Abstract
Chilling has a critical role in the growth and development of perennial plants. The chilling requirement (CR) for dormancy breaking largely depends on the species. However, global warming is expected to negatively affect chilling accumulation and dormancy release in a wide range of perennial plants. Here, we used Chimonanthus praecox as a model to investigate the CR for dormancy breaking under natural and artificial conditions. We determined the minimum CR (570 chill units, CU) needed for chilling-induced dormancy breaking and analyzed the transcriptomes and proteomes of flowering and non-flowering flower buds (FBs, anther and ovary differentiation completed) with different CRs. The concentrations of ABA and GA3 in the FBs were also determined using HPLC. The results indicate that chilling induced an upregulation of ABA levels and significant downregulation of SHORT VEGETATIVE PHASE (SVP) and FLOWERING LOCUS T (FT) homologs at the transcript level in FBs when the accumulated CR reached 570 CU (IB570) compared to FBs in November (FB.Nov, CK) and nF16 (non-flowering FBs after treatment at 16 °C for −300 CU), which suggested that dormancy breaking of FBs could be regulated by the ABA-mediated SVP-FT module. Overexpression in Arabidopsis was used to confirm the function of candidate genes, and early flowering was induced in 35S::CpFT1 transgenic lines. Our data provide insight into the minimum CR (570 CU) needed for chilling-induced dormancy breaking and its underlying regulatory mechanism in C. praecox, which provides a new tool for the artificial regulation of flowering time and a rich gene resource for controlling chilling-induced blooming.
Similar content being viewed by others
Introduction
Bud dormancy is a protective strategy for perennials to suspend growth for survival from adverse environmental conditions, which leads to the temporal insensitivity of plants to growth-promoting signals and the cessation of meristem activities1. Only when dormancy is released can plants resume growth under favorable environmental conditions1.
Photoperiod and temperature are the two central environmental signals that have a crucial role in the seasonal cycling of growth and dormancy2,3,4. The photoperiod is known to govern the cessation of perennial growth5. In autumn, short days (SDs) induce growth cessation of the shoot apical meristem (SAM), followed by the formation of buds and induction of dormancy before winter6,7,8. In contrast to photoperiod-controlled entry into dormancy, temperature primarily controls the duration of dormancy, and prolonged cold exposure induces dormancy breaking and bud break9,10,11. Interestingly, there is another situation in which low temperature (LT) can also cause growth cessation and control dormancy induction, regardless of photoperiod conditions12,13.
Phytohormones are known to have a crucial role in regulating the bud dormancy cycle of perennials14, and the complex process is mainly subject to the antagonistic regulation of gibberellins (GAs) and abscisic acid (ABA)4,15. GAs act as downstream targets of SDs in dormancy regulation16 and work in promoting dormancy release and bud break at a high level17,18. In contrast to GAs, ABA exerts antagonistic effects on dormancy regulation. SDs induces the accumulation of ABA and the expression of ABA signaling components to trigger dormancy establishment. High levels of ABA have been shown to maintain dormancy19. In contrast, ABA contents decrease towards dormancy release6,20,21,22,23, accompanied by a gradual increase in the GA level24.
Although the molecular mechanism underlying dormancy regulation is complicated and remains to be further understood, several critical regulators involved in this process have been described so far. In particular, the importance of dormancy regulated by ABA and the components downstream or upstream of ABA/GA have been identified.
The dormancy-associated MADS-box (DAM), a gene orthologous to the floral repressor SVP of Arabidopsis25, has been identified in the nondormant evergrowing mutant of peach26. This gene belongs to the STMADS11 subfamily of MADS-box genes, and its counterpart SVP transcriptionally inhibits the expression of FT in Arabidopsis27. DAMs showed distinct expression patterns in response to different environmental signals. Both SD and short-term LT treatments induce the upregulation of DAMs28,29, while a prolonged chilling requirement (CR) causes the reduction of DAM expression30. DAM homologs have also been characterized in other perennial species and have an essential role in the control of bud dormancy31,32,33,34,35. Overexpression of the DAM gene conferred early growth cessation and bud set in Japanese apricot36 and delayed bud break in apple37,38. PpDAM1 of pear upregulated the expression of the ABA biosynthesis enzyme 9-cis-epoxy-carotenoid dioxygenase (NCED) by binding to the CArG motif in the promoter of PpNCED3, which has an essential role in the regulation of bud dormancy39. Recent studies have provided insights into the roles of the poplar SVP-LIKE (SVL) gene in dormancy regulation. This gene negatively regulates FT1 to affect bud break19. It also regulates NCED3 expression directly, resulting in a high accumulation of ABA towards bud dormancy. Low temperature reduces ABA accumulation and represses the expression of SVL to promote bud break19. The overexpression of SVL was reported to suppress the dormancy defects of the abi1-1 mutant in hybrid aspen19. Moreover, ABA mediates SD-induced SVL expression during bud dormancy16. In response to SDs, SVL induces the expression of CALLOSE SYNTHASE 1 (CALS1), which mediates plasmodesmata (PD) closure to promote dormancy16. ABA can also induce the expression of CALS1 by suppressing the expression of PICKLE (PKL), where dormancy defects can be restored in abi1-1 plants16,20,40. ABA accumulation also contributes to PD closure to prevent dormancy release by restricting the transport of FT20. In contrast, the GA pathway is a downstream target of SVL in temperature-controlled bud break19. In addition, GA-mediated callose hydrolyzation helps the passage of FT to promote dormancy release17. The variation in FT gene expression levels during dormancy and after dormancy release is consistent with the observation that decreased expression of FT is induced by SD19,34,41, while LT upregulates the transcription of FT in breaking dormancy17,42. By comparison, PaFT has been shown to display increased expression levels in response to LT/SD in London plane trees43. Moreover, PaFTL, another FT orthologous gene in London plane trees, showed low and variable expression levels during dormancy and under LT/SD conditions44.
The FT gene functions as an upstream regulator of the AP1/FUL genes. In the SAM, the transcription of floral meristem identity genes AP1/FUL was stimulated by the FT/FD complex to promote flowering45,46. Interestingly, three AP1/FUL-like genes were observed to work upstream of FT homologs in rice47. The expression levels of AP1 were observed to increase before dormancy release in Ziziphus jujuba48. Growth cessation was induced by SD-induced downregulation of Like-AP1 (LAP1), an ortholog of the floral meristem identity gene APETALA1 (AP1) in hybrid aspen49. However, similar to LAP1 decreased expression of PlacFL2 (FUL-like gene, an ancestor of AP1) was observed to control growth cessation mediated by SD downstream of CO/FT50.
C. praecox (wintersweet) is a crucial deciduous shrub that originated in China and is well known for its fragrant flowers and high ornamental value51. Adequate bud dormancy, followed by adequate bud breaking, is critical for its flowering and ornamental value. Warm winters, presumably due to global warming, have severe adverse effects on bud dormancy (and bud breaking, flowering, and plant performance) of wintersweet. Similar adverse effects have been observed in many other important plants, particularly horticultural crops such as apple (Malus domestica)30,35,37,52,53,54,55, blueberry, peach (Prunus persica)56, Japanese pear (Pyrus pyrifolia)57,58,59, and Prunus spp.60,61,62 In this study, we used wintersweet as a model to gain a comprehensive understanding of the fundamental molecular mechanism of floral bud dormancy in plants.
Materials and methods
Plant materials and chilling treatment
Eighteen-year-old potted C. praecox ‘Concolor’ plants and 15-year-old ‘Concolor’ and ‘Grandiflorus’ plants were grown on the campus of Southwest University (106°43E, 29°83 N, Beibei District, Chongqing City, China). Plants were exposed to forcing conditions at 12, 14, or 16 °C with different chilling requirement (CR) under SD conditions (8/16 h of light/dark) and 8000 lux illumination intensity in climate chambers (RDN-1000D-4, China) for a predetermined period from 8 Nov 2018 (average air temperature below 14 °C for three consecutive days). Richardson chill units (CU) for C. praecox ‘Concolor’ flower buds (FBs) were calculated according to the “UTAH Model”63,64. When the CU reached −300, 0, 150, 300, 450, and 570 (i.e., 16 °C for 600 h; 14 °C for 600 h; 12 °C for 300, 600, 900, and 1140 h, respectively), the treated plants were transferred to 14 °C for phenotype analysis. The following FBs were harvested and stored at −80 °C for sequencing and/or quantitative real-time PCR (qRT-PCR) analysis: FBs collected in April, May, and November (FB.Apr, FB.May, FB.Nov); FBs exposed to chilling conditions at 12 °C for 150, 300, 450 CU (FB150, FB300, FB450) and 570 CU (IB570, FBs initiate blooming); and nF16 (non-flowering FBs collected after treatment at 16 °C for −300 CU). Wintersweet plants cultured in open fields were used for the analysis of the influence of CR on chilling-induced dormancy breaking. The temperature data from Beibei meteorological station 57511# (106°27E, 29°51N, Beibei District, Chongqing City, China) were provided by the Chongqing Meteorological Bureau.
