Abstract
Flavonols and proanthocyanidins (PAs) are the main pigments in the black spines of cucumber (Cucumis sativus) fruit, and CsMYB60 is a key regulator of the biosynthesis of flavonols and PAs. However, in cucumber, the tissue distribution pattern of flavonols and PAs and the mechanism of their biosynthesis regulated by CsMYB60 remain unclear. In this study, we clarified the tissue-specific distribution of flavonoids and the unique transcriptional regulation of flavonoid biosynthesis in cucumber. CsMYB60 activated CsFLS and CsLAR by binding to their promoters and directly or indirectly promoted the expression of CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein, resulting in the formation of complexes of these four proteins to increase the expression of Cs4CL and interact with CsTATA-box binding protein to regulate the expression of CsCHS, thereby regulating the biosynthesis of flavonols and PAs in cucumber. Our data provide new insights into the molecular mechanism of flavonoid biosynthesis, which will facilitate molecular breeding to improve fruit quality in cucumber.
Similar content being viewed by others
Introduction
Flavonols and proanthocyanidins (PAs), which are polyphenolic compounds synthesized through the flavonoid biosynthetic pathway, are widely distributed in plants and have a variety of biological functions, including UV light protection, pigmentation, and anti-herbivore and phytopathogen resistance activities1,2,3,4,5,6, in addition to presenting many benefits for human health7,8,9. Flavonols play pivotal roles in modulating reactive oxygen species induced by abscisic acid to control the stomatal aperture10 and protect plants against UV light11,12. More importantly, flavonols significantly contribute to protection against cardiovascular diseases because of their well-recognized antioxidant, anti-inflammatory, and vasorelaxant actions13. PAs play roles in the coloration of plant seeds and in plant adaptation to biotic and abiotic stresses2,14,15. Moreover, PAs and their monomeric building blocks (catechin and epicatechin) can act as potential antioxidants showing beneficial effects on human health by protecting against free radical-mediated injury and cardiovascular diseases7,16. In addition, PAs also contribute to the taste of multiple beverages and fruits, for instance, tea, wine, and fruit juices17,18. Therefore, it is of great importance to clarify the function of flavonols and PAs and the molecular mechanism of their biosynthesis to improve plant stress resistance and fruit quality in cucumber (Cucumis sativus).
During plant development, the accumulation of flavonols and PAs is restricted to specific cell types and organs. In white clover (Trifolium repens), PAs and their flavan-3-ol precursors were specifically stained with dimethylaminocinnamaldehyde (DMACA) reagent and localized to the epidermal cell layer of the floral organs, showing an adaxial–abaxial asymmetric pattern19. PA accumulation is also frequently found specifically in the trichomes on plant organs at relatively low levels20. In Arabidopsis thaliana, PAs are synthesized and accumulate only in the seed coat and localize to endothelial cells derived from the inner integument21, whereas flavonols are found in all tissues22. Flavonols accumulate mainly in the skins and flowers of grape berries and in the leaves and stems. No detectable amounts of flavonols have been identified in the seeds and pulp of Shiraz berries and Chardonnay23,24. In cucumber, flavonols and PAs are the main pigments of the black spines6, but their distribution pattern in other tissues is unknown.
The biosynthetic pathways of flavonols and PAs have been well clarified by studying pigment-deficient mutants of model species25,26. In addition to structural genes, many transcription factors (TFs, which act as modulators of gene expression at the transcriptional level through sequence-specific DNA binding or protein–protein interactions during chromatin remodeling) are required for flavonol and PA biosynthesis27,28. The key enzyme 4-coumarate: CoA ligase (4CL) provides the precursors for lignin and flavonoid biosynthesis29. The first committed step in flavonoid biosynthesis, which is a major pathway of plant secondary metabolism, is catalyzed by chalcone synthase (CHS)30. Flavonol synthase (FLS) catalyzes the synthesis of flavonols from dihydroflavonols31. An alternative PA monomer (catechin) is thought to be produced from leucoanthocyanidin by leucoanthocyanidin reductase (LAR). LAR genes have been isolated from several plant species, and the activity associated with PA accumulation has been characterized32,33,34. In almost all plant species studied so far, the structural genes of the flavonoid biosynthetic pathway are mainly regulated by certain TFs or by a MYB-bHLH-WD40 (MBW) complex at the transcriptional level35,36,37,38. In Arabidopsis, AtMYB11, AtMYB12, and AtMYB111 regulate AtFLS1 and early steps in the control of flavonol biosynthesis, while an MBW complex activates later flavonoid biosynthetic genes to control the production of anthocyanins and PAs22,30,39. MBW regulatory complexes also control PA biosynthesis in strawberry fruits37. MtPAR is an MYB transcription factor that has a positive regulatory effect on PA biosynthesis in Medicago truncatula40. In cucumber, an R2R3-MYB gene, CsMYB60 was mapped as the best candidate for the B (black spine) locus in a 50 kb region on chromosome 441,42. Although CsMYB60 is a known key regulatory gene in the biosynthesis of flavonols and PAs responsible for the coloration of black spines6, it is still unclear how CsMYB60 regulates the structural genes of the flavonol and PA biosynthetic pathways.
Here, we clarified the tissue-specific distribution of flavonols and PAs and the unique mechanism of flavonol and PA biosynthesis regulated by CsMYB60 in cucumber. CsMYB60 was shown to directly activate CsFLS and CsLAR, two key genes for flavonol and PA biosynthesis, respectively. Additionally, CsMYB60 activated Cs4CL transcription through a complex composed of CsMYB60 and its interacting proteins CsbHLH42 (CsaV3_6G037080.1) and/or CsMYC1 (CsaV3_6G000530.1), CsWD40 (CsaV3_1G031140.1), and CsTATA-box binding protein (CsaV3_1G041520.1) and simultaneously promoted the expression of CsCHS by interacting with CsTATA-box binding protein, thereby regulating the biosynthesis of flavonols and PAs in cucumber.
