Abstract
NAC (NAM, ATAF1/2, and CUC2) transcription factors play important roles in fruit ripening and quality. The watermelon genome encodes 80 NAC genes, and 21 of these NAC genes are highly expressed in both the flesh and vascular tissues. Among these genes, ClNAC68 expression was significantly higher in flesh than in rind. However, the intrinsic regulatory mechanism of ClNAC68 in fruit ripening and quality is still unknown. In this study, we found that ClNAC68 is a transcriptional repressor and that the repression domain is located in the C-terminus. Knockout of ClNAC68 by the CRISPR-Cas9 system decreased the soluble solid content and sucrose accumulation in mutant flesh. Development was delayed, germination was inhibited, and the IAA content was significantly decreased in mutant seeds. Transcriptome analysis showed that the invertase gene ClINV was the only gene involved in sucrose metabolism that was upregulated in mutant flesh, and expression of the indole-3-acetic acid-amido synthetase gene ClGH3.6 in the IAA signaling pathway was also induced in mutant seeds. EMSA and dual-luciferase assays showed that ClNAC68 directly bound to the promoters of ClINV and ClGH3.6 to repress their expression. These results indicated that ClNAC68 positively regulated sugar and IAA accumulation by repressing ClINV and ClGH3.6. Our findings provide new insights into the regulatory mechanisms by which NAC transcription factors affect fruit quality and seed development.
Similar content being viewed by others
Introduction
Fruit ripening is a complex process, and multiple metabolic changes, including changes that affect sugar, organic acids, and pigments, occur during fruit ripening and postharvest storage1. The network of underlying molecular mechanisms that directly regulate fruit ripening and determine fruit quality has been revealed; this network includes hormones and signaling pathways, transcriptional regulators, and other regulatory elements2. Watermelon (Citrullus lanatus) is one of the most popular fruits3, and it is important to explore the intrinsic mechanism regulating metabolic changes to improve fruit quality.
Numerous studies have shown that enzymes involved in sugar metabolism, including sucrose synthase (Sus), sucrose phosphate synthase (SPS), and invertases (INVs), play important roles in regulating sucrose accumulation in fruit4. Among these enzymes, Sus or INVs catalyze the hydrolysis of sucrose into fructose and glucose, and the activity of these two enzymes shows a negative correlation with sugar content5. Recently, studies revealed that transcription factors mediate the regulation of these important enzymes to determine the sugar content in seeds and fruit. In maize, the endosperm-specific transcription factor Opaque2 (O2) was found to activate the expression of sus1 and sus2 and result in a higher soluble sugar content in the o2 mutant6. In pitaya, HpWRKY3 was found to activate the expression of HpINV2 and HpSuSy1 to regulate sugar accumulation, but there was no transgenic evidence to further determine this relationship7. In our previous study, we found that the total sugar and sucrose contents increased, while the expression of Sus and acid INV showed a significant negative correlation with sucrose content during watermelon fruit ripening8. However, the regulators of Sus and acid INV during watermelon fruit and seed development have not been well illustrated.
Plant hormones and hormone signal transduction pathways play important roles in regulating seed germination, plant growth, development, and fruit ripening9. In cucumber, the IAA content was found to show a positive correlation with fruit size10. In peach, suppression of PpYUC11 was found to decrease the IAA level and result in increased fruit fitness11. Another important IAA-amino synthetase (GH3) gene in the IAA signaling pathway catalyzes the binding of amino acids to IAA and results in inactive IAA and a reduction in the free IAA content12. In apple, overexpression of MsGH3.5 was found to result in dwarf plants and fewer adventitious roots, and the IAA and cytokinin contents were significantly reduced13. In kiwifruit, silencing AcGH3.1 by VIGS increased fruit firmness during kiwifruit postharvest storage. Moreover, the IAA content was increased during seed development, and a higher IAA content affected seed size14. In soybean, the IAA content was found to be significantly higher in variants with larger seeds than in variants with smaller seeds14. In rice, overexpression of OsYUC2 was found to increase grain yield by increasing the IAA level15. However, how the IAA and IAA signaling pathways regulate fruit and seed development in watermelon is still unknown.
NACs (NAM, ATAF1/2, and CUC2) constitute one of the largest plant-specific transcription factor families. NACs are involved in regulating processes of plant development, including root elongation, leaf senescence, and fruit ripening16,17. Overexpression of MdNAC42 in apple calli was found to activate the expression of flavonoid pathway genes and induce anthocyanin accumulation18. In tomato, overexpression of SlNAP2 was found to accelerate leaf senescence by increasing ABA biosynthesis genes and the expression of chlorophyll degradation genes19. Furthermore, although the sugar content was found to be significantly reduced in OE lines and induced in knockout lines, the mechanism of SlNAP2 in the regulation of sugar metabolism is still unknown19. Most previous studies focused on NAC-mediated regulation of seed germination via the ABA pathway, while a few studies suggested that NAC regulates seed germination through the IAA pathway. In rice, overexpression of OsNAC2 was found to inhibit root elongation by affecting the expression of auxin- and cytokinin-responsive genes20. In Arabidopsis, the NAC transcription factor NTM2 was found to negatively regulate seed germination under salt stress by upregulating IAA30 expression21. In watermelon, 80 NAC transcription factors and 21 ClNACs were found to be expressed in both vascular tissues and fruit, and these 21 ClNACs might regulate watermelon fruit ripening or quality22. Among these NAC genes, ClNAC68 (Cla97C03G059250, Cla019693 in version 1 of the watermelon genome) was found to be highly expressed in the later stages, and its expression was significantly higher in flesh than in rind. These results indicated that ClNAC68 might play an important role in regulating watermelon fruit ripening. However, the intrinsic mechanism of ClNACs in the regulation of fruit ripening, fruit quality, and seed development is still unknown.
In this study, we characterized the NAC gene ClNAC68 and explored its function in fruit and seeds. Knockout of ClNAC68 by the CRISPR-Cas9 system decreased the fruit sugar content and delayed seed maturation. The seed germination rate and root elongation were also decreased in mutant lines, and exogenous IAA treatment partially reversed the inhibition. The biological function and the regulatory network of ClNAC68 were comprehensively analyzed, and the findings revealed that ClNAC68 plays a critical role in sugar accumulation and seed development.
