Abstract
It is hypothesized that the genome-wide genic markers may increase the prediction accuracy of genomic selection for quantitative traits. To test this hypothesis, a set of candidate gene-based markers for yield and grain traits-related genes cloned across the rice genome were custom-designed. A multi-model, multi-locus genome-wide association study (GWAS) was performed using new genic markers developed to test their effectiveness for gene discovery. Two multi-locus models, FarmCPU and mrMLM, along with a single-locus mixed linear model (MLM), identified 28 significant marker-trait associations. These associations revealed novel causative alleles for grain weight and pleiotropic associations with other traits. For instance, the marker YD91 derived from the gene OsAAP3 on chromosome 1 was consistently associated with grain weight, while the gene has a significant effect on grain yield. Furthermore, nine genomic selection methods, including regression-based and machine learning-based models, were used to predict grain weight using a leave-one-out five-fold cross-validation approach to optimize the genomic selection model with genic markers. Among nine prediction models, Kernel Hilbert Space Regression (RKHS) is the best among regression-based models, and Random Forest Regression (RFR) is the best among machine learning-based models. Genomic prediction accuracies with and without GWAS significant markers were compared to assess the effectiveness of markers. The rapid decreases in prediction accuracy upon dropping GWAS significant markers indicate the effectiveness of new genic markers in genomic selection. Apart from that, the candidate gene-based markers were found to be more effective in genomic selection programs for better accuracy.

Similar content being viewed by others
Introduction
The primary and ultimate goal in any rice breeding program is to develop high-yielding varieties with acceptable-quality grains to meet the global demand for food and achieve nutritional security (Xu et al., 2014). Among the many correlated traits contributing to overall yield in rice, grain weight, measured as the weight of one thousand grains, is considered the most critical (Weng et al., 2008). Grain weight improvement in rice requires special attention as it is quantitatively inherited and is influenced by the growing environment. Rice breeders have made significant efforts for decades to improve the trait through phenotype-based selection (Ahmar et al., 2020; Chen et al., 2021; Li et al., 2021). However, the genetic gain achieved through phenotype-based selection approaches is negligible (Chakraborti et al. 2021). A revolution in DNA marker technology provided new vistas for improvement of quantitative traits like grain weight by acting as surrogates to select in early generations (Katara et al., 2021). With the application of molecular markers and linkage mapping approaches, several determinants of grain weight have been identified and fine mapped by several rice research teams (Huang et al., 2013; Zhang et al., 2016; Gao et al., 2016; Feng et al., 2018). In recent times, genome-wide association studies (GWAS) have proven promising for identifying major and minor causative alleles for complex traits like grain weight. Upon identification of major and minor quantitative trait loci (QTL), a few consensus genomic regions or meta-QTL regions accommodating QTL cumulatively contributing to grain weight have also been identified (Anilkumar et al., 2022b). While the grain weight trait is inherited through multiple minor genes distributed across all 12 rice chromosomes, marker-assisted selection (MAS) is ineffective to improve this trait significantly (Chen et al., 2021). However, genomic selection (GS) or genomic prediction, a complement to MAS for improving complicated quantitative characteristics like grain weight, shows promise in crops like rice (Ahmadi et al., 2021; Anilkumar et al., 2022a; Azharudheen et al., 2022).
Rapid advancements in genotyping platforms have facilitated the effective implementation of GS in field crops such as rice to accelerate genetic gain for quantitative traits (Meuwissen et al., 2001; Bernardo and Yu, 2007). The performance of an individual is predicted using genome-wide distributed markers that capture the impacts of major and minor alleles/genes contributing to the complex trait based on its genomic-estimated breeding values (GEBVs). Research on plant breeding has demonstrated that GS increases prediction accuracy compared to pedigree-based selection for traits with low to moderate heritability (Crossa et al., 2010, 2014; Hickey and Gorjanc, 2012; Pérez-Rodríguez et al., 2012; Xu et al., 2018). Hence, GS accounts for a greater proportion of the genetic variation of the trait as compared to MAS, where only major alleles associated with the trait are used for selection and introgression. With the implementation of GS, which saves resources and time, it is now possible to predict an individual’s genetic worth in an early generation by estimating additive effects for the targeted trait (Beyene et al., 2015; Vivek et al., 2017). In this connection, focusing on the prediction of genotypes using GS will also help in the identification of superior parents for hybridization and combining more desirable alleles for a complex trait. Broadly, two approaches in GS have been identified based on their utility. The first approach uses individuals as random effects, and uses genetic relationships among these individuals and a variance-covariance matrix from the predicted population to estimate the breeding values following an iterative method (Li et al., 2018). This is often called the direct method of predicting the breeding values of an individual. The second approach, which is also called the indirect approach, is where marker effects are estimated on a reference group and then the breeding values of individuals in the testing set are estimated based on the marker effect combined with genotype information from the testing set (Misztal and Legarra, 2017).
In order to improve the accuracy of prediction, several GS models have been developed and tested for suitability under different circumstances. Based on the broad statistical background of the models, two classes of models have been identified. First, parametric models such as genomic best linear unbiased prediction (GBLUP) (VanRaden, 2008), Bayesian models including Bayes A, Bayes B, and Bayes C (González-Recio and Forni, 2011), the partial least squares (PLS) model (Geladi and Kowalski, 1986), and the least absolute shrinkage and selection operator (LASSO) model (Tibshirani, 1996). The second class includes nonparametric models such as random forest approaches (Svetnik et al. 2003), neural network methods (NN) (Ehret et al., 2015), support vector machine models (SVM) (Maenhout et al., 2007), and a semi-nonparametric model, kernel Hilbert space regression (RKHS) (de los Campos et al., 2010). Further, these models are classified as regression-based and machine-learning-based models. The predictive efficiency of these models is tested by many researchers across different crops using empirical and simulation datasets (Riedelsheimer et al., 2012; Howard et al., 2014; Wang et al., 2015). Only a limited amount of literature is available for comparison of the prediction accuracy of these models in rice (Onogi et al., 2015; Spindel et al., 2015; Grenier et al., 2015; Dreisigacker et al., 2021). Some genomic prediction programs conducted on hybrid rice resulted in the identification of promising candidates based on genomic estimated breeding values (GEBVs) derived from different models (Spindel et al., 2015; Grenier et al., 2015). Thus, genomic prediction is the most reliable approach in rice breeding to achieve maximum genetic gain.