Construction of libraries and RNA-seq
A total of 24 independent RNA-Seq libraries from the FBs of eight groups, with three biological replicates for each group, were constructed and sequenced: FB.Apr, FB.May, FB.Nov (Fig. 1b, f), nF16 (Fig. 1c, g), FB150, FB300, FB450 (phenotypes just like nF16, photos not shown here), and IB570 (Fig. 1d, e). Briefly, a TRIzol reagent kit (Invitrogen, Carlsbad, CA, USA) was used to extract total RNA from each sample. Poly (A) mRNA was purified and fragmented into short lengths. The obtained fragments were used as templates, and first-strand cDNA synthesis was conducted using random hexamer primers, followed directly by second-strand cDNA synthesis. Next, the synthesized cDNA fragments were purified using a QIAquick PCR extraction kit (Qiagen, German); then, they were end-repaired, and poly(A) tails were added followed by ligation to Illumina sequencing adapters. These generated fragments were purified by agarose gel electrophoresis and then enriched by PCR amplification. Finally, the constructed cDNA libraries were sequenced on an Illumina HiSeq™ 4000 platform at Gene Denovo Biotechnology Co., Guangzhou, China. We deposited our sequencing data set in the NCBI Sequence Read Archive (SRA) under accession number PRJNA613935.
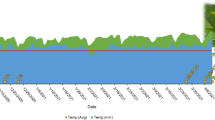
a Investigation under natural conditions in 2018–2019. The CR value was 5.5 on 5 Nov 2018. Blooming ratios of 12.8% and 13.6% occurred when CR accumulation reached 570 CU on 24 Dec 2018 in FBs from C. praecox ‘Grandiflorus’ and ‘Concolor’, respectively. b–h Artificial chilling-induced dormancy breaking and samples for omics analysis. b Potted C. praecox of eighteen-year-old in November as a control. c Plants subjected to −300 CU chilling requirement (CR) at 16 °C; FBs did not fully open and gradually dropped, showing the same phenotype as FBs subjected to 150, 300, and 450 CU at 12 °C (FB150, FB300, and FB450, photos not shown here). d Flowering plants subjected to 570 CU at 12 °C. e FBs initiated blooming after exposure to 570 CR (IB570). f FBs in Nov (FB.Nov). g Non-flowering FBs at 16 °C subjected to −300 CU (nF16). h Open flower stage after exposure to 570 CU (OF570). FB.Apr, FB.May, FB.Nov, FB150, FB300, FB450, nF16, and IB570 were used for RNA- and/or TMT-sequencing
Gene annotation, WGCNA, Venn diagrams, trends, and network analysis
Raw sequence reads were trimmed and checked for quality. The high-quality clean reads were subjected to de novo transcriptome assembly using Trinity65. Basic annotation of unigenes was performed based on the NCBI Nr, KOG, SWISS-PROT, and KEGG databases with E-values below 1e-5 as a threshold. Blast2GO software was employed to analyze the GO annotations of each unigene66. Subsequently, the functional classification of unigenes was conducted using WEGO software67,68. The number of reads per kilobase per million mapped reads (RPKM) was calculated and normalized to present the abundances of each unigene66. We used the edgeR package (version 3.12.1) to determine significant differentially expressed genes (DEGs) between two compared groups with a fold change threshold of 2 and a false discovery rate (FDR) threshold of 0.05. The calculated DEGs were then statistically enriched according to GO functions and KEGG pathways with an FDR value <0.05.
In addition, coexpressed modules and hub genes were detected by the Weighted Gene Co-expression Network Analysis (WGCNA, v1.47) package in R69 based on pairwise correlations between genes due to their similar expression profile. After filtering of the genes, the gene expression values from 70,458 genes were imported into WGCNA to construct co-expression modules using the automatic network construction function blockwiseModules (TOMType = ‘signed’, mergeCutHeight = 0.15, minModuleSize = 50, power = 12). Total connectivity and intramodular connectivity (function softConnectivity, connet = abs (cor(datExpr, use = “p”))^6, Alldegrees1 = intramodularConnectivity (connet, moduleColors)), kME (for modular membership, also known as eigengene-based connectivity, datKME = signedKME(datExpr, MEs, outputColumnName = “MM.”)) and kME-P-values were calculated for the 30,038 genes, which were clustered into 17 correlated modules. The other 57 genes were outliers (gray module) and are not presented. For those DEGs, we used hierarchically clustered heat maps to investigate their relative expression levels in different samples.
Venn diagrams were employed to calculate the DEG set in various comparisons. Trend analysis was performed using Short Time-series Expression Miner software (STEM)70. The networks were visualized using Cytoscape_v3.0.0.
Protein extraction, protein digestion, and TMT labeling
The samples (FB.Apr, FB.May, FB.Nov, nF16, and IB570) were used for protein extraction. The total proteins from each sample were extracted using the cold acetone method71. The concentration of total extracted protein was measured using a BCA protein assay kit, and protein quality was confirmed by SDS-PAGE. Proteins were then tryptically digested quickly and entirely into peptides with sequence-grade modified trypsin (Promega, Madison, WI). After trypsin digestion, the peptide samples were centrifuged, drained by vacuum, and then reconstituted in 500 mM TEAB (triethylammonium bicarbonate). Each peptide mixture sample was labeled with various TMT 10-plex isobaric labeling tags (ThermoFisher Scientific, MA, USA) according to the manufacturer’s instructions by incubating at room temperature for 2 h. Subsequently, the labeled peptide mixture was redissolved and fractionated using high-pH separation. For each sample, twelve separated fractions were collected and dried for subsequent identification.
LC–MS/MS, data, and functional analysis
Each collected peptide fraction was resuspended, separated, and analyzed on an Easy-nLC 1000 system (ThermoFisher Scientific, MA, USA) connected to an Orbitrap Fusion Tribrid mass spectrometer (ThermoFisher Scientific, MA, USA) equipped with an online nanoelectrospray ion source by Gene Denovo Biotechnology Co., Ltd., Mascot Distiller (version 2.6) was employed to extract tandem mass spectra, deconvolute the data, and deisotope charge states. The Mascot search engine (version 2.3.0.2, Matrix Science, London, UK) was used to identify and quantify the peptide/protein. In this study, the mascot database was set up based on the C. praecox reference transcriptome. The parameter settings for protein identification and quantitation were as described in a previous study72. We deposited our data set in iProX/ProteomeXchange under accession numbers IPX0001890001/PXD017101.
All identified proteins were annotated by searching against the GO, KOG, and KEGG databases. The significant enrichment of GO functions and KEGG pathways was determined using the hypergeometric test within differentially expressed proteins (fold change ≥ 1.2) with P ≤ 0.05. In addition, correlation analysis was performed for each comparison. To provide a valuable framework for a better understanding of the functional clustering of proteins, the protein–protein interaction network was constructed using the String program (http://www.stringdb.org/) with a confidence score threshold of 0.7. For some specific proteins, cluster heat maps were also used to reveal hierarchical clusters in the data matrices.
Verification of RNA-seq data by qPCR and phylogenetic and functional analyses of CpFT1 in Arabidopsis
To verify the reliability of RNA-seq data and determine the expression pattern of key DEGs related to chilling-induced dormancy breaking, qPCR was performed on a Bio-Rad CFX96 system using specific primers (Supplementary Table S1)51. The correlations between RNA-Seq and qRT-PCR were performed using the Spearman method, and the significance was analyzed with the Kruskal–Wallis test in R. The phylogenetic tree was constructed using neighbor-joining (NJ) bootstrap analysis (1000 replications) with the maximum composite likelihood model for DNA performed using MEGA6.0. The plant overexpression vector 35S::CpFT1 was transformed into wild-type Arabidopsis thaliana (Col-0) using the floral-dip method73. Finally, the transgenic plants, eight homozygous individuals from four different lines, were obtained by Kan-resistance screening for seeds, DNA identification of the T1 generation, and qRT-PCR analysis of T2 generation homozygotes of transgenic Arabidopsis using the primers listed in Supplementary Table S1. Three homozygous lines with different expression levels (high, medium, and low) were selected for follow-up analysis and phenotypic observations73. The significance of differences was analyzed by Student’s t-test.
Quantification of ABA and GA by HPLC
To understand the endogenous hormone levels in chilling-induced dormancy, the FB samples, FB.Nov, FB150/300/570, IB570, LB570, and WP570, potted for artificial chilling experiments from C. praecox ‘Concolor’ were quantified by high-performance liquid chromatography-mass spectrometry (HPLC-MS) analysis of citrate-buffered acetone extracts in three biological replicates. The plant hormones (ABA and GA3) were analyzed qualitatively using HPLC and the relative retention time by comparison with reference standards. The ABA and GA3 isolates had retention times of 8.99 and 11.34 min, respectively.
Results
Chilling requirement for dormancy breaking
Based on paraffin sectioning, floral organ formation in wintersweet, including tepal, stamen, and pistil primordia differentiation, was completed in April and May. Under natural conditions, wintersweet dormancy breaking occurred when CR reached 570 CU on 24 Dec 2018, and C. praecox ‘Grandiflorus’ and ‘Concolor’ had bloom ratios (BRs) of 12.8% and 13.6%, respectively. By 27 and 28 Dec 2018, the BR reached 32.9% and 32.5% under the accumulated CR of 627 CU and 649 CU, respectively, followed by 54.9% and 50.9% on 2 and 4 Jan 2019 under the accumulated CR of 768 CU and 815 CU, respectively (Fig. 1a).