Results
Distribution patterns of flavonols and PAs in white-spined and black-spined cucumbers
The main pigments of the black spines of cucumber have been identified as flavonols and PAs6. Hence, we sought to determine the patterns and contents of flavonols and PAs in different tissues between white-spined and black-spined cucumber lines. Therefore, we performed DMACA staining for the analysis of PA contents in different tissues. The results indicated that large amounts of PAs specifically accumulated in black spines, and there were also detectable amounts in the pericarp, pulp, seed coat, and male flowers of the black-spined cucumber line, while there were detectable amounts of PAs only in the pericarp, pulp, and seed coat of the white-spined line (Fig. 1a, b). Catechin contents of approximately 43.8, 10.9, 5.9, 5.0, 2.0, and 1.2 μg per gram fresh weight (gFW) were detected in the spines, pericarp, pulp, seed coat, male flowers, and roots, respectively, of the black-spined cucumber, while the contents in the white-spined cucumber were only 2.0, 1.5, and 1.0 μg/gFW in the pericarp, pulp, and seed coat, respectively (Fig. 1b).
a DMACA (dimethylaminocinnamaldehyde) staining of black-spined and white-spined cucumber lines. b Quantification of soluble PAs from (a) based on spectrophotometric absorbance. c DMACA staining of cucumber black spines at different developmental stages. d Quantification of soluble PAs from (c) based on spectrophotometric absorbance. e, f HPLC analysis of flavonols in different tissues of black-spined and white-spined cucumber lines. Absorbance was monitored at 365 nm. Peaks A–E represent five kinds of flavonols: quercetin-3-O-rutinoside-7-O-glucose (A), kaempferol-3-O-rutinoside-7-O-glucoside (B), isorhamnetin-3-O-rutinoside-7-O-glucoside (C), kaempferol-3-O-rutinoside (D), and isorhamnetin-3-O-rutinoside (E). R, root; St, stem; YL, young leaf; MF, male flower; Sp, spine at 12 days after anthesis (DAA); T, tendril; Pe, pericarp; Pu, pulp; Se, seed; SC, seed coat. B-4, black spine at 4 days before anthesis (DBA); B0, black spine at 0 DBA; B4, black spine at 4 DAA; B8, black spine at 8 DAA; B12, black spine 12 at DAA; B16, black spine at 16 DAA
The coloration of black spines occurs gradually and is complete at 12 days after anthesis (DAA); the accumulation of flavonols is consistent with the coloration process of black spines6. Here, our data showed that PA accumulation presented the same trend as the coloration of black spines (Fig. 1c, d). These data further strengthened the conclusion that flavonols and PAs were the main pigments determining the coloration of black spines6 (Fig. 1c, d).
In contrast to the PA distribution pattern, flavonols were distributed in all tested tissues of black-spined cucumber, but high levels of flavonols accumulated only in the male flowers and leaves of white-spined cucumber (Figs. 1e, f and S1). Five flavonols were verified: quercetin-3-O-rutinoside-7-O-glucose, kaempferol-3-O-rutinoside-7-O-glucoside, isorhamnetin-3-O-rutinoside-7-O-glucoside, kaempferol-3-O-rutinoside, and isorhamnetin-3-O-rutinoside6 (Figs. 1e, f and S1).
Overexpression of CsMYB60 increases the accumulation of flavonols and PAs in different tissues, especially in spines
As the best candidate gene for the B locus41, CsMYB60 has been verified to be a key regulator of the biosynthesis of flavonols and PAs in black spines through a transient expression assay6. Moreover, CsMYB60 shares the highest similarity among its homologous proteins with AtMYB111, which functions in flavonoid biosynthesis30 (Fig. S2). To obtain additional genetic evidence and more information about the function of CsMYB60 in the biosynthesis of flavonols and PAs, we constitutively expressed CsMYB60 in Xintaimici with white fruit spines (wild type, WT), which is a north China-type cucumber inbred line. A total of six independent first-generation transgenic lines (T1, OX1-6) were obtained, and CsMYB60 expression levels were significantly higher in all six T1 lines compared with WT and even compared with the black-spined line RNS9 (except for OX6) (Fig. 2f).
a Cucumber ovaries at 0 DBA. b Cucumber fruits at 4 DAA. c The spine color of cucumber fruits at 4 DAA from the Xintaimici (wild type, WT) and OX-1 lines observed under a stereoscopic microscope. d Cross-section of cucumber fruits at 4 DAA from the WT and OX-1 lines observed under a stereoscopic microscope. e The trichome color of male flower receptacles at 0 days before anthesis (DBA) from the WT and OX-1 lines observed under a stereoscopic microscope. f Relative expression levels of CsMYB60 in the WT, RNS9, and 35S:CsMYB60 transgenic lines. The young leaves of the WT, RNS9, and 35S:CsMYB60 transgenic lines were used for qRT-PCR analysis. g Relative expression levels of CsMYB60 in different tissues from WT, RNS9, and OX-1. R, young root at 10 days after sowing; S, young stem; YL, young leaf; MF, male flower at 0 DBA; O, ovary at 0 DBA; T, young tendril obtained from plants after 8 weeks in the soil. The cucumber β-actin gene was used as an internal control for normalization, and three biological replicates were performed for these experiments. Error bars indicate SE. Different letters indicate significant differences (P < 0.05). DBA, days before anthesis; DAA, days after anthesis. Scale bars: 0.5 cm in (a) and (b); 1 mm in (c)–(e)
Based on the CsMYB60 expression level and the phenotype of black spines, OX1 was selected from the black-spined transformants for further studies (Figs. 2a, b and S3). OX1 showed a similar process of pigment accumulation in the fruit spines and a similar pattern in the flower trichomes to RNS9 (Figs. 2a–e and S3). Quantitative real-time RT-PCR (qRT-PCR) analysis showed that the CsMYB60 expression levels were higher in various tested tissues of OX1 than in those of WT, similar to the pattern observed in RNS9 (Fig. 2g).
To verify the identity of the accumulated pigments as flavonols and PAs and their distribution patterns in OX1, T2 nontransgenic and transgenic siblings isolated from T1 selfing progenies of OX1 were used for the analyses of flavonols and PAs via high-performance liquid chromatography (HPLC) and DMACA staining. The flavonol contents were significantly increased to different extents in all tested tissues of the transgenic siblings, showing similar patterns to those in RNS9 (Figs. 1e, S1, S4a, b, and S5); similarity to RNS9 was also observed for the PA contents, which were significantly increased in most tested tissues of the transgenic siblings compared to nontransgenic siblings (Figs. 1b and S4c). Moreover, we examined the expression of key genes encoding enzymes responsible for flavonoid biosynthesis by real-time PCR analysis in the transgenic and nontransgenic lines. CsMYB60 and five structural genes, Cs4CL, CsCHS, CsFLS, CsDFR, and CsLAR, were significantly upregulated in the transgenic siblings (Fig. S4d).
