Abstract
Population genetic studies of symbiotic anthozoans have been historically challenging because their endosymbioses with dinoflagellates have impeded marker development. Genomic approaches like reduced representation sequencing alleviate marker development issues but produce anonymous loci, and without a reference genome, it is unknown which organism is contributing to the observed patterns. Alternative methods such as bait-capture sequencing targeting Ultra-Conserved Elements are now possible but costly. Thus, RADseq remains attractive, but how useful are these methods for symbiotic anthozoan taxa without a reference genome to separate anthozoan from algal sequences? We explore this through a case-study using a double-digest RADseq dataset for the sea anemone Bartholomea annulata. We assembled a holobiont dataset (3854 loci) for 101 individuals, then used a reference genome to create an aposymbiotic dataset (1402 loci). For both datasets, we investigated population structure and used coalescent simulations to estimate demography and population parameters. We demonstrate complete overlap in the spatial patterns of genetic diversity, demographic histories, and population parameter estimates for holobiont and aposymbiotic datasets. We hypothesize that the unique combination of anthozoan biology, diversity of the endosymbionts, and the manner in which assembly programs identify orthologous loci alleviates the need for reference genomes in some circumstances. We explore this hypothesis by assembling an additional 21 datasets using the assembly programs pyRAD and Stacks. We conclude that RADseq methods are more tractable for symbiotic anthozoans without reference genomes than previously realized.
Similar content being viewed by others
Introduction
The study of intraspecific genetic diversity across broad geographic space (i.e., population genetics and phylogeography) can shed light on the historical and contemporary processes responsible for the generation and maintenance of biodiversity within species and ecosystems (Arbogast 2001; Avise 2009; Knowles 2009; Avise et al. 2016). Intraspecific genetic surveys demarcate barriers and avenues of dispersal, identify cryptic species, and, with increasing dataset sizes and statistical approaches, estimate important demographic parameters such as effective population size, divergence time, migration rates, and historical changes in population size among others (e.g., Knowles 2009; Carstens et al. 2013; Pelletier and Carstens 2014; Pante et al. 2015; Avise et al. 2016; Smith et al. 2017).
Although population genetics and phylogeography have a long history in marine systems (e.g., Reeb and Avise 1990; Bowen et al. 1992, 1994), cnidarians in the class Anthozoa (i.e., stony corals, sea anemones, soft corals, zoanthids, and corallimorpharians), which form the backbone of coral reefs and a major component of its biodiversity, have been historically challenging to work with at the population genetic level. In addition to large range sizes and the logistical difficulties of sampling underwater, mitochondrial DNA barcodes (mtDNA), the molecular marker of choice for metazoan population genetic studies since the advent of Sanger sequencing, evolve too slowly in most anthozoans to be useful for intraspecific studies (e.g., Shearer et al. 2002; Daly et al. 2010; Allio et al. 2017). Further, the overwhelming majority of tropical anthozoans found on coral reefs form endosymbioses with photosynthetic dinoflagellates in the family Symbiodinaceae, which allows these animals to thrive in oligotrophic habitats (Muscatine et al. 1981; Rowan and Powers 1991; Gates and Edmunds 1999; Baker 2003; Santos 2016). In field-collected samples, contamination from symbiodiniaceans is unavoidable and resulting DNA extractions harbor a mix of anthozoan, dinoflagellate, and other microorganism DNA (termed “holobiont” DNA). The combination of slowly evolving mtDNA and dinoflagellate contamination complicated the development of molecular markers suitable for population level questions (Shearer et al. 2005). Until the recent development of bait-capture probes targeting Ultra Conserved Element (UCE) and exon loci were developed for Anthozoa (Quattrini et al. 2018; Cowman et al. 2020; Erickson et al. 2020; McFadden et al. 2021), no universal molecular markers suitable for shallow evolutionary questions existed, although their development had remained a goal (e.g., Reitzel et al. 2013; Brugler et al. 2018). Thus, most intraspecific genetic studies of tropical anthozoans relied on species-specific microsatellite loci to make population-level inferences (e.g., Baums et al. 2005, Andras et al. 2013; Foster et al. 2012; Rippe et al. 2017; Titus et al. 2017a; Nakabayashi et al. 2019).
At present, bait-capture sequencing remains expensive and only recently tested at the population level for anthozoans (Erickson et al. 2020; Glon et al. 2021; Grinblat et al. 2021). Species-specific microsatellite markers remain logistically burdensome to develop and provide fewer independent loci. An alternative approach is reduced representation sequencing (e.g., RADseq, ddRADseq, GBS, 2bRAD, ezRAD), which can generate data from thousands of single nucleotide polymorphisms (SNPs) and unlinked loci, and provide genome-scale data for non-model organisms, alleviating many of the issues with marker development that have stunted the study of anthozoans and other marine invertebrates (reviewed by Reitzel et al. 2013). In other systems, RADseq approaches have been particularly powerful at the population level, allowing for greater statistical and explanatory power into complex evolutionary and demographic histories (e.g., Carstens et al. 2012; Excoffier et al. 2013; McCormack et al. 2013). However, marine scientists studying symbiotic anthozoans must still contend with symbiodiniacean contamination in genomic sequence data because there are no simple or reliable ways to completely separate symbiont and host DNA before sequencing. Post-sequencing bioinformatic approaches are the most accessible avenue to separate symbiont from host DNA sequences and create anthozoan-only datasets, but these create additional hurdles that are functionally insurmountable for most taxa.