Materials and Methods
Plant transformation and plant growth conditions
To investigate the function of ClNAC68, the CRISPR/Cas9 vector pBSE401 was used to generate gene knockout plants for further analysis. Two target sites were designed on the website (https://www.addgene.org/crispr/) and incorporated into forward and reverse PCR primers (DT1 and DT2, respectively) as described by Tian et al.23 (Supplemental Table S1). Then, the pBSE401-ClNAC68 construct was transferred into Agrobacterium tumefaciens strain EHA105 and used for transformation. Plant transformation was performed according to the method described by Tian et al.23 PCR was used to identify the homozygous mutant and Cas9-free generations. The primers used are listed in Table S1. All seedlings of mutant lines and WT lines were grown in a plastic greenhouse under natural growing conditions. The phenotypes of the mutant and WT lines were observed at 18 and 26 days after pollination (DAP).
Subcellular localization
The full coding sequence (CDS) of the gene without the stop codon was inserted into the pYBA1332 vector, which contained a GFP tag. The expression construct and empty vector were transferred into watermelon leaf protoplasts. The methods used for protoplast extraction and transformation were described in our previous study24. After 24 h of transformation, the protoplasts were incubated with DAPI solution (Sigma, USA; 1 mg/mL) in the dark for 5 min and observed by confocal microscopy (Zeiss, LSM700 laser-scanning confocal microscope, Germany).
Transcriptional assay
For the yeast two-hybrid assay, the full-length NAC domain fragment and transcription domain fragment of ClNAC68 were cloned into the pGBKT7 vector (Table S1). All constructs were transformed into the yeast strain Y2H-Gold and cultured in SD-T and SD-TH selective media according to the manufacturer’s instructions (Clontech, USA) at 28 °C for three to five days. pGBKT7-P53 and pGBKT7 were used as the positive and negative controls, respectively. The experiment was performed in triplicate.
For measurement of transient luciferase activity, different fragments of ClNAC68 were inserted into the GAL4DB vector. The empty GAL4DB vector and herpes simplex virus 16 (VP16) were used as the negative and positive controls, respectively. Then, these constructs functioned as effectors and were transiently transformed into watermelon leaf protoplasts with a luciferase reporter vector. Renilla luciferase was used as the internal control. The plasmid ratio used was pTRL:reporter:ClNAC68/none/VP16) = 1:6:6. Furthermore, these constructs were coexpressed with VP16 in watermelon leaf protoplasts with the reporter and internal control, and the plasmid ratio used was pTRL:reporter:ClNAC68/actin/none:VP16 = 1:6:6:6, following the method described in a previous study by Wei et al.25. The watermelon actin gene Cla97C02G026960 was used as the noninteractive control. The dual-luciferase activity was measured according to the manufacturer’s instructions (Vazyme Biotech, China, Nanjing). The experiment was performed with six biological replicates.
Measurement of sugar content and invertase activity
The brix of WT and clnac68 mutant fruits was measured with a hand-held ATC-1E refractometer (ATAGO, Tokyo). Sugar content was measured by Majorbio (http://www.majorbio.com/) based on the Thermo ICS5000 platform. Invertase activity was measured as described in our previous study8. Three biological replicates were performed.
Measurement of phytohormone content
Free IAA and ABA were measured with an ESI-HPLC-MS system. IAA and ABA standards (Sigma, USA) were used to determine the retention time and mass spectra. Free IAA and ABA were measured following the method described in our previous study26 in a biological mass spectrometry laboratory at China Agricultural University. Three biological replicates were performed.
Seed germination and root elongation assays
For the seed germination assay, WT and clnac68 mutant seeds were germinated under normal and exogenous IAA and ABA treatment. The seeds were soaked for 4 h in 50 °C water and were then transferred into culture dishes with filter paper containing 5 mL of distilled water, 10 μM IAA (Sigma, USA), and 10 μM ABA (Sigma, USA). All seeds were induced to germinate at 28 °C in the dark and were observed at 24, 36, 48, 60, 72, 84, and 96 h to calculate the seed germination rate (germinated seeds/total seeds). Sixty seeds were used for each treatment condition, and three biological replicates were performed. For the root elongation assay, after 48 h of germination, germinated seeds were selected to measure the root length. All seeds were grown vertically on filter paper containing 5 mL of distilled water or 10 μM IAA (Sigma, USA). After 5 days, the root lengths of WT and clnac68 mutant seeds were quantified.
Transcriptome analysis and qRT-PCR analysis
Intact leaves in the middle of watermelon vines and healthy roots were collected from 10 to 34 days after the pollination stage. The center flesh and seeds were collected and immediately frozen in liquid nitrogen for further use. Three seedlings or fruits were collected for each analysis.
For transcriptome analysis, 12 RNA libraries were built with a TruSeqTM RNA Sample Prep Kit (Illumina, USA) and sequenced on the Illumina HiSeq 4000 platform using the paired-end 150 bp read mode. Gene expression levels were normalized to transcripts per million reads (TPM) values. The differentially expressed genes (DEGs) (p-adjust < 0.05 and | log2FC | > = 1) between the clnac68 mutant lines and WT lines were selected for further research. The data were analyzed on the free online platform of the Majorbio I-Sanger Cloud Platform (www.i-sanger.com).
For qRT-PCR analysis, cDNA was reverse transcribed with FastKing gDNA Dispelling RT SuperMix (Tiangen Biotech, Beijing, China). qRT-PCR was performed in a LightCycler 480 RT-PCR system (Roche, Switzerland) with specific primers (Supplemental Table S1). The watermelon actin gene Cla97C02G026960 was used as the internal control.
Electrophoretic mobility shift assay (EMSA)
The full-length ClNAC68 sequence without a stop codon was fused to an MBP tag and inserted into the vector pMCSG7-MBP-His, and the ClNAC68 protein was purified by His-agarose affinity chromatography. The biotin-labeled probes were the promoters of ClINV or ClGH3.6 containing NAC binding sites, and in the mutant probes, the NAC core binding sites ‘CGTG(A)’ were replaced with continuous ‘AAAA’ sequences. The nonbiotin-labeled segments of the same sequences were used as competitors (Supplemental Table S1). EMSA was performed according to the protocol of the Light Shift Chemiluminescent EMSA Kit (Thermo Fisher Scientific, Shanghai, China). The MBP protein was used as the negative control.