CsMYB60 directly activates CsFLS and CsLAR by binding to their promoters
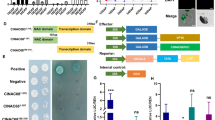
Since CsMYB60 promotes the expression of Cs4CL, CsCHS, CsFLS, CsDFR, and CsLAR, which are the key genes in flavonoid biosynthesis (Figs. 3a and S4d), we wanted to know how CsMYB60 regulates these genes. Therefore, yeast one-hybrid (Y1H) assays were performed to determine whether CsMYB60 binds to the promoters of these genes. The Y1H results showed that CsMYB60 could directly bind to the promoters of CsFLS and CsLAR, but not the promoters of Cs4CL, CsCHS, and CsDFR (Fig. 3b). Transient expression assays with the β-glucuronidase reporter gene (GUS) in the leaves of tobacco (Nicotiana benthamiana) showed that CsMYB60 could activate the expression of CsFLS and CsLAR in vivo (Fig. 3c).
a Simplified diagram of the flavonoid biosynthetic pathway. CHS, chalcone synthase; CHI, chalcone isomerase; C4H cinnamate 4-hydroxylase; 4CL, 4-coumaroyl CoA ligase; DFR, dihydroflavanol 4-reductase; F3H, flavanone 3-hydroxylase; F3′H, flavonoid 3′-hydroxylase; FLS, flavonol synthase; PAL, phenylalanine ammonia lyase; GST, glutathione S-transferase; LAR, leucoanthocyanidin reductase. b CsMYB60 could bind to the promoters of CsFLS and CsLAR in a yeast one-hybrid assay. c CsMYB60 activated the expression of GUS driven by CsFLS and CsLAR, respectively, in transient tobacco leaf assays. Three biological replicates were performed for the gene expression analyses. Error bars represent SE. Statistical significance: *P < 0.05, **P < 0.01. d Schematic diagram of CsMYB60 binding cis-elements in the promoters of CsFLS and CsLAR. The translational start site (ATG) is shown at position 0. e–g GST-CsMYB60 could directly bind to the motifs of the CsFLS and CsLAR promoters in EMSAs. The GST protein was incubated with the labeled probe in the first lane to serve as a negative control. 10- and 20-fold excesses of unlabeled probes were added for competition. The mutant probe contained two nucleotide mutation
To identify the cis-elements to which CsMYB60 could bind, different fragments from the CsFLS and CsLAR promoters were tested in Y1H assays. We identified one cis-element (CAGTTG) in the CsFLS promoter, which belonged to the MYB-core type I element43,44, and two cis-elements (TGGTTG and ACCCAC) in the CsLAR promoter, which were a MYB-core type II element43 and an AC-rich element22 (Figs. 3d and S6). These three cis-elements were further proven to be bound by CsMYB60 through EMSAs (Fig. 3e–g).
CsMYB60 physically interacts with CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein and directly or indirectly activates their expression
CsMYB60 greatly induced the expression of Cs4CL and CsCHS (Fig. S4d) but could not bind to their promoters (Fig. 3b). To search for CsMYB60-interacting partners that could bind to the Cs4CL and CsCHS promoters, we employed CsMYB60 as bait to screen a cucumber cDNA library via yeast two hybrid (Y2H) assays, and 12 putative interacting proteins were obtained (Table 1). Then, the interactions between these 12 proteins and CsMYB60 were confirmed via Y2H assays (Figs. 4a and S7). Among the 12 putative interacting proteins, we focused on the four transcription factors CsbHLH42 (CsaV3_6G037080.1), CsMYC1 (CsaV3_6G000530.1), CsWD40 (CsaV3_1G031140.1), and CsTATA-box binding protein (CsaV3_1G041520.1), which were repeatedly identified when we screened the Y2H library. Then, we confirmed the interactions of CsMYB60 with CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein in bimolecular fluorescence complementation (BiFC) and pull-down assays (Fig. 4b–f).
a Yeast two-hybrid assays. b Bimolecular fluorescence complementation (BiFC) assays. c–f Pull-down assays validating the interaction of CsMYB60 with CsbHLH42 (c), CsMYC1 (d), CsWD40 (e), and CsTATA-box binding protein (f). g Relative expression levels of CsMYB60, CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein in spines at 4 days before anthesis in OX-1 and its nontransgenic siblings. The cucumber β-actin gene was used as an internal control. Three biological replicates were performed for the gene expression analyses. Error bars represent standard errors (SE). Statistical significance: *P < 0.05, **P < 0.01
If CsMYB60 interacts with CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein in vivo, they must be expressed at the same time and in the same place. To verify this, we analyzed the spatiotemporal expression patterns of these five genes. The qRT-PCR analysis demonstrated that they were all expressed at higher levels in black spines (from RNS9) than in white spines (from RNS8), at least in 4 days before anthesis (Fig. S8). We also found that there were higher expression levels of these five genes in the black spines of the transgenic siblings of OX-1 than in the white spines of the nontransgenic siblings of OX-1 at 4 days before anthesis, even though their expression levels showed large differences (Fig. 4g).
The data indicated that CsMYB60 was expressed earlier than the CsMYC1, CsbHLH42, CsTATA-box, and CsWD40 genes (Fig. S8), and these four genes were all upregulated in the transgenic lines overexpressing CsMYB60 (Fig. 4g). Therefore, we speculated that CsMYB60 might regulate their expression. Using Y1H assays, we found that CsMYB60 could directly bind to the promoters of CsbHLH42 and CsTATA-box binding protein but not the promoters of CsMYC1 and CsWD40 (Fig. S9).
CsMYB60 can form complexes with its interaction partners to activate the expression of Cs4CL and CsCHS
CsMYB60 activated the expression of the key genes Cs4CL and CsCHS but did not directly bind to their promoters, indicating that the mode of activation was indirect (Figs. 3b and S4d). To test whether CsMYB60 regulates the expression of Cs4CL and CsCHS through its interacting proteins, we performed Y1H assays. We found that CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein could directly bind to the promoter of Cs4CL, but only CsTATA-box binding protein could bind to the promoter of CsCHS (Fig. 5a).