For population genetic studies using transcriptomic approaches, anthozoan and dinoflagellate DNA can be parsed bioinformatically with relative ease, as the assembled transcripts are long and conserved enough to map to published genomic resources (e.g., Davies et al. 2016; Kenkel and Matz 2017; Kenkel et al. 2018). However, the reduced representation sequencing approaches most commonly used in population-level studies produce short (50–100 bp), anonymous loci that are expected to be recovered largely from non-coding regions. Currently available anthozoan reference genomes are thus of limited use to separate dinoflagellate from anthozoan loci bioinformatically unless the reference species is closely related to the focal taxa (e.g., congeneric, conspecific). Likewise, the currently available genomic resources for Symbiodinaceae also are of limited use for subtracting symbiont sequences from reduced representation datasets because of the genetic diversity within its members: long considered to belong to a single genus (Symbiodinium), these photosymbiotic dinoflagellates are now recognized to represent 7–15 genus-level lineages (LaJeunesse et al. 2018). Diversification within Symbiodinaceae began in the mid-Jurassic (~165 mya) and genetic distances between many genera are on par with order-level divergences in other dinoflagellate taxa (Rowan and Powers 1992; Santos 2016; LaJeunesse et al. 2018). Thus, like anthozoan reference genomes, any symbiodiniacean reference genome used to disambiguate endosymbiont and host DNA would also need to be very closely related to the specific dinoflagellate found within the focal anthozoan species to effectively identify dinoflagellate sequences within reduced representation datasets.
These issues have long been recognized, and researchers employing reduced representation sequencing approaches for symbiotic anthozoans that do not have access to conspecific reference genomes have gone to great lengths to eliminate endosymbionts from their resulting datasets. Some have mapped anonymous loci to conspecific or congeneric transcriptomes and used only the resulting protein-coding datasets for interspecific phylogenetic reconstruction and hybridization studies among closely related taxa (Combosh and Vollmer 2015; Forsman 2017; Johnston et al. 2017; Iguchi et al. 2019). Bongearts et al. (2017) used pre- and post-sequencing methods and employed a subtraction library approach, spinning down homogenized tissue in an effort to remove dinoflagellate cells prior to DNA extraction and creating a separate reduced representation dinoflagellate reference library. Porro et al. (2020) compared the total yield of RADseq loci and genetic clustering results from symbiont free epidermal tissue with symbiont-rich tentacle tissue in the snakelocks anemone Anemonia viridis. Others, such as Leydet et al. (2018), targeted anthozoan RADseq loci by including a congeneric, aposymbiotic, species in their library prep and sequencing to act as a de facto reference library. Regardless of approach, each of these studies recognized the importance of removing symbiodiniacean sequences from their reduced representation datasets, acknowledging that successful interpretation of patterns or population parameters requires knowing the extent to which each organism is contributing to the observed patterns.
While contamination represents a problem in theory, in practice, it remains unclear how important it is to account for and remove 100% of endosymbiotic dinoflagellate loci from reduced representation datasets. Are reference genomes or other complicated pre- and post-sequencing approaches always required to obtain anthozoan-only datasets that lead to robust population genomic inferences? We explore this question through a case study of previously published double-digest restriction-site associated DNA (ddRADseq) sequence data for the corkscrew sea anemone Bartholomea annulata (see Titus et al. 2019a), a common and ecologically important member of coral reef communities throughout the Tropical Western Atlantic (Huebner and Chadwick 2012; O’Reilly and Chadwick 2017, O’Reilly et al. 2018; Titus and Daly 2017; Titus et al. 2017a, 2017b, 2017c). Similar to what had been done for coral species with reference genomes (Shinzato et al. 2011, 2015; Devlin-Durante and Baums 2017; Drury et al. 2017; Rosser et al. 2017; Cunning et al. 2018; van Oppen et al. 2018), Titus et al. (2019a) used the genome of the sea anemone Exaiptasia diaphana (formerly Aiptasia pallida; see Grajales and Rodriguez 2014; Baumgarten et al. 2015; ICZN 2017), a closely related species from the same family (Aiptasiidae; see Grajales and Rodríguez 2016), to create an aposymbiotic ddRADseq dataset. Here we explore the necessity and impact of screening reads from photosymbionts by comparing the aposymbiotic population-level data of Titus et al. (2019a) with the raw, unfiltered, holobiont ddRADseq data from which it was derived. We compare the spatial genetic structure of the aposymbiotic and holobiont datasets throughout the region and use coalescent simulation and model selection to understand whether the two datasets can be interpreted as having the same demographic histories and parameter estimates (i.e., effective population size, migration rate, divergence times). We discuss the alternative population genomic reconstructions and their implications for future studies on symbiotic anthozoans.
Methods
Dataset assembly
We obtained raw ddRAD sequence data, produced by Titus et al. (2019a), from NCBI Sequence Read Archive (SRA) BioProject ID PRJNA542967. These data represent 123 individuals of Bartholomea annulata collected throughout their known geographic range in the Tropical Western Atlantic, and from localities separated by known phylogeographic barriers (Fig. 1a; Supplementary Tables S1 and S2; reviewed by DeBiasse et al. 2016). Sample collection, processing, ddRADseq library preparation, and sequencing are detailed by Titus et al. (2019a) and are broadly representative of other ddRADseq protocols used for anthozoans, including the use of enzymes (Psti-HF and EcoRI-HF), size selection (400–800 bp), sequencing platform (Illumina HiSeq 2500), PCR cycles (18 cycles), maximum number of shared polymorphic sites in a locus to detect potential paralogs (max_shared_Hs_locus = 0.5), and sequence length (single-end 100 bp sequencing). Demultiplexed raw sequence data were downloaded from NCBI and assembled de novo using the program pyRAD v3.0.66 (Eaton 2014) because of its broad flexibility to adjust parameter settings and ability to handle indels. Briefly, we set the clustering threshold (Wclust) to 0.90 to assemble reads into loci and required a minimum coverage depth of seven to call a locus (Mindepth). We required a locus to be present in 75% of all individuals, allowing some missing data in our final dataset. Finally, individuals with low sequencing coverage (<500,000 reads; N = 22 individuals), were removed from the dataset resulting in a final dataset of N = 101 individuals (Supplementary Tables S1 and S2).