GS assumes at least one marker will always be in linkage disequilibrium with at least one of the genes or QTL associated with the target trait (Sorrells, 2015). Hence, using genome-wide dense markers for prediction is considered a prelude. Advances in sequencing technology reduced the genotyping cost, which allowed the abundant use of single nucleotide polymorphic (SNP) markers in different crops. However, considering genome-wide coverage of markers is preferred over marker type (Juliana et al., 2019), on the other hand, single sequence repeats (SSR) are considered highly informative and can retrieve more genome information with modest genome coverage (Vieira et al., 2016; Anilkumar et al., 2022c). Comparing the efficiency of retrieving genome information, a few hundred of SSR markers can perform on par with thousands of SNP markers (Vieira et al., 2016). If the SSR markers are derived from well-characterized gene sequences (candidate gene SSR or cgSSR), they can offer additional chances of increasing the accuracy of predictions (Molla et al., 2019). Nayak et al. (2022) reported the significance of using candidate gene derived SSR (cgSSR) markers for the identification of genomic regions for grain size traits in rice. Hence, exploiting genome-wide gene-derived markers is expected to increase the precision of predictions for quantitative traits (Singh et al., 2018).
In order to determine the effectiveness of genome-wide candidate gene-derived SSR markers for exploiting GS in rice breeding, a set of genome-wide cgSSR markers was deployed on a diverse population to evaluate the performance of different GS models for predicting rice grain weight. To effectively identify makers associated with grain weight, GWAS was performed using one single locus model and two multi-locus models. To enhance the accuracy of phenotype data, the population was evaluated over two seasons, and the best linear unbiased predictors across seasons were used for prediction purposes. The hypothesis of the study was to evaluate the effectiveness of new cgSSR markers for prediction-based rice breeding, and it was tested by performing a genome-wide association study and genomic predictions for grain weight using different models. Further, the influence of trait-associated markers on genomic prediction accuracy was also investigated. The results of this study have significant implications for practical rice breeding.
Materials and methods
Experimental genetic material
A total of 189 ‘indica’ rice genotypes that comprised 60 advanced breeding lines, 123 varieties released across different eastern and north-eastern states of India, and six germplasm accessions collected from eastern states of India were used in the study (Supplementary File S1). Many of these experimental genotypes are either released or targeted for cultivation in different ecologies, including irrigated, upland, and lowland rice-growing conditions. Seeds of released varieties and germplasm were collected from the ICAR-National Rice Research Institute gene bank, and advanced breeding materials from the Crop Improvement Division of ICAR- NRRI, Cuttack, India, were utilized. After collecting the experimental material, it was maintained by selecting true-to-type panicles from a single plant per genotype, similar to nucleus seed maintenance (Sahu et al. 2020), for two cropping seasons before the start of experimentation.
Experimental site and phenotyping
The experimental genotypes were planted in a randomized complete block design with three replications at the experimental plots of ICAR-NRRI, Cuttack, India (20.4537° N, 85.9338° E). The field experimentation was conducted initially during the wet season of 2020 and repeated during the wet season of 2021. Each genotype was planted in a 4 m2 area with 15 cm between plants and 20 cm between rows to establish the optimum crop stand. Appropriate agronomic practices were followed to raise a healthy crop. At maturity, the grains from each genotype were harvested separately, placed in cloth bags, and dried under the sun for two days. Further, the seeds were oven dried to reduce the moisture content to 12 ± 1 %, which is the recommended moisture content before storage for rice grains. Five random samples of one thousand grains were drawn from each genotype in each replication and weighed on a precision analytical balance to record the thousand-grain weight (TGW).
Candidate gene-based marker designing
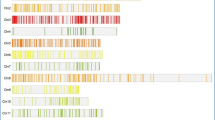
We selected genes after an extensive review of reports on candidate genes for yield-related traits and grain dimension traits identified and validated through RT-PCR, map-based cloning, or transgenic approaches. Every care was taken to cover all 12 rice chromosomes while considering the genes for marker development. Further, one round of confirmation on the functional characters of selected genes was done with the help of the OGRO database (Yamamoto et al., 2012) and Oryzabase (Kurata and Yamazaki, 2006). The gene sequences for selected candidate genes were retrieved from the rice genome annotation project database (Ouyang et al., 2007) (http://rice.plantbiology.msu.edu/) and the National Centre for Biotechnology Information (NCBI) gene bank database (https://www.ncbi.nlm.nih.gov/genbank/). The online SSR identification tool (Temnykh et al., 2001) was utilized to select potential repeat sequences to develop microsatellites from the gene sequences. Primer 3 software (Rozen and Skaletsky, 1999) was used to identify and pick the potentially polymorphic microsatellites from gene sequences. All the standard measures of primer length (20–25 bp), annealing temperature (55–60 °C), GC content (40–60), and the amplifiable size ranged between 100 bp and 300 bp were considered while designing the primers. The detailed procedure for developing cgSSR markers is shown in Supplementary Fig. S1. A total of 1200 cgSSR markers were derived from different genic sequences across all 12 rice chromosomes. A subset of these markers was tested in genome-wide association studies (GWAS) for plant-type characters (Azharudheen et al. (2022)) and grain characters (Nayak et al. 2022) and was found effective in the identification of significant QTL. From 1200 cgSSR markers, markers with proper amplification, polymorphism, and distribution on all chromosomes and bi-allelic on a subset of genotypes in the study population were considered for use in the present study. In the present study, a subset of 254 highly polymorphic markers distributed across all 12 chromosomes of rice (Supplementary Fig. S2) was used for GWAS and genomic prediction (GP) for thousand-grain weight.
Genomic DNA isolation and genotyping
The genomic DNA of all the genotypes in the population was isolated from the young leaves following the Cetyl Trimethyl Ammonium Bromide (CTAB) method (Doyle and Doyle, 1987). The quality of isolated DNA was tested using electrophoresis using 0.8% agarose and the quantity was measured using a Nanodrop spectrophotometer (Thermo Fisher Scientific, USA). A total of 307 SSR markers, including 254 cgSSR markers and 53 random SSR (rSSR) markers, were used for genotyping the population (Supplementary Fig. S2). The polymerase chain reaction mixture is constituted with 1 μL of each forward and reverse primer of the marker, 1 μL of 20 ng/μL genomic DNA, 4 μL standard premix, and 3 μL of nuclease free water to make up the final volume of 10 μL. Following initial denaturation at 94 °C for 4 minutes, 40 cycles of denaturation at 94 °C for 40 seconds, annealing temperature was adjusted depending on the primer for 40 seconds, primer elongation at 72 °C for 2 minutes, and final extension at 72 °C for 7 minutes, and genomic DNA was amplified using a 384 well thermocycler (Agilent technologies® Surecycler 8800). Amplicons were separated using 3.5% agarose electrophoresis and documented with Zenith (Gel.Pro CCD gel doc, Biozen Laboratories, India) documentation set up. Gel images were scored using CLIQS software (Totallab®) to reduce the manual scoring errors.