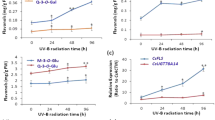
a Yeast-one-hybrid assays. b Schematic diagrams of the effector constructs and GUS reporter gene constructs driven by the Cs4CL promoter or CsCHS promoter. cGUS activity assays in tobacco leaves showing that CsMYB60 promoted the expression of Cs4CL. dGUS activity assays showing that CsTATA-box binding protein promoted the expression of CsCHS and that CsMYB60 increased CsTATA-box binding protein activation in tobacco leaves. e Relative GUS expression in the transient expression assays in (d). fGUS activity assays showing that CsTATA-box binding protein promoted the expression of CsCHS and that CsMYB60 increased CsTATA-box binding protein activation in cucumber cotyledons. gGUS relative expression in the transient expression assays in (f). Three biological replicates were performed for the gene expression analyses. Error bars represent SE. Statistical significance: *P < 0.05, **P < 0.01
Transient expression assays in tobacco leaves or cucumber cotyledons showed that CsMYB60 could activate the expression of Cs4CL and CsCHS through its interacting proteins, which could directly bind to the Cs4CL and CsCHS promotors (Fig. 5). Although CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein could promote the expression of GUS driven by the Cs4CL promoter to a certain extent, when CsMYB60 was cotransfected with these proteins, GUS expression was increased in tobacco leaves. When the five proteins were coinjected, GUS expression reached a maximum6 (Fig. 5c). CsMYB60 could effectively increase the expression of GUS driven by the CsCHS promoter in cucumber cotyledons but had no obvious effect in tobacco leaves, suggesting that CsMYB60 also regulates CsCHS indirectly (Fig. 5d–g). Then, we found that CsTATA-box binding protein could activate the expression of GUS in cucumber cotyledons and tobacco leaves (Fig. 5d–g), and that the GUS expression level was significantly increased upon cotransfection with 35S:CsMYB60 and 35S:CsTATA-box (Fig. 5d–g).
CsMYC1 physically interacts with CsbHLH42, CsWD40, and itself
MBW complexes regulate flavonoid biosynthesis in numerous plant species, such as Arabidopsis, strawberry, and petunia37,45,46. To compare CsbHLH42, CsMYC1, and CsWD40 with the bHLH and WD40 TFs involved in flavonoid synthesis from other species, two phylogenetic trees were constructed (Fig. S10). The results showed that the CsbHLH42 and CsMYC1 proteins shared high similarities with bHLH TFs that regulate the biosynthesis of flavonoids in other species (Fig. S10a), whereas the CsWD40 protein shared low similarity with WD40 TFs from other species (Fig. S10b).
The Y2H results showed that CsMYC1 directly interacted with CsbHLH42, CsWD40, and itself but not with CsTATA-box binding protein (Fig. 6a). The results were further confirmed by BiFC (Fig. 6b) and pull-down assays (Fig. 6c–e).
CsbHLH42 and CsMYC1 can induce the biosynthesis of anthocyanins and PAs in Arabidopsis
Although interacting relationships have been identified among these proteins, it is necessary to explore their biological functions. Arabidopsis was used for genetic transformation to clarify the biological functions of CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein (Figs. S11b and S12a). The visible pigmentation in the cotyledons and hypocotyls was significantly increased only in the CsbHLH42- and CsMYC1-overexpressing transgenic lines and not in the CsWD40- and CsTATA-box-overexpressing transgenic lines compared with the control (Fig. S11a). The intense pigmentation phenotype in the cotyledons and hypocotyls of the CsbHLH42- and CsMYC1-overexpressing lines is expected to result from the increases in anthocyanin and PA accumulation (Fig. S11c, d). However, neither anthocyanin nor PA contents were altered in the CsWD40- and CsTATA-box-overexpressing lines compared with the control (Fig. S12b, c).
Endogenous structural genes, including AtPAL, AtCHS, AtDFR, and AtANS, etc. were coordinately upregulated in the transgenic lines overexpressing CsbHLH42 or CsMYC1 (Fig. S11e). In addition, CsMYC1 and CsbHLH42 could interact with the MYB TFs AtPAP1 and AtTT2, which are involved in the regulation of flavonoid biosynthesis in Arabidopsis (Fig. S12d).
Discussion
CsMYB60 is the B gene that controls the biosynthesis of flavonols and PAs in cucumber
As an economic trait, the spine color of cucumber fruit was investigated in a classical genetics study one century ago47. Thereafter, it was confirmed that black spines were dominant over white spines and that this trait was controlled by a single gene, the B gene48. Although the B locus was found early last century47,48, its mapping and cloning were delayed for quite some time due to a lack of genomic information. In 2013, the B locus was finely mapped to a 50 kb genomic DNA region containing an R2R3-MYB TF-encoding gene, CsMYB6041,42. The best candidate gene for the B locus seemed to be CsMYB60, which controls the coloration of the orange mature fruit skins and black spines of cultivated cucumber41,42. Recent studies supported this hypothesis6,49. Moreover, the present study further confirmed this hypothesis using transgenic lines overexpressing CsMYB60 (Fig. 2). In addition, it has been clarified that the main pigments of black spines are flavonols and PAs6. In addition to black spines and orange mature fruit skins, flavonols and PAs accumulated in other tissues at different levels (Figs. 1 and S1). Moreover, the distribution patterns of flavonols and PAs were similar to those in the black-spined inbred line (Figs. 1, S1, S4, and S5). Therefore, CsMYB60 is the B gene that regulates the biosynthesis of flavonols and PAs not only in black spines and orange mature fruit skins but also in other tissues in cucumber.
Flavonols and PAs that exist in the seeds, bark, leaves, and fruit of various plants; not only do these compounds play important roles in many biological processes such as plant growth and development and resistance to insects and diseases17,50, but they also greatly benefit human health8,9. Flavonols and PAs showed high accumulation in the black spines and orange mature fruit skins of the black-spined inbred lines but presented low accumulation in other tissues6,41 (Figs. 1 and S1). This spatiotemporal pattern of flavonols and PAs in cucumber does not benefit human health because when cucumbers are consumed fresh, the spines are washed off, and the fruit skins contain very low levels of flavonols and PAs at the commercial stage6 (Fig. 1). The present study showed that the contents of flavonols and PAs could be increased in the fruit pericarp and pulp (Fig. S4a–c). Therefore, this study provides a theoretical basis, target genes, and technical support for the design of molecular breeding strategies to improve the fruit quality of cucumber.