A Map of sampling localities throughout the Tropical Western Atlantic. PA Bocas del Toro, Panama, CC Cayos Cochinos, Honduras, UT Utila, Honduras, MX Mahahual, Mexico, FT Ft. Lauderdale, Florida, UK Upper Keys, Florida, MK Middle Keys, Florida, LK Lower Keys, Florida, BH Eleuthera, Bahamas, SAN San Salvador, Bahamas, ST St. Thomas, US Virgin Islands, BR Barbados, CU Curacao, BD Bermuda. Red dashed lines denote previously recovered major phylogeographic breaks in the region. B Genetic clustering results (K = 2) for the aposymbiotic Bartholomea annulata RADseq dataset. C Genetic clustering results (K = 2) for the holobiont Bartholomea annulata RADseq dataset. D Genetic clustering results (K = 2) for the unmapped Bartholomea annulata RADseq dataset. Similarity metric calculated by pong represents the average pairwise percentage of individuals with identical cluster assignment probabilities across aposymbiotic, holobiont, and unmapped datasets.
After running pyRAD to completion, the remaining loci represented our “holobiont” dataset, sequences that were, putatively, an unknown combination of anemone and endosymbiont DNA. We then re-created the anemone-only “aposymbiotic” dataset from Titus et al. (2019a) by mapping polymorphic loci from the holobiont dataset to the genome of the closely related Exaiptasia diaphana (see Baumgarten et al. 2015) to identify anemone-only sequences. Mapping of loci was conducted using BLAST and an 85% sequence similarity threshold after Titus et al. (2019a) to allow for substitutions because we were not using a conspecific reference. All remaining loci that did not map to the Exaiptasia genome were put in a third “unmapped” dataset. As a final data check, we created local BLAST databases by downloading publicly available endosymbiotic dinoflagellate genomes: Symbiodinium micradriaticum (Aranda et al. 2016 as Symbiodinium micradriaticum “Clade A”), Breviolum minutum (Shoguchi et al. 2013 as Symbiodinium minutum “Clade B”), Cladocopium goreaui (Liu et al. 2018, as Symbiodinium goreaui “Clade C”), and Fugacium kawagutii (Liu et al. 2018, as Symbiodinium kawagutii “Clade F”). Using BLAST, we mapped both our holobiont and apoysymbiotic datasets to the symbiodiniacean genomes to see if we could identify any symbiodiniacean sequences in the holobiont data and confirm that no loci in our aposymbiotic dataset mapped to both symbiodiniacean and Exaiptasia genomes. Lastly, we used BLAST to map our holobiont dataset to the genome of the distantly related starlet sea anemone, Nematostella vectensis (Putnam et al. 2007), to gauge the extent to which intra-order (Actiniaria) genomic resources could be used to effectively identify anemone-only 100 bp ddRADseq loci. All scripts for mapping and parsing anemone from symbiodiniacean DNA, along with full details and instructions for using them, can be found in Titus et al. (2019a), Dryad (https://doi.org/10.5061/dryad.c6f51c0), and GitHub (github.com/pblischak/Bann_spdelim).
Population genetic structure
We used classic population genetic approaches to infer structure in B. annulata populations from across the Tropical Western Atlantic. First, we used the Bayesian clustering program Structure v2.3.4 (Pritchard et al. 2000). For holobiont, aposymbiotic, and unmapped datasets, we collapsed bi-allelic data into haplotypes at each locus as detailed by Titus et al. (2019a). This allowed us to incorporate more information from each locus into our analyses when more than one SNP was present in a 100 bp locus. Structure analyses were conducted using the admixture model, correlated allele frequencies, and sample locality information. Each MCMC chain for each value of K was run with a burn-in of 1 × 105 generations and sampling period of 2 × 105 generations. We initially conducted two separate Structure analyses for the holobiont and aposymbiotic datasets. First, we conducted three iterations of a broad range of K values (1–6) to gain an initial snapshot of the data across the region. In both initial analyses we used the peak ln Pr(D|K) and the ∆K (Evanno et al. 2005) to inform the selection of the best K value. We then re-ran Structure using a narrower range of K values (1–4) but with more iterations (n = 10). Each MCMC chain for each value of K was run with a burn-in of 1 × 105 generations and sampling period of 2 × 105 generations. Again, we used ln Pr(D|K) and ∆K to select the best value of K.
Next, we conducted an analysis of molecular variance (AMOVA) in Arlequin v.3.5 (Excoffier and Lischer 2010) to test for hierarchical partitioning of genetic diversity across the region. Following our Structure results (see Results), we partitioned samples into Eastern and Western populations. We tested for hierarchical structure among sample localities (φST), among sample localities within a region (φSC), and between regions (φCT). We used Arlequin v3.5 to calculate distance matrices using the number of different alleles per locus, assessing statistical significance with 10,000 permutations. For each locality, we generated genetic diversity summary statistics and calculated pairwise φST values to test for differentiation among sample localities. All calculations were conducted for the aposymbiotic and for the holobiont datasets which have been deposited in Dryad (https://doi.org/10.5061/dryad.kkwh70s6p).
Demographic modeling selection and parameter estimation
While broad-scale patterns of spatial genetic structure may be robust to some levels of dinoflagellate contamination in reduced representation sequencing datasets, we expect that demographic model selection approaches that make inferences regarding patterns of demographic history and that generate important population parameter estimates (i.e., effective population sizes, migration rates), should be more sensitive to the incorporation of data from taxa with different evolutionary histories because different demographic processes can lead to the same genetic structuring. Thus, for each data set, we conducted demographic model selection using the allele frequency spectrum (AFS) and coalescent simulations in the program fastsimcoal2 (FSC2; Excoffier et al. 2013). FSC2 uses coalescent simulations to calculate the composite likelihood of arbitrarily complex demographic models under a given AFS; best-fit model(s) can be selected using the Akaike information criterion (AIC). We developed 12 demographic models (Fig. 2) for each dataset, all variants of a two-population isolation-migration model as Structure delimited K = 2 as the best clustering scheme (see Fig. 1b–d). Models differed in the directionality of gene flow, population size changes following divergence, and in patterns of secondary contact following divergence. Genetic clusters in Structure were largely partitioned East and West in the TWA, and 25 individuals from each putative population (50 individuals total; Supplementary Table S3) were randomly selected to generate two-population, joint-folded AFS.