Statistical analysis
Phenotype analysis
The thousand grain weight recorded on five samples in each replication for each genotype was subjected to META-R software estimation of the best linear unbiased prediction (BLUP) (Alvarado et al., 2020). The BLUPs were estimated for both years separately and also across the years to shrink the phenotype data recorded over samples, replications, and years to minimize the experimentation error (Piepho et al., 2008). The BLUP values were considered for the estimation of descriptive statistics of the population with respect to grain weight using RStudio version 1.4.17. The measures of central tendency, along with third- and fourth- degree statistics, skewness, and kurtosis, were also estimated to ensure the best suitability of the population for further genome-wide association analysis and genomic prediction. The Shipiro-Wilk’s ‘p’ was estimated to make sure that the grain weight is normally distributed in the population.
Analysis of allele diversity and population structure
The allelic diversity, polymorphic information content, and allele frequency for cgSSR and rSSR markers in the study population were estimated using PowerMarker V3.25 (Liu and Muse, 2005). Population structure was assessed using four different approaches. First, principle component analysis (PCA) was performed to identify possible subgroups based on marker allelic compression using the ‘factoextra’ package in R software (Kassambara and Mundt, 2017). Second, STRUCTURE software (Pritchard et al., 2000) with 100000 burn-in periods was performed to identify subpopulations within the population. Subpopulations within the population were identified following ΔK value (Evanno et al., 2005) obtained by harvesting the STRUCTURE results in structure harvester, an online tool (Earl and vonHoldt, 2012). Third, genetic distance between genotypes obtained by the Neighbor-Joining method using DARwin software was depicted as an unrooted tree with clusters, developed using the iTOL software (Letunic and Bork, 2021). Fourth, genetic relatedness between individuals in the population was calculated following the VanRaden kinship algorithm (VanRaden, 2008) and represented as a familial relatedness heatmap using the GAPIT package in R software (Lipka et al., 2012).
GWAS and GP analysis
Genome-wide association analysis was performed between thousand grain weight BLUP values estimated across years and genome-wide marker information generated on the panel. Analysis was performed using the GAPIT package in the R software following three different models: (1) single locus mixed linear model (MLM), (2) multi-locus random mixed linear model (mrMLM) and (3) Fixed and random model Circulating Probability Unification (FarmCPU) model, were used to test the efficiency of new cgSSR markers in identifying associations between traits and markers. The MLM and FarmCPU models were accessed directly from GAPIT package, while mrMLM v4.0.2 was used to analyze mrMLM model (Zhang et al., 2020). Multi-locus models were used to overcome the limited efficiency of single-locus models (MLM) in detecting the marginal QTL effect in polygenic backgrounds (Wang et al., 2016). Multi-locus models scan the genome in multiple dimensions and simultaneously estimate marker effects to detect the association (Cui et al., 2018). The GAPIT package helps in the identification of marker-trait associations while simultaneously correcting the population structure and kinship of the population (Yu et al., 2006). Considering the importance of candidate gene-derived markers, threshold marker P < 0.05 (raw p-values instead of corrected p-values) in MLM and FarmCPU and, LOD value >3 in mrMLM was considered for the identification of significant marker-trait associations. Further, considering the multi-season evaluation data, to account for QTL-by-environment interactions, the three-variance multi-locus random effect mixed linear model (IIIVmrMLM) (Li et al., 2022a) was performed following the methodology described in (Li et al. 2022b).
Genomic prediction analysis was performed by developing seven regression-based models [Bayes A, Bayes B, Bayes C, Bayes ridge regression (BRR), LASSO, ridge regression BLUP (rrBLUP), and RKHS] and two machine learning models [support vector regression (SVR) and random forest regression (RFR)]. The differentiating features of these models are listed in Supplementary File S2. The basic model considered for developing these models is described as the following:
where, Y = vector of n observations, X = matrix of genotype information, β = vector of fixed effects, m = number of markers, Zk = vector of genotype indicator variable, γk = vector of marker effects, and ε = vector of residual error.
The genomic predictions were performed using the G2P package in R software (https://github.com/cma2015/G2P). The genomic predictions were obtained by following the 5-fold cross validation approach. The complete population was divided into 5 mutually exclusive subsets. Following the leave-one-out approach, one subset was used as a testing set and the remaining 4 subsets as training sets. With ten inclusive iterations, prediction values were obtained for all the genotypes in the population. Prediction accuracy was calculated following Pearson’s correlation coefficients between predicted and observed phenotype for all the models, and the model with highest prediction accuracy was used to calculate GEBVs of genotypes in the population for thousand grain weight. In parallel, prediction analysis was also performed by excluding markers identified as significantly associated with trait by different models in GWAS. This provides the effect of identified significant loci and also the effectiveness of associated markers on prediction accuracy.
Results
Phenotype variation
A wide range of significant grain weight variation was observed over different samples and across the years. During the wet season of 2020, the grain weight ranged between 11.06 g and 32.30 g, with a mean thousand grain weight of 23.20 g. During the wet season of 2021, the minimum grain weight was 11.70 g and the maximum was 32.25 g, with an average of 22.86 g. The grain weight across year BLUP values ranged between 11.67 g and 31.86 g, following an average of 23.18 g (Table 1). The average broad sense heritability coefficient of the trait across three datasets was considerably high (h2 = 0.78). There was however, no significant difference in grain weight, as indicated by a modest difference in sample variance and standard error estimated over the years. Skewness, a third-degree statistic, was negligible while the kurtosis, a fourth-degree statistic, was platykurtic for all the environments. Third and fourth degree statistics supported the quantitative inheritance of the trait involving minor genes with additive effects. The estimate of Shapiro-Wilk’s ‘P’ values was non-significant, indicating a normal distribution of grain weight variation in the population. Variation in grain weight in different type of genetic material in the population was compared and found a significant difference between material types, further, breeding lines and released varieties showed higher grain weight compared to germplasm accessions in the population. Similarly, grain weight variation was compared between genotypes grouped under different cultivation ecologies and no significant difference was noticed in mean grain weight (Supplementary Fig. S3). Material-wise differences in grain weight may be attributed to the intended selection of genotypes for higher grain weight in breeding material and released varieties, and negligible differences among genotypes of different ecologies indicate a lack of relationship between growing ecology and grain weight.
