and tumours
Characterization of myeloid subpopulations in the colon LP of untreated C57BL/6J mice and in “spontaneously” formed tumours obtained from 5-month-old ApcMin/+ mice confirmed the presence of all myeloid subpopulations described in Fig. 1, although with a clear increase in the proportion of the tumoural CD11b+F4/80− neutrophil fraction (up to 70% of total CD11b+ cells) (Supplementary Fig. 1).
MHCIIlow macrophages accumulate during tumour progression
A previous study reported that F4/80hi tissue-resident macrophages in the colon LP progressively disappear with increasing age17. We therefore analysed the myeloid cell subsets present in the fetal colon (embryonic day 19.5) and compared them to those present in the colon LP of young and old mice (from 1 week to 12 months). In contrast to Bain et al.,17 our analysis clearly found that all three macrophage fractions (I–III) were maintained across all age groups with only minor fluctuations detected in their frequency (Fig. 2a). The predominant tissue-resident macrophage subset in the fetal colon was the F4/80hiMHCIIlow fraction. However, as a function of age this macrophage subset almost disappeared to leave mainly F4/80hiMHCIIhi cells to represent the colon tissue-resident macrophages (fraction I) (Fig. 2b). This decrease in MHCIIlow cell representation suggests that the LP environment gradually changes with age, perhaps due to the establishment of a mature microbiota and/or a tonic increase in inflammatory mediators17.
Ageing and tumour progression influence the ratio between F4/80hiMHCIIhi and F4/80hiMHCIIlow subpopulations. a F4/80hi tissue-resident macrophages persist in the colon LP after birth into adulthood, as shown by the representative flow cytometry dot plots and bar chart. Error bars represent the s.e.m. b The frequency of colon F4/80hiMHCIIlow cells rapidly declines after birth. Representative dot plots of F4/80hiMHCIIhi and F4/80hiMHCIIlow subpopulations obtained from fetal colon (E19.5) and from colons of mice aged 2 days, 1, 3, 4, 6 and 8 weeks and 12 months. The bar chart represents the age-dependent ratio between F4/80hiMHCIIhi (blue) and F4/80hiMHCIIlow (red) subpopulations in the colon LP. E19.5: n = 1 (pool of 15 embryos obtained from 3 different pregnant mice); day 2: n = 2 (each group pool of 5 mice); 1 week: n = 2 (each group pool of 5 mice); 3 weeks: n = 6 mice; 4 weeks: n = 5 mice; 6 weeks: n = 4; 8 weeks: n = 7 and 12 months: n = 4 mice. Error bars represent the s.e.m. c Representative flow cytometry analysis (left panel) and mean percentage of F4/80hiMHCIIhi and F4/80hiMHCIIlow subpopulations (right panel) in tumours of different sizes (0.5–6.0 mm in diameter) obtained from ApcMin/+ mice. 0.5 mm: n = 3; 1 mm: n = 3; 2 mm: n = 4; 3–4 mm: n = 3; 5–6 mm: n = 4. Error bars represent the s.e.m. Scale bars: 2.5 mm. Gating strategy is shown in Supplementary Fig. 4
Given the high frequency of F4/80hiMHCIIlow cells in colon polyps (Fig. 1b), we assessed whether there was an association between tumour progression and the frequency of MHCIIlow macrophages. Therefore, for this analysis we opted for the spontaneous tumour model of ApcMin/+ mice, which generate progressively adenomas up to 5–6 mm in size that is rarely achieved in the DSS-accelerated tumour model. Indeed, we observed an increasing abundance of this cell fraction in conjunction with tumour progression until MHCIIlow macrophages became a dominant cell type among tumour-resident macrophages (Fig. 2c). This observation clearly suggests that the tumour microenvironment favours the differentiation or maintenance of MHCIIlow macrophages.
Intratumoural macrophages are independent from CCR2+ monocytes
Given that circulating Ly6Chi monocytes express high levels of CCR2 receptor25, we decided to examine the presence of this receptor on tissue-resident and monocyte-derived macrophages in colon LP and adenoma polyps. The aim of this experiment was to determine the potential origins of these cells. In the LP, consistent with being descendants of classical monocytes, Ly6ChiMHCII− (P1), Ly6ChiMHCIIhi (P2) and Ly6C−MHCII+ (P3) cells in fraction II subset had the highest and most homogeneous expression of CCR2 (Fig. 3a)17, whereas tissue-resident F4/80hiMHCIIhi and F4/80hiMHCIIlow macrophages were highly heterogeneous in their CCR2 expression levels, from very high to very low levels (Fig. 3a and Supplementary Fig. 2). As expected, numbers of monocytes and monocyte-derived cells (P1, P2 and P3) in fraction II were severely diminished in Ccr2−/− mice (Fig. 3b). In addition, MHCIIhi tissue-resident macrophages were also mostly CCR2 dependent, whereas MHCIIlow cells were not, which is likely a reflection of their markedly lower CCR2 expression level (Fig. 3b).
Lamina propria and intratumoural colon F4/80hiCD11b+ macrophage subsets show differential CCR2-driven monocyte dependence. a Fluorescence-activated cell sorting histograms showing CCR2 expression profiles on distinct colon LP and intratumoural myeloid cell subpopulations: fraction I consists of MHCIIlow and MHCIIhi cells, and fraction II consists of monocytes (P1) and monocyte-derived macrophages (P2 and P3). b Myeloid cell profiling in WT and Ccr2−/− colon LP. Representative flow cytometry analysis with F4/80 and CD11b-expressing myeloid subpopulations, obtained from the colon LP of WT and Ccr2−/− mice. Fractions I and II were further dissected for MHCII and Ly6C expression (left panel). The absolute numbers of eosinophils, F4/80hiMHCIIhi and F4/80hiMHCIIlow tissue-resident macrophages and P1–P3 subpopulations are shown (right panel). The bar chart represents the mean number of mice in each group and the error bars represent the s.e.m. (WT; n = 7 and Ccr2−/−; n = 14). c Myeloid cell profiling in WT and Ccr2−/− colon tumours as described above for LP cells. Bar charts show the mean±s.e.m. of absolute numbers of eosinophils, F4/80hiMHCIIhi and F4/80hiMHCIIlow tissue-resident macrophages and P1–P3 subpopulations obtained from ApcMin/+ (n = 8) and ApcMin/+Ccr2−/− (n = 9) mice. Statistical significance was determined using an unpaired Student's t-test. **P<0.001; ***P < 0.0001; ns, not significant. Gating strategy is shown in Supplementary Fig. 4
Interestingly, we found that tumour-resident MHCIIhi and MHCIIlow macrophages both expressed relatively low levels of CCR2 compared to their LP counterparts and, consistently, their numbers did not change in Ccr2−/− mice. Conversely, fraction II cells (P1, P2 and P3) in polyps behaved exactly like their WT counterparts (Fig. 3a, c).