The mechanism of flavonol and PA biosynthesis regulated by CsMYB60 in cucumber
The regulation of flavonoid biosynthesis has been studied in depth, especially in the model plant A. thaliana30,51. Key structural genes and numerous transcription factors are required for flavonol and PA biosynthesis. Since the first plant MYB regulating anthocyanin biosynthesis was identified in maize (Zea mays)52, a growing body of evidence has demonstrated that MYB TFs are major regulators of flavonoid biosynthesis in diverse species53,54. In Arabidopsis, three closely related R2R3-MYB proteins, MYB11, MYB12, and MYB111, transcriptionally regulate flavonol biosynthesis by directly activating the early biosynthetic genes (EBGs) CHS, CHI, F3H, and FLS, whereas the MBW complex activates the late biosynthetic genes (LBGs) DFR, LODX, and ANR, leading to the production of PAs and anthocyanins30. The seed-specific accumulation of PAs requires the activity of the MBW complex composed of TT2, TT8, and TTG1, and anthocyanin biosynthesis is regulated by the MBW complex consisting of one R2R3-MYB TF (PRODUCTION OF ANTHOCYANIN PIGMENTS (PAP1), PAP2, MYB113, or MYB114), one bHLH protein (TT8, GLABROUS3 (GL3), or ENHANCER OF GLABRA3 (EGL3)) and TTG130.
Although CsMYB60 is known to be a key regulator of flavonoid biosynthesis6, it is still unclear how CsMYB60 transcriptionally regulates the structural genes of the biosynthetic pathway of flavonoids in cucumber. In this study, we demonstrated the mechanism whereby flavonol and PA biosynthesis is regulated by CsMYB60 (Fig. 7). The expression levels of some structural genes involved in flavonoid biosynthesis were increased in the 35S:CsMYB60 transgenic lines. For example, the expression levels of CsCHS, CsFLS, CsLAR, and Cs4CL in the transgenic lines were significantly higher than those in their nontransgenic siblings (Fig. S4d). CHS catalyzes the first committed step in flavonoid biosynthesis, and FLS and LAR are essential enzymes for the biosynthesis of flavonols and PAs, respectively, in plants30,31,32. However, these three key genes in flavonoid biosynthesis did not change white spines into black spines in a transient expression system, but Cs4CL did, which suggests that Cs4CL is the limiting gene for flavonoid biosynthesis in cucumber6. However, CsCHS, CsFLS, and CsLAR must work in coordination with Cs4CL (Fig. S4d); moreover, this coordination could be executed by CsMYB60 (Fig. 7). As a homolog of AtMYB11, 12, and 111 (Fig. S2), which directly regulate EBGs30, CsMYB60 could directly activate the expression of CsFLS and CsLAR (Figs. 3 and S6). CsMYB60 did not directly promote the expression of CsCHS, a key gene in flavonoid biosynthesis (Fig. 5e), but did increase the direct activation activity of CsTATA-box binding protein toward CsCHS through the protein interaction between CsMYB60 and CsTATA-box binding protein (Figs. 4a, b, f and 5). In addition to the EBGs and LBGs involved in flavonoid biosynthesis, CsMYB60 could indirectly regulate Cs4CL6, the gene responsible for the last step of the general phenylpropanoid metabolism pathway, through its interacting proteins CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein, which can directly bind to the Cs4CL promotor to activate its expression (Figs. 4 and 5). Beyond the MBW complex, we identified a new function of the CsTATA-box binding protein in the regulation of flavonoid biosynthesis, which has never been reported previously to our knowledge. Therefore, there might be a unique regulatory mechanism for flavonoid biosynthesis in cucumber involving CsMYB60 as a key regulator. Although MYBs are common key factors in the regulation of flavonoid accumulation in plants, most species present distinct regulatory mechanisms53,54. However, further studies are needed to clarify what kind of complexes are formed during the activation of Cs4CL and what other factors take part in the fine-tuning of flavonoid biosynthesis in cucumber, which will reveal TFs that may be used as tools for metabolic engineering for flavonoid production in cucumber fruits.
CsMYB60 could activate the expression of CsFLS and CsLAR by directly binding to their promoters. Additionally, CsMYB60 directly or indirectly activated the expression of CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein, whose products then interacted with CsMYB60 to form the complex regulating the expression of Cs4CL, and CsMYB60 interacted with CsTATA-box binding protein to promote the expression of CsCHS. Blue, orange, red, green, and purple ellipses represent CsMYB60, CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein, respectively
Materials and methods
Plant materials and growth conditions
White-spined (RNS8) and black-spined (RNS9) cucumber inbred lines were used for flavonol and PA analyses6. The cucumber inbred line Xintaimici (wild type, WT) and the Arabidopsis Columbia-0 ecotype were used for genetic transformation. Two generations of cucumbers per year were grown in a solar greenhouse at the experimental station of Shandong Agricultural University, Tai’an, China.
Construction of the CsMYB60 overexpression vector and genetic transformation
The complete CsMYB60 coding sequence was inserted into the BamHI and SacI sites of the pCAMBIA1300 vector to form the 35S:CsMYB60 construct. Subsequently, the 35S:CsMYB60 vector was transformed into Xintaimici55. Transgenic plants were screened on hygromycin-containing medium, and the presence of the transgene was confirmed by PCR with genomic DNA as the template.
Spatial and temporal expression analysis of CsMYB60 by qRT-PCR
Total RNA was isolated from cucumber and Arabidopsis plants using an RNAprep pure Plant Kit (TianGen, Beijing, China) following the manufacturer’s instructions. Subsequently, the RNA was reverse transcribed using the PrimeScript® 1st Strand cDNA Synthesis Kit (Takara, Japan). qRT-PCR was performed using the UltraSYBR Mixture (with ROX I; Cwbiotech) in the iCycler iQ5 system (BioRad, CA, USA). The results were normalized to those of the cucumber ACTIN gene. Three biological replicates were performed for each analysis. The primers used in this study are provided in Table S1.