Allele diversity and population structure
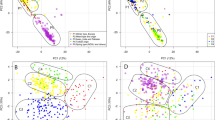
All 189 individuals in the population were genotyped with 254 cgSSR markers and 53 rSSR markers. A total of 905 alleles were amplified in the population, with a range of two to six alleles per marker. Major allele frequencies ranged between 0.37 and 0.99, and gene diversity explained by the markers ranged between 0.01 and 0.66. The informativeness of the markers was tested by estimating polymorphic information content (PIC), which is the function of allele frequency in the population. The PIC value estimated ranged between 0.01 and 0.59, and a PIC value >0.5 was considered significantly higher (Supplementary File S3). The parameters estimated for the population were compared between cgSSR and rSSR markers and are presented in Table 2. The comparison of PIC values and genetic diversity explained by two marker types is presented in Fig. 1. The cgSSRs were found more informative than rSSR based on average PIC value. Different approaches are used to delineate population structure in order to identify the presence of three subpopulations within the population. A total of three subpopulations were identified within the population by PCA estimate based grouping (Fig. 2A), STRUCTURE analysis (Fig. 2B), unrooted Neighbor-Joining cluster analysis (Fig. 2C), and genetic relatedness based heatmap analysis (Fig. 2D). The relatedness matrix developed with the VanRaden kinship algorithm depicted as a heatmap indicates low to moderate genetic relationships between individuals in the population.
A Principle component analysis grouped the panel to three clusters. B Structure analysis determined the presence of three subpopulations within the panel C Phylogenetic analysis classified the panel into three clusters and D Heatmap of relationship coefficients among individuals of the panel determined the presence of three subpopulations in the panel.
GWAS for grain weight
A total of 28 significant marker-trait associations (MTA) for across year BLUP values of grain weight were identified with three different models. The QTL identified by different methods were different; however, few were consistent between methods (Table 3 and Fig. 3). Only two MTAs were identified with a single locus model (MLM) alone, one on chromosome 1 and the other on chromosome 7. Together, MLM and mrMLM identified 12 significant MTAs on different chromosomes. Similarly, the multi-locus model, FarmCPU alone identified 12 significant MTAs on various chromosomes. However, all three models together identified a major QTL on chromosome 1 with 8.57% phenotypic variance explained (PVE) in MLM, 10.54% PVE in mrMLM and 11.53% PVE in FarmCPU models. The marker, YD91, associated with this QTL was derived from OsAAP3, an amino acid transporter gene important for higher grain yield located at 119391 bp on chromosome 1. Similarly, a QTL identified by RM5672 by two multi-locus methods explained higher phenotypic variance located at 6379987 bp on chromosome 7. Multi-environment joint GWAS analysis with 3VmrMLM was performed to detect QTL-by-Environment interactions. There were no QTL-by-environment interactions recorded in the present study (Fig. 3D). Among 28 QTL identified in the study, only 5 were identified by rSSR markers and remaining 23 QTL were by cgSSR markers.
In Manhattan plots, x-axis represents 12 rice chromosomes and –log(p) values on Y-axis indicates significant associations. Quantile-Quantile plots show deviation of observed –log(p) values from that of expected –log(p) values for significant marker trait associations. A Manhattan and Q-Q plots for FarmCPU, B Manhattan and Q-Q plots for mrMLM, C Manhattan and Q-Q plots for MLM models and D Manhattan plot for QTL-by-Environment interactions.
Genomic predictions
Genome-wide marker-based prediction for grain weight was performed with nine different models, which included regression-based parametric models and machine learning based non-parametric models. The predictions were performed following a five-fold cross validation approach with a leave-one-out strategy. Fold-wise cross validation predictions presented in Supplementary File S4. Among different models used to predict the phenotype, RFR, a machine learning model, produced the highest accuracy of prediction compared to other models, and the least accuracy of prediction was found in the LASSO model (Supplementary Fig. S4A and Fig. 4). Prediction accuracies were estimated by dropping GWAS significant markers and random markers to see the impact of marker number and corresponding markers effects. Prediction accuracy was drastically reduced upon excluding GWAS significant markers from the prediction analysis (Supplementary Fig. S4B and Fig. 4). The prediction accuracies with genome-wide markers were greater than 0.6, which is considered higher prediction accuracy for a quantitative trait. With the exclusion of significant markers from GWAS, prediction accuracy dropped significantly and reached below 0.5 in the case of the LASSO model. Even though, random SSR markers almost double the number of GWAS significant markers, dropping random SSR markers did not altered the prediction accuracies significantly (Fig. 4). Hence, proved the reduction of prediction accuracies after dropping GWAS significant marker was not merely due to marker numbers but due to marker effects. The GEBV values were estimated using the RFR model and compared with observed phenotypic values. A linear relationship between calculated GEBVs from the RFR model and observed phenotypes was observed (Supplementary Fig. S5). The GEBVs of genotypes differed significantly in different material types, and fewer differences were observed for the cultivation ecology based classification of genotypes (Fig. 5). The observed grain weights of genotypes with the top 10% GEBVs and the rest of the population were compared. The top 10% genotypes with higher GEBVs recorded significantly higher observed grain weights than the rest of the population.
Discussion
For improving a quantitative trait, it is important to understand the genetics and deploy it with a proper breeding strategy in the breeding program. Enhancing the genetic gain for a quantitative trait in the breeding material is an ultimate goal in crop improvement. Selecting genetic stocks that carry a higher number of positive alleles for a trait plays a significant role in achieving higher rewards in crop improvement. The genomic selection approach helps in selecting genotypes with higher genetic potential for a trait by estimating GEBV using genome-wide marker effects (Bernardo and Yu, 2007). Instead of using random genomic markers for GS, utilizing candidate gene derived markers for a quantitative trait increases the probability of selecting genetic stock with higher genetic potential for the trait (Sah et al., 2022). Hence, in the present study, the effects of new cgSSR markers derived from well characterized yield, yield-related traits, and grain morphology-related traits were evaluated for their utility for GWAS and genomic selection for grain weight in rice.
The cgSSR marker performed equally efficient in explaining population parameters as that of the rSSR markers used in the study. The number of alleles available in the population explained the abundance of genetic diversity in the population with limited gene outflow and this was in agreement with previous reports obtained using rSSR and RAPD markers (Rahman et al., 2007; Raju et al., 2016). The PIC values of the new cgSSR markers were slightly higher than those of rSSR markers, indicating their usefulness in QTL identification and genomic predictions (Fig. 1A). Apart from that, new cgSSR markers explained more genetic variation in the population than rSSR markers, suggesting their effectiveness in genetic analyses (Fig. 1B). The reports of Molla et al. (2019) and Nayak et al. (2022) support the use of cgSSR markers for genetic dissection of quantitative traits. On the other hand, these cgSSR markers along with rSSR markers explained the relatedness of individuals in the study population by delineating population structure. A total of three subpopulations were identified as having moderate genetic relatedness. The results were on par with the earlier reports (Zhang et al., 2013; Wang et al., 2014) in rice. Proper understanding of population structure is a prerequisite for the discovery of causative loci for quantitative traits and also for genomic prediction studies.