Measurement of transient dual-luciferase activity
The full-length ClNAC68 sequence was inserted into the pGreenII 62-SK vector as an effector, and the promoters of ClINV and ClGH3.6 containing NAC binding sites were inserted into pGreenII 0800-LUC as the reporter (Supplemental Table S1). All these constructs were transformed into Agrobacterium tumefaciens strain GV3101, and the effector was transfected with different reporters into tobacco leaves. The pGreenII 62-SK empty vector was used as the negative control. Luciferase activity was measured following the protocol of the Dual-Luciferase Reporter Assay Kit (Vazyme Biotech, Nanjing, China). Five biological replicates were performed.
Statistical analysis
The data were analyzed by Student’s t test, and the significance of differences relative to WT plants or negative controls was evaluated using GraphPad 8.0.
Results
ClNAC68 was highly expressed in flesh and proved to be a transcriptional repressor
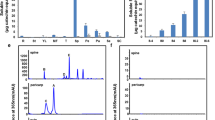
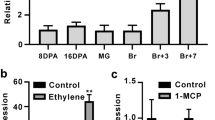
A previous study showed that there are 80 NAC transcription factors encoded in the watermelon genome and that 21 ClNACs are expressed in both the vascular tissues and fruit22. Among these NAC genes, ClNAC68 (Cla97C03G059250, Cla019693 in version 1 of the watermelon genome) was highly expressed at 18, 26, and 34 DAP, and its expression was significantly higher in flesh than in rind. We further tested the expression of ClNAC68 in different tissues and found that the expression of ClNAC68 was highest in flesh and lowest in leaves (Fig. 1A) and was higher in cultivated watermelon (97103) than in wild watermelon (PI296341-FR) (Fig. 1B). The ClNAC68-GFP fusion protein was detected by confocal microscopy and was observed to be localized in the nucleus (Fig. 1C).
A. The expression pattern of ClNAC68 in different tissues. DAP days after pollination, FPKM Fragments Per Kilobase per Million mapped reads. B The expression pattern of ClNAC68 in cultivated and wild watermelons during fruit ripening. C. ClNAC68 was localized in the nucleus. pYBA1332 was used as the negative control and is indicated as CK. Bar = 10 μm. D Schematic representation of different lengths of ClNAC68. Full-length ClNAC68: ClNAC681–319, N-terminus of ClNAC68: ClNAC681–197, and C-terminus of ClNAC68: ClNAC68198–319. E Growth performance of yeast cells containing ClNAC681–319, ClNAC681–197, and ClNAC68198–319 on SD-T medium and SD-TH medium containing X-α-gal. pGBKT7-P53 and pGBKT7 were used as the positive and negative controls, respectively. F Schematic representation of different effectors, reporters, and internal controls. G Transcriptional regulatory activity of ClNAC68. The GALDB-VP16 and GALDB vectors were used as the positive and negative controls, respectively. Five independent replicates were performed. The asterisks denote significance compared with the negative control by two-way ANOVA: *p < 0.05, **p < 0.01. H Effects of different lengths of ClNAC68 on VP16-mediated LUC gene expression. The GALDB-VP16 and GALDB vectors were used as the positive and negative controls, respectively. Watermelon ClActin was used as the noninteractive control. Five independent replicates were performed. The asterisks denote significance compared with VP16 by two-way ANOVA: *p < 0.05, **p < 0.01, ***p < 0.001
To test the transcriptional activity of ClNAC68, we generated N- and C-terminal ClNAC68 constructs containing the NAC domain (aa 1–197) and transcription domain (aa 198–319), respectively (Fig. 1D). Then, Y2H and dual-luciferase reporter assays were performed. For the Y2H assay, all yeast cells containing BD-ClNAC681–319, BD-ClNAC681–197, BD-ClNAC68 198–319, or the positive or negative control survived in SD-T medium, while only those containing the positive control survived and appeared blue in SD-Trp/-His medium containing X-α-gal. No other yeast cells survived (Fig. 1E). These results indicated that ClNAC68 had no transcriptional activation activity. Then, we performed a dual-luciferase reporter assay to further detect its transcriptional activity in watermelon protoplasts. Plasmids were constructed as shown in Fig. 1F. The relative luciferase activities of full-length ClNAC681–319 and the transcription domain ClNAC68198–319 were significantly lower than those of the negative control, while there was no significant difference in relative luciferase activity between the NAC domain ClNAC681–197 and the negative control (Fig. 1G). Furthermore, these three constructs were individually coexpressed with VP16, and both the full-length ClNAC681–319 and the transcription domain ClNAC68198–319 had significant inhibitory effects on VP16-induced gene expression, while the NAC domain ClNAC681–197 and the negative control had no inhibitory effects (Fig. 1H). These results indicated that ClNAC68 is a transcriptional repressor and that its transcriptional repression domain is located in the C-terminus.
Knockout of ClNAC68 decreased sugar accumulation in flesh
To explore how ClNAC68 regulates fruit ripening and quality, we knocked out ClNAC68 by the CRISPR-Cas9 system (clnac68) in the cultivated inbred watermelon line ZZJM. The clnac68-13 mutant line showed a 1 bp insertion in the first target site and a 7 bp deletion in the second target site. The clnac68-51 mutant line showed a 5 bp deletion in the first target site and a 6 bp deletion in the second target site (Fig. 2A). Both the mutant lines exhibited loss of function of ClNAC68.
A Gene editing of clnac68-13 and clnac68-51. B Phenotype of WT and clnac68 mutant fruits at 18 DAP and 26 DAP. Bar = 10 cm. C Brix of WT and clnac68 mutant fruits at 18 DAP and 26 DAP. D–K Fructose D, glucose E, fucose F, galactose G, sucrose H, raffinose I, maltose J, and lactose K contents in WT and clnac68 mutant fruits at 26 DAP. The asterisks denote significance compared with WT fruits at different developmental stages by two-way ANOVA: *p < 0.05, **p < 0.01, ***p < 0.001
There was no significant difference in plant growth performance between the WT and clnac68 lines during plant development and fruit ripening. At 18 DAP and 26 DAP, the flesh color showed no significant difference between the WT fruits and clnac68 mutant fruits (Fig. 2B). The brix of clnac68 mutant fruits was significantly lower than that of WT fruits at both 18 DAP and 26 DAP, but the difference in brix at 18 DAP was less significant than that at 26 DAP (Fig. 2C). GC/MS analysis showed that among fructose, glucose, fucose, galactose, sucrose, raffinose, maltose, and lactose, only fructose, glucose, and sucrose exhibited a significantly lower content in both the clnac68-13 and clnac68-51 mutant flesh than in the corresponding WT flesh at 26 DAP (Fig. 2D-K). These results indicated that ClNAC68 positively regulated the accumulation of sugars, especially sucrose, fructose, and glucose, at the mature fruit ripening stage.