Each model is a two-population isolation-migration (IM) model that varies in the degree and directionality of gene flow and effective population size. Models are as follows: a isolation only, b isolation with population size changes following divergence, c IM model with symmetric migration, d IM model with symmetric migration and population size changes, e IM model with migration from the Western to Eastern population, f IM model with migration from the Eastern to Western population, g IM model with symmetric migration between populations immediately following divergence followed by more contemporary isolation, h IM model with isolation immediately following divergence, followed by secondary contact and symmetric migration, i IM model with migration from the Western to Eastern population immediately following divergence, followed by more contemporary isolation, j IM model with isolation immediately following divergence followed by secondary contact and migration from the Western to Eastern population, k IM model with migration from the Eastern to Western population immediately following divergence, followed by more contemporary isolation, and l IM model with isolation immediately following divergence followed by secondary contact and migration from the Eastern to Western population. Model 6 was selected by Akaike Information Criterion as the best fit demographic model for both holobiont and aposymbiotic datasets.
Two-population, joint-folded AFS were generated from pyRAD output files and previously published python scripts (see Titus et al. 2019a for detailed explanations on AFS building). We repeated the AFS building procedure 10 times for each dataset to account for variation during model selection and to calculate confidence intervals on our parameter estimates (Satler and Carstens 2017; Smith et al. 2017; Titus et al. 2019a). Each simulation analysis in FSC2 (i.e., each AFS replicate per model; 12 models x 10 replicates) was repeated 50 times and we selected the run with the highest composite likelihood for each AFS replicate and model. The best-fit model was then calculated using the AIC and model probabilities following Burnham and Anderson (2002). To scale parameter estimates into real values, we used the substitution per site per generation mutation rate of 4.38 × 10−8 calculated for tropical anthozoans (Prada et al. 2016) and a generation time of 1 year for B. annulata (Jennison 1981; O’Reilly et al. 2018). All analyses were conducted on the Oakley cluster at the Ohio Supercomputer Center (http://osc.edu). All allele frequency spectrums, models, and replicates for conducing FSC2 analyses for holobiont and aposymbiotic datasets are deposited in Dryad (https://doi.org/10.5061/dryad.kkwh70s6p).
Comparative de novo assembly analyses
Variation in de novo RADseq assembly programs and parameters (e.g., clustering and missing data thresholds) can impact the number of recovered loci in resulting datasets (e.g., O’Leary et al. 2018). To assess how this variation may affect the number of holobiont and aposymbiotic loci recovered in our B. annualata dataset, as well as how many loci map to available Symbiodineacean genomes, we constructed 21 additional datasets using pyRAD v3.0.66 and Stacks v2.55 (Catchen et al. 2013). In pyRAD, we varied both the clustering threshold parameter to assemble reads into loci (Wclust = 0.90 or 0.85) and the maximum amount of missing data permitted at a locus before it could be incorporated into the final dataset (25 or 50%). In addition to the original dataset assembled above (Wclust = 0.90 & locus presence = 25%) these parameters resulted in pyRAD datasets with the following parameter values: 0.90_50, 0.85_25, and 0.85_50%.
Similarly, we used the program Stacks v.2.55 to create de novo RADseq datasets (using denovo_map.pl pipeline) analogous to those we created in pyRAD. In Stacks, we set the minimum sequencing coverage depth to seven (-m 7) and varied the number of mismatches allowed between sequences to assemble reads into loci (-M = 10 or 15). Given our 100 bp sequences, this allowed us to create a clustering threshold analogous to pyRAD (0.90 or 0.85). We also varied maximum amount of missing data permitted at a locus before it could be incorporated into the final dataset (25 or 50%). These parameters resulted in Stacks datasets with the same parameter values as in pyRAD: 0.90_25, 0.90_50, 0.85_25, and 0.85_50%.
For each new dataset, resulting holobiont loci were mapped against the Exaiptasia genome, all available dinoflagellate genomes, and the Nematostella genome using BLAST as above to compare the number of recovered holobiont, aposymbiotic, and symbiont loci for each dataset and parameter setting. Results from these BLAST searches were then used to create “aposymbiotic” and “unmapped” Structure files to compare genetic clustering patterns for each of the 21 new datasets. Genetic clustering patterns for K = 2 clusters was explored using a discriminant analysis of principal components method (DAPC; Jombart et al. 2010) in the adegenet package (Jombart and Ahmed, 2011) in R v3.5.0 (R Core Team 2015). This approach was taken to reduce the overall computation time. In adegenet, the best clustering scheme was assessed using the K-means method, setting the maximum K = 10, and retaining all principal components. The genetic data were transformed using principal component analyses (PCA) and linear discriminant analyses were performed on the retained principal components (no more than 50% were retained to avoid overfitting). The number of retained PCs were determined using the optimum alpha score. We then assigned each individual to a genetic unit at K = 2 according to its maximum membership probability.
To quantify similarities in the genetic cluster plots across datasets and assembly programs, we used pong (Behr et al. 2016). pong uses genetic cluster output (e.g., individual membership assignments, Q-matrices) to calculate a pairwise similarity metric across multiple cluster matrices. Pong’s pairwise similarity metric is based on the Jaccard index, which we used to quantitatively highlight how variation in assembly program, parameter setting, and dataset type impacts individual membership assignments in resulting cluster bar plots.
Results
Dataset assembly
Raw ddRADseq data from Titus et al. (2019a) comprised 186.1 million sequence reads across 123 individuals; 175 million reads passed quality control filtering in pyRAD and were retained to create the final dataset. Accounting for individuals with low sequence reads (<500,000 reads) resulted in a final intraspecific dataset of 101 individuals (Supplementary Tables S1 and S2). Requiring a locus to be present in at least 75% of all individuals resulted in a final holobiont data set of 11,331 SNPs distributed across 3854 loci. After mapping these loci to the Exaiptasia diaphana genome, we retained 1402 loci in the final aposymbiotic dataset and 2452 loci in the unmapped dataset. Only 59 of the 3854 holobiont ddRADseq loci (~1.5%) mapped to Symbiodiniaceae genomes (Supplementary Table S4), but this confirms the presence of at least some symbiont DNA in our holobiont dataset. Of these, 58 mapped to the S. microadriaticum genome (formerly Clade A) and one mapped to the C. goreaui genome (formerly Clade C; Supplementary Table S4). Only five loci from the holobiont dataset mapped to the starlet sea anemone N. vectensis genome (Supplementary Table S5). Holobiont and aposymbiotic datasets are available on Dryad (https://doi.org/10.5061/dryad.kkwh70s6p).