Among 28 significant marker-trait associations identified using three different approaches, a greater number of associations resulted from multi-locus models. Considering the importance of the markers used in genomic selection, the probability threshold of significant QTLs was set loosely in genome-wide association studies. Only a few significant MTAs were identified with very low explained phenotypic variance in single-locus method. Moreover, MTAs with higher PVE% were identified mostly using multi-locus models, indicating the efficiency of these models in association analysis. Among the 28 markers associated with thousand grain weight, only five markers were rSSR and the remaining 23 markers were cgSSRs derived from different yield-related genes and grain morphological trait-controlling genes in rice. These results suggest the efficiency of cgSSR markers in the identification of multiple causative loci for grain weight. One major QTL, qTGW1.4, identified by all three models with a maximum PVE of 11.53%, was associated with marker YD91 derived from the OsAAP3 gene on chromosome 1. The gene is reported to have a significant role in amino acid transporter activity that improves grain yield (Lu et al., 2018), and regulating arginine transporter activity that enhances grain yield and nitrogen use efficiency (Wei et al., 2021), respectively. Another major QTL, qTGW7.6, identified by two multi-locus models, was identified by rSSR marker RM5672 on chromosome 7. The allelic association of YD91 indicated 242 bp allele associated with higher grain weight and 300 bp allele associated with lower grain weight (Fig. 6A). Similarly, for marker RM5672, 175 bp alleles were associated with higher grain weight and 182 bp alleles with lower grain weight (Fig. 6B). Many of the cgSSR markers that established association with the TGW trait were derived from yield and grain related genes in rice. For instance, marker YD27, derived from the SMG1 gene, was identified to be associated with grain weight by two multi-locus models. The gene SMG1 is reported to influence the grain size in rice (Duan et al., 2014). The marker trait associations identified using cgSSR markers have significant scope in combining multiple causative alleles for grain weight in rice. These results also unraveled the efficacy of new cgSSR markers in the genetic dissection of a quantitative trait like grain weight in rice. The multi-environment joint GWAS analysis results no QTL-by-Environment interactions owing to evaluation of association of panel in the same location and same season over years, suggesting the low environmental variance and high narrow sense heritability of the phenotype data recorded. However, the size of association mapping population was only 189 and the number of markers was also limited in the present study. This may be one important reason for a few trait-marker associations, especially, the conclusion in this study should be further validated.
The marker YD91 identified to be associated with grain weight by all three models used in the study with 8.57–11.53% PVE and 242 bp allele of the marker was found to be associated with higher grain weigh whereas 300 bp allele with lower grain weight. Similarly, marker RM5672 found to be associated with grain weight as identified by two multi-locus models (mrMLM and FarmCPU) with 10.64 and 11.02% PVE. The 175 bp allele of the marker was more commonly associated with high grain weight, whereas 182 bp allele associated with relatively lower grain weight.
The predicted grain weight differed with different models of genomic selection suggesting the differences in predictive ability of various models. The differences in predictive accuracy of different models suggest the continuous improvement of genomic selection models over time to optimize the computational efficiency and accuracy of predictions (Robertsen et al., 2019). Many previous studies reported that no single model can be used for all quantitative traits; however, model optimization for specific traits is critical (Yu et al., 2022). From the present study, the RKHS model was found superior among regression-based models and the RFR model among machine learning models for prediction and selection of grain weight trait in rice. The prediction accuracies with repeat sequence (SSR/cgSSR) markers were on par with that obtained from SNP markers for grain weight in rice by Yu et al. (2022). Further, reduced prediction accuracies upon excluding GWAS significant markers evidenced the effectiveness of new cgSSR markers in trait identification and prediction-based breeding programs. Thus, trait-associated markers should be considered for genomic selection in plant breeding programs for higher prediction accuracy (He et al., 2019). The GEBVs estimated using the RFR model showed a linear relationship with observed grain weight, indicating the effectiveness of the model in predicting the performance of genotypes. Higher GEBVs of breeding lines followed by released varieties and lower GEBVs of germplasm accessions suggest the intended selection of genotypes for higher grain weight in breeding programs. Further, genotypes with top 10% GEBV values were found with higher observed grain weight compared to the rest of the population. This is not only evidence the accuracy of the prediction model but also suggest its utility in increasing genetic gain.
This investigation aimed to understand the effectiveness of candidate gene-derived markers for genomic selection in plant breeding programs. At the same time, we explored the appropriate model for the discovery of genomic regions for grain weight and their effect on the selection of candidates for future breeding through genome-wide marker effects. The superior performance of multi-locus models in the identification of all possible casual alleles for rice grain weight has significant scope in dissecting other quantitative traits. Similarly, superior prediction performance of machine learning models over Bayesian models eases the determination of complex interactions between predictors and responses by providing better outputs (Sandhu et al., 2021). For quantitatively inherited traits, Bayesian models might not be effective as several minor alleles contribute to their expression. Hence, multi-locus models for allele discovery and machine learning models with candidate gene-derived markers would ensure better rewards in genomic selection in plant breeding programs for improving quantitative traits.
Conclusion
A set of 307 markers, including 254 cgSSR and 53 rSSR markers, were used on a population of 189 genotypes to discover the causative alleles for thousand grain weight in rice. The results identified a total of 28 significant marker-trait associations following single and multi-locus GWAS approaches. Among the markers associated with the trait, 23 out of 28 markers were cgSSR markers, and only 5 rSSR markers were associated. The results suggested the effectiveness of the cgSSR marker in the discovery phase of breeding to identify genomic regions for quantitative traits. Further, genotypic information was utilized to predict the grain weight using genomic selection models. Among nine models used in the study, RFR, a machine learning model, was found to be highly effective with greater prediction accuracies for the trait. The results of this study demonstrated the optimization of the genomic selection model for grain weight could be used in genomic selection-based rice breeding programs. Apart from that, the candidate gene-based markers like cgSSRs were found to be more effective in genomic selection in plant breeding programs for better accuracy.
Data availability
The dataset used in the present study is deposited in Dryad, which can be accessed at https://doi.org/10.5061/dryad.931zcrjq6.