For C. praecox ‘concolor’, from 18 Dec 2018 to 14 Jan 2019, the bloom ratio continued to increase from 1.3% to 75.1% under a cumulative CR from 485 to 1026 CU. To further clarify the CR for chilling-induced dormancy breaking, plants in November (FB.Nov, Fig. 1b) were treated at 12 °C or 16 °C with different CRs under the same light/dark period and light intensity conditions in the climate cabinet. After the CU reached −300, 150, 300, 450, and 570, the plants were transferred to 14 °C for phenotype observation. The results showed that only FBs under LT and SDs (8/16 h of light/dark at 12 °C) with a CR of 570 CU (IB570, Fig. 1e) could accomplish dormancy breaking and fully open (Fig. 1d, h). The FBs of nF16 (Fig. 1c, g) as well as FB150, FB300, and FB450 at 12 °C showed the same morphological phenotype as FB.Nov (CK), i.e., they could not expand and open normally, and the FBs gradually fell off.
Sequencing, assembly, and annotation of the C. praecox reference transcriptome
To obtain a reference transcriptome from C. praecox FBs, RNA-seq libraries were constructed using RNA samples including five FBs under different CR conditions (nF16, FB150, FB300, FB450, and IB570) and three FBs collected in April, May, and November (FB.Apr, FB.May, and FB.Nov). A total of 100,553 contigs were generated using Trinity65, with total residues of 82,113,056 bp and an N50 of 1,252 bp. The average length of each transcript was 816 bp, the shortest sequence was 201 bp, and the longest was 16,864 bp. A total of 36,779 unigenes were annotated, of which 36,565 (36.4%), 24,758 (24.6%), 21,727 (21.6%), and 13,556 (13.5%) unigenes showed significant similarity to known genes in the NR, Swissprot, KOG, and KEGG databases, respectively. Approximately 27.8% (10,226) of unigenes could be assigned to a homolog in all four databases (Supplementary Fig. S1a), and the distributions of E-values in each database are shown in Supplementary Fig. S1b. A large number of unigenes in C. praecox showed close identities to the genes in other plant species. The numbers of homologous genes in the top 10-hit species are shown in Supplementary Fig. S1c. The highest number of C. praecox homologous genes (9537, 26.1%) was identified in a basal eudicot, Nelumbo nucifera, suggesting a closer phylogenetic relationship between C. praecox and N. nucifera74.
Enrichment analysis of DEGs based on GO, KOG, and KEGG
Gene ontology (GO) enrichment analysis was carried out using a threshold value (P ≤ 0.05), and the primary biological functions of DEGs were classified into biological process (BP, 25,627; 48.19%), molecular function (MF, 16,250; 30.56%), and cellular components (CC, 11,302; 21.25%) (Supplementary Fig. S2a and Supplementary Table S2). Among the annotated DEGs of IB570/FB.Nov, the BP category contained the majority of GO annotations (1,515/3,335 up-/downregulated), followed by CC (868/1,843 up-/downregulated) and MF (710/1,514 up-/downregulated), in which developmental processes and rhythmic processes contained 50/120 and 1/3 up-/downregulated GO annotations, while in the case of IB570/nF16, the BP category contained 2009/1650 up-/downregulated annotations, in which the developmental process contained 68/54 up-/downregulated annotations (Supplementary Fig. S2b and Supplementary Table S3). The IB570/FB.Nov and IB570/nF16 clusters included 314 and 330 genes, respectively, and were enriched in GO categories related to floral development, organ development, and secondary metabolite production (Table 1 and Supplementary Table S4). IB570/FB.Nov or IB570/nF16 GO enrichment of the BP category contained 19 or 16 flower development (GO:0009908) annotations and 18 or 15 floral organ development (GO:0048437) annotations (Table 1 and Supplementary Table S4).
In a further analysis, the GO annotations were subjected to a search against the Clusters of Orthologous Group for eukaryotic complete genomes (KOG) database for functional classification and prediction. Based on sequence homology, 35,354 unique sequences were assigned a KOG functional classification. These sequences were classified into 25 KOG categories, denoting involvement in cellular processes, signal transduction, metabolism, and other processes (Supplementary Fig. S2c and Supplementary Table S5). To further determine the involvement of metabolic pathways in the flower opening process, we predicted the 132 KEGG pathways represented by all the assembled unigenes. The map with the highest unigene representation was the metabolic pathway (ko01100) with 2953 (39.08%) unigenes. Starch and sucrose metabolism (ko00500), plant hormone signal transduction (ko04075), and circadian rhythm (ko04712) contained 373 (4.94%), 326 (4.31%), and 72 (0.95%) unigenes ranked 7th, 10th, and 60th, respectively, of the total 132 pathways, including PHYC, PRR73, GI, FT1, and FT2 (Supplementary Table S6). The top 20 KEGG pathway enrichments in IB570/FB.Nov are shown in Supplementary Fig. S2d (Supplementary Table S7).
Correlation of differentially expressed genes (DEGs) and WGCNA
Compared with FB.Nov, nF16, and FB150/300/FB450, there were 9,884/11,817, 10,987/6,815, 20,011/36,130, 20,585/36,602, and 19,623/38,169 up-/downregulated genes in IB570, respectively (Supplementary Fig. S1d). There were 167/158 upregulated and downregulated proteins in IB570/FB.Nov and 25/50 in IB570/nF16 (Supplementary Fig. S1e).
Principal component analysis (PCA) within the R package DESeq275 and the average linkage cluster tree analysis within the WGCNA R package69,76 were carried out to screen for outlying libraries. The PCA revealed that the libraries were segregated horizontally (PC1) based on FB.Nov, nF16, and FB150/300/450 samples. Vertical segregation (PC2) clearly differentiated flowering (IB570) and non-flowering samples under different CR values, including FB.Nov, nF16, and FB150/300/450 (Fig. 2a). The Pearson correlation among three biological replicates of one sample was 0.98–1, while that among FB150, FB300, and FB450 was 0.87–0.99 and that between FB150/300/450 and IB570 was 0.66–0.72 (Fig. 2b). All the biological replicates had a strong correlation higher than 0.98 among three replicates of one sample, including libraries FB300-3 and FB450-1 (Fig. 2b).
a PCA plot of the 18 RNA-seq libraries. b Pearson correlation of the 18 RNA-seq samples. Red, dashed rectangles are placed around outlying libraries. c Module-tissue association. d Module-group relationship. Each row corresponds to a module. The major tree branches constitute 17 modules labeled by different colors. Each row corresponds to a module. The number of genes in each module is indicated on the left. The columns correspond to FB.Nov, nF16, FB150/300/450, and IB570
Chilling-induced dormancy breaking-related network modules and gene expression patterns were identified by WGCNA and STEM. The linkage cluster tree revealed that libraries FB300-3 and FB450-1 did not group with their corresponding biological replicates (Supplementary Fig. S3a). Non-flowering samples under different CR values (FB150, FB300, and FB450) at 12 °C were clustered in the same group (Supplementary Fig. S3a). They are shown in the dendrogram, in which each leaf on the branch is one gene and each branch on the tree constitutes a module (Supplementary Fig. S3b and Supplementary Table S8).
The 17 module eigengenes for the 17 distinct modules were each correlated with the CR phenotype due to the eigengenes’ CR phenotype-specific expression profiles (Fig. 2c, d). The CR combined with different temperature treatments and/or distinct phenotype networks incorporated 17 clusters (labeled by different colors) of coexpressed genes from a total of 30,038 unigenes based on WGCNA (Fig. 2d). A total of 7,042 hub genes were scanned from the module with kME > 0.95 and P < 10e−8. The module eigengene can be considered representative of the module’s gene expression profile for the first principal component of a given module. Notably, based on module-group relationship analysis (Fig. 3d and Supplementary Table S8), 7 out of 17 co-expression modules comprised genes that were highly expressed between a flowering tissue type (IB570) under 570 CU and non-flowering ones (FB150/300/450 or nF16), such as green, light cyan, cyan, dark green, turquoise, dark magenta (r ≥ 0.5, P ≤ 0.05), and especially red (r = 1, P = 1e−17). Among these modules, the correlation coefficients of turquoise-green, turquoise-cyan, green-dark green, and turquoise-light cyan ranged from −0.85 to −0.72 (p = 7e−6 ~ 8e−4) and that of dark green-light cyan was −0.51 (p = 0.03), while in the case of green-cyan, cyan-red, green-red, red-dark magenta, and green-light cyan, the correlation coefficients were between 0.5 and 0.76 (p = 0.04 ~ 3e−04; Supplementary Fig. S3c, correlation coefficients and P-values are marked in blue; Supplementary Table S8, the correlation coefficient is represented in a different color). Heat maps of the modules showing the relative normalized RPKM were generated to visualize the gene expression patterns over time, in which the largest of the modules was turquoise, containing 15,221 (50.7%) genes, followed by cyan, light cyan, pink, blue, green, dark green, and red (Figs. 2c, d, 4f–g and Supplementary Fig. S3d–i).