A previous study proved that IAA and ABA are important hormones that positively regulate fruit ripening and sugar accumulation26,27. Therefore, we measured the contents of free IAA and ABA in flesh at 18 DAP and 26 DAP, respectively. Free IAA and ABA were significantly lower at both the 18 DAP and 26 DAP in clnac68 mutant fruit than in WT fruit, and the difference in free IAA and ABA at 26 DAP was more significant than the difference at 18 DAP (Figure S1A and B). These results indicated that ClNAC68 might regulate watermelon fruit sweetness and ripening by mediating the contents of free IAA and ABA.
Knockout of ClNAC68 delayed seed maturation and inhibited seed germination and root elongation
Although there no significant difference in fruit ripening was found between the WT and clnac68 mutant fruits, the maturation of clnac68 mutant seeds was obviously delayed. At 18 DAP, the seeds of the WT line started to undergo coloration, while those of the clnac68 mutant lines remained yellow (Fig. 3A). At 26 DAP, the seed color of WT lines became completely black, while clnac68 mutant seeds were unevenly colored (Fig. 3B). The length of mutant seeds was also shorter than that of WT seeds (Figure S2A), while the seed width was not significantly different (Figure S2B). The hundred-seed weight of clnac68 mutant seeds was significantly lower than that of WT seeds (Fig. 3C). Due to the significantly different seed phenotype between the WT and clnac68 mutant lines, we measured hormone levels in seeds of WT and clnac68 mutant lines at 26 DAP and found that the IAA and ABA contents were also significantly lower in clnac68 mutant seeds (Fig. 3D and E). Then, we further tested the seed germination rate and root elongation performance. After 72 h of germination, WT seeds had completely germinated, while the germination rate of clnac68 mutant seeds was significantly lower, only 60%. clnac68 mutant seeds had completely germinated after 96 h of germination (Fig. 3F). To further confirm the effect of IAA and ABA on mutant seed germination, we measured the seed germination rates of the WT and clnac68 mutant lines after treatment with exogenous IAA and ABA. The seed germination rate of the clnac68 lines after exogenous IAA treatment was significantly higher than that after control (H2O) treatment, which indicated that IAA could partially restore seed germination in the clnac68 mutant (Fig. 3G). However, ABA treatment delayed seed germination in both the WT and clnac68 lines, and clnac68 mutant seeds started germinating after 60 h and germinated much more slowly than WT seeds (Fig. 3H).
Knockout of ClNAC68 inhibited seed germination. A–B Phenotype of WT and clnac68 mutant seeds at 18 DAP A and 26 DAP B. Bar=1 cm. C Hundred-seed weight of WT and clnac68 mutant lines. D–E Free IAA D and ABA E contents in WT and clnac68 mutant seeds at 26 DAP. F Seed germination rate of WT and clnac68 mutant lines under normal conditions. G–H. Seed germination rate of WT and clnac68 mutant lines under treatment with 10 μM IAA G or 10 μM ABA H. The asterisks denote significance compared with WT plants at different developmental stages by two-way ANOVA: *p < 0.05, **p < 0.01, ***p < 0.001
Regarding root elongation, the comparison between the WT and clnac68 mutant lines showed that after 2 days of treatment, there was no significant difference in root length between the WT line and clnac68 mutant lines (Fig. 4A). After 5 days of treatment, the root length in the clnac68 mutant lines was significantly lower than that in the WT line (Fig. 4B and C). Furthermore, the free IAA content in roots of the clnac68 mutant lines was significantly lower than that in the WT line (Fig. 4D) but the ABA content showed no significant difference between the WT and clnac68 mutant lines (Fig. 4E). Then, we further evaluated root elongation under treatment with exogenous free IAA. The root length of both the WT and clnac68 mutant lines was significantly increased after treatment with exogenous free IAA but the root length of the clnac68 mutant lines was significantly shorter than that of the WT line (Fig. 4F-H). These results indicated that ClNAC68 positively regulates seed germination and root elongation via the increased free IAA content.
Knockout of ClNAC68 inhibited root elongation. A–B Phenotype of WT and clnac68 mutant root elongation at two DAT A and five DAT B. C Root length (cm) of the WT and clnac68 mutant lines at two DAT and five DAT. D–E. Free IAA D and ABA E contents in roots in the WT and clnac68 mutant lines at two DAT and five DAT. F–G Phenotype of WT and clnac68 mutant root elongation under treatment with 10 μM IAA at two DAT F and five DAT G. H Root length (cm) of the WT and clnac68 mutant lines at two DAT and five DAT under H2O or IAA treatment. The asterisks denote significance compared with WT plants at different developmental stages by two-way ANOVA: *p < 0.05, **p < 0.01, ***p < 0.001
ClNAC68 binds directly to the invertase gene ClINV and represses its expression
To further explore the regulatory mechanism of ClNAC68, transcriptome analysis was performed on flesh and seeds at 26 DAP, which showed more significant differences in the clnac68 mutant. The ripening-related differentially expressed genes (DEGs) (p-adjust < 0.05 and | log2FC | >= 1) between the WT and clnac68-13 lines were selected to explore the potential target gene of ClNAC68. There were 501 DEGs (195 upregulated and 306 downregulated) at 26 DAP in clnac68-13 mutant fruits (Figure S3A), and there were 958 DEGs (527 upregulated and 431 downregulated) at 26 DAP in seeds (Figure S3B). Of these DEGs, 83 were found in both the fruits and seeds at 26 DAP, and 835 DEGs and 418 DEGs were specifically expressed in seeds and flesh, respectively, at 26 DAP (Figure S3C). KEGG analysis showed that the DEGs in flesh were clustered mainly into the energy metabolism, carbohydrate metabolism, and signal transduction pathways, while the DEGs in seeds were mainly clustered into the amino acid metabolism, biosynthesis of other secondary metabolites, carbohydrate metabolism, and signal transduction pathways (Figure S3D and E).