Taken together, these data assert that the tumour microenvironment can modify the properties of MHCIIhi tissue-resident macrophages such that they also become CCR2 independent like MHCIIlow cells in the LP and hence are presumably able to maintain themselves without monocyte replenishment.
Turnover of colon lamina propria myeloid cells
We utilized the KitMerCreMer/R26 fate mapping mouse5—where yellow florescence protein (YFP) expression can be induced in early BM progenitors—to monitor the cell population turnover rates driven by the BM input in colon LP myeloid cell populations. After injecting adult mice with tamoxifen, the labelling index of tissue-resident macrophages (F4/80hiMHCIIlow and MHCIIhi), monocytes (P1), monocyte-derived macrophages (P2–P3 fractions), eosinophils and neutrophils was analysed at different time points over a period of 5 months. As anticipated, all tested myeloid populations, with the exception of tissue-resident macrophages, were rapidly labelled with YFP and reached a plateau by 1–2 weeks after the final tamoxifen injection. These data suggest rapid replacement of myeloid cell populations by BM-derived cells (Fig. 4).
Intestinal tissue-resident macrophages exhibit a slower, gradual replacement by bone marrow-derived cells compared to other gut myeloid cells. a Schematic representation of the adult fate mapping protocol using KitMerCreMer/R26 mice. Mice aged 6 weeks were injected with tamoxifen five times and groups of 4–8 animals were sacrificed 1, 2, 3, 4, 7 and 20 weeks later. b Representative flow cytometry analysis indicating the labelling efficiency of distinct colon myeloid cell populations as defined in Fig. 1: MHCIIlow-expressing and MHCIIhi-expressing tissue-resident macrophages, monocytes and monocytes-derived macrophages (P1–P3), eosinophils and neutrophils. Neutrophils acted as internal controls for labelling efficiency and the tracings are from the same mouse. Gating strategy is shown in Supplementary Fig. 4. c The bar chart represents the mean percentage of yellow florescence protein-positive (YFP+) cells after normalization to the percentage of YFP+ neutrophils. The error bars represent the s.e.m.
Although it is widely considered that gut F4/80hiMHCIIhi cells lose their prenatal origin and are replaced by CCR2+ monocytes in adulthood16, our data clearly demonstrated that they are in fact replaced by BM-derived monocytes only very slowly, if at all. Our fate mapping data indicated that it took several months to fully replace F4/80hiMHCIIhi cells, whereas MHCIIlow macrophages exhibited minimal labelling (Fig. 4), suggesting that these tissue-resident macrophages retained their original fetal seed population for a long period.
Intratumoural resident macrophages expand by self-renewal
We noted that F4/80hiMHCIIlow tumour-resident macrophages in particular seemed to expand in polyps and thus wanted to determine if this effect was due to increased self-renewal or the recruitment of new cells. Again, our adult fate mapping analyses in KitMerCreMer/R26ApcMin/+ mice showed that the recruitment of new cells to DSS-accelerated adenomas was not increased and that the population turnover rate of F4/80hiMHCIIlow cells was comparable to that of the LP (Fig. 5a). A decrease in YFP labelling was observed in tumour-resident F4/80hiMHCIIhi cells when compared to their LP counterparts. This is probably due to the fact that the labelling index of macrophages in polyps was “frozen” to the level corresponding to the time for their entrapment in this niche that became independent from new inputs (Fig. 4 and Fig. 5a).
Identification of self-renewing colon macrophage subsets in healthy and tumour tissue. a Dextran sodium sulphate (DSS)-induced intestinal adenoma formation was combined with adult fate mapping using the KitMerCreMer/R26 and KitMerCreMer/R26ApcMin/+ mice (schematic representation shown in upper panel). Representative flow cytometry analysis illustrates the colon myeloid cell subset labelling efficiency. Gating strategy is shown in Supplementary Fig. 4. The bar chart represents the mean percentage of yellow fluorescence protein-positive (YFP+) cells after normalization to the percentage of YFP+ neutrophils±s.e.m. A total of 15 KitMerCreMer/R26ApcMin/+ and 20 KitMerCreMer/R26 6–8-week-old mice were subjected to DSS treatment. Statistical significance was determined by two-way ANOVA followed by Bonferroni test; ***P < 0.001. For reasons of clarity, the non-significant differences between groups were not indicated. b Embryonic fate mapping confirms the prenatal origins of intestinal LP and tumoural tissue-resident F4/80hiMHCIIhi and F4/80hiMHCIIlow macrophages. Pregnant KitMerCreMer/R26 and KitMerCreMer/R26ApcMin/+ mice (E10.5) were administered tamoxifen by intraperitoneal injection, as schematically illustrated in the upper panel. The offspring were then subjected to DSS treatment at 8 weeks old and sacrificed 4 weeks later for tissue and cell isolation. Representative flow cytometry analysis shows the labelling efficiency of each myeloid cell subset analyzed. Gating strategy is shown in Supplementary Fig. 4. The bar chart represents the mean percentage of YFP+ cells after normalization to the percentage of YFP+ neutrophils. The error bars represent the s.e.m. A total of 4 KitMerCreMer/R26ApcMin/+ and 6 KitMerCreMer/R26 mice were analysed. E, embryonic day; P1, monocytes; P2 and P3, monocyte-derived macrophages. Statistical significance was determined by two-way ANOVA followed by Bonferroni test. For reasons of clarity, the non-significant differences between groups were not indicated
Of note, when tamoxifen was administered during embryogenesis (Fig. 5b, upper panel), all myeloid cell subpopulations—including tissue-resident macrophages in the healthy LP and colon tumours—exhibited similar YFP tagging patterns (Fig. 5b, lower panel). This result was expected, as all of these cells are the progeny of classical haematopoietic stem cells, as previously described5.