A total of 161 genes out of 701 selected genes (including 168 hub genes, containing 136 TFs, among which 45 TFs were hub TFs) were scanned for further gene network analysis by Cytoscape (Supplementary Table S9). Seventy-six DEGs out of the 161 genes belonged to 11 modules, the top five of which were red, light cyan, dark green, pink, and blue, with 19, 16, 9, 8, and 7 DEGs, respectively. There were 14 hub genes, and those with the top 5 highest total and intramodular connectivity were GBE3, FUL-L1, ARF10, PIF3, and LAX3 (Supplementary Table S9). Each colored circle or square (node) represents one gene. A larger node size indicates greater connectivity within the network. Unigene (U) ID in orange represents the hub gene (kME > 0.95, and P < 10E−8)
WGCNA can also be employed to construct gene networks in which each node represents a gene and the connecting lines (edges) between genes represent co-expression correlations. Hub genes are those that show the most connections in the network (kME > 0.95, and P < 10E−8). The FT homolog CpFT1 (Unigene0043357) was correlated with CpSVP1 and CpPP2C-6 (Unigene0096511, R = 0.59 and Unigene0045734, R = 0.7; Fig. 3; Supplementary Table S9), which were found in gene sets 2397 and 1628 (Fig. 4a, d and Supplementary Table S10).
a Venn diagram of DEGs from IB570/FB.Nov, IB570/nF16, IB570/FB150, IB570/FB300, and IB570/FB450. b, c Trend analysis of FB150, FB300, FB450 and IB570 (b) or FB.Nov, nF16, FB450, and IB570 (c) from the 2397 intersection in a. d Venn diagram of DEGs from FB300/FB150 and FB450/FB300, the 2,397 DEGs intersection in a, 2311 DEGs from profiles 10, 9, 19, and 6 in b, and 1700 DEGs from profiles 10, 8, 9, 3, and 6 in the trend analysis in c. e Heat map of normalized RPKM scaled by the Z-score method for 461 annotated DEGs out of 1,628 DEGs intersection in d, including CYP707A1, FT1, and SPL9. f–g Eigengene expression pattern of dark green (f) and red (g) modules
Venn diagrams, trends, and heat map analysis
To filter preferential chilling-induced dormancy breaking-related DEGs, the 2,397 DEGs intersection of IB570/FB.Nov, IB570/nF16, IB570/FB150, IB570/FB300, and IB570/FB450 was identified using a Venn diagram (Fig. 4a). To reflect the major trends and the key transitional states among different CR tissues (Fig. 4b) and diverse temperature-treated samples (Fig. 4c), 2,397 selected DEGs were assigned to different profiles by STEM, and four and six profiles (profiles 10, 9, 19, and 6 from the trend analysis of FB150, FB300, FB450, and IB570; profiles 10, 8, 9, 3, 12, and 6 from FB. Nov, nF16, FB450, and IB570) were significantly enriched (Fig. 4b, c). After excluding the union of FB300/FB150 and FB450/FB300, together with 2,397-intersection (Fig. 4a), 2,311 DEGs (profiles 10, 9, 19, and 6 in Fig. 4b), and 1,700 DEGs (profiles 10, 8, 9, 3, and 6 in Fig. 4c) for another Venn analysis, 1,628 DEGs intersection was selected for further analysis (Fig. 4d). A heat map depicting the expression of 461 annotated DEGs (Supplementary Table S10) was constructed. Beyond the flowering samples of IB570, non-flowering samples of FB150/FB300/FB450 and nF16, and FB.Nov (CK) clustered together (Fig. 4e). Downregulation or upregulation of IB570 compared with FB.Nov, nF16, FB150, FB300, and FB450 was observed in clusters I (such as CYP707A1, FT1) and II-1 (PDCB5), which showed a positive correlation with eigengene expression in the dark green or red module from WGCNA, such as CBL7 and KEGG significantly enriched FT1 in the circadian rhythm-plant pathway (Supplementary Fig. S4), or Unigene0000644 LEA3 and Unigene0094227 SPL9. The COR from the red module showed a negative correlation with its eigengene expression (Fig. 4e–g).
Proteome/transcriptome association analysis
To identify the key genes for dormancy breaking and to complement the transcriptome results, we performed a comparative proteome survey by using TMT. To identify the scenarios that were in action, we compared the log2 Fold Change between IB570 and FB.Nov or nF16 samples in the transcriptome and proteome data sets from IB570, FB.Nov, and nF16. In group IB570/FB.Nov, 8, 162, 12, 243, 3,726, 256, 16, 267, and 23 transcripts/proteins were annotated in quadrants 1–9, compared with 2, 213, 3, 85, 4,040, 51, 5, 212, and 2 in the case of IB570/nF16. Of the 4,713 proteins with both mRNA and protein values, 20.9% (987 proteins) or 14.3% (673 proteins) showed at least 2- or 1.2-fold differences in mRNA and/or protein levels according to the log2 (ratio) from IB570/FB.Nov or IB570/nF16. A scatter plot of the nine-quadrant association analysis was then subdivided into sectors, and each was assessed for gene ontology terms that showed a statistically significant increase in protein levels only for FB.Nov or nF16 (quadrant 4), in mRNA only for IB570 (quadrant 2), in protein only for IB570 (quadrant 6), and in both protein and mRNA for IB570 (quadrant 3) (Fig. 5a, b). Interestingly, the majority (25.9% or 256 proteins; quadrant 6) showed significant fold changes (IB570/FB.Nov) only in protein abundance and not in mRNA, implying that the increase in most proteins in IB570 was independent of synthesis (that is, transcription and/or translation) (Fig. 5a), whereas in the case of IB570/nF16, the majority (1.8% or 85 proteins; quadrant 4) suggested downregulation of most proteins in IB570 (Fig. 5b).
a, b Scatter plot of 9-quadrant associate analyses of mRNA and proteins from log2 FC (IB570/FB.Nov) and (IB570/nF16). Number 1–9, quadrant NO. The number of points in each quadrant is shown in parentheses. Unigene ID highlighted 24 or 14 transcripts and/or proteins. c KOG functional classification of differentially expressed proteins (fold change ≥ 1.2). d, e Heat map analysis of 24 (d) or 14 (e) normalized RPKM values scaled by the Z-score method from RNA and proteins in FB.Nov, nF16, and IB570
A total of 7,344 TMT proteins were classified into 25 KOG categories, denoting involvement in RNA processing and modification, cell wall/membrane/envelope biogenesis, and other processes (Fig. 5c and Supplementary Table S11). Labels in the nine-quadrant association analysis diagram and heat map analysis were obtained using 24 and 14 selected unigenes and the predicted proteins from IB570/FB.Nov or IB570/nF16. Protein expression was upregulated in IB570 cells compared with FB.Nov, such as Unigene0096025 CML26, Unigene0068304 FPA, and Unigene0056062 FY from quadrant 6, Unigene0068303/Unigene0031805 FPA from quadrant 9, which was downregulated in Unigene0068565 PHYC, Unigene41519 EXLA1 from quadrant 4, and Unigene0084969 TPS1 and Unigene0022050 HSP70 from quadrant 7 (Fig. 5a, d). In group IB570/nF16, protein expression was downregulated in Unigene0069339 COR2 from quadrant 4 (Fig. 5b, e). At the mRNA level, Unigene0059859 SAPK7 expression in IB570 was significantly higher than that in FB.Nov and nF16 by ca. 1.61- and 1.28-fold, respectively (Fig. 5).
Verification and phylogenetic analysis of DEGs during dormancy breaking
The transcript abundance of 12 DEGs was analyzed to verify the reliability of the RNA-Seq data using qRT-PCR (Fig. 6a). Linear regression analysis showed an overall correlation coefficient (R) = 0.75 and 0.76 for IB570/FB.Nov and IB570/nF16, respectively, which indicates a good correlation between the qRT-PCR results and transcription profile, with corresponding Spearman correlations of R = 0.52–0.97 (Fig. 6a, b). The relative expression level in IB570 was significantly downregulated compared with that in FB.Nov and nF16, respectively, such as COR2, SVP1, SVP2, CO, FT1, SOC1a, FUL-like1, SEP1, and CWINV1; among them, COR2, CO, FUL-L1, and CWINV1 were also downregulated in IB570 compared with FB300 and FB450, while significant upregulation of IAA16 was detected in IB570 compared to other samples (Fig. 6a).
Phylogenetic analysis of flower development-related DEGs showed that Unigene0096511 CpSVP1 and Unigene0042050 CpSVP2 clustered closer to SVP in Arabidopsis and PtSVL in Populus than to Japanese apricot, peach, or pear DAMs, which comprise a larger subfamily of MADS-box genes, SVP/AGL24; Unigene0025911 CpAGL6 and Unigene0046232 CpFUL-L1 formed a clade with AGL6 and FUL from the AP1/SEP/AGL6 superclade of MADS-box genes (Supplementary Fig. S5).