KEGG enrichment analysis was used to further investigate the DEGs in carbohydrate metabolism, including those related to sugar, coloration, metabolism, and hormone biosynthesis, in flesh. The top three enriched pathways were ‘galactose metabolism’, ‘starch and sucrose metabolism,’ and ‘pentose and glucuronate interconversions’ (Fig. 5A). These results indicated that ClNAC68 participated in sugar metabolism regulation in flesh. There were 7 DEGs involved in starch and sucrose metabolism: two invertase genes (beta-fructofuranosidase) involved in regulating sucrose degradation5, three amylase genes, an alpha-glucosidase involved in regulating hydrolysis of polysaccharides28, and a beta-glucosidase involved in regulating the free ABA content by catalyzing the conjugation of Glc to ABA (to form ABA-GE)29,30. Heatmap analysis of these DEGs showed that only one invertase gene, Cla97C10G201810, was significantly upregulated in clnac68 mutant fruits and that all the other genes were significantly downregulated in clnac68 mutant fruits (Fig. 5B). Cla97C10G201810 belongs to the cell wall invertase family, and we found that cell wall invertase activity was significantly higher in clnac68 mutant fruits (Fig. 5C). These results were consistent with the lower sucrose content in clnac68 mutant fruits. Furthermore, ClNAC68 was found to be a transcriptional repressor, and ClNAC68 might mediate the upregulation of Cla97C10G201810 (an invertase gene) expression and invertase activity to control sugar accumulation.
A KEGG enrichment analysis of DEGs involved in carbohydrate metabolism. B Heatmap analysis of DEGs involved in starch and sucrose metabolism in WT and clnac68 mutant fruits. C Invertase activity in WT and clnac68 mutant fruits at 26 DAP. The error bars show the standard deviations of three independent replicates. D EMSAs showed that ClNAC68 bound to the ClINV promoter. P1 and P2 denote the two different binding sites. MP1 and MP2 denote binding of the mutant probe to P1 and P2. Competitor denotes non-biotin-labeled probes. The untagged MBP protein incubated with biotin-labeled probes was used as the negative control. E. Schematic representation of effector and reporter vectors. F The transient dual-luciferase assay showed that ClNAC68 significantly repressed ClINV expression. Five independent replicates were performed. The asterisks denote significance compared with the negative control by two-way ANOVA: *p < 0.05, **p < 0.01, ***p < 0.001
To test this hypothesis, EMSA and dual-luciferase assays were performed. The motifs (T/A)(T/G)(A/G/C)CGT(G/A)(T/A) were previously reported as NAC transcription factor binding sites31. We analyzed the promoter (2 kb upstream of the start codon) of ClINV, and two binding sites were found. The EMSA showed that ClNAC68 bound to all the cis-elements in ClINV and that the level of bound complex decreased as the concentration of competitors increased, while ClNAC68 could not bind to the mutant probes. MBP could not bind to these probes (Fig. 5D). In the transient dual-luciferase assay, pGreenII 62-SK-ClNAC68 was used as the effector, and proClINV-LUC was used as the reporter (Fig. 5E). The LUC/REN ratio in leaves coexpressing ClNAC68 and proClINV-LUC was significantly lower than the LUC/REN ratio in mock control leaves (Fig. 5F). These results indicated that ClNAC68 directly binds to the promoter of ClINV and represses its expression.
Sugars Will Eventually be Exported Transporters (SWEET) were identified as novel sugar transporters and were characterized as candidate genes for maintaining sugar homeostasis in sink tissues. In this study, we analyzed SWEET gene expression in clnac68 mutant fruits. Nine SWEET genes were found, and three SWEETs were significantly downregulated in the clnac68 mutant lines (Figure S4).
ClNAC68 directly binds to the IAA deactivator ClGH3.6 and represses its expression
In this study, we found that the free IAA and ABA contents in both the flesh and seeds were significantly lower in clnac68 mutant lines. Thus, we focused on the DEGs in the related hormone biosynthesis and signaling pathways. KEGG enrichment analysis showed eight DEGs involved in the metabolism of terpenoids and polyketides; among these, one and three DEGs in flesh and seeds, respectively, were clustered into carotenoid biosynthesis (including ABA biosynthesis); two and five DEGs were clustered into diterpenoid biosynthesis (including gibberellin biosynthesis); and three and one DEGs were clustered into brassinosteroid biosynthesis. However, no DEGs were clustered into tryptophan metabolism (including IAA biosynthesis) in either flesh or seeds. Due to the reductions in free IAA and ABA, we analyzed the DEGs involved in ABA biosynthesis, and we found that the expression of the key ABA biosynthesis gene Cla97C07G137260, encoding 9-cis-epoxycarotenoid dioxygenase (NCED), was significantly lower in the flesh of clnac68 mutant fruits than in that of WT fruits at 26 DAP (Figure S5A). In seeds, the expression of ABA biosynthesis genes, including the violaxanthin de-epoxidase (VDE) gene Cla97C11G216330, NCED gene Cla97C06G117340, and ABA catabolic gene CYP707A Cla97C07G137840, was significantly lower in the clnac68 mutant lines (Figure S5B). Given that ClNAC68 was identified as a transcriptional repressor, these lower expression levels indicated that the DEGs involved in ABA metabolism are not the target genes of ClNAC68.
Given that no DEGs were involved in IAA biosynthesis, we further analyzed the plant hormone signaling pathways in both the flesh and seeds to understand how ClNAC68 regulates the content of free IAA. The DEGs in flesh and seeds were clustered mainly into the IAA signaling pathway. In flesh, four DEGs were upregulated, including auxin transporter (AUX1), auxin response factor (ARF), GH3, and auxin-responsive protein (AUX/IAA), while a gene encoding an auxin-responsive family protein (SAUR) was downregulated (Fig. 6A). In seeds, two GH3 genes and a SAUR gene were upregulated in clnac68 mutant seeds (Fig. 6B). Among these DEGs, only Cla97C05G096220, encoding GH3.6, was upregulated in both seeds and flesh. GH3.6 is a deactivator of IAA, and the expression of GH3.6 showed a negative correlation with the free IAA content32. Therefore, we predicted that ClNAC68 might regulate the free IAA content by repressing GH3.6 expression.