We found clear evidence, however, for increased self-renewal of F4/80hi macrophages that could explain their capacity to expand in polyps. Firstly, tumour-resident macrophages were clearly in an active phase of the cell cycle unlike those in the LP, as indicated by the expression of the Ki-67 cellproliferation marker (Fig. 6a). Secondly, RNA-sequencing (RNA-seq) analyses of cell-sorted, purified tumour-resident F4/80hi (both MHCIIhi and MHCIIlow) cells showed a clear upregulation of several cell cycle regulators, including cyclin-dependent kinase Cdk1, cyclin-A2 and -B2, E2f2, NEK family of serine/threonine kinases and various cell division cycle and mini-chromosome maintenance proteins family members, in comparison to their LP counterparts (Fig. 6b). Interestingly, we could not find any evidence for active cell cycling of F4/80hiMHCIIhi and F4/80hiMHCIIlow cells in the LP (Fig. 6a, b). This effect could be because F4/80hiMHCIIlow macrophages gradually disappear in the adult LP and probably do not vigorously self-renew, and that F4/80hiMHCIIhi cells are predominantly maintained in the LP by recruited BM-derived cells and, therefore, do not depend on in situ self-renewal.
Tissue-resident macrophages in polyps expand by self-renewal and undergo metabolic reprogramming. a Proliferative activity of the LP and tumoural tissue-resident macrophage subpopulations. Intracellular expression levels of Ki-67 in adult F4/80hiMHCIIhi and F4/80hiMHCIIlow cells, obtained from the colon LP of wild-type mice and colon tumours from dextran sodium sulphate-treated ApcMin/+ mice. Representative dot plot and bar chart representing data obtain for colon LP (n = 5 mice) and tumours (n = 10 mice). Error bars represent the s.e.m. Statistical significance was determined by two-way ANOVA followed by Bonferroni test; **P < 0.01; ***P < 0.001; ns, not significant. b, c Myeloid subpopulations were sorted from pooled colon LP and tumours of 13-15 mice in three independent experiments in an RNA-seq study: Ly6ChiMHCII− monocytes, Ly6GhiF4/80−CD11b+ neutrophils, F4/80hiMHCIIhi and F4/80hiMHCIIlow tissue-resident macrophage subpopulations. Gating strategy is shown in Supplementary Fig. 4. b Heat map and clustering of cell cycle-associated transcripts enriched in intratumoural F4/80hiMHCIIhi and F4/80hiMHCIIlow subsets. The heat map was generated with log2 transformed RPKM values and with the row/gene median subtracted. c Heat map and clustering of glycolysis- and urea cycle-associated transcripts enriched in intratumoural F4/80hiMHCIIhi and F4/80hiMHCIIlow subsets. The heat map was generated with log2 transformed FPKM values and with the row/gene median subtracted. d Quantitative PCR of Arg-1 expression levels in the LP and intratumoural F4/80hiMHCIIhi and F4/80hiMHCIIlow subpopulations. The qPCR results shown are representative of sorted macrophage subpopulations obtained from pooled LP and tumours of 13-15 mice. Error bars represent the s.e.m. Statistical significance was determined by two-way ANOVA followed by Bonferroni test; ***P<0.001 e Representative flow cytometry histograms (upper panels) and bar charts (lower panel) showing the intracellular mean of fluorescence intensity (MFI) of ARG1 expression in colon LP and intratumoural F4/80hiMHCIIhi and F4/80hiMHCIIlow cells. Error bars represent the s.e.m. Statistical significance was determined by two-way ANOVA followed by Bonferroni test; **P<0.01; ***P < 0.001; ns, not significant. f Differential expression of distinct metalloproteinases (Mmp2, Mmp9 and Mmp12) in LP and intratumoural F4/80hiMHCIIhi and F4/80hiMHCIIlow subsets measured by qPCR. Error bars represent the s.e.m. Statistical significance was determined by two-way ANOVA followed by Bonferroni test; *P<0.05; ***P < 0.001; ns, not significant. g Transcriptome analysis of distinct colon LP and tumoural myeloid cell populations. Hierarchical clustering of the LP and intratumoural monocytes (P1), neutrophils and tissue-resident macrophage (F4/80hiMHCIIhi and MHCIIlow) subsets
Tumour microenvironment alters resident macrophage phenotype
Besides the upregulation of genes involved in cell proliferation, macrophages in polyps clearly adopted a distinct metabolic signature. Similar to cancer cells, which can alter their metabolism in aerobic glycolysis and production of lactate (so-called Warburg effect) macrophages in polyps increased the transcript levels of key glycolytic genes, such as ENO1, GAPDH, TPI1, PGAM and LDHA, with the F4/80hiMHCIIlow macrophages expressing the highest levels (Fig. 6c). In addition, our RNA-seq found that both tissue-resident subsets displayed enhanced transcripts of ARG1, typical for M2-polarized and tumour-associated macrophages (TAMs)26,27, with MHCIIlow macrophages being the major producers of ARG1 within tumours (Fig. 6c–e). The expression of other urea cell cycle genes, such as Arginase-2 (ARG2) and Arginosuccinate synthetase 1 (AAS1), was increased in tumour-resident macrophages as well (Fig. 6c). Moreover, several transcripts of matrix metalloproteases (MMPs), such as MMP2, MMP9 and MMP12, involved in the degradation of extracellular matrix and promotion of metastasis were upregulated in both subpopulations of tissue-resident macrophages when compared to their LP counterparts (Fig. 6f), suggesting a possible contribution of these cells in tumour progression.