Concentration of ABA and GA and expression analysis
The concentration of ABA in FBs increased significantly from FB.Nov to FB150 and FB300 reached a peak in FB570, then experienced a sustained decline from IB570 to LB570 and WP570. A significant decrease in the concentration of GA3 was detected from FB.Nov to FB150 and FB300, followed by an increase to FB570, and then a decreasing trend, similar to that of ABA, from IB570 to LB570 and WP570 (Fig. 7a). Through the analysis of the RNA-seq data and qPCR, we found that hormone-related genes changed greatly in the process of chilling-induced dormancy breaking, especially the genes related to ABA and GA biosynthesis and signal transduction pathways (Figs. 7b and Supplementary Fig. S6).
a Concentration of plant hormones (ABA and GA3) in samples under different artificial chilling treatments at different stages. b Heat map of 26 candidate genes based on the log2 (fold change) of RPKM scaled by the Z-score method. B, blue; DGn, dark green; DT, dark turquoise; LC, light cyan; P, pink; R, red; T, turquoise. Black bold italics, yellow bold/white bold numbers represent no significant difference, significant up-/downregulation at a fold change ≥ 1.5 (FDR ≤ 0.05). c, d Relative expression of ABA- and GA-related DEGs in FBs under natural conditions in 2019–2020. Different letters indicate significant differences (P < 0.05) in FBs on 15-Nov, 24-Dec, 29-Dec (567 CU), 2-Jan (nDP stage, 608CU), and 7-Jan (DP and IB stages) based on a t-test
The following was also observed: significant upregulation of the abscisic acid receptor Unigene0111683 PYL4 in IB570/FB.Nov and IB570/FB150, followed by the downregulation of protein phosphatase, Unigene0045734 PP2C-6, in IB570 compared with that in FB.Nov, nF16, and FB150/300/450, followed by the upregulation of serine/threonine-protein kinase SnRK2, Unigene0040392 SRK2E in IB570/FB150, IB570/FB300, and IB570/FB450 and Unigene0059859 SAPK7 in IB570/FB.Nov (Fig. 7b). Significantly enriched abscisic acid-insensitive 5 (ABI5, Unigene0029725, and Unigene0038509) and Unigene0010266 ABI3 showed downregulation in pairwise comparisons, except for Unigene0038509 ABI5–3 in IB570/FB300 and IB570/nF16, with no significant difference (Fig. 7b and Supplementary Fig. S6). Unigene0061565/Unigene0019450 CYP707A1 was downregulated in five pairwise comparisons, except for Unigene0019450 in IB570/FB.Nov. The relative expression of CpPP2C-6 (Unigene0045734), CpABI3 (Unigene0010266), and CpABI5-1/2/3 (Unigene0029724, Unigene0029725, Unigene0038509) in FBs on 29 Dec 2019 (567 CU, 1.42% BR) was ca. 6.3-, 25.7-fold, and 13.1-/19.4-/3-fold lower, respectively, than that on 15 November (0 CU). The expression of Unigene0061565 CYP707A1 in nDP (nearly to the displayed petal stage, 2-Jan) was significantly reduced by ~2.1-fold compared to that in FBs on 29-Dec (Fig. 7c).
Two gibberellin receptor GIDs (Unigene0076024 GID1C, Unigene0000456 GID2-1) and transcription factor Unigene0026500/Unigene0037263 PIF3 were downregulated in IB570 compared with FB.Nov and/or nF16. Downregulation of Unigene0026500/Unigene0037263 PIF3 was detected in IB570 cells compared to FB150/300/450 cells (Fig. 7b). The expression of Unigene0076024/Unigene000456 GID1C/GID2-1 and Unigene0037263 PIF3 in FBs on 29 Dec was ca. 3.8-/2.8-fold and 6.3-fold lower, respectively, than that on 15 Nov (Fig. 7d and Supplementary Fig. S6).
The expression of CpSVP1, CpSVP2, CpFT1, CpSOC1a, and CpFUL-L1 in FBs on 24-Dec (475 CU, BR of 0.4%), 29-Dec 2019 (567 CU, BR of 1.42%), 2-Jan (nDP stage, 608CU, BR of 6.94%) and 7-Jan 2020 (DP, displayed petal stage, 690CU, BR of 35.83%) showed significant downregulation compared to that in FBs on 15-Dec-2019 (Fig. 8).
Functional analysis of CpFT1
CpFT1 (accession number: MT565392) is 1,060 bp in length and has an open reading frame (ORF) of 525 bp, which encodes a predicted polypeptide of 174 aa with a 5′/3′-UTR of 346 and 189 bp (Fig. 9a). In the phylogenetic tree, CpFT1 clustered close to Hd3a in Oryza, TaFT in wheat, and ZCN15 in maize (Fig. 9b). The ectopic expression results indicated that the time of bolting, the first flower opening, and the first fruit emerging in the 35S::CpFT1 transgenic lines (OE4–5, OE6–5, OE5–7) were significantly earlier than those in the wild-type (WT; Fig. 9c, d). The number of rosette leaves of transgenic lines (average 8.22) was lower than that of the WT (average 12.47) (Fig. 9e). The results of qRT-PCR for different 35S::CpFT1 lines and WT showed that the heterologous expression of CpFT1 in Arabidopsis upregulated the expression levels of SOC1, LFY, AP1, and SEP3, which resulted in early flowering in Arabidopsis (Fig. 9f). Therefore, it is inferred that the overexpression of CpFT1 can promote flowering in Arabidopsis.
a The DNA and protein sequence of CpFT1. b Phylogenetic tree based on NJ bootstrap analysis. c, d Phenotype of the 35S::CpFT1/Col-0 T2 generation. e Numbers of rosette leaves of transgenic lines compared with WT. f Relative expression analysis in ectopic expression lines of CpFT1. The blue underline shows the forward and reverse primers for the 35S::CpFT1 vector construction
Discussion
Prolonged exposure to cold temperatures, also known as chilling, regulates dormancy release, including bud break or bud flush77. In Prunus mume, floral meristem (FM) differentiation is completed in summer, while blooming occurs until the next early spring56. In the case of wintersweet, paraffin sections showed that FM formed from March to early April, followed by tepal primordium differentiation from the end of April to early May, followed by stamen and pistil primordium in mid- and late May. The development of anthers stagnated from June to August, resumed in September, and entered the dormant period from October (Supplementary Fig. S7). Under natural conditions, FB experienced cold induction from mid- to late November and opened fully in late December to January in Chongqing.
P. mume cultivars ‘Nanko’ and ‘Ellching’, from temperate Japan and subtropical Taiwan, require ~500 and 300 “chilling hours” (CHs), respectively, to break FB dormancy78. Peach cultivars with CRs ranging from fewer than 50 CU to more than 1000 CU have been developed and used for breeding and cultivation worldwide56. The results from our experiment showed that all the wintersweet plants in the field or potted plants underwent the CR for dormancy breaking, reaching 570 CU. The FBs expanded and fully opened, although temperature fluctuation may result in the CR value for chilling-induced dormancy breaking in wintersweet cultured in the field being slightly higher than that in the artificial climate chamber under LT (12 °C) and SDs conditions. Chilling below 7 °C is not the necessary condition for dormancy breaking in wintersweet. In Guangzhou City, Guangdong Province, the FBs of wintersweet cultured in the open field did not fully open (just like nF16), similar to the nonchilled peach FBs, which were almost undeveloped even at 20–23 °C for a few months. Early work was based on the sum of hours at a temperature <7.2 °C (chilling hours), which were added throughout the estimated period of dormancy in peach79. The “UTAH Model” proposes records on the number of “chill units”, establishing a different chilling contribution for different temperature ranges63, which is more suitable for chilling requirement analysis in chilling-induced dormancy breaking in wintersweet.
Previously, a group of genes encoding MADS-box TFs called DAMs, phylogenetically related to A. thaliana SVP and AGL24, was recruited to regulate the dormancy cycle in perennial woody plants80. For example, a negative correlation between the PpDAM5 and PpDAM6 transcript levels of lateral vegetative buds and the bud burst ratios and flower organ expansion rate was found in the low- and high-chill field cultivars of P. persica81. After organ differentiation, a prolonged period of artificial chilling is necessary for the reduction of the PpDAM5 and PpDAM6 transcript levels in the FBs, which are recruited in flower organ enlargement29,82. Six PmDAM genes in P. mume showed downregulation following prolonged artificial chilling exposure. A certain CR may be necessary for the downregulation of PmDAM4 to PmDAM6 in high-chill ‘Nanko’56. The distinct changes in PmDAM4 to PmDAM6 expression may contribute to the different CRs for dormancy release in ‘Ellching’ and ‘Nanko’36. PpDAM proteins downregulated the expression of PpFT2 during dormancy release in P. pyrifolia ‘Suli’34. In addition, MdDAMs showed seasonal mRNA fluctuation patterns and were downregulated by artificial cold exposure30. In hybrid aspen, SVL has been shown to participate in the regulation of both the entry and release of dormancy as mediators of temperature-controlled bud break16,19,77. Although high similarity was found among SVL, SVP, and DAM, the SVL clusters were closer to the SVP than to the DAM genes19. Phylogenetic analysis of CpSVP1/2 in C. praecox showed the same results as poplar SVL (Fig. S5). The downregulation of SVL in buds of hybrid aspen by prolonged exposure to LT was similar to the downregulation of the other DAM genes associated with bud break28,33,83. SVL negatively regulates FT1, one of two poplar FT paralogs (FT1 and FT2), which have divergent functionally84. FT has an important role in flower initiation and participates in the regulation of dormancy in perennial trees8,85. FT1 is highly upregulated by chilling in dormant vegetative buds84. In our study, CpFT1 was significantly enriched in the KEGG circadian rhythm-plant pathway and associated with chilling-induced dormancy breaking (Fig. 4d, e). However, it was unexpected that the expression trend of CpFT1 was highly consistent with that of CpSVP1, other than the negative regulation of FT by SVP in other species. One possible explanation for this result is that there is no FLC in the C. praecox genome. The repression of FT expression by SVP may depend on the FLC that can interact with SVP86. The transcription profiling of the chilling requirement for bud break in apples revealed strong differential expression in FLC-like and MADS AFFECTING FLOWERING (MAF); the upregulation of FLC-like showed a remarkable induction towards dormancy release55. Recently, it was reported that PavDAM1/5 could interact with PavSOC1 in vivo and in vitro and coregulate flower development in sweet cherries (Prunus avium)87. In wintersweet, CpSVP1 may interact with CpSOC1a to positively regulate CpFT1, so the downregulation of CpSVP1 and CpSOC1a results in the downregulation of CpFT1 during dormancy breaking (Fig. 10).