References
Ahmadi N, cao T-V, Frouin J, Norton GJ, Price AH (2021) Genomic prediction of arsenic tolerance and grain yield in rice: contribution of trait-specific markers and multi-environment models. Rice Sci 28:268–278. https://doi.org/10.1016/j.rsci.2021.04.006
Ahmar S, Gill RA, Jung K-H, Faheem A, Qasim MU, Mubeen M et al. (2020) Conventional and molecular techniques from simple breeding to speed breeding in crop plants: recent advances and future outlook. Int J Mol Sci 21:2590. https://doi.org/10.3390/ijms21072590
Alvarado G, Rodríguez FM, Pacheco A, Burgueño J, Crossa J, Vargas M et al. (2020) META-R: A software to analyze data from multi-environment plant breeding trials. Crop J 8:745–756. https://doi.org/10.1016/j.cj.2020.03.010
Anilkumar C, Sah RP, Muhammed ATP, Sunitha NC, Behera S, Marndi BC et al. (2022a) Genomic selection in rice: current status and future prospects. in: genomic selection in plants a guide for breeders, CRC Press, Taylor and Francis group, United States, pp 68–82
Anilkumar C, Sah RP, Muhammed Azharudheen TP, Behera S, Singh N, Prakash NR et al. (2022b) Understanding complex genetic architecture of rice grain weight through QTL-meta analysis and candidate gene identification. Sci Rep. 12:13832. https://doi.org/10.1038/s41598-022-17402-w
Anilkumar C, Sunitha NC, Devate NB, Ramesh S (2022c) Advances in integrated genomic selection for rapid genetic gain in crop improvement: a review. Planta 256:1–20. https://doi.org/10.1007/s00425-022-03996-y
Azharudheen M, Molla K, Anilkumar C, Sah R (2022) Advanced technologies for climate-smart breeding. In: Bhattacharyya, P., Chakraborty, K., Molla, K. A., Poonam, A., Bhaduri, D., Sah, R. P., Paul, S., Hanjagi, P. S., Basana-Gowda, G. and Swain, P. (Eds.) (2022). Climate Resilient Technologies for Rice based Production Systems in Eastern India. ICAR-National Rice Research Institute, Cuttack, Odisha, India, pp 408
Azharudheen M, Nayak A, Behera S, Anilkumar C, Marndi BC, Moharana D et al. (2022) Genome-wide association analysis for plant type characters and yield using cgSSR markers in rice (Oryza sativa L.). Euphytica 218:1–13. https://doi.org/10.1007/s10681-022-03021-z
Bernardo R, Yu J (2007) Prospects for genomewide selection for quantitative traits in maize. Crop Sci 47:1082–1090. https://doi.org/10.2135/cropsci2006.11.0690
Beyene Y, Semagn K, Mugo S, Tarekegne A, Babu R, Meisel B et al. (2015) Genetic gains in grain yield through genomic selection in eight bi-parental maize populations under drought stress. Crop Sci 55:154–163. https://doi.org/10.2135/cropsci2014.07.0460
Chakraborti M, Anilkumar C, Verma R, Fiyaz RA, Raj R, Patra B et al. (2021) Rice breeding in India: eight decades of journey towards enhancing the genetic gain for yield, nutritional quality, and commodity value. ORYZA- Int J Rice 58:69–88. https://doi.org/10.35709/ory.2021.58.spl.2
Chen K, Łyskowski A, Jaremko Ł, Jaremko M (2021) Genetic and molecular factors determining grain weight in rice. Front Plant Sci 12:605799. https://doi.org/10.3389/fpls.2021.605799
Crossa J, Campos Gdelos, Pérez P, Gianola D, Burgueño J, Araus JL et al. (2010) Prediction of genetic values of quantitative traits in plant breeding using pedigree and molecular markers. Genetics 186:713–724. https://doi.org/10.1534/genetics.110.118521
Crossa J, Pérez P, Hickey J, Burgueño J, Ornella L, Cerón-Rojas J et al. (2014) Genomic prediction in CIMMYT maize and wheat breeding programs. Heredity 112:48–60. https://doi.org/10.1038/hdy.2013.16
Cui Y, Zhang F, Zhou Y (2018) The application of multi-locus GWAS for the detection of salt-tolerance loci in rice. Front Plant Sci 9:1464. https://doi.org/10.3389/fpls.2018.01464
de los Campos G, Gianola D, Allison DB (2010) Predicting genetic predisposition in humans: the promise of whole-genome markers. Nat Rev Genet 11:880–886. https://doi.org/10.1038/nrg2898
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15. (Eds.)
Dreisigacker S, Crossa J, Pérez-Rodríguez P, Montesinos-L OA, Mondal S, Crespo-Herrera L et al. (2021) Implementation of genomic selection in the CIMMYT Global Wheat Program, findings from the past 10 years. Crop Breed 3:e210005. https://doi.org/10.20900/cbgg20210005
Duan P, Rao Y, Zeng D, Yang Y, Xu R, Zhang B et al. (2014) SMALL GRAIN 1, which encodes a mitogen-activated protein kinase kinase 4, influences grain size in rice. Plant J 77:547–557. https://doi.org/10.1111/tpj.12405
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Ehret A, Hochstuhl D, Gianola D, Thaller G (2015) Application of neural networks with back-propagation to genome-enabled prediction of complex traits in Holstein-Friesian and German Fleckvieh cattle. Genet Sel Evol 47:22. https://doi.org/10.1186/s12711-015-0097-5
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Feng B, Chen K, Cui Y, Wu Z, Zheng T, Zhu Y et al. (2018) Genetic dissection and simultaneous improvement of drought and low nitrogen tolerances by designed QTL pyramiding in rice. Front Plant Sci 9:306. https://doi.org/10.3389/fpls.2018.00306
Gao F, Zeng L, Qiu L, Lu X, Ren J, Wu X et al. (2016) QTL mapping of grain appearance quality traits and grain weight using a recombinant inbred population in rice (Oryza sativa L.). J Integr Agric 15:1693–1702. https://doi.org/10.1016/S2095-3119(15)61259-X
Geladi P, Kowalski BR (1986) Partial least-squares regression: a tutorial. Anal Chim Acta 185:1–17. https://doi.org/10.1016/0003-2670(86)80028-9
González-Recio O, Forni S (2011) Genome-wide prediction of discrete traits using bayesian regressions and machine learning. Genet Sel Evol 43:7. https://doi.org/10.1186/1297-9686-43-7
Grenier C, Cao T-V, Ospina Y, Quintero C, Châtel MH, Tohme J et al. (2015) Accuracy of genomic selection in a rice synthetic population developed for recurrent selection breeding. PLOS ONE 10:e0136594. https://doi.org/10.1371/journal.pone.0136594
He L, Xiao J, Rashid KY, Jia G, Li P, Yao Z, Wang X, Cloutier S, You FM (2019) Evaluation of genomic prediction for pasmo resistance in flax. Int J Mol Sci 20(2):359. https://doi.org/10.3390/ijms20020359