Of note, although hierarchical clustering between distinct colon myeloid cells clearly identified F4/80hiMHCIIhi and F4/80hiMHCIIlow as demarcated populations, “high/low” pairs always formed more closely related couples in tumours or in the LP than “low/low” or “high/high” comparisons between samples. Although these data suggested a close relationship between F4/80hiMHCIIhi and F4/80hiMHCIIlow cells (Fig. 6g), ingenuity pathway analysis (IPA®) revealed that multiple pathways involved in glycolysis, gluconeogenesis, urea cycle, hypoxia-inducible factor-1 and colorectal cancer metastasis signalling were significantly upregulated in F4/80hiMHCIIlow cells (P < 0.05 by right-tailed Fisher’s exact test, Supplementary Fig. 3, red arrows). Clearly, tumour-resident F4/80hiMHCIIlow cells not only accumulate, but also undergo a robust metabolic reprogramming during tumour progression.
CSF1 maintains and expands pro-tumoural resident macrophages
CSF1 is a crucial growth factor for macrophage proliferation and survival, and supports the self-maintenance of tissue-resident macrophages26,27,28,29. Here, we compared the levels of CSF1 in the colon LP and colon adenomas and detected a progressive increase in the amounts of CSF1 as a function of polyp size (Fig. 7a). In addition, when the effects of CSF1 were neutralized upon exposure to an anti-CSF1 receptor neutralizing antibody in tumour-bearing ApcMin/+ mice, we observed a strong reduction in the number of both F4/80hiMHCIIlow and F4/80hiMHCIIhi tumour-resident macrophages, whereas the numbers of monocytes and monocyte-derived macrophages were unperturbed (Fig. 7b). All together, these data strongly suggest that F4/80hi-resident macrophages exposed to high levels of CSF1 (and likely other tumour niche factors) in the tumour microenvironment gain the ability to self-renew within the tumour.
Pro-tumoural F4/80hi resident macrophages depend on CSF1. a Intra-tumoural CSF1 levels gradually increase with tumour progression. Colon LP and polyps of different sizes (ranging from 0.5 mm to 6 mm in diameter) were processed for quantitative PCR analysis of Csf1 expression. Data were normalized to β-actin and are expressed as fold change in Csf1 expression from 0.5 mm tumours. b Anti-CSF1 receptor (CSF1R) antibody depletes intratumoural F4/80hiMHCIIlow and F4/80hiMHCIIhi macrophages. The 12-week-old dextran sodium sulphate (DSS)-treated ApcMin/+ mice were injected intraperitoneally with rat IGg2a or anti-CSF1R blocking antibody (400 μg/mouse) at days −4 and −1 before collection of the colons, as shown in the schematic representation (upper panel). Polyps were processed and cells were analysed by flow cytometry for the presence of different myeloid cell populations defined in Fig. 1. Gating strategy is shown in Supplementary Fig. 4. The bar charts show the means±s.e.m. of absolute numbers of F4/80hiMHCIIhi and F4/80hiMHCIIlow tissue-resident macrophage subpopulations, monocytes (P1), monocyte-derived macrophages (P2–P3) and eosinophils (control mice: n = 6 and anti-CSF1R Ab injected mice: n = 5). Statistical significance was determined using an unpaired Student’s t-test. ***P < 0.0001; ns, not significant. c CSF1R blockade but not CCR2 deficiency reduces tumour burden in ApcMin/+ mice. Polyp counts in the colon of ApcMin/+ (n = 9) and Ccr2−/−ApcMin/+ mice (n = 9) (upper panel) and anti-rat IgG2a injected ApcMin/+ (n = 11) and anti-CSF1R Ab injected ApcMin/+ mice (n = 9) (lower panel). Statistical significance was determined using an unpaired Student’s t-test. *P < 0.05 and **P < 0.001
To evaluate the pro-tumoural potential of different types of tumour-resident macrophages, polyp numbers and their sizes were assessed in Ccr2−/−ApcMin/+ and anti-CSF1R injected ApcMin/+ mice. Clearly, CCR2 deficiency, although strongly reduced monocytes and monocyte-derived macrophage P2 and P3 populations, had no effect on colonic polyp numbers in Ccr2−/−ApcMin/+ mice compared to their ApcMin/+ littermates in the DSS-accelerated tumour model (Fig. 7c, upper panel). In contrast, depletion of tumour-resident F4/80hi macrophages lowered significantly the polyp counts in the colon (P < 0.05 by unpaired Student's t-test) and the tumours formed were of smaller size (P < 0.001 by unpaired Student's t-test) suggesting their pro-tumoural role in established tumours (Fig. 7c, lower panel).
Discussion
The macrophages found resident in the majority of tissues are established during fetal development and are able to locally self-maintain in adulthood without any further input from BM haematopoietic progenitors4. In tissues exposed to a low-grade inflammation, however, such as the intestine and skin, or to mechanical stress, such as the heart, the maintenance of these cells relies not on self-renewal, but on a continuous replenishment by infiltrating BM-derived CCR2-dependent Ly6Chi circulating monocytes12,14,17.
Here we confirmed that in adult mice the vast majority of colon LP macrophages are derived from CCR2-dependent BM progenies. These macrophages included F4/80hiMHCIIhi LP-resident macrophages (fraction I) in addition to a classical monocyte-derived series of F4/80intCD11b+ cells (fraction II). We found one exception to this rule, whereby LP-resident F4/80hiMHCIIlow macrophages that could be detected in high numbers only in young mice were also clearly found in adult Ccr2−/− mice. Interestingly, our fate mapping experiments showed that LP-resident F4/80hi macrophages exhibited a very slow turnover rate compared to other myeloid populations in the LP. In particular, F4/80hiMHCIIlow macrophages showed minimal labelling after adult mouse tamoxifen administration, which suggests that they are a fetal-derived population that gradually disappear with age. This phenomenon is perhaps expected due to increased competition to occupy the niche by monocyte-derived cells, which can then efficiently differentiate into F4/80hiMHCIIhi macrophages17.