Population genetic structure
Genetic clustering analysis in Structure resolved similar patterns across the TWA for aposymbiotic, holobiont, and unmapped datasets. As in Titus et al. (2019a), K = 2 was selected by Structure using both lnP(K) and ∆K as the best clustering scheme for the aposymbiotic dataset (Fig. 1b; Supplementary Table S6). Diversity was largely binned into Western and Eastern partitions, but with admixture (Fig. 1b). The most notable genetic break was that between the Lower Keys (LK) and Eleuthera, Bahamas (BH), sample localities in close proximity and bisected by the Florida Straits (Fig. 1). The holobiont dataset recovered similar geographic partitioning, but Structure selected K = 3 as the best partitioning scheme using lnP(K) and ∆K (Fig. 1; Supplementary Fig. S1; Supplementary Table S7). However, the ∆K values between K = 2 and K = 3 are so similar this likely reflects a lack of significant biological difference between the two clustering models, a result seen in other studies (e.g., Abalaka et al. 2015; Leydet and Hellberg 2015). The additional genetic cluster did not illuminate any unrecovered geographic partitioning across the region beyond what was recovered by a K = 2 partitioning scheme (Supplementary Fig. S1), and ∆K values for K = 2 and K = 3 were very similar (Supplementary Table S7). The West-East genetic break across the TWA, with admixture, is still largely resolved in the holobiont dataset with the most notable break again between the LK and BH sample localities (Fig. 1c). The K = 3 result may just simply be because including more data in the analysis led to a higher probability of shared alleles in the dataset. Finally, K = 3 was selected as the best partitioning scheme for the unmapped dataset, which, like the holobiont dataset, showed similar genetic partitioning across the region and highly similar ∆K values between K = 2 and K = 3 (Fig. 1d; Supplementary Fig. S1; Supplementary Table S8).
Population genetic analyses in Arlequin reflect nearly identical results for the holobiont and aposymbiotic datasets. AMOVA results indicate low, but significant, population genetic structure at all hierarchical levels for both datasets, and both datasets have similar patterns of genetic variation at each hierarchical level (Table 1). Similarly, pairwise φST values calculated by Arlequin were low but significant among many sample localities for both datasets (Supplementary Table S9). Genetic diversity summary statistics for both datasets were virtually indistinguishable across all sample localities (Table 2).
Demographic model selection
Coalescent modeling in FSC2 returned identical model selection results between aposymbiotic and holobiont datasets (Table 3). For both, the best-fit model as chosen by the Akaike Information Critereon (AIC) is model 6, an IM model with unidirectional gene flow from East to West (Fig. 2). According to Akaike model weights, model 6 received >0.70 of the support (Table 3) in both the aposymbiotic and holobiont datasets. A secondary contact model (Model 10; Fig. 2) with isolation immediately after divergence, followed by secondary contact and unidirectional West-East gene flow, received the next highest amount of support according to AIC, although the Akaike weight differed between the datasets, with the holobiont dataset having a clearer preference for this model over the next best one, compared to the aposymbiotic dataset (Table 3). The increased power of the holobiont dataset to discriminate between the 2nd and 3rd best models (based on Akaike weights) points to the importance of dataset size for distinguishing between alternative models.
Parameter values and confidence intervals for effective population size, divergence time, and migration rate estimated from FSC2 simulations were overlapping between aposymbiotic and holobiont datasets (Table 4). For both datasets, FSC2 estimated that Eastern populations of B. annulata had greater effective population sizes than Western populations, and that the per-generation migration rate was low (Table 4). Divergence time estimates varied more than other parameter values but still had overlapping confidence intervals. The aposymbiotic dataset had an estimated a mean divergence time between Eastern and Western populations at ~39,000 ybp, whereas the holobiont dataset had an estimated a mean divergence time between populations at ~58,000 ybp.
Comparative de novo assembly analyses
Variation in clustering and missing data thresholds across de novo pyRAD and Stacks assembly programs revealed similarities and important differences in the number of recovered holobiont, aposymbiotic, and symbiont loci. Broadly, missing data thresholds had greater impact on dataset size and the total number of symbiont loci recovered by both programs than the clustering parameters (Table 5). For both pyRAD and Stacks datasets, increasing the missing data threshold to 50% roughly doubled the number of recovered holobiont loci (Table 5). In contrast, decreasing the clustering threshold from 0.90 to 0.85 resulted in more modest loci increases in pyRAD and slightly decreased the total number of holobiont loci recovered in Stacks (Table 5). The proportion of confirmed aposymbiotic loci recovered from holobiont datasets remained consistent across assembly programs and parameter settings, ranging from 0.33–0.42 of the total loci (Table 5).
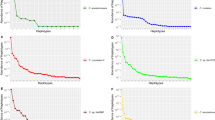
The greatest differences between programs and parameter settings are in the number of recovered symbiont loci (Table 5). The program pyRAD positively identified more symbiont loci than Stacks across all parameter settings, but no more than 4.6% of holobiont loci mapped to Symbiodneaceae genomic resources in any dataset (Table 5). As above, missing data thresholds had a far greater impact on the recovery of symbiont loci than clustering threshold. At the most conservative missing data threshold (≤25%), pyRAD datasets never recovered more than 200 symbiont loci, while Stacks datasets did not recover more than 10 symbiont loci (Table 5). Excluding the Stacks 85_25 dataset, which had only one locus map to a symbiont genome, all other datasets had loci that mapped to at least two symbiont genera (Table 6). The genera Symbiodinium and Cladocopium were the closest match for the most commonly recovered symbiont loci, but, interestingly, Cladocopium only appeared at meaningful levels when the missing data threshold was raised to 50% (Table 6). Similarly, the genera Breviolum and Fugacium, although never accounting for more than 14 total loci in any dataset, also only appeared in the RADseq datasets when the missing data threshold was raised to 50% (Table 6). Lowering the clustering threshold did not have a similar impact on the presence of symbiont loci. The number of recovered Breviolum, Cladocopium, and Fugacium loci remained almost identical across both pyRAD and Stacks when the clustering thresholds were lowered from 0.90 to 0.85 while keeping the missing data thresholds the same (Table 6). The number of recovered Symbiodinium loci did increase with lower clustering thresholds, but only in pyRAD (Table 6). Finally, the number of anemone loci that mapped to the distantly related Nematostella genome remained extremely low across all assembly programs and parameter settings (Table 5).