Hickey JM, Gorjanc G (2012) Simulated data for genomic selection and genome-wide association studies using a combination of coalescent and gene drop methods. G3 Genes|Genomes|Genet 2:425–427. https://doi.org/10.1534/g3.111.001297
Howard R, Carriquiry AL, Beavis WD (2014) Parametric and nonparametric statistical methods for genomic selection of traits with additive and epistatic genetic architectures. G3 Genes|Genomes|Genet 4:1027–1046. https://doi.org/10.1534/g3.114.010298
Huang R, Jiang L, Zheng J, Wang T, Wang H, Huang Y et al. (2013) Genetic bases of rice grain shape: so many genes, so little known. Trends Plant Sci 18:218–226. https://doi.org/10.1016/j.tplants.2012.11.001
Juliana P, Montesinos-López OA, Crossa J, Mondal S, González Pérez L, Poland J et al. (2019) Integrating genomic-enabled prediction and high-throughput phenotyping in breeding for climate-resilient bread wheat. Theor Appl Genet 132:177–194. https://doi.org/10.1007/s00122-018-3206-3
Kassambara A, Mundt F (2017) Factoextra: extract and visualize the results of multivariate data analyses. R Package Version 1:337–354
Katara J, Chidambaranathan P, Devanna BN, Verma R, Anilkumar C, Patra B et al. (2021) Genomics assisted breeding: The need and current perspective for rice improvement in India. Oryza- Int J Rice 58:61–68. https://doi.org/10.35709/ory.2021.58.spl.1
Kurata N, Yamazaki Y (2006) Oryzabase. An Integrated biological and genome information database for rice. Plant Physiol 140:12–17. https://doi.org/10.1104/pp.105.063008
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296. https://doi.org/10.1093/nar/gkab301
Li B, Zhang N, Wang Y-G, George AW, Reverter A, Li Y (2018) Genomic prediction of breeding values using a subset of SNPs identified by three machine learning methods. Front Genet 9:237. https://doi.org/10.3389/fgene.2018.00237
Li G, Tang J, Zheng J, Chu C (2021) Exploration of rice yield potential: Decoding agronomic and physiological traits. Crop J 9:577–589. https://doi.org/10.1016/j.cj.2021.03.014
Li M, Zhang YW, Zhang ZC, Xiang Y, Liu MH, Zhou YH, Zuo JF, Zhang HQ, Chen Y, Zhang YM (2022a) A compressed variance component mixed model for detecting QTNs and QTN-by-environment and QTN-by-QTN interactions in genome-wide association studies. Mol Plant 15:630–650. https://doi.org/10.1016/j.molp.2022.02.012
Li M, Zhang YW, Xiang Y, Liu MH, Zhang YM (2022b) IIIVmrMLM: The R and C++ tools associated with 3VmrMLM, a comprehensive GWAS method for dissecting quantitative traits. Mol Plant 15:1251–1253. https://doi.org/10.1016/j.molp.2022.06.002
Lipka AE, Tian F, Wang Q, Peiffer J, Li M, Bradbury PJ et al. (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399. https://doi.org/10.1093/bioinformatics/bts444
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Lu K, Wu B, Wang J, Zhu W, Nie H, Qian J et al. (2018) Blocking amino acid transporter OsAAP3 improves grain yield by promoting outgrowth buds and increasing tiller number in rice. Plant Biotechnol J 16:1710–1722. https://doi.org/10.1111/pbi.12907
Maenhout S, De Baets B, Haesaert G, Van Bockstaele E (2007) Support vector machine regression for the prediction of maize hybrid performance. Theor Appl Genet 115:1003–1013. https://doi.org/10.1007/s00122-007-0627-9
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829. https://doi.org/10.1093/genetics/157.4.1819
Misztal I, Legarra A (2017) Invited review: efficient computation strategies in genomic selection. Animal 11:731–736. https://doi.org/10.1017/S1751731116002366
Molla KA, Azharudheen TPM, Ray S, Sarkar S, Swain A, Chakraborti M et al. (2019) Novel biotic stress responsive candidate gene based SSR (cgSSR) markers from rice. Euphytica 215:17. https://doi.org/10.1007/s10681-018-2329-6
Nayak AK, Anilkumar C, Behera S, Rameswar Prasad Sah, Lavanya GR, Kumar A et al. (2022) Genetic dissection of grain size traits through genome-wide association study based on genic markers in rice. Rice Sci 29:462
Onogi A, Ideta O, Inoshita Y, Ebana K, Yoshioka T, Yamasaki M et al. (2015) Exploring the areas of applicability of whole-genome prediction methods for Asian rice (Oryza sativa L.). Theor Appl Genet 128:41–53. https://doi.org/10.1007/s00122-014-2411-y
Ouyang S, Zhu W, Hamilton J, Lin H, Campbell M, Childs K et al. (2007) The TIGR Rice Genome Annotation Resource: improvements and new features. Nucleic Acids Res 35:D883–D887. https://doi.org/10.1093/nar/gkl976
Pérez-Rodríguez P, Gianola D, González-Camacho JM, Crossa J, Manès Y, Dreisigacker S (2012) Comparison between linear and non-parametric regression models for genome-enabled prediction in wheat. G3 Genes|Genomes|Genet 2:1595–1605. https://doi.org/10.1534/g3.112.003665
Piepho HP, Möhring J, Melchinger AE, Büchse A (2008) BLUP for phenotypic selection in plant breeding and variety testing. Euphytica 161:209–228. https://doi.org/10.1007/s10681-007-9449-8
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Rahman SN, Islam MS, Alam MS, Nasiruddin KM (2007) Genetic polymorphism in rice (Oryza sativa L.) through RAPD analysis. Indian J Biotechnol 6:224–229
Raju BR, Mohankumar MV, Sumanth KK, Rajanna MP, Udayakumar M, Prasad TG et al. (2016) Discovery of QTLs for water mining and water use efficiency traits in rice under water-limited condition through association mapping. Mol Breed 36:35. https://doi.org/10.1007/s11032-016-0457-z
Riedelsheimer C, Technow F, Melchinger AE (2012) Comparison of whole-genome prediction models for traits with contrasting genetic architecture in a diversity panel of maize inbred lines. BMC Genom 13:452. https://doi.org/10.1186/1471-2164-13-452
Robertsen CD, Hjortshøj RL, Janss LL (2019) Genomic selection in cereal breeding. Agronomy 9:95. https://doi.org/10.3390/agronomy9020095
Rozen S, Skaletsky H (1999) Primer3 on the WWW for general users and for biologist programmers. In: Misener S, Krawetz SA (eds) Bioinformatics Methods and Protocols, Methods in Molecular Biology TM. Humana Press: Totowa, NJ, pp 365–386
Sah RP, Behera S, Dash SK, Azharudheen TPM, Meher J, Kumar A et al. (2022) Unravelling genetic architecture and development of core set from elite rice lines using yield-related candidate gene markers. Physiol Mol Biol Plants 28:1217–1232. https://doi.org/10.1007/s12298-022-01190-8