Similarly to the gut, an analogous tissue-resident macrophage subpopulation that expresses low levels of MHCII was recently characterized in the skin dermis—another tissue exposed to commensal microbes and under tonic, low-level inflammation30. Consistent with the colon tissue-resident macrophages, the dermal F4/80hiMHCIIhi fraction is clearly BM derived, whereas the recruitment of the F4/80hiMHCIIlow subset in the dermis occurs mainly during embryogenesis and rapidly declines during adulthood5,12.
To date, the developmental origins of intestinal TAMs and their maintenance in colon adenomas have not been extensively studied. As macrophages have become potential targets for cancer therapy31, we therefore decided to study the dynamics of myeloid populations in colon adenomas. In breast cancer13, glioma32 and lung metastases33 TAMs can arise from Ly6Chi circulating monocytes. Their influx and positioning in the tumour is mediated by chemokines (e.g., CCL2), growth factors (e.g., CSF1) and complement components31,34. In addition, a recent study demonstrated that local proliferation of TAMs contributes to their accumulation in spontaneous mammary tumours24. Many have speculated that in cancers exposed to microbes, such as colon cancer, the recruitment of macrophages and their maintenance might differ from cancers located in “sterile” sites, such as breast cancers31.
Here, we have analysed in detail the origins and maintenance of macrophage subpopulations in colon tumours obtained from ApcMin/+ mice, which is a widely used animal model for intestinal tumourigenesis35, and focused on both subsets of tissue-resident F4/80hi macrophages in the colon LP.
The lymphoid and myeloid composition of colon adenomas of ApcMin/+ mice differed from that of the colon LP. In the LP, we found that lymphoid cells (mainly B cells) were the predominant cell type, whereas in adenomas the majority consisted of myeloid cells and were almost devoid of lymphocytes and, notably, eosinophils that were the most common LP myeloid cells. Neutrophils and monocytes were strongly increased in adenomas, but interestingly also F4/80hiMHCIIlow macrophages. These MHCIIlow cells are usually a minor population in LP, but clearly “flourished” in the tumour environment in our model to such an extent that in large polyps they became the dominant population among F4/80hi cells. Our fate mapping studies and analyses of Ccr2−/− mice showed that MHCIIlow macrophages expanded in situ in adenomas by self-renewal, without any recruitment of new cells. This finding was also supported by our RNA-seq and Ki-67 analyses showing that MHCIIlow macrophages in polyps were actively cycling, unlike macrophages found in the colon LP. Similarly, the tumour microenvironment was also conducive for the self-renewal of F4/80hiMHCIIhi macrophages. This subpopulation of macrophages was also actively cycling and had become independent from CCR2-dependent inputs from the BM. The tumour milieu, however, did not favour monocyte differentiation to macrophages. In adenomas, monocytes (P1) got “stuck” at the beginning of the “waterfall” maturation pathway to macrophages (P3), which resulted in the F4/80hi cells becoming the dominant macrophage population. This effect has also been reported in the context of mammary cancers13. Our data clearly suggest that unique characteristics of the tumour niche can trigger self-renewal of F4/80hi macrophages. CSF1 is certainly a key component of the tumour niche to manipulate macrophage self-renewal, but other cues can also contribute to this process and, therefore, reconstruction of the tumour niche would be very illuminating.
According to the “M1/M2 paradigm”, mainly based on in vitro polarization experiments, pro-inflammatory M1macrophages display anti-tumoural characteristics, whereas alternatively activated M2-macrophages are generally considered pro-tumoural. Both populations can co-exist in the tumour, although M2-like MHCIIlow TAMs are claimed to enrich in hypoxic and M1-like MHCIIhi TAMs in normoxic areas22,36. Due to the complex environment of the gut, intestinal tissue-resident macrophages do not follow the rigid M1–M2 classification37 and “classic” M1 and M2 features are often shared by the same cells20,38. Similar mixed M1/M2 phenotype is observed in macrophages of colon tumours, since both F4/80hi MHCIIhi and MHCIIlow populations express the classical M2 marker ARG1, but also many glycolytic genes typical for M1 macrophages, although the F4/80hiMHCIIlow-resident macrophage subset shows always the highest transcript levels. As observed in other tumour models22,36,39, F4/80hiMHCIIlow cells, with their augmented metabolic switch together with their progressive predominance during tumour development, represent a pro-tumoural macrophage subpopulation also in intestinal adenomas.
Macrophages are not only crucial mediators of many pathologies (ranging from infectious to metabolic diseases and cancers) but also influence the outcome of anticancer therapeutics, such as chemotherapy, radiotherapy and even immunotherapy40,41,42. As such, these cells are becoming attractive candidates for therapeutic intervention. Manipulation of macrophage numbers43 and/or reprogramming their phenotype44 are techniques currently used to target macrophages in a therapeutic context. A better understanding of their ontogeny in different tumours, such as the data presented here for colon cancer, will facilitate the development of successful innovative macrophage-based therapeutic approaches.
Methods
Mice
KitMerCreMer/R26 mice were generated in our laboratory as previously described5. Ccr2−/− (B6.129S4-Ccr2tm1lfc/J)45, C57BL/6J and ApcMin/+ (C57BL/6J-ApcMin/J)46 mice were obtained from The Jackson Laboratory (USA). ApcMin/+ mice were backcrossed with KitMerCreMer/R26 mice or Ccr2−/− mice to obtain ApcMin/+KitMerCreMer/R26 and Ccr2−/−ApcMin/+ mice, respectively. Mice were specific-pathogen-free (SPF) and maintained under a 12 h light/dark cycle with standard chow diet provided ad libitum. Both male and female mice (6–8 weeks old) were used for all experiments and were equally distributed within experimental and control groups.
All transgenic mice were bred and housed under SPF conditions in the Nanyang Technological University animal facility. This study was carried out in strict accordance with the recommendations of the NACLAR (National Advisory Committee for Laboratory Animal Research) guidelines under the Animal & Birds (Care and Use of Animals for Scientific Purposes) Rules of Singapore. The protocol was approved by the Institutional Animal Care and Use Committee of the Nanyang Technological University of Singapore.