Hormone homeostasis has an important role in bud dormancy4. Two core hormones, ABA and GA, antagonistically regulate bud dormancy status. In hybrid aspen, a prolonged chilling signal reduces ABA levels and triggers a reduction in SVL expression, resulting in the upregulation of GA biosynthesis and FT1 expression, which promotes bud break19. In P. mume, the ABA/GA ratio was reported to steadily decline during the dormancy release process60. In the present investigation, however, the ABA content significantly increased in FB570 compared with FB.Nov under artificial treatment, while the content of GA3 decreased (Fig. 7a), implying that the ABA/GA3 ratio significantly increased during chilling-induced dormancy breaking. During the process of FB enlargement and blossoming, the content of GA3 in FBs was much higher than that of ABA, which is consistent with observations in tree peony88,89. In support of this, we found a significant downregulation of the ABA 8-hydrolase gene (CYP707A1) and upregulation of the ABA synthesis gene NCED in FB570 compared to FB.Nov and FB150-450. Moreover, significant changes in the expression of the genes related to the ABA signaling pathway were also detected between FB570 and FB.Nov. For instance, the genes encoding ABA receptors (PYL4) and SnRK2 kinase were significantly upregulated, while the ABA coreceptor PP2C was evidently downregulated in FB570, which is consistent with their roles in the ABA signaling pathway and the regular responses to a high ABA content (Fig. 10). Upon ABA signaling, PYR/PYL receptors interact with PP2Cs, thus disrupting the PP2C-SnRK2 interactions, which release SnRK2 kinases from PP2C-mediated inhibition, allowing SnRK2s to phosphorylate and activate downstream transcription factors such as ABI/ABFs90. The downregulation of the ABA pathway could result in the downregulation of SVL expression upon exposure to prolonged chilling in hybrid aspen19. In wintersweet, however, the downregulation of CpSVP1 was accompanied by an increased ABA content, which may be explained as follows: although the ABA content increased during CR accumulation in wintersweet, ABI3 and ABI5 homologs were significantly downregulated in the FBs during the process of CR accumulation (Fig. 10), which is consistent with the results in other species during chilling-induced dormancy breaking, where the ABA content usually decreased. Recently, it was reported that ABA-responsive PpyABF3 can directly bind to the ABRE element in the promoter of PpyDAM3 and activate its expression to promote bud dormancy in Asian pear (P. pyrifolia)91. In addition, SOC1 was identified to be a direct downstream target of ABF3/4 in Arabidopsis, and the induction of SOC1 by ABA was hampered in abf3 abf4 mutants92. In wintersweet, the expression patterns of CpSVP1 and CpSOC1a were consistent with those of the ABF homologs (CpABI3 and CpABI5s), suggesting that CpABI3 and CpABI5s transcription factors may positively regulate the expression of CpSVP1 and CpSOC1a (Fig. 10).
In C. praecox, the reactivation of blossoming (dormancy breaking followed by enlargement and blooming) in the FBs was induced by LT combined with SD conditions when the CR reached 570 CU. The hormone accumulation patterns in the FBs during chilling-induced dormancy breaking in C. praecox, that is, up- and downregulation of ABA and GA, are opposite to the situation in other species such as Populus during bud break. This may have resulted from the fact that bud break in Populus is different from FB dormancy breaking in C. praecox, which undergoes two different biological processes, vegetative bud growth as well as FB enlargement and blooming.
AP1/FUL-like genes have been reported to work downstream of FT-like genes or to be direct targets in other species46,93,94,95. In winter wheat and barley, similar regulation strategies of flower arrest and promotion to Arabidopsis were found, but different policies were exploited, including at least four critical regulators for vernalization-induced flowering, such as VRN1 coding for AP1/FUL-like, VRN2 for CO-like, VRN3 for FT, and PPD1 for PRR796,97,98,99. VRN2 and/or VEGETATIVE TO REPRODUCTIVE TRANSITION2 (VRT2, SVP orthologous gene) repressed VRN1 expression in wheat100,101. VRN1 was proven to work upstream of VRN3 (WFT) by binding to the CArG-box in the promoter region102; OsMADS14, OsMADS15, and OsMADS18 (AP1/FUL-like genes) can directly upregulate Hd3a and RFT1 (FT homologs) in rice47. Before bud hibernation, the FUL-like ortholog PlacFL2 from the basal eudicot Platanus declined, implying that it may control SD-mediated dormancy and growth cessation50. Similar to LAP1, a tree ortholog of AP1 participated in SD-mediated seasonal growth cessation downstream of the CO/FT module in hybrid aspen49. In this study, CpSVP1 participated in the reduction of the SVP-mediated promotion of CpFT1 and subsequently gave rise to CpFUL-L1. In addition, CpFUL-L1 could give rise to FT expression levels in 35S::CpFUL-L1/Col-0 transgenic Arabidopsis lines with an early flowering phenotype compared to WT (unpublished data).
In summary, we propose that the extended cold exposure signal results in the upregulation of the ABA content in FBs under SD and LT of 12 °C with a CR of 570 CU. An increase in ABA may be a result of the upregulated NCED and downregulated CYP707A1. ABA as a signal was accepted by PYL4, and ABA-PYL complexes can inhibit PP2C, which suppresses SnRK2 function through dephosphorylation. Phosphorylated SnRK2 can activate RAV1, which then inhibits the expression of ABI3 and ABI5s. SD and short-term LT treatments induced the upregulation of SVP1, while prolonged CR caused the reduction of SVP1/2, which combined with SOC1a to induce FT1 expression and reduction of GA, subsequently inducing the breaking of dormancy in C. praecox.
References
Rohde, A. & Bhalerao, R. P. Plant dormancy in the perennial context. Trends Plant Sci. 12, 217–223 (2007).
Singh, R. K., Svystun, T., AlDahmash, B., Jonsson, A. M. & Bhalerao, R. P. Photoperiod- and temperature-mediated control of phenology in trees—a molecular perspective. N. Phytol. 213, 511–524 (2017).
Olsen, J. E. Light and temperature sensing and signaling in induction of bud dormancy in woody plants. Plant Mol. Biol. 73, 37–47 (2010).
Cooke, J. E., Eriksson, M. E. & Junttila, O. The dynamic nature of bud dormancy in trees: environmental control and molecular mechanisms. Plant Cell Environ. 35, 1707–1728 (2012).
Petterle, A., Karlberg, A. & Bhalerao, R. P. Daylength mediated control of seasonal growth patterns in perennial trees. Curr. Opin. Plant Biol. 16, 301–306 (2013).
Ruttink, T. et al. A molecular timetable for apical bud formation and dormancy induction in poplar. Plant Cell 19, 2370–2390 (2007).
Rinne, P. L., Kaikuranta, P. M. & van der Schoot, C. The shoot apical meristem restores its symplasmic organization during chilling-induced release from dormancy. Plant J. 26, 249–264 (2001).
Bohlenius, H. et al. CO/FT regulatory module controls timing of flowering and seasonal growth cessation in trees. Science 312, 1040–1043 (2006).
Maurya, J. P. & Bhalerao, R. P. Photoperiod- and temperature-mediated control of growth cessation and dormancy in trees: a molecular perspective. Ann. Bot. 120, 351–360 (2017).
Heide, O. M. Dormancy release in beech buds (Fagus-Sylvatica) requires both chilling and long days. Physiol. Plant. 89, 187–191 (1993).
Heide, O. M. Interaction of photoperiod and temperature in the control of growth and dormancy of Prunus species. Sci. Hortic. 115, 309–314 (2008).
Heide, O. M. & Prestrud, A. K. Low temperature, but not photoperiod, controls growth cessation and dormancy induction and release in apple and pear. Tree Physiol. 25, 109–114 (2005).
Tanino, K. K., Kalcsits, L., Silim, S., Kendall, E. & Gray, G. R. Temperature-driven plasticity in growth cessation and dormancy development in deciduous woody plants: a working hypothesis suggesting how molecular and cellular function is affected by temperature during dormancy induction. Plant Mol. Biol. 73, 49–65 (2010).
Liu, J. & Sherif, S. M. Hormonal orchestration of bud dormancy cycle in deciduous woody perennials. Front. Plant Sci. 10, 1136 (2019).