Yeast two-hybrid (Y2H) screening and confirmation
Yeast (Saccharomyces cerevisiae) two-hybrid (Y2H) assays were performed as described in the manufacturer’s instructions (Clontech, Mountain View, CA, USA). The CsMYB60 CDS was inserted into the bait vector pGBKT7. Subsequently, the CDSs of CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein, etc. were cloned into the prey vector pGADT7. All of the constructs were transformed into the yeast strain AH109 Gold using the lithium acetate method. Subsequently, the cells were cultured on synthetic defined (SD) medium lacking Trp and Leu (SD/-Trp-Leu) at 28 °C. Putative transformants were transferred to medium lacking Leu, Trp, His, and adenine (SD/-Leu-Trp-His-Ade) with 3-AT and X-a-gal.
Electromobility shift assays
EMSAs were performed as previously described56. CsMYB60 was cloned into the vector pGEX4T-1. The production of the recombinant CsMYB60-GST protein was induced in BL21 cells. Subsequently, the recombinant protein was purified using glutathione Sepharose beads (Thermo Scientific, Waltham, MA, USA). Three oligonucleotide probes for the CsFLS and CsLAR promoters were labeled using an EMSA probe biotin labeling kit (Beyotime, Shanghai, China). The EMSAs were carried out using an EMSA kit (Thermo Scientific) following the manufacturer’s instructions. Excess unlabeled oligonucleotides were also added to the reactions. The primers used in this study are provided in Table S1.
Pull-down assays
The CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein open reading frames were introduced into the PET-32a vector containing a histidine (HIS) tag sequence. The ORF of CsMYB60 was inserted into the PGEX-4T-1 plasmid containing a glutathione S-transferase (GST) tag sequence. Then, the recombinant plasmids were transformed into Escherichia coli BL21 (DE3) (TransGen Biotech, China) to induce the formation of fusion proteins. The pull-down assays were performed following the instructions of the Pierce GST Spin Purification Kit (Thermo Scientific). Then, bound proteins were detected by immunoblotting with anti-HIS and anti-GST antibodies.
BiFC assays
The full-length CsbHLH42, CsMYC1, CsWD40, CsTATA-box binding protein, and CsMYB60 genes were cloned into the plasmids pSPYCE-35S and pSPYNE-35S. Agrobacterium tumefaciens LBA4404 cells were transformed with the recombinant plasmids, and different combinations of Agrobacterium strains were cotransformed into tobacco leaves. After 48 h of coinfiltration, YFP fluorescence was detected using a confocal microscope with excitation at 488 nm.
PA staining and determination
The DMACA reagent [1% (w/v) in ethanol:6 M HCl (1:1, v/v)] was used to stain different tissues of cucumber for 1 h. Tissue samples of equal weight were ground with liquid nitrogen and suspended in a 70% (v/v) acetone aqueous solution containing 0.1% (w/v) ascorbic acid; this procedure was repeated three times. The corresponding PA content of each tissue was extracted by the method described previously57.
Yeast one-hybrid (Y1H) assays
The promoter fragments (2000 bp) of Cs4CL, CsFLS, and CsLAR were inserted into the pHIS2 plasmid. The CsbHLH42, CsMYC1, CsWD40, CsTATA-box binding protein, and CsMYB60 CDSs were separately inserted into the pGADT7 vector (Clontech) to obtain the constructs AD-CsMYB60, AD-CsbHLH42, AD-CsMYC1, AD-CsWD40, and AD-CsTATA-box, respectively. Yeast strain Y187 cells harboring different combinations of recombinant plasmids were examined on SD/-Leu-Trp-His medium with an optimal concentration of 3-AT.
Transient GUS activity assays
Cucumber cotyledons or tobacco leaves were used to conduct transient GUS activity assays. The promoters of CsFLS, CsLAR, CsCHS, and Cs4CL were inserted into pCAMBIA1300-GUS to activate the GUS reporter gene. The ORFs of CsMYB60, CsbHLH42, CsMYC1, CsWD40, and CsTATA-box binding protein were separately cloned into the pCAMBIA1300 plasmid to obtain the 35S:CsMYB60, 35S:CsbHLH42, 35S:CsMYC1, 35S:CsWD40, and 35S:CsTATA-box recombinant plasmids. The different combinations were injected into cucumber cotyledons (8-day-old) or tobacco leaves (5-week-old) for Agrobacterium-mediated transformation. The injected tobacco or cucumber seedlings were grown for approximately 2 days under normal conditions. GUS activity was measured as previously described6,58.
HPLC analysis of flavonols extracted from cucumber under optimum conditions
For the analysis of flavonols in various tissues of cucumber, 0.3 g tissue was ground in liquid nitrogen and extracted with MeOH:H2O (50:50 v/v) at 4 °C for 2 h with constant shaking, followed by sonication for 30 min at room temperature. The mixture was subsequently centrifuged at 12,000 rpm for 10 min. After centrifugation, the supernatant was collected, filtered through a 0.45 µm filter and stored at 4 °C until chromatographic analysis. The flavonol extracts obtained from cucumber were analyzed using an Agilent 1200 liquid chromatographic (LC) system equipped with an autosampler, quaternary pump, and diode-array detector (DAD) (Agilent Technologies, Waldrom, Germany). The sample (20 µL) was injected into the HPLC-ESI-QToF and separated in a Superspher RP C18 column (4 × 250 mm2, 4 µm; Agilent Technologies). Elution was performed via gradient elution with solvent systems A (TFA:FA:H2O/0.1:2:97.9) and B (TFA:FA:CAN:H2O/0.1:2:35:62.9) using the following conditions: 0–12 min, 30–48% B; 12–22 min, 48–55% B; 22–35 min, 55–60% B; 35–45 min, 60–30% B, with a flow rate of 0.8 mL/min. The ultraviolet–visible light detector wavelength was monitored at 365 nm.
Determination of the total anthocyanin content
The total anthocyanin contents of the samples were extracted with the HCl–methanol method. A 0.5 g sample was ground in liquid nitrogen and then incubated with 5 mL of cold 1% (v/v) methanol–HCl in the dark for 24 h. The absorbance of the solution was measured at 650, 620, and 530 nm with a UV-1600 spectrophotometer. The anthocyanin content was calculated using the following formula: OD = (A530 − A620) − 0.1(OD650 − OD620).
Data availability
The data that support the results are included within the article and its Supplementary Information. Other relevant materials are available from the corresponding author upon reasonable request.