Spontaneous and chemical-induced intestinal tumourigenesis
ApcMin/+ mice spontaneously develop macroscopically detectable adenomas mainly in the small intestine and only few in the colon in aged mice35. Therefore, for all our experiments (with exception of the experiments shown in Fig. 2 and Supplementary Fig. 1) we opted for a chemically induced colitis model which selectively enhances the development of adenomas in the colon in a relatively short time of 5 weeks. Mice were terminated when showing symptoms of anaemia in combination with weight loss and/or other signs of physical discomfort.
In the “spontaneous” tumour model, mice were analysed at the age of 6–7 months, whereas in case of chemically induced tumourigenesis, 6–8-week-old female and male ApcMin/+ mice (and correspondent control C57BL/6J mice) were treated with 1.5 or 1.25% (w/v) DSS (50,000 Da; MP Biomedicals, Santa Ana, CA, USA), respectively, in the drinking water for 1 week and sacrificed 4 weeks later at an age of 3–4 months.
Colorectal polyp counts
Polyps in each colon were macroscopically counted and categorized as >2 mm (large) and <2 mm (small). Colon tumours were measured ex vivo with the help of a sliding caliper.
Tamoxifen-inducible fate mapping mouse model
KitMerCreMer/R26 and ApcMin/+KitMerCreMer/R26 mice were used for cell fate mapping to delineate the ontogeny of intratumoural macrophages. Then, 80 mg tamoxifen (Sigma, T5648; Sigma-Aldrich, St. Louis, MO, USA) was dissolved in 6 ml corn oil (C8267; Sigma) and a total of 4 mg tamoxifen per mouse was administered for five consecutive days by gavage for adult labelling or by intraperitoneal injection in pregnant mothers (embryonic day 10.5) for embryonic labelling. After tamoxifen administration, c-kit+ BM-derived haematopoietic stem cells (HSCs) in ApcMin/+KitMerCreMer/R26 mice were labelled with yellow fluorescent protein (YFP+) and all cells deriving from YFP+ BM-HSCs would maintain YFP expression. This effect enables to distinguish whether adult intratumoural macrophages are derived from adult BM-definitive haematopoiesis or from embryonic haematopoiesis. Adult and fetal labelling experimental protocols are shown in Fig. 5.
Myeloid cell isolation
LP myeloid cells were isolated as previously described47. Briefly, after CO2 killing of the mice, the colon was opened and rinsed with phosphate-buffered saline (PBS) to remove the luminal contents. To remove the epithelium, the colon was incubated in 25 ml PBS with 1.3 mM EDTA under shaking conditions at 37 °C for 1 h. After incubation, the colon was washed in 2% Iscove’s modified Dulbecco’s medium (IMDM) to remove the EDTA solution, and then minced. The colon pieces were digested in 2% IMDM containing 1 mg/ml Collagenase D (Roche, Switzerland) under shaking conditions at 37 °C for 1.5 h. The digested tissue was then gently mashed through a 150 μm cell strainer. The leukocyte population was enriched using a 70/40% Percoll gradient (GE Healthcare Life Science, Chicago, IL, USA). Low-density cells at the interface were harvested and processed further for flow cytometry.
Tumours were manually cut, minced and digested in 2% IMDM containing 1 mg/ml Collagenase D and 20 U/ml DNase I (Life Technologies, Carlsbad, CA, USA) under shaking conditions at 37 °C for 1 h. Digested tissue was subsequently mashed through a cell strainer to obtain a cell suspension. The leukocyte population was enriched by 35% Percoll and further processed for flow cytometry and, if required, cell sorting using a FACSAria cell sorter (BD Biosciences, San Jose, CA, USA). Gating strategy used for sorting is shown in Supplementary Fig. 4.
Transcriptomics analysis by RNA-sequencing
Total RNA was extracted using the ArcturusTM PicoPureTM RNA Isolation Kit (Cat. No. KIT0214, Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol. All mouse RNAs were analysed using an Agilent Bioanalyser (Agilent, Santa Clara, CA, USA) for quality assessment; the RNA Integrity Number range was 6.5–10, with a median of 9.2. Complementary DNA (cDNA) libraries were prepared from 2 ng total RNA starting material and 1 μl of a 1:50,000 dilution of ERCC RNA Spike in Controls (Cat. No. 4456740, Ambion® Thermo Fisher Scientific) using the SMARTSeq v2 protocol48 with the following modifications: (1) addition of 20 µM TSO; and (2) use of 250 pg cDNA with 1/5 reaction of Illumina Nextera XT kit (Cat. No. FC-131–1024, Illumina, San Diego, CA, USA). The length distribution of the cDNA libraries was monitored using a DNA High Sensitivity Reagent Kit on the Perkin Elmer Labchip (Cat. No. CLS760672, Perkin Elmer, Waltham, MA, USA). All eight samples were subjected to an indexed paired-end sequencing run of 2 × 51 cycles on an Illumina HiSeq 2500 system (Illumina) under rapid run mode (17 samples/lane).
The paired-end reads were mapped to the Mouse GRCm38/mm10 reference genome using the STAR alignment tool49. Mapped reads were summarized to the gene level using featureCounts (V1.5.0-P1) software50 and with GENCODE gene annotation51. Genes with an average number of reads per sample <10 in all cell subpopulations were filtered out from subsequent analyses. For differentially expressed gene (DEG) analysis, the limma/voom pipeline was used as recommended by the MicroArray Quality Control (MAQC) project52 as one of the best performing RNA-seq data analysis pipelines. Comparisons between different cell populations were performed using limma and DEGs were selected with Benjamini–Hochberg adjusted P-values < 0.05. Hierarchical clustering and principal component analysis were performed with Log2 transformed value of RPKM (Reads Per Kilobase of transcript per Million mapped reads). All analyses were carried out in R version 3.1.2 (URL http://www.R-project.org/).