A–B Heatmap of DEGs in the IAA signaling pathway in WT and ClNAC68 fruits A and seeds B. C–D. Relative expression of ClGH3.6 in WT and clnac68 mutant fruits C and seeds D at 18 and 26 DAP. ClActin was used as the internal control. The error bars show the standard deviations of three independent replicates. E EMSAs showed that ClNAC68 bound to the ClGH3.6 promoter. MP denotes the mutant probe, and Competitor denotes the nonbiotin-labeled probes. The untagged MBP protein incubated with biotin-labeled probes was used as the negative control. F The transient dual-luciferase assay showed that ClNAC68 significantly repressed ClGH3.6 expression. Five independent replicates were performed. Empty pGreen62SK vector was used as the negative control. The asterisks denote significance compared with the negative control by two-way ANOVA: *p < 0.05, **p < 0.01, ***p < 0.001
To assess the relationships between ClNAC68 and IAA signaling genes, we analyzed the promoters of these upregulated genes and found a NACRS in the promoter of ClGH3.6 but not in the other genes. The relative expression of ClGH3.6 was significantly higher in both the flesh and seeds at 18 DAP and 26 DAP (Fig. 6C and D). EMSA showed that ClNAC68 could bind to the NACRS in the ClGH3.6 promoter and that the binding activity decreased with an increase in the probe concentration, while ClNAC68 could not bind to the mutant probe, and the negative control could not bind to the NACRS (Fig. 6E). Further transient dual-luciferase assays were performed in tobacco, and the results showed that in the presence of ClNAC68, the LUC/REN ratio was significantly lower than the LUC/REN ratio in the mock control (Fig. 6F). These results indicated that ClNAC68 can directly bind to the promoter of ClGH3.6 and suppress its expression. In summary, these results demonstrated that ClNAC68 increases the accumulation of free IAA in both the flesh and seeds by directly repressing the expression of the IAA deactivator ClGH3.6.
Discussion
NAC is one of the largest gene families in plants and is involved in stress tolerance regulation33, plant development, and fruit ripening16,34. In banana, MaNAC1 and MaNAC2 are transcriptional repressors and negatively regulate MaERF11 to induce ethylene accumulation and fruit ripening35. Most previous studies focused on mechanisms of development and ripening, and studies of how NAC transcription factors regulate fruit quality are limited. In maize, ZmNAC34 functions as a transcriptional repressor, and overexpression of ZmNAC34 decreases the soluble solid content and starch content by downregulating the expression of starch biosynthesis-related genes36. In this study, we characterized the transcriptional repressor ClNAC68, which was highly expressed in sweet flesh (Fig. 1). Knockout of ClNAC68 resulted in reduced sugar accumulation, and the total contents of sugar, sucrose, fructose, and glucose were significantly lower in clnac68 mutant fruits, especially at 26 DAP (Fig. 2). Cla97C10G201810, belonging to the cell wall invertase family, is directly regulated by ClNAC68 (Fig. 5). A previous study showed that silencing lycopersicum invertase 5 (LIN5), a cell wall invertase in tomato, reduced brix and the fructose content but increased sucrose accumulation37. In clnac68 mutant fruits, ClINV activity was increased, and this increase might induce sucrose degradation and result in a decrease in the total sugar content and sucrose accumulation. These results indicated that ClNAC68 regulates the sugar content by repressing invertase activity in watermelon. Sucrose is transported from photosynthetic leaf sources to nonphotosynthetic sink organs by sucrose transporters (SUTs) and is then hydrolyzed by invertases and sucrose synthases to maintain sink strength38. Significant changes in the metabolic enzymes involved in sugar metabolism directly affect sweetness in fruits39, and sugar transporters control sucrose allocation40. SWEET, as a sugar transporter, directly determines the sugar content in fruits41. In this study, three SWEETs were significantly downregulated in the clnac68 mutant lines. Lower sweet activity might result in lower glucose and fructose contents (Figure S4).
Plant hormones play important roles in the regulation of plant growth, development, and fruit ripening2. In our previous study, we found that the ABA content increased during watermelon fruit ripening, and that treatment with exogenous ABA or an ABA inhibitor could induce or inhibit fruit ripening, respectively26. The ABA content was significantly lower in clnac68 mutant fruits, and transcriptome data showed that the expression of the ABA biosynthesis gene NCED (Cla97C07G137260) was significantly lower in mutant fruits (Figure S5A). Moreover, Cla97C08G153160 encodes a beta-glucosidase and is homologous to AtBG1, which catalyzes the formation of ABA-GE to activate ABA and was significantly downregulated in mutant fruits (Figure S5A). These results indicated that ClNAC68 may affect the ABA content by downregulating ClNCED and ClBG1. An increasing number of studies have shown that NAC transcription factor can directly bind to the ABA biosynthesis gene NCED to regulate ABA accumulation. Given that ClNAC68 was found to be a transcriptional repressor, the downregulated ClNCED and ClBG1 were not its direct targets in this study. More comprehensive research is needed to explore the interaction mechanism between ClNAC68 and other regulators of ABA biosynthesis in the future.
The fruit ripening process is regulated by the metabolism of multiple hormones and by multiple signaling pathways. In contrast to the positive regulatory function of ABA in fruit ripening, the function of IAA in the regulation of fruit ripening is still under debate. Previous studies showed that the IAA content decreased during fruit ripening and affected fruit acidity and sugar accumulation42,43, while several studies showed that the IAA content increased during fruit ripening and peaked at the mature stage in peach27. In this study, we found that the content of free IAA increased during fruit ripening and seed maturation and was significantly lower in both the fruits and seeds of the clnac68 mutants (Figs. 3 and 4). Transcriptome analysis showed several DEGs in the IAA signaling pathway, but the function of IAA signaling in the regulation of fruit ripening is still unclear. VcIAA27 plays a negative role in the IAA signaling pathway44, while PpIAA19 expression increases during fruit ripening to change fruit shape45. The other key component, ARF, also showed opposite functions during fruit ripening: overexpression of SlARF10 accelerated fruit ripening and induced sugar accumulation46; however, overexpression of SlARF4 delayed fruit ripening42. Both the Cla97C06G119080, encoding Aux/IAA, and Cla97C05G099830, encoding ARF, were significantly upregulated in clnac68 mutant fruits, but how they regulate watermelon fruit ripening is still unknown. Further studies are needed to confirm the functions of IAA and IAA signaling in the regulation of watermelon fruit ripening.
Seed development and seed size are also affected by the levels of multiple hormones. In a previous study, the free IAA content was found to be increased during development in both the variant with larger seeds and the variant with smaller seeds, and the free IAA content was significantly higher in the variant with larger seeds14. In this study, seed maturation was delayed and the seed size was decreased in the clnac68 mutants. We suspected that the lower free IAA content in clnac68 mutant seeds might result in this phenotype. Due to delayed seed maturation, the germination and root elongation of clnac68 mutant seeds were significantly decreased compared with those of WT seeds. The expression of two GH3s (Cla97C05G096220 and Cla97C07G138770) involved in IAA signaling was increased in mutant seeds, while no DEGs involved in IAA biosynthesis were identified in either flesh or seeds. Further EMSA and transient dual-luciferase assays showed that ClNAC68 can directly bind to the NACRS in the promoter of ClGH3.6 (Fig. 6). These results indicated that ClNAC68 mediates the negative regulation of the IAA deactivator ClGH3.6 to increase the free IAA content in seeds, and these results were supported by a previous study20.