While missing data thresholds appeared to have the greatest impact on both assembly programs in terms of the number of recovered holobiont, aposymbiotic, and symbiont loci, the resulting genetic structure plots revealed important differences in the way these programs and parameters resolve genetic structure in B. annulata (Fig. 3). In pyRAD, the clustering parameter (Wclust) appears to have the most impact on the genetic structure across holobiont, aposymbiotic, and unmapped datasets, while missing data thresholds appear to have the most impact on genetic structure in datasets produced by Stacks (Fig. 3). At high clustering thresholds (0.90), genetic structure plots are highly similar across the Caribbean for B. annulata regardless of missing data parameters or whether the dataset is comprised of holobiont, aposymbiotic, or unmapped loci (Figs. 1 and 3). The program pong calculated similarity metrics >77% when comparing individual cluster membership assignments across holobiont, aposymbiotic, and unmapped loci (Fig. 3). However, when genetic clustering thresholds are relaxed (Wclust = 0.85), only the aposymbiotic structure plots show consistency with those produced at higher cluster thresholds. Both the py85_25 and py85_50 holobiont and unmapped datasets resolved genetic structures for B. annulata that were discordant with all aposymbiotic datasets in pyRAD and with holobiont and unmapped datasets produced with a Wclust threshold of 0.90 (Fig. 3). Similarity metrics generated by pong reflected this discordance and resulting genetic cluster plots had average pairwise similarities of <70% (Fig. 3).
Genetic clusters were detected using discriminant analysis of principal components (DAPC). Datasets varied by the clustering threshold parameter to assemble reads into loci (0.90 or 0.85 sequence similarity) and the maximum amount of missing data allowed before a locus could be incorporated into the final dataset (25 or 50%). Genetic cluster plots with the prefix “py” were produced in pyRAD and plots labeled with the prefix “st” were produced in Stacks. Similarity metrics for each assembly program and parameter setting were calculated by pong and represent the average pairwise percentage of individuals with identical cluster assignment probabilities across aposymbiotic, holobiont, and unmapped datasets.
For datasets produced by Stacks, genetic cluster plots are remarkably consistent across holobiont, aposymbiotic, and unmapped data within a given parameter setting and had the highest pairwise similarity scores produced by pong (all > 84%, Fig. 3). However, between datasets, missing data appears to produce the largest differences in genetic structure plots (Fig. 3). More genetic structure was recovered in Stacks datasets when the missing data threshold was reduced to ≤50% (Fig. 3) than when left at the more conservative missing data threshold of ≤25%.
In addition to variation between datasets within an assembly program, the most variation in genetic structure appeared between assembly programs (Fig. 3). The strongest biogeographic signal occurred in datasets produced by pyRAD, which resolved the Florida Straits as the strongest genetic break in the region. Genetic structure plots from Stacks datasets showed much less resolution, but structure plots from datasets where the missing data threshold was ≤50% did loosely recover genetic structuring across the Florida Straits as well (Fig. 3). All files and raw data are available on Dryad (https://doi.org/10.5061/dryad.kkwh70s6p).
Discussion
Marker development and symbiodiniacean contamination has represented a substantial hurdle for researchers working on tropical anthozoans (e.g., Shearer et al. 2005; Bongearts et al. 2017; Leydet et al. 2018). These issues have been presumed to remain for reduced representation sequencing approaches, which may be why these methods have not been leveraged in symbiotic anthozoans to the extent that they have for other taxa. However, based on the analyses we conduct here, we find broadly similar interpretations from our main holobiont and aposymbiotic datasets produced by reduced representation sequencing. In our primary analyses, we expected that more than doubling our dataset (~1400 vs 3800 loci) and including putative symbiont loci would lead to major differences in our population genetic results. The ~2400 unmapped loci from our holobiont dataset that did not map to the E. diaphana genome have nearly identical genetic clustering results to the aposymbiotic dataset even though the additional loci represent some combination of anemone and symbiont sequences. Because each anthozoan tentacle cell can contain multiple symbiodiniacean cells, photosymbiont nuclei can potentially outnumber anemone nuclei in some tissue layers and thus constitute a significant source of potential contamination (Muscatine et al. 1998). Therefore, although only 59 loci are confirmed as from symbionts, our holobiont dataset could contain more dinoflagellate sequences than B. annulata sequences. If even half of the 2400 unmapped loci from our holobiont dataset were from members of Symbiodiniaceae, we would expect this to greatly influence our holobiont analyses. This should be especially true of our parameter estimates and genetic diversity summary statistics, which should be the most sensitive to the incorporation of sequence data from multiple species with different evolutionary histories. That we recover largely indistinguishable population genetic results with completely overlapping diversity indices, summary statistics, and parameter estimates leads us to hypothesize that we have very few symbiodiniacean loci in our holobiont dataset, and that most symbiodineacean sequences were filtered out by pyRAD during dataset assembly. Our additional 21 comparative datasets produced through pyRAD and Stacks, along with the corresponding genetic cluster analyses, provide additional support for this hypothesis and important insight into dataset assembly parameters for future studies. Ultimately, our results and observations suggest that neither reference genomes, nor complicated pre- and post-sequencing approaches, are necessary to make robust population genetic inferences using reduced representation sequencing approaches for at least some symbiotic anthozoans. Our direct test of this hypothesis provides a posteriori validation that many previous studies analyzing coral holobiont DNA using these sequencing approaches (e.g., Quattrini et al. 2019; Porro et al. 2020) were acquiring their signal from the coral host.