Sahu RK, Patnaik S, Sah RP (2020) Quality seed production in rice. In: NRRI Research Bulletin No. 27, ICAR-National Rice Research Institute, Cuttack 753006, Odisha, India, p 58
Sandhu K, Patil SS, Pumphrey M, Carter A (2021) Multitrait machine- and deep-learning models for genomic selection using spectral information in a wheat breeding program. Plant Genome 14:e20119. https://doi.org/10.1002/tpg2.20119
Singh AK, Chaurasia S, Kumar S, Singh R, Kumari J, Yadav MC et al. (2018) Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat. BMC Plant Biol 18:249. https://doi.org/10.1186/s12870-018-1476-1
Sorrells ME (2015) Genomic selection in plants: empirical results and implications for wheat breeding. In: Advances in wheat genetics: from genome to field, Springer, Tokyo, pp 401–409
Spindel J, Begum H, Akdemir D, Virk P, Collard B, Redoña E et al. (2015) Genomic Selection and Association Mapping in Rice (Oryza sativa): Effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines. PLOS Genet 11:e1004982. https://doi.org/10.1371/journal.pgen.1004982
Svetnik V, Liaw A, Tong C, Culberson JC, Sheridan RP, Feuston BP (2003) Random forest: a classification and regression tool for compound classification and QSAR modeling. J Chem Inf Comput Sci 43:1947–1958. https://doi.org/10.1021/ci034160g
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch S (2001) Computational and experimental analysis of microsatellites in Rice (Oryza sativa L.): Frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452. https://doi.org/10.1101/gr.184001
Tibshirani R (1996) Regression shrinkage and selection via the Lasso. J R Stat Soc: Ser B (Methodol) 58:267–288. https://doi.org/10.1111/j.2517-6161.1996.tb02080.x
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423. https://doi.org/10.3168/jds.2007-0980
Vieira MLC, Santini L, Diniz AL, Munhoz C, de F (2016) Microsatellite markers: what they mean and why they are so useful. Genet Mol Biol 39:312–328. https://doi.org/10.1590/1678-4685-GMB-2016-0027
Vivek BS, Krishna GK, Vengadessan V, Babu R, Zaidi PH, Kha LQ et al. (2017) Use of genomic estimated breeding values results in rapid genetic gains for drought tolerance in maize. Plant Genome 10: plantgenome2016 07:0070. https://doi.org/10.3835/plantgenome2016.07.0070
Wang C, Yang Y, Yuan X, Xu Q, Feng Y, Yu H et al. (2014) Genome-wide association study of blast resistance in indica rice. BMC Plant Biol 14:311. https://doi.org/10.1186/s12870-014-0311-6
Wang S-B, Feng J-Y, Ren W-L, Huang B, Zhou L, Wen Y-J et al. (2016) Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci Rep. 6:19444. https://doi.org/10.1038/srep19444
Wang X, Yang Z, Xu C (2015) A comparison of genomic selection methods for breeding value prediction. Sci Bull 60:925–935. https://doi.org/10.1007/s11434-015-0791-2
Wei Q, Yan Z, Xiong Y, Fang Z (2021) Altered expression of OsAAP3 influences rice lesion mimic and leaf senescence by regulating arginine transport and nitric oxide pathway. Int J Mol Sci 22:2181. https://doi.org/10.3390/ijms22042181
Weng J, Gu S, Wan X, Gao H, Guo T, Su N et al. (2008) Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res 18:1199–1209. https://doi.org/10.1038/cr.2008.307
Xu S, Zhu D, Zhang Q (2014) Predicting hybrid performance in rice using genomic best linear unbiased prediction. Proc Natl Acad Sci USA 111:12456–12461. https://doi.org/10.1073/pnas.1413750111
Xu Y, Wang X, Ding X, Zheng X, Yang Z, Xu C et al. (2018) Genomic selection of agronomic traits in hybrid rice using an NCII population. Rice 11:32. https://doi.org/10.1186/s12284-018-0223-4
Yamamoto E, Yonemaru J, Yamamoto T, Yano M (2012) OGRO: The Overview of functionally characterized Genes in Rice online database. Rice 5:26. https://doi.org/10.1186/1939-8433-5-26
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF et al. (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208. https://doi.org/10.1038/ng1702
Yu P, Ye C, Li L, Yin H, Zhao J, Wang Y et al. (2022) Genome-wide association study and genomic prediction for yield and grain quality traits of hybrid rice. Mol Breed 42:16. https://doi.org/10.1007/s11032-022-01289-6
Zhang D, Zhang H, Qi Y, Wang M, Sun J, Ding L et al. (2013) Genetic structure and eco-geographical differentiation of cultivated Hsien rice (Oryza sativa L. subsp. indica) in China revealed by microsatellites. Chin Sci Bull 58:344–352. https://doi.org/10.1007/s11434-012-5396-4
Zhang H-W, Fan Y-Y, Zhu Y-J, Chen J-Y, Yu S-B, Zhuang J-Y (2016) Dissection of the qTGW1.1 region into two tightly-linked minor QTLs having stable effects for grain weight in rice. BMC Genet 17:98. https://doi.org/10.1186/s12863-016-0410-5
Zhang YW, Tamba CL, Wen YJ, Li P, Ren WL, Ni YL, Gao J, Zhang YM (2020) mrMLM v4. 0.2: an R platform for multi-locus genome-wide association studies. Genom Proteom Bioinforma 18:481–487. https://doi.org/10.1016/j.gpb.2020.06.006
Acknowledgements
Authors are thankful to ICAR-National Rice Research Institute for financial support to complete this research.
Author information
Authors and Affiliations
Contributions
CA: Conceptualization, methodology, formal analysis, writing original draft. MA: Developed new cgSSR markers, methodology. RS: methodology, genomic resources, review & editing. NS: Data curation and formal analysis. BD: Tabulation, review & editing. BM: Resources, project administration. BP: Resources, writing-review & editing
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor Yuan-Ming Zhang.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Anilkumar, C., Muhammed Azharudheen, T.P., Sah, R.P. et al. Gene based markers improve precision of genome-wide association studies and accuracy of genomic predictions in rice breeding. Heredity 130, 335–345 (2023). https://doi.org/10.1038/s41437-023-00599-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-023-00599-5
This article is cited by
-
Genetic dissection of green pod yield in dolichos bean, an orphan vegetable legume, using new molecular markers
Journal of Applied Genetics (2024)