Flow cytometry
Single-cell suspensions were then stained and subsequently analysed by a BD Fortessa 5 laser flow cytometer (BD Bioscience). Data were analysed using a FlowJo software (TreeStar, Ashland, OR, USA). The following antibodies were used: APC/Cy7-labelled anti-CD11b (clone: M1/70, 1:600) and BV605-labelled anti-Ly6C (HK1.4, 1:600) were purchased from Biolegend (San Diego, CA, USA). PE-labelled anti-EMR1 (also known as F4/80) (clone: BM8, 1:600), PE/Cy7-labelled anti-CD45.2 (clone: 104, 1:600), APC-labelled anti-CD11c (clone: N418, 1:600) and eFluor®450-labelled anti-MHCII (clone:M5/114.15.2, 1:800) were obtained from eBioscience (San Diego, CA, USA). FITC-labelled polyclonal sheep anti-human/mouse ARG1 (1:20) and APC-labelled anti-CCR2 (Cat. No. FAB5538A) (1:20) were obtained from R&D System (Minneapolis, MN, USA), FITC-labelled anti-Mouse/Rat Ki-67 (clone SolA15, 1:40) from Miltenyi Biotech (Bergisch-Gladbach, Germany) and BUV395-labelled anti-Ly6G (clone: 1A8, 1:600) from BD (San Diego, CA, USA)
Anti-CSF1R antibody treatment
Twelve-week-old DSS-treated ApcMin/+ mice were injected intraperitoneally with rat Ig2a isotype control (Biolegend) or anti-CSF1R blocking antibody (Clone AFS98, BioXCell, West Lebanon, NH, USA) (400 μg/mouse) at days −4 and −1 before collection of the colons.
Quantitative PCR (qPCR) analysis
cDNA was generated using SuperScript III Reverse Transcriptase (Cat. No. 18080093, Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Quantitative real-time PCR was then performed using the FAST 2× qPCR Master mix (PrecisionFAST-SY, Primerdesign Ltd, Cambridge, UK). Reactions were run on a real-time qPCR system (Illumina, San Diego, CA, USA). Samples were normalized to β-actin, and data represent the mean of triplicate analyses and are displayed as a fold change from F4/80hiMHCIIhi LP macrophages unless otherwise stated. The primer sequences were as follows: Arg1; Fwd: gaatctgcatgggcaacc, Rev: gaatcctggtacatctgggaac; Mmp2; Fwd: taacctggatgccgtcgt, Rev: ttcaggtaataagcacccttgaa; Mmp9; Fwd: catccagtatctgtatggtcgtg, Rev: gctgtggttcagttgtggtg; Mmp12; Fwd: gctgctcccatgaatgaca, Rev: aagcattgcacacggttgt; Csf1; Fwd: ggtggaactgccagtatagaaag, Rev: tcccatatgtctccttccataaa; β-actin Fwd: aaggccaaccgtgaaaagat, Rev: gtggtacgaccagaggcatac.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 6 software (GraphPad Software, La Jolla, CA, USA). All values are expressed as the mean±s.e.m as indicated in the legend. Samples were analysed by unpaired Student’s t-test (two-tailed) or Bonferroni two-way analysis of variance (ANOVA). A P-value of <0.05 was considered to be statistically significant.
Data availability
The accession number for the RNA-seq data reported in this paper is GEO: GSE90153. Original flow cytometry data are deposited in the NTU Open Access Data Repository DR-NTU (https://researchdata.ntu.edu.sg/dataset.xhtml?persistentId=doi:10.21979/N9/EQXCRF). All other data are available from the authors upon reasonable request.
References
Varol, C., Mildner, A. & Jung, S. Macrophages: development and tissue specialization. Annu. Rev. Immunol. 33, 643–675 (2015).
Gross, M., Salame, T. M. & Jung, S. Guardians of the gut - murine intestinal macrophages and dendritic cells. Front. Immunol. 6, 254 (2015).
Ginhoux, F. & Guilliams, M. Tissue-resident macrophage ontogeny and homeostasis. Immunity 44, 439–449 (2016).
Perdiguero, E. G. & Geissmann, F. The development and maintenance of resident macrophages. Nat. Immunol. 17, 2–8 (2016).
Sheng, J., Ruedl, C. & Karjalainen, K. Most tissue-resident macrophages except microglia are derived from fetal hematopoietic stem cells. Immunity 43, 382–393 (2015).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013).
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013).
Ginhoux, F., Lim, S., Hoeffel, G., Low, D. & Huber, T. Origin and differentiation of microglia. Front Cell Neurosci. 7, 45 (2013).
Tamoutounour, S. et al. Origins and functional specialization of macrophages and of conventional and monocyte-derived dendritic cells in mouse skin. Immunity 39, 925–938 (2013).
Franklin, R. A. et al. The cellular and molecular origin of tumor-associated macrophages. Science 344, 921–925 (2014).
Lavine, K. J. et al. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc. Natl. Acad. Sci. USA 111, 16029–16034 (2014).
Calderon, B. et al. The pancreas anatomy conditions the origin and properties of resident macrophages. J. Exp. Med. 212, 1497–1512 (2015).
Bain, C. C. et al. Resident and pro-inflammatory macrophages in the colon represent alternative context-dependent fates of the same Ly6Chi monocyte precursors. Mucosal Immunol. 6, 498–510 (2013).
Bain, C. C. et al. Constant replenishment from circulating monocytes maintains the macrophage pool in the intestine of adult mice. Nat. Immunol. 15, 929–937 (2014).
Smythies, L. E. et al. Mucosal IL-8 and TGF-beta recruit blood monocytes: evidence for cross-talk between the lamina propria stroma and myeloid cells. J. Leukoc. Biol. 80, 492–499 (2006).
Tamoutounour, S. et al. CD64 distinguishes macrophages from dendritic cells in the gut and reveals the Th1-inducing role of mesenteric lymph node macrophages during colitis. Eur. J. Immunol. 42, 3150–3166 (2012).
Zigmond, E. et al. Macrophage-restricted interleukin-10 receptor deficiency, but not IL-10 deficiency, causes severe spontaneous colitis. Immunity 40, 720–733 (2014).
Shouval, D. S. et al. Interleukin-10 receptor signaling in innate immune cells regulates mucosal immune tolerance and anti-inflammatory macrophage function. Immunity 40, 706–719 (2014).
Movahedi, K. et al. Different tumor microenvironments contain functionally distinct subsets of macrophages derived from Ly6C(high) monocytes. Cancer Res. 70, 5728–5739 (2010).