Our interpretation of these patterns, and resulting hypothesis, rests on several observations surrounding basic anthozoan and symbiodiniacean biology, the manner in which de novo RADseq assembly programs assemble orthologous loci, and how these factors likely interact to determine whether symbiont loci will be removed from or incorporated into resulting RADseq datasets. First, many tropical anthozoans, including B. annualata, have flexible symbiont associations that involve diverse lineages of Symbiodiniaceae (e.g., Santos 2016; Silverstein et al. 2012), which in hospite, are haploid gametophytes (Santos and Coffroth 2003). Members of the same host species can harbor different genera and species of Symbiodiniaceae (previously called Clades and Types of Symbiodinium) in different habitats, across broad geographic space, and within the same individual or colony (Baker 2003; Silverstein et al. 2012; Santos 2016). Thus, across a population of anthozoan hosts, it is not uncommon to find some individuals harboring a clonal haploid population of a single species of symbiont, others hosting multiple species within the same genus, and other individuals hosting two or more symbiont genera simultaneously (reviewed by Wham and LaJeunesse 2016). The flexibility of these associations means that even within an individual host, the identity of the symbiont can be switched altogether, or if a host harbors multiple species or genera simultaneously, the relative abundance of these symbionts can be shuffled (e.g., Mieog et al. 2007). Second, the genetic divergences between genera of Symbiodiniaceae are comparable to order-level differences in other dinoflagellates, representing divergences as old as the mid-Jurassic (LaJeunesse et al. 2018; Santos 2016). Given an adequate ecological or geographic distribution, many sets of field collected samples will therefore harbor multiple genera and species of Symbiodiniaceae that have been diverged for tens of millions of years. These aspects of anthozoan and symbiodineacean biology are important to consider because they will interact with reduced representation assembly programs, and likely result in symbiont loci being filtered out of resulting datasets.
Reduced representation datasets, as outlined above, should be comprised of short (50–100 bp sequences), largely non-coding DNA fragments. When de novo SNP-calling programs assemble DNA sequence fragments and call orthologous loci, the user specifies a series of important parameters that are well-documented to impact the number of recoverable loci (reviewed by O’Leary et al. 2018). Two of these parameters, the locus clustering threshold and the missing data threshold, are particularly germane to our hypothesis that symbiont loci often get filtered out during data set assembly. The locus clustering threshold uses a sequence similarity threshold to assemble orthologous loci within individuals. Overly stringent thresholds lead to over-splitting orthologous sequences into separate loci, while relaxed thresholds lead to RADseq programs over-clustering paralogous loci and potentially introducing artefactual SNPs (Catchen et al. 2013; Eaton 2014; O’Leary et al. 2018).
Missing data thresholds specify the percentage of individuals in which an orthologous locus must be present before it can be included in the final dataset. Factors such as library preparation and sequencing depth can lead to allele dropout in RADseq datasets (Catchen et al. 2013; Eaton 2014; O’Leary et al. 2018), but so can biologically meaningful factors, like evolutionary relatedness among samples within a dataset (Harvey et al. 2015; Rubin et al. 2012). If samples are distantly related, RADseq programs may have difficulty finding mutationally-conserved, orthologous loci that meet missing data thresholds. This problem underscores the rationale behind why phylogenomic studies do not use reduced representation sequencing to resolve ordinal relationships. These methods typically do not produce enough orthologous loci across taxa that have diverged over deep evolutionary timescales to generate well resolved phylogenies (Harvey et al. 2015; Rubin et al. 2012). Reinforcing this point, in our primary B. annulata dataset, only five loci mapped to the genome of the starlet anemone Nematostella vectensis, and no more than 16 total loci mapped to N. vectensis even under the most relaxed assembly parameters. This is likely because B. annulata and N. vectensis are distant relatives that share a last common ancestor ~500 mya (McFadden et al. 2021; Quattrini et al. 2020; Rodríguez et al. 2014; Titus et al. 2019b). Thus, our ddRADseq approach would be severely limited for understanding evolutionary relationships among Actiniaria at this scale.
Sampling and sequencing an anthozoan species at the population level regularly results in sampling and sequencing symbiodineacean diversity at the genus level. Thus, moderately conservative assembly parameters may be enough to filter out the majority of symbiodiniacean sequences because the program cannot find enough mutationally-conserved, orthologous loci across the genetically divergent Symbiodiniaceae hosted by the focal anthozoan to meet the missing data thresholds. For example, imagine a ddRADseq dataset consisting of 10 individuals of an anthozoan from Florida that harbor a monoclonal haploid population of symbionts from the genus Symbiodinium, and 10 individuals from Bermuda that harbor a monoclonal haploid population of symbionts from the genus Breviolum. The genetic divergence between Symbiodinium and Breviolum is large enough (pairwise distance for LSU DNA = 0.37, estimated divergence ~170 mya: LaJeunesse et al. 2018) that few non-coding orthologous DNA sequences from Symbiodinium and Breviolum would be retained under a pyRAD missing data threshold requiring a locus to be present in 75% of all individuals. Thus, the loci that would be retained in the final dataset would primarily be from the host anthozoan, which represents intraspecific diversity at shallower evolutionary timescales, compared to their symbionts. Under this hypothetical scenario where anthozoan hosts harbor monoclonal haploid symbionts, any symbiont loci that do end up in the final dataset should, in theory, be 1) from highly conserved genomic regions and 2) treated as fixed homozygous loci due assembly programs assuming the focal taxa are diploid and thus lead to heterozygote deficiencies. Alternatively, if resulting symbiont loci in the final dataset are returned as heterozygous, it could indicate that low clustering thresholds are over-clustering paralogous loci giving the appearance of diploid genomes. In our analyses, no symbiont locus that was present in any final dataset mapped to multiple symbiont genomes, suggesting that our sequencing and assembly approaches were not recovering highly conserved symbiodineacean loci.