References
Pourcel, L., Routaboul, J. M., Cheynier, V., Lepiniec, L. & Debeaujon, I. Flavonoid oxidation in plants: from biochemical properties to physiological functions. Trends Plant Sci.12, 29–36 (2007).
Buer, C. S., Imin, N. & Djordjevic, M. A. Flavonoids: new roles for old molecules. J. Integr. Plant Biol.52, 98–111 (2010).
Barbehenn, R. V. & Constabel, C. P. Tannins in plant–herbivore interactions. Phytochemistry72, 1551–1565 (2011).
Emiliani, J., Grotewold, E., Falcone Ferreyra, M. L. & Casati, P. Flavonols protect Arabidopsis plants against UV-B deleterious effects. Mol. Plant6, 1376–1379 (2013).
Henry-Kirk, R. A. et al. Solar UV light regulates flavonoid metabolism in apple (Malus x domestica). Plant Cell Environ.41, 675–688 (2018).
Liu, M. et al. CsMYB60 is a key regulator of flavonols and proanthocyanidans that determine the colour of fruit spines in cucumber. J. Exp. Bot.70, 69–84 (2019).
Bagchi, D. et al. Free radicals and grape seed proanthocyanidin extract: importance in human health and disease prevention. Toxicology148, 187–197 (2000).
Qin, C. X., Chen, X., Hughes, R. A., Williams, S. J. & Woodman, O. L. Understanding the cardioprotective effects of flavonols: discovery of relaxant flavonols without antioxidant activity. J. Med. Chem.51, 1874–1884 (2008).
Bondonno, N. P. et al. Flavonoid intake is associated with lower mortality in the Danish Diet Cancer and Health Cohort. Nat. Commun.10, 3651 (2019).
Watkins, J. M., Chapman, J. M. & Muday, G. K. Abscisic acid-induced reactive oxygen species are modulated by flavonols to control stomata aperture. Plant Physiol.175, 1807–1825 (2017).
Ryan, K. G., Swinny, E. E., Markham, K. R. & Winefield, C. Flavonoid gene expression and UV photoprotection in transgenic and mutant Petunia leaves. Phytochemistry59, 23–32 (2002).
Czemmel, S. et al. Transcriptome-wide identification of novel UV-B- and light modulated flavonol pathway genes controlled by VviMYBF1. Front. Plant Sci.8, 1084 (2017).
Leo, C. H. & Woodman, O. L. Flavonols in the prevention of diabetes-induced vascular dysfunction. J. Cardiovasc. Pharmacol.65, 532–544 (2015).
Mellway, R. D., Tran, L. T., Prouse, M. B., Campbell, M. M. & Constabel, C. P. The wound-, pathogen-, and ultraviolet B-responsive MYB134 gene encodes an R2R3 MYB transcription factor that regulates proanthocyanidin synthesis in Poplar. Plant Physiol.150, 924–941 (2009).
Wang, L. et al. The transcription factor MYB115 contributes to the regulation of proanthocyanidin biosynthesis and enhances fungal resistance in poplar. N. Phytol.215, 351–367 (2017).
Koleckar, V. et al. Condensed and hydrolysable tannins as antioxidants influencing the health. Mini-Rev. Med. Chem.8, 436–447 (2008).
Dixon, R. A., Xie, D. Y. & Sharma, S. B. Proanthocyanidins-a final frontier in flavonoid research? N. Phytol.165, 9–28 (2005).
Bogs, J., Jaffe, F. W., Takos, A. M., Walker, A. R. & Robinson, S. P. The grapevine transcription factor VvMYBPA1 regulates proanthocyanidin synthesis during fruit development. Plant Physiol.143, 1347–1361 (2007).
Abeynayake, S. W., Panter, S., Mouradov, A. & Spangenberg, G. A high-resolution method for the localization of proanthocyanidins in plant tissues. Plant Methods7, 13 (2011).
Pang, Y. et al. A WD40 repeat protein from Medicago truncatula is necessary for tissue-specific anthocyanin and proanthocyanidin biosynthesis but not for trichome development. Plant Physiol.151, 1114–1129 (2009).
Debeaujon, I. et al. Proanthocyanidin-accumulating cells in Arabidopsis testa: regulation of differentiation and role in seed development. Plant Cell15, 2514–2531 (2003).
Xu, W. et al. Complexity and robustness of the flavonoid transcriptional regulatory network revealed by comprehensive analyses of MYB-bHLH-WDR complexes and their targets in Arabidopsis seed. N. Phytol.202, 132–144 (2014).
Souquet, J. M., Labarbe, B., Le, Guernevé, C., Cheynier, V. & Moutounet, M. Phenolic composition of grape stems. J. Agric. Food Chem.48, 1076–1080 (2000).
Downey, M. O. & Rochfort, S. Simultaneous separation by reversed-phase high-performance liquid chromatography and mass spectral identification of anthocyanins and flavonols in Shiraz grape skin. J. Chromatogr. A1201, 43–47 (2008).
Shirley, B. W. et al. Analysis of Arabidopsis mutants deficient in flavonoid biosynthesis. Plant J.8, 659–671 (1995).
Lepiniec, L. et al. Genetics and biochemistry of seed flavonoids. Annu. Rev. Plant Biol.57, 405–430 (2006).
Hichri, I. et al. Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J. Exp. Bot.62, 2465–2483 (2011).
James, A. M. et al. Poplar MYB115 and MYB134 transcription factors regulate proanthocyanidin synthesis and structure. Plant Physiol.174, 154–171 (2017).
Sun, H. et al. Positive selection drives adaptive diversification of the 4-coumarate: CoA ligase (4CL) gene in angiosperms. Ecol. Evol.5, 3413–3420 (2015).
Li, S. Transcriptional control of flavonoid biosynthesis. Plant Signal. Behav.9, e27522 (2014).
Czemmel, S. et al. The grapevine R2R3-MYB transcription factor VvMYBF1 regulates flavonol synthesis in developing grape berries. Plant Physiol.151, 1513–1530 (2009).
Bogs, J. et al. Proanthocyanidin synthesis and expression of genes encoding leucoanthocyanidin reductase and anthocyanidin reductase in developing grape berries and grapevine leaves. Plant Physiol.139, 652–663 (2005).
Yuan, L. et al. Molecular cloning and characterization of PtrLAR3, a gene encoding leucoanthocyanidin reductase from Populus trichocarpa, and its constitutive expression enhances fungal resistance in transgenic plants. J. Exp. Bot.63, 2513–2524 (2012).