Hao, X. et al. Comprehensive transcriptome analyses reveal differential gene expression profiles of Camellia sinensis Axillary Buds at para-, endo-, ecodormancy, and bud flush stages. Front. Plant Sci. 8, 553 (2017).
Singh, R. K., Miskolczi, P., Maurya, J. P. & Bhalerao, R. P. A tree ortholog of SHORT VEGETATIVE PHASE floral repressor mediates photoperiodic control of bud dormancy. Curr. Biol. 29, 128–133.e2 (2019).
Rinne, P. L. et al. Chilling of dormant buds hyperinduces FLOWERING LOCUS T and recruits GA-inducible 1,3-beta-glucanases to reopen signal conduits and release dormancy in Populus. Plant Cell 23, 130–146 (2011).
Zhuang, W. et al. Comparative proteomic and transcriptomic approaches to address the active role of GA4 in Japanese apricot flower bud dormancy release. J. Exp. Bot. 64, 4953–4966 (2013).
Singh, R. K. et al. A genetic network mediating the control of bud break in hybrid aspen. Nat. Commun. 9, 4173 (2018).
Tylewicz, S. et al. Photoperiodic control of seasonal growth is mediated by ABA acting on cell-cell communication. Science 360, 212–215 (2018).
Wang, D. et al. Expression of ABA metabolism-related genes suggests similarities and differences between seed dormancy and bud dormancy of peach (Prunus persica). Front. Plant Sci. 6, 1248 (2015).
Li, J. et al. Abscisic acid (ABA) promotes the induction and maintenance of pear (Pyrus pyrifolia White Pear Group) flower bud endodormancy. Int. J. Mol. Sci. 19, https://doi.org/10.3390/ijms19010310 (2018).
Wang, Z. et al. Comparative transcriptome analysis identifies differentially expressed genes between normal and late-blooming Siberian apricot. J. Forest. Res. 30, 2277–2288 (2018).
Wen, L. H. et al. Expression analysis of ABA- and GA-related genes during four stages of bud dormancy in Japanese apricot (Prunus mume Sieb. et Zucc). J. Hortic. Sci. Biotechnol. 91, 362–369 (2016).
Hartmann, U. et al. Molecular cloning of SVP: a negative regulator of the floral transition in Arabidopsis. Plant J. 21, 351–360 (2000).
Bielenberg, D. G. et al. Sequencing and annotation of the evergrowing locus in peach [Prunus persica (L.) Batsch] reveals a cluster of six MADS-box transcription factors as candidate genes for regulation of terminal bud formation. Tree Genet. Genomes 4, 495–507 (2008).
Lee, J. H. et al. Role of SVP in the control of flowering time by ambient temperature in Arabidopsis. Genes Dev. 21, 397–402 (2007).
Li, Z., Reighard, G. L., Abbott, A. G. & Bielenberg, D. G. Dormancy-associated MADS genes from the EVG locus of peach [Prunus persica (L.) Batsch] have distinct seasonal and photoperiodic expression patterns. J. Exp. Bot. 60, 3521–3530 (2009).
Yamane, H., Ooka, T., Jotatsu, H., Sasaki, R. & Tao, R. Expression analysis of PpDAM5 and PpDAM6 during flower bud development in peach (Prunus persica). Sci. Hortic. 129, 844–848 (2011).
Porto, D. D. et al. Structural genomics and transcriptional characterization of the dormancy-associated MADS-box genes during bud dormancy progression in apple. Tree Genet. Genomes 12, 46 (2016).
Leida, C., Conesa, A., Llacer, G., Badenes, M. L. & Rios, G. Histone modifications and expression of DAM6 gene in peach are modulated during bud dormancy release in a cultivar-dependent manner. N. Phytol. 193, 67–80 (2012).
Saito, T. et al. Histone modification and signalling cascade of the dormancy-associated MADS-box gene, PpMADS13-1, in Japanese pear (Pyrus pyrifolia) during endodormancy. Plant Cell Environ. 38, 1157–1166 (2015).
Ubi, B. E. et al. Molecular cloning of dormancy-associated MADS-box gene homologs and their characterization during seasonal endodormancy transitional phases of Japanese pear. J. Am. Soc. Hortic. Sci. 135, 174–182 (2010).
Niu, Q. et al. Dormancy-associated MADS-box genes and microRNAs jointly control dormancy transition in pear (Pyrus pyrifolia white pear group) flower bud. J. Exp. Bot. 67, 239–257 (2016).
Kumar, G. et al. Comparative phylogenetic analysis and transcriptional profiling of MADS-box gene family identified DAM and FLC-like genes in apple (Malusx domestica). Sci. Rep. 6, 20695 (2016).
Sasaki, R. et al. Functional and expressional analyses of PmDAM genes associated with endodormancy in Japanese apricot. Plant Physiol. 157, 485–497 (2011).
Wu, R. et al. SVP-like MADS box genes control dormancy and budbreak in apple. Front. Plant Sci. 8, 477 (2017).
Yamane, H. et al. Overexpression of Prunus DAM6 inhibits growth, represses bud break competency of dormant buds and delays bud outgrowth in apple plants. PLoS ONE 14, e0214788 (2019).
Tuan, P. A., Bai, S., Saito, T., Ito, A. & Moriguchi, T. Dormancy-associated MADS-box (DAM) and the abscisic acid pathway regulate pear endodormancy through a feedback mechanism. Plant Cell Physiol. 58, 1378–1390 (2017).
Zhang, H., Bishop, B., Ringenberg, W., Muir, W. M. & Ogas, J. The CHD3 remodeler PICKLE associates with genes enriched for trimethylation of histone H3 lysine 27. Plant Physiol. 159, 418–432 (2012).
Ito, A. et al. Physiological differences between bud breaking and flowering after dormancy completion revealed by DAM and FT/TFL1 expression in Japanese pear (Pyrus pyrifolia). Tree Physiol. 36, 109–120 (2016).
Ruonala, R., Rinne, P. L., Kangasjarvi, J. & van der Schoot, C. CENL1 expression in the rib meristem affects stem elongation and the transition to dormancy in Populus. Plant Cell 20, 59–74 (2008).
Zhang, J. et al. The FLOWERING LOCUS T orthologous gene of Platanus acerifolia is expressed as alternatively spliced forms with distinct spatial and temporal patterns. Plant Biol. 13, 809–820 (2011).
Cai, F. et al. Identification and characterisation of a novel FT orthologous gene in London plane with a distinct expression response to environmental stimuli compared to PaFT. Plant Biol. 21, 1039–1051 (2019).
Abe, M. et al. FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309, 1052–1056 (2005).
Wigge, P. A. et al. Integration of spatial and temporal information during floral induction in Arabidopsis. Science 309, 1056–1059 (2005).
Kobayashi, K. et al. Inflorescence meristem identity in rice is specified by overlapping functions of three AP1/FUL-like MADS box genes and PAP2, a SEPALLATA MADS box gene. Plant Cell 24, 1848–1859 (2012).
Meir, M. et al. Dormancy release and flowering time in Ziziphus jujuba Mill., a “direct flowering” fruit tree, has a facultative requirement for chilling. J. Plant Physiol. 192, 118–127 (2016).
Azeez, A., Miskolczi, P., Tylewicz, S. & Bhalerao, R. P. A tree ortholog of APETALA1 mediates photoperiodic control of seasonal growth. Curr. Biol. 24, 717–724 (2014).
Zhang, S. et al. Identification and characterization of FRUITFULL-like genes from Platanus acerifolia, a basal eudicot tree. Plant Sci. 280, 206–218 (2019).
Li, Z. et al. Floral scent emission from nectaries in the adaxial side of the innermost and middle petals in Chimonanthus praecox. Int. J. Mol. Sci. 19, 3278 (2018).
Kumar, G. et al. Chilling affects phytohormone and post-embryonic development pathways during bud break and fruit set in apple (Malus domestica Borkh.). Sci. Rep. 7, 42593 (2017).
Takeuchi, T. et al. RNA-sequencing analysis identifies genes associated with chilling-mediated endodormancy release in apple. J. Am. Soc. Hortic. Sci. 143, 194–206 (2018).
Falavigna, Vd. S. et al. Differential transcriptional profiles of dormancy-related genes in apple buds. Plant Mol. Biol. Rep. 32, 796–813 (2013).
Porto, D. D. et al. Transcription profiling of the chilling requirement for bud break in apples: a putative role for FLC-like genes. J. Exp. Bot. 66, 2659–2672 (2015).
Yamane, H. Regulation of bud dormancy and bud break in Japanese apricot (Prunus mume Siebold ^|^amp; Zucc.) and peach [Prunus persica (L.) Batsch]: a summary of recent studies. J. Jpn. Soc. Hortic. Sci. 83, 187–202 (2014).
Bai, S. et al. Transcriptome analysis of Japanese pear (Pyrus pyrifolia Nakai) flower buds transitioning through endodormancy. Plant Cell Physiol. 54, 1132–1151 (2013).
Liu, G. et al. Transcriptomic analysis of ‘Suli’ pear (Pyrus pyrifolia white pear group) buds during the dormancy by RNA-Seq. BMC Genomics 13, 700 (2012).
Nishitani, C. et al. Transcriptome analysis of Pyrus pyrifolia leaf buds during transition from endodormancy to ecodormancy. Sci. Hortic. 147, 49–55 (2012).