Franklin, R. A. & Li, M. O. Ontogeny of tumor-associated macrophages and its implication in cancer regulation. Trends Cancer 2, 20–34 (2016).
Tymoszuk, P. et al. In situ proliferation contributes to accumulation of tumor-associated macrophages in spontaneous mammary tumors. Eur. J. Immunol. 44, 2247–2262 (2014).
Shi, C. & Pamer, E. G. Monocyte recruitment during infection and inflammation. Nat. Rev. Immunol. 11, 762–774 (2011).
Tushinski, R. J. et al. Survival of mononuclear phagocytes depends on a lineage-specific growth factor that the differentiated cells selectively destroy. Cell 28, 71–81 (1982).
Tagliani, E. et al. Coordinate regulation of tissue macrophage and dendritic cell population dynamics by CSF-1. J. Exp. Med. 208, 1901–1916 (2011).
Hume, D. A., Pavli, P., Donahue, R. E. & Fidler, I. J. The effect of human recombinant macrophage colony-stimulating factor (CSF-1) on the murine mononuclear phagocyte system in vivo. J. Immunol. 141, 3405–3409 (1988).
Stanley, E. R. & Chitu, V. CSF-1 receptor signaling in myeloid cells. Cold Spring Harb. Perspect. Biol. 6, a021857 (2014).
Belkaid, Y. & Tamoutounour, S. The influence of skin microorganisms on cutaneous immunity. Nat. Rev. Immunol. 16, 353–366 (2016).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61 (2014).
Pyonteck, S. M. et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat. Med. 19, 1264–1272 (2013).
Qian, B. et al. A distinct macrophage population mediates metastatic breast cancer cell extravasation, establishment and growth. PLoS ONE 4, e6562 (2009).
Bonavita, E. et al. PTX3 is an extrinsic oncosuppressor regulating complement-dependent inflammation in cancer. Cell 160, 700–714 (2015).
Moser, A. R. et al. ApcMin: a mouse model for intestinal and mammary tumorigenesis. Eur. J. Cancer 31A, 1061–1064 (1995).
Van Overmeire, E. et al. M-CSF and GM-CSF receptor signaling differentially regulate monocyte maturation and macrophage polarization in the tumor microenvironment. Cancer Res. 76, 35–42 (2016).
Bain, C. C. & Mowat, A. M. Macrophages in intestinal homeostasis and inflammation. Immunol. Rev. 260, 102–117 (2014).
Weber, B., Saurer, L., Schenk, M., Dickgreber, N. & Mueller, C. CX3CR1 defines functionally distinct intestinal mononuclear phagocyte subsets which maintain their respective functions during homeostatic and inflammatory conditions. Eur. J. Immunol. 41, 773–779 (2011).
Wang, B. et al. Transition of tumor-associated macrophages from MHC class II(hi) to MHC class II(low) mediates tumor progression in mice. BMC Immunol. 12, 43 (2011).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
Ries, C. H. et al. Targeting tumor-associated macrophages with anti-CSF-1R antibody reveals a strategy for cancer therapy. Cancer Cell 25, 846–859 (2014).
Mantovani, A. & Allavena, P. The interaction of anticancer therapies with tumor-associated macrophages. J. Exp. Med. 212, 435–445 (2015).
Engblom, C., Pfirschke, C. & Pittet, M. J. The role of myeloid cells in cancer therapies. Nat. Rev. Cancer 16, 447–462 (2016).
Schultze, J. L. Reprogramming of macrophages–new opportunities for therapeutic targeting. Curr. Opin. Pharmacol. 26, 10–15 (2016).
Boring, L. et al. Impaired monocyte migration and reduced type 1 (Th1) cytokine responses in C-C chemokine receptor 2 knockout mice. J. Clin. Invest. 100, 2552–2561 (1997).
Moser, A. R., Pitot, H. C. & Dove, W. F. A dominant mutation that predisposes to multiple intestinal neoplasia in the mouse. Science 247, 322–324 (1990).
Muzaki, A. R. et al. Intestinal CD103(+)CD11b(-) dendritic cells restrain colitis via IFN-gamma-induced anti-inflammatory response in epithelial cells. Mucosal Immunol. 9, 336–351 (2016).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Harrow, J. et al. GENCODE: producing a reference annotation for ENCODE. Genome Biol. 7(Suppl. 1), S4.1–S4.9 (2006).
SEQC/MAQC-III Consortium. A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium. Nat. Biotechnol. 32, 903–914 (2014).
Acknowledgements
The authors would like to thank Monika Tetlak for providing excellent mouse management. The authors would also like to thank Insight Editing London for proofreading the manuscript prior to submission. This work was supported by MOE2014-T2-1-011 and MOE2016-T2-1-012 Ministry of Education Tier2 grants to C.R.
Author information
Authors and Affiliations
Contributions
Conceptualization: C.R.; methodology: I.S., J.S., S.F., J.L. and F.Z.; investigation: I.S., J.S. and Q.C.; formal analysis: I.S.; bioinformatic analysis: K.D. and M.P.; writing (original draft): C.R.; writing (review and editing): C.R. and K.K.; visualization: I.S.; funding acquisition: C.R.; supervision: J.S., K.K. and C.R.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Soncin, I., Sheng, J., Chen, Q. et al. The tumour microenvironment creates a niche for the self-renewal of tumour-promoting macrophages in colon adenoma. Nat Commun 9, 582 (2018). https://doi.org/10.1038/s41467-018-02834-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-02834-8
This article is cited by
-
Monocyte-macrophages modulate intestinal homeostasis in inflammatory bowel disease
Biomarker Research (2024)
-
Modulation of the tumor microenvironment and mechanism of immunotherapy-based drug resistance in breast cancer
Molecular Cancer (2024)
-
The roles of tissue resident macrophages in health and cancer
Experimental Hematology & Oncology (2024)
-
Elucidating immunological characteristics of the adenoma-carcinoma sequence in colorectal cancer patients in South Korea using a bioinformatics approach
Scientific Reports (2024)
-
Tissue-resident macrophages are major tumor-associated macrophage resources, contributing to early TNBC development, recurrence, and metastases
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.