Liao, L. et al. Molecular characterization of genes encoding leucoanthocyanidin reductase involved in proanthocyanidin biosynthesis in apple. Front. Plant Sci.6, 243 (2015).
Nesi, N., Jond, C., Debeaujon, I., Caboche, M. & Lepiniec, L. The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell13, 2099–2114 (2001).
Mano, H., Ogasawara, F., Sato, K., Higo, H. & Minobe, Y. Isolation of a regulatory gene of anthocyanin biosynthesis in tuberous roots of purple-fleshed sweet potato. Plant Physiol.143, 1252–1268 (2007).
Schaart, J. G. et al. Identification and characterization of MYB-bHLH-WD40 regulatory complexes controlling proanthocyanidin biosynthesis in strawberry (Fragaria x ananassa) fruits. N. Phytol.197, 454–467 (2013).
Zhang, B. & Schrader, A. TRANSPARENT TESTA GLABRA 1-dependent regulation of flavonoid biosynthesis. Plants6, 65 (2017).
Stracke, R. et al. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J.50, 660–677 (2007).
Verdier, J. et al. MtPAR MYB transcription factor acts as an on switch for proanthocyanidin biosynthesis in Medicago truncatula. Proc. Natl. Acad. Sci. U.S.A.109, 1766–1771 (2012).
Li, Y., Wen, C. & Weng, Y. Fine mapping of the pleiotropic locus B for black spine and orange mature fruit color in cucumber identifies a 50 kb region containing a R2R3-MYB transcription factor. Theor. Appl. Genet.126, 2187–2196 (2013).
Zhao, P., Li, Q., Li, J., Wang, L. & Ren, Z. Genome-wide identification and characterization of R2R3MYB family in Solanum lycopersicum. Mol. Genet. Genomics289, 1183–1207 (2014).
Ishida, T. et al. Arabidopsis TRANSPARENT TESTA GLABRA2 is directly regulated by R2R3 MYB transcription factors and is involved in regulation of GLABRA2 transcription in epidermal differentiation. Plant Cell19, 2531–2543 (2007).
Wang, N. et al. The proanthocyanidin-specific transcription factor MdMYBPA1 initiates anthocyanin synthesis under low-temperature conditions in red-fleshed apples. Plant J.96, 39–55 (2018).
Baudry, A. et al. TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsisthaliana. Plant J.39, 366–380 (2004).
Quattrocchio, F. et al. PH4 of Petunia is an R2R3 MYB protein that activates vacuolar acidification through interactions with basic-helix-loop-helix transcription factors of the anthocyanin pathway. Plant Cell18, 1274–1291 (2006).
Wellington, R. Mendelian inheritance of epidermal characters in the fruit of Cucumis sativus. Science38, 61 (1913).
Strong, W. J. Breeding experiments with the cucumber (Cucumis sativus L.). Sci. Agric.11, 333–346 (1931).
Zhang, C., Win, K. T., Kim, Y. C. & Lee, S. Two types of mutations in the HEUKCHEEM gene functioning in cucumber spine color development can be used as signatures for cucumber domestication. Planta250, 1491–1504 (2019).
Pollastri, S. & Tattini, M. Flavonols: old compounds for old roles. Ann. Bot.108, 1225–1233 (2011).
Xu, W., Dubos, C. & Lepiniec, L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci.20, 176–185 (2015).
Paz-Ares, J., Ghosal, D., Wienand, U., Peterson, P. A. & Saedler, H. The regulatory c1 locus of Zea mays encodes a protein with homology to myb proto-oncogene products and with structural similarities to transcriptional activators. EMBO J.6, 3553–3558 (1987).
Naing, A. H. & Kim, C. K. Roles of R2R3-MYB transcription factors in transcriptional regulation of anthocyanin biosynthesis in horticultural plants. Plant Mol. Biol.98, 1–18 (2018).
Ma, D. & Constabel, C. P. MYB repressors as regulators of phenylpropanoid metabolism in plants. Trends Plant Sci.24, 275–289 (2019).
Wang, H. et al. Antisense suppression of cucumber (Cucumis sativus L.) sucrose synthase 3 (CsSUS3) reduces hypoxic stress tolerance. Plant Cell Environ.37, 795–810 (2014).
An, J. P. et al. Apple bZIP transcription factor MdbZIP44 regulates ABA-promoted anthocyanin accumulation. Plant Cell Environ.41, 2678–2692 (2018).
Wang, N. et al. MYB12 and MYB22 play essential roles in proanthocyanidin and flavonol synthesis in red-fleshed apple (Malus sieversii f. niedzwetzkyana). Plant J.90, 276–292 (2017).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: betaglucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J.6, 3901–3907 (1987).
Acknowledgements
This work was supported by fundings from the National Natural Science Foundation of China (31672170 and 31872950), the Natural Science Foundation of Shandong Province (JQ201309), the Shandong “Double Tops” Program (SYL2017YSTD06), and the “Taishan Scholar” Foundation of the People’s Government of Shandong Province (ts20130932). The authors thank Professor Yujin Hao (College of Horticultural Science and Engineering, Shandong Agricultural University) for the gift of the pSPYNE and pSPYCE vectors.
Author information
Authors and Affiliations
Contributions
Z.R. conceived the project. J.L. and Z.R. designed the experiments. J.L., Q.L., J.H., C.Z., and M.L. performed the experiments. J.L. and Z.R. wrote and revised the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, J., Luan, Q., Han, J. et al. CsMYB60 directly and indirectly activates structural genes to promote the biosynthesis of flavonols and proanthocyanidins in cucumber. Hortic Res 7, 103 (2020). https://doi.org/10.1038/s41438-020-0327-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-0327-z
This article is cited by
-
Identification of LsPIN1 gene and its potential functions in rhizome turning of Leymus secalinus
BMC Genomics (2022)
-
Comprehensive analysis of cucumber C-repeat/dehydration-responsive element binding factor family genes and their potential roles in cold tolerance of cucumber
BMC Plant Biology (2022)
-
A light-responsive transcription factor SmMYB35 enhances anthocyanin biosynthesis in eggplant (Solanum melongena L.)
Planta (2022)
-
Molecular research progress and improvement approach of fruit quality traits in cucumber
Theoretical and Applied Genetics (2021)