Zhang, Z. et al. Transcriptome profiles reveal the crucial roles of hormone and sugar in the bud dormancy of Prunus mume. Sci. Rep. 8, 5090 (2018).
Kanjana, W. et al. Transcriptome analysis of seed dormancy after rinsing and chilling in ornamental peaches (Prunus persica (L.)) Batsch. BMC Genomics 17, 575 (2016).
Habu, T. et al. 454-Pyrosequencing of the transcriptome in leaf and flower buds of Japanese apricot (Prunus mume Sieb. et Zucc.) at different dormant stages. J. Jpn. Soc. Hortic. Sci. 81, 239–250 (2012).
Richardson, E. A., Seeley, S. D. & Walker, D. R. A model for estimating the completion of rest for redhaven and elberta peache trees. HortScience 9, 331–332 (1974).
Li, J. et al. PpCBFs selectively regulate PpDAMs and contribute to the pear bud endodormancy process. Plant Mol. Biol. 99, 575–586 (2019).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Ye, J. et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res. 34, W293–W297 (2006).
Ye, J. et al. WEGO 2.0: a web tool for analyzing and plotting GO annotations, 2018 update. Nucleic Acids Res. 46, W71–W75 (2018).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Ernst, J. & Bar-Joseph, Z. STEM: a tool for the analysis of short time series gene expression data. BMC Bioinformatics 7, 191 (2006).
Zhang, C. W. et al. Transcriptomic and proteomic analyses provide new insights into the regulation mechanism of low-temperature-induced leafy head formation in Chinese cabbage. J. Proteom. 144, 1–10 (2016).
Wu, L. Y. et al. Complementary iTRAQ proteomic and transcriptomic analyses of leaves in tea plant (Camellia sinensis L.) with different maturity and regulatory network of flavonoid biosynthesis. J. Proteome Res. 18, 252–264 (2019).
Li, Z., Zeng, S., Li, Y., Li, M. & Souer, E. Leaf-like sepals induced by ectopic expression of a SHORT VEGETATIVE PHASE (SVP)-like MADS-box gene from the basal eudicot epimedium sagittatum. Front. Plant Sci. 7, 1461 (2016).
Byng, J. W. et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 181, 1–20 (2016).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Langfelder, P. & Horvath, S. Fast R. Functions for robust correlations and hierarchical clustering. J. Stat. Softw. 46, http://www.ncbi.nlm.nih.gov/pubmed/23050260 (2012).
Busov, V. B. Plant development: dual roles of poplar SVL in vegetative bud dormancy. Curr. Biol. 29, R68–R70 (2019).
Yamane, H. et al. Differential expression of dehydrin in flower buds of two Japanese apricot cultivars requiring different chilling requirements for bud break. Tree Physiol. 26, 1559–1563 (2006).
Weinberger, J. H. Chilling requirements of peach varieties. Proc. Am. Soc. Hortic. Sci. 56, 122–128 (1950).
Falavigna, V. D. S., Guitton, B., Costes, E. & Andres, F. I want to (Bud) break free: the potential role of DAM and SVP-like genes in regulating dormancy cycle in temperate fruit trees. Front. Plant Sci. 9, 1990 (2018).
Yamane, H. et al. Comparative analyses of dormancy-associated MADS-box genes, PpDAM5 and PpDAM6, in low- and high-chill peaches (Prunus persica L.). J. Jpn. Soc. Hortic. Sci. 80, 276–283 (2011).
Yamane, H. et al. Expressional regulation of PpDAM5 and PpDAM6, peach (Prunus persica) dormancy-associated MADS-box genes, by low temperature and dormancy-breaking reagent treatment. J. Exp. Bot. 62, 3481–3488 (2011).
Horvath, D. Common mechanisms regulate flowering and dormancy. Plant Sci. 177, 523–531 (2009).
Hsu, C. Y. et al. FLOWERING LOCUS T duplication coordinates reproductive and vegetative growth in perennial poplar. Proc. Natl Acad. Sci. USA 108, 10756–10761 (2011).
Abelenda, J. A., Navarro, C. & Prat, S. From the model to the crop: genes controlling tuber formation in potato. Curr. Opin. Biotechnol. 22, 287–292 (2011).
Mateos, J. L. et al. Combinatorial activities of SHORT VEGETATIVE PHASE and FLOWERING LOCUS C define distinct modes of flowering regulation in Arabidopsis. Genome Biol. 16, 31 (2015).
Wang, J. et al. Dormancy-associated MADS-Box (DAM) genes influence chilling requirement of sweet cherries and co-regulate flower development with SOC1 gene. Int. J. Mol. Sci. 21, https://doi.org/10.3390/ijms21030921 (2020).
Xue, J. et al. Elucidation of the mechanism of reflowering in tree peony (Paeonia suffruticosa) ‘Zi Luo Lan’ by defoliation and gibberellic acid application. Plant Physiol. Biochem. 132, 571–578 (2018).
Guan, Y.-r et al. Effect of exogenous GA(3) on flowering quality, endogenous hormones, and hormone- and flowering-associated gene expression in forcing-cultured tree peony (Paeonia suffruticosa). J. Integr. Agric. 18, 1295–1311 (2019).
Ali, A., Pardo, J. M. & Yun, D. J. Desensitization of ABA-signaling: the swing from activation to degradation. Front. Plant Sci. 11, 379 (2020).
Yang, Q. et al. ABA-responsive ABRE-BINDING FACTOR3 activates DAM3 expression to promote bud dormancy in Asian pear. Plant Cell Environ. 43, 1360–1375 (2020).
Hwang, K., Susila, H., Nasim, Z., Jung, J. Y. & Ahn, J. H. Arabidopsis ABF3 and ABF4 transcription factors act with the NF-YC complex to regulate SOC1 expression and mediate drought-accelerated flowering. Mol. Plant 12, 489–505 (2019).
Komiya, R., Ikegami, A., Tamaki, S., Yokoi, S. & Shimamoto, K. Hd3a and RFT1 are essential for flowering in rice. Development 135, 767–774 (2008).
Teper-Bamnolker, P. & Samach, A. The flowering integrator FT regulates SEPALLATA3 and FRUITFULL accumulation in Arabidopsis leaves. Plant Cell 17, 2661–2675 (2005).
Li, C. & Dubcovsky, J. Wheat FT protein regulates VRN1 transcription through interactions with FDL2. Plant J. 55, 543–554 (2008).
Turner, A., Beales, J., Faure, S., Dunford, R. P. & Laurie, D. A. The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310, 1031–1034 (2005).
Yan, L. et al. Positional cloning of the wheat vernalization gene VRN1. Proc. Natl Acad. Sci. USA 100, 6263–6268 (2003).
Yan, L. et al. The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 303, 1640–1644 (2004).
Yan, L. et al. The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc. Natl Acad. Sci. USA 103, 19581–19586 (2006).
Kane, N. A. et al. TaVRT-2, a member of the StMADS-11 clade of flowering repressors, is regulated by vernalization and photoperiod in wheat. Plant Physiol. 138, 2354–2363 (2005).
Kane, N. A. et al. TaVRT2 represses transcription of the wheat vernalization gene TaVRN1. Plant J. 51, 670–680 (2007).
Tanaka, C. et al. Direct interaction between VRN1 protein and the promoter region of the wheat FT gene. Genes Genet. Syst. 93, 25–29 (2018).
Acknowledgements
This work was supported by grants from the Natural Science Foundation of Chongqing (No. cstc2020jcyj-msxmX1014), Fundamental Research Funds for the Central Universities (No. XDJK2020B059), National Natural Science Foundation of China (Grant No. 31971711), and Chongqing education committee project (CY200210, S202010635221). We are grateful to Professor Guofeng Liu from Guangzhou Institute of Forestry and Landscape Architecture, Professor Zhanao Deng from University of Florida, and Dr. Yan Xia from Southwest University (SWU) for critical editing of the manuscript; Chengzhi Deng from Chongqing Meteorological Bureau for providing the meteorological data; Lingying Zhu and Chaorui Wang from the high school affiliated with SWU for participating in part of the work; and all the colleagues in our laboratory for technical assistance.
Author information
Authors and Affiliations
Contributions
Z.L., S.S., and M.L. designed the research. Z.L. analyzed the sequencing data, performed graphic drawing, and editing. Z.L. and W.Z. wrote the article. N.L., Y.J., and C.W. performed the chilling requirement and HPLC experiments. N.L. performed qPCR and CpFT1 gene function analyses. J.M designed the qPCR primers.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Z., Liu, N., Zhang, W. et al. Integrated transcriptome and proteome analysis provides insight into chilling-induced dormancy breaking in Chimonanthus praecox. Hortic Res 7, 198 (2020). https://doi.org/10.1038/s41438-020-00421-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-00421-x
This article is cited by
-
Integrated transcriptome and proteome analysis provides insights into CpFPA1 for floral induction in Chimonanthus praecox (Magnoliidae) without FLC in genome
Plant Cell Reports (2024)
-
Functional characterisation of five SVP genes in grape bud dormancy and flowering
Plant Growth Regulation (2022)
-
Identification of key gene networks controlling vernalization development characteristics of Isatis indigotica by full-length transcriptomes and gene expression profiles
Physiology and Molecular Biology of Plants (2021)