ABA induces seed dormancy and inhibits seed germination47, and we found that the ABA content was significantly lower in clnac68 mutant seeds than in WT seeds. The DEGs in the ABA biosynthesis pathway, including NCED and VDE, were significantly downregulated in the clnac68 mutant, while CYP707A, involved in ABA catabolism, was also significantly downregulated in mutant seeds. Given that ClNAC68 was found to be a transcriptional repressor, these downregulated DEGs were not the target of ClNAC68. A previous study showed that this NAC transcription factor positively regulates ABI5 expression to increase ABA sensitivity and affect seed germination48. In this study, neither WT nor clnac68 mutant seeds showed hypersensitivity to exogenous ABA treatment, and no DEGs in the ABA signaling pathway were identified in clnac68 mutant seeds. These results might indicate that ClNAC68 mainly regulates seed development via the IAA signaling pathway and then affects seed germination and root elongation. How NAC regulates ABA biosynthesis and signaling in watermelon is still unknown, and more research is needed to explore the regulatory mechanism in both flesh and seeds.
In summary, ClNAC68 was found to be a transcriptional repressor that was highly expressed in flesh. Knockout of ClNAC68 significantly reduced the sucrose, fructose, and glucose contents. ClNAC68 directly inhibited ClINV expression and invertase activity to prevent sucrose degradation. Furthermore, ClNAC68 might regulate the free IAA content by directly suppressing ClGH3.6 expression in seeds, resulting in seed maturation and germination. These findings revealed that ClNAC68 plays an important role in sugar metabolism and the IAA signaling pathway and provide new insight for modifying breeding practices to improve the fruit quality of watermelon.
Change history
18 December 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41438-021-00710-z
References
Pott, D. M., Vallarino, J. G. & Osorio, S. Metabolite changes during postharvest storage: effects on fruit quality traits. Metabolites. 10, 187 (2020).
Chen, T., Qin, G. & Tian, S. Regulatory network of fruit ripening: current understanding and future challenges. New Phytol. https://doi.org/10.1111/nph.16822 (2020).
Guo, S. et al. Resequencing of 414 cultivated and wild watermelon accessions identifies selection for fruit quality traits. Nat. Genet. 51, 1616–1623 (2019).
Duran-Soria, S., Pott, D. M., Osorio, S. & Vallarino, J. G. Sugar signaling during fruit ripening. Front. Plant Sci. 11, 564917 (2020).
Manoochehri, H. et al. A review on invertase: Its potentials and applications. Biocatalysis Agric. Biotechnol. 25, 101599, https://doi.org/10.1016/j.bcab.2020.101599 (2020).
Deng, Y., Wang, J., Zhang, Z. & Wu, Y. Transactivation of Sus1 and Sus2 by Opaque2 is an essential supplement to sucrose synthase-mediated endosperm filling in maize. Plant Biotechnol. J. 18, 1897–1907 (2020).
Wei, W. et al. Pitaya HpWRKY3 is associated with fruit sugar accumulation by transcriptionally modulating sucrose metabolic genes HpINV2 and HpSuSy1. Int. J. Mol. Sci. 20, 1890 (2019).
Liu, J. et al. Dynamic characteristics of sugar accumulation and related enzyme activities in sweet and non-sweet watermelon fruits. Acta Physiologiae Plant. 35, 3213–3222 (2013).
McAtee, P., Karim, S., Schaffer, R. & David, K. A dynamic interplay between phytohormones is required for fruit development, maturation, and ripening. Front. Plant Sci. 4, 79 (2013).
Liu, X. et al. Cucumber fruit size and shape variations explored from the aspects of morphology, histology, and endogenous hormones. Plants-Basel. 9, 772 (2020).
Tatsuki, M. et al. Insertion of a transposon-like sequence in the 5 ‘-flanking region of the YUCCA gene causes the stony hard phenotype. Plant J. 96, 815–827 (2018).
Staswick, P. E. et al. Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17, 616–627 (2005).
Zhao, D. et al. Overexpression of MsGH3.5 inhibits shoot and root development through the auxin and cytokinin pathways in apple plants. Plant J. 103, 166–183 (2020).
Monpara, J. K., Chudasama, K. S. & Thaker, V. S. Role of Phytohormones in soybean (Glycine max) seed development. Russian J. Plant Physiol. 66, 992–998 (2019).
Han, X. L., Zhao, F. Y., Wang, Z. L., Che, X. & Cui, G. C. The Role of OsYUCCA2 in auxin synthesis and promotion of rice growth and development. Russian J. Plant Physiol. 67, 1018–1027 (2020).
Zhu, M. et al. A new tomato NAC (NAM/ATAF1/2/CUC2) transcription factor, SlNAC4, functions as a positive regulator of fruit ripening and carotenoid accumulation. Plant Cell Physiol. 55, 119–135 (2014).
Olsen, A. N., Ernst, H. A., Lo Leggio, L. & Skriver, K. NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci. 10, 79–87 (2005).
Zhang, S. et al. A novel NAC transcription factor, MdNAC42, regulates anthocyanin accumulation in red-fleshed apple by interacting with MdMYB10. Tree Physiol. 40, 413–423 (2020).
Ma, X. et al. The NAC transcription factor SlNAP2 regulates leaf senescence and fruit yield in tomato. Plant Physiol. 177, 1286–1302 (2018).
Mao, C. et al. OsNAC2 integrates auxin and cytokinin pathways to modulate rice root development. Plant Biotechnol. J. 18, 429–442 (2020).
Park, J. et al. Integration of auxin and salt signals by the NAC transcription factor NTM2 during seed germination in Arabidopsis. Plant Physiol. 156, 537–549 (2011).
Lv, X. et al. Global expressions landscape of NAC transcription factor family and their responses to abiotic stresses in Citrullus lanatus. Sci. Rep. 6, 30574 (2016).
Tian, S. et al. Efficient CRISPR/Cas9-based gene knockout in watermelon. Plant Cell Rep. 36, 399–406 (2017).
Ren, Y. et al. Localization shift of a sugar transporter contributes to phloem unloading in sweet watermelons. N. Phytologist 227, 1858–1871 (2020).
Wei, W. et al. The Alfin-like homeodomain finger protein AL5 suppresses multiple negative factors to confer abiotic stress tolerance in Arabidopsis. Plant J. 81, 871–883 (2015).
Wang, Y. et al. Abscisic acid pathway involved in the regulation of watermelon fruit ripening and quality trait evolution. PLos ONE. 12, e0179944 (2017).
Liu, N. Effects of IAA and ABA on the immature peach fruit development process. Horticultural Plant J. 5, 145–154 (2019).
Hossain, M. A., Rana, M. M., Kimura, Y. & Roslan, H. A. Changes in biochemical characteristics and activities of ripening associated enzymes in mango fruit during the storage at different temperatures. Biomed. Res. Int. 2014, 232969 (2014).
Liang, B. et al. Overexpression of the persimmon abscisic acid beta-glucosidase gene (DkBG1) alters fruit ripening in transgenic tomato. Plant J. https://doi.org/10.1111/tpj.14695 (2020).
Zhang, S. H. et al. Enzymatic and functional analysis of beta-glucosidase FaBG1 during strawberry fruit ripening. J. Horticultural Sci. Biotechnol. 89, 733–739 (2014).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell. 165, 1280–1292 (2016).
Bernales, M. et al. Expression of two indole-3-acetic acid (IAA)-amido synthetase (GH3) genes during fruit development of raspberry (Rubus idaeus Heritage). Sci. Horticulturae. 246, 168–175 (2019).
Hu, H. et al. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc. Natl Acad. Sci. USA. 103, 12987–12992 (2006).
Zhu, F. et al. A NAC transcription factor and its interaction protein hinder abscisic acid biosynthesis by synergistically repressing NCED5 in Citrus reticulata. J. Exp. Bot. 71, 3613–3625 (2020).
Shan, W. et al. MaXB3 Modulates MaNAC2, MaACS1, and MaACO1 stability to repress ethylene biosynthesis during banana fruit ripening. Plant Physiol. 184, 1153–1171 (2020).
Peng, X. et al. A maize NAC transcription factor, ZmNAC34, negatively regulates starch synthesis in rice. Plant Cell Rep. 38, 1473–1484 (2019).
Zanor, M. I. et al. RNA interference of LIN5 in tomato confirms its role in controlling brix content, uncovers the influence of sugars on the levels of fruit hormones, and demonstrates the importance of sucrose cleavage for normal fruit development and fertility. Plant Physiol. 150, 1204–1218 (2009).
Braun, D. M., Wang, L. & Ruan, Y.-L. Understanding and manipulating sucrose phloem loading, unloading, metabolism, and signalling to enhance crop yield and food security. J. Exp. Bot. 65, 1713–1735 (2014).
Nookaraju, A. et al. Molecular approaches for enhancing sweetness in fruits and vegetables. Sci. Horticulturae. 127, 1–15 (2010).
Baker, R. F., Leach, K. A. & Braun, D. M. SWEET as sugar: new sucrose effluxers in plants. Mol. Plant. 5, 766–768 (2012).
Zhang, Z. et al. VvSWEET10 mediates sugar accumulation in grapes. Genes. 10, 255 (2019).
Sagar, M. et al. SlARF4, an auxin response factor involved in the control of sugar metabolism during tomato fruit development. Plant Physiol. 161, 1362–1374 (2013).
Fenn, M.A. & Giovannoni, J.J. Phytohormones in fruit development and maturation. Plant Mol. Biol. https://doi.org/10.1111/tpj.15112 (2020).
Hou, Y. et al. Identification and functional characterization of the Aux/IAA gene VcIAA27 in blueberry. Plant Signal. Behav. 15, 1700327 (2020).
Ding, Y. et al. Over-expression of peach PpIAA19 in tomato alters plant growth, parthenocarpy, and fruit shape. J. Plant Growth Regul. 38, 103–112 (2019).
Yuan, Y. et al. SIARF10, an auxin response factor, is involved in chlorophyll and sugar accumulation during tomato fruit development. J. Exp. Bot. 69, 5507–5518 (2018).
Zhao, H. et al. ABI5 modulates seed germination via feedback regulation of the expression of thePYR/PYL/RCARABA receptor genes. N. Phytologist. 228, 596–608 (2020).
Wang, J. et al. CsATAF1 positively regulates drought stress tolerance by ABA-dependent pathway and promoting ROS scavenging in cucumber. Plant Cell Physiol. https://doi.org/10.1093/pcp/pcy030 (2018).
Acknowledgements
This work was financially supported by the Collaborative Innovation Center of BAAFS (KJCX201907-2), Beijing Science & Technology Program (D171100007617001), National Key R&D Program of China (2018YFD0100703), National Natural Science Foundation of China (31930096 and 1902034), Beijing Natural Science Foundation (6204038 and 6202010), and Beijing Scholar Program (BSP026 and YBSP019).
Author information
Authors and Affiliations
Contributions
J.-F.W., Y.-P.W., H.-Y.Z., S.-G.G., and Y. X. designed the research; J.-F.W., Y.-P.W., S.-G.G., Y.-T.Y., J.Z., Y.R., M.-Y.L., S.-W. T., and Y. Z. performed the experiments; J.-F.W., Y.-P. W., Y.-T.Y., and J.Z. analyzed the data; G.-Y.G., H.-Y.Z., S.-G.G., and Y.X. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Wang, Y., Zhang, J. et al. The NAC transcription factor ClNAC68 positively regulates sugar content and seed development in watermelon by repressing ClINV and ClGH3.6. Hortic Res 8, 214 (2021). https://doi.org/10.1038/s41438-021-00649-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-021-00649-1
This article is cited by
-
Telomere-to-telomere Citrullus super-pangenome provides direction for watermelon breeding
Nature Genetics (2024)
-
Applications of CRISPR/Cas genome editing in economically important fruit crops: recent advances and future directions
Molecular Horticulture (2023)
-
CRISPR/Cas9 mediated editing of phytoene desaturase gene in squash
Journal of Plant Biochemistry and Biotechnology (2023)
-
Identification of a stable major sucrose-related QTL and diagnostic marker for flavor improvement in peanut
Theoretical and Applied Genetics (2023)
-
Correction: The NAC transcription factor ClNAC68 positively regulates sugar content and seed development in watermelon by repressing ClINV and ClGH3.6
Horticulture Research (2021)