In other scenarios where the composition of symbionts within host anthozoans are more complicated, other outcomes could be expected. If individuals of a target anthozoan harbor mixtures of different genotypes from the same symbiodineacean species, different species from the same genus, or different genera altogether then the relative abundance and relatedness of each symbiont lineage will likely be important for determining how/if symbiont loci are retained in the final dataset. Under lower clustering thresholds, loci from closely related symbionts could be over-clustered and treated as diploid; if more than two symbiont lineages are present and cluster together at a locus, then it would violate the diploid assumptions of the assembly programs and the loci would be removed. If a single lineage of symbiont dominated, but background levels of other lineages were present, then missing data thresholds would be important in determining if these loci were ultimately retained.
In our comparative de novo analyses using both pyRAD and Stacks, we can begin to see preliminary support for our hypothesis and how these factors can interact to either include or filter out symbiont loci from our B. annulata datasets. We know previously that B. annulata hosts multiple members of Symbiodiniaceae throughout its range: Symbiodinium in Bermuda and Florida and Cladocopium in Florida, Mexico, and Panama (Grajales et al. 2015; see LaJeunesse et al. 2018 for updated taxonomy). Loci from our holobiont dataset primarily mapped to the published genome of Symbiodinium in both pyRAD and Stacks, although at much greater levels in pyRAD. Once missing data thresholds were relaxed to 50%, Cladocopium loci appeared, along with low numbers of loci that also mapped to Brevioulum and Fugacium, confirming the occurrence of four symbiont genera in the B. annulata we collected. This pattern was importantly recovered from datasets produced by both assembly programs. The number and diversity of symbiont loci did not increase to nearly the same degree in pyRAD when clustering thresholds were relaxed, and in fact decreased in the program Stacks, highlighting the potential of missing data thresholds to filter out symbiont loci.
Clustering thresholds do appear particularly important when constructing datasets in pyRAD. Here we saw consistency across genetic cluster plots at a clustering threshold of 0.90, regardless of missing data values, but substantial differences between holobiont, aposymbiotic, and unmapped genetic cluster plots when clustering thresholds were relaxed to 0.85. pyRAD is often touted for its ability to handle indels, and thus outperforms other RADseq assembly programs when datasets harbor interspecific diversity (Eaton 2014). This may have contributed to the greater number of symbiont loci that made it into final pyRAD datasets, compared to Stacks. The genetic cluster plots for aposymbiotic datasets produced in pyRAD at a clustering threshold of 0.85 were identical to those produced at higher clustering thresholds, suggesting that symbiont loci in the corresponding holobiont and unmapped datasets were driving the observed discordance. In Stacks, genetic cluster plots were remarkably consistent across holobiont, aposymbiotic, and unmapped datasets within a given set of parameters. We did not see a corresponding set of parameters in Stacks where the aposymbiotic genetic cluster plots were significantly different from their corresponding holobiont or unmapped genetic cluster plot results.
Our comparative de novo analyses between pyRAD and Stacks also highlight important differences in the B. annulata population genetic structure recovered by both assembly programs. While some similarities exist, our pyRAD datasets picked up much more biogeographic structure than our Stacks dataset, regularly recovering a genetic break across the Florida Straits between the Florida Keys and the Bahamas. At more relaxed missing data thresholds, this pattern began to emerge in Stacks as well, but was much less clear. Another interesting difference was the consistency of the population genetic structure recovered in the aposymbiotic datasets in both programs across different assembly parameters. In pyRAD, the genetic structure plots for B. annulata are virtually identical across aposymbiotic datasets regardless of the clustering or missing data thresholds, highlighting an underlying consistency of pyRAD to recover the same signal. In Stacks, however, even though genetic cluster plots were consistent within a given set of parameters, unlike pyRAD, the pattern of genetic structure did change within the aposymbiotic datasets between parameter settings. This suggests that the program, rather than an underlying biological signal from the animal, has an important impact on population genetic results. An alternative interpretation is that our B. annulata RADseq dataset lacks a strong enough biogeographic signal to be recovered by both datasets. A less admixed or heterogeneous dataset may not encounter as much variation between assembly programs as we see here. This variation between programs, however, should not overshadow the more important implications from these analyses, which is the consistency of Stacks, and to a lesser degree pyRAD, to recover the same genetic signal across holobiont, aposymbiotic, and unmapped datasets within a given set of assembly parameters.
The framework and experimental design of our study, effectively a single-species phylogeographic study that spans the entire range of our focal taxon, is representative of many studies that examine the spatial and demographic history of a given species at the population level. Although the degree to which symbiotic anthozoans are specific to a particular lineage of Symbiodiniaceae is unresolved, evidence is overwhelming that these associations are often spatially and temporally variable, particularly in stony corals, where much of this research has focused (e.g., Silverstein et al. 2012). Thus, given a broad sampling scheme with respect to geography and habitat, de novo assembly programs may act as de facto filtering programs for symbiodiniaceans in many reduced representation datasets produced from symbiotic anthozoans. Resulting datasets will be overwhelmingly comprised of anthozoan DNA loci and any remaining symbiont loci may simply be genetic “noise”.
Of course, our case study is not without important caveats. Because we do not have a B. annulata reference genome, we cannot account for 100% of all ddRADseq loci that belong to the host. That the reference genome is from a species in the same family, rather than more closely related, is the most likely explanation for why ~2400 loci remain uncharacterized in our holobiont dataset. These loci are simply not shared between E. diaphana and B. annulata, and so are not included in the aposymbiotic dataset. Similarly, although mapping our reads to genomic resources from members of Symbiodiniaceae confirms we do have some dinoflagellate sequence data in our holobiont dataset (at least ~1.5% of all loci), we do not have genomes for the exact genus and species of symbiodiniacean in our B. annulata samples. Further, the choice of restriction enzymes used in our double-digest library preparation could impact the loci recovered. We used two six-base pair cutting enzymes, and it is unclear how using more frequent cutting enzymes, or simply conducting a single restriction digest, would affect the results. ddRAD approaches may be more prone to allele dropout than single enzyme digestions because the likelihood of mutation at restriction sites increases with as the number of restriction enzymes used to digest DNA increases. Follow-up studies that do have access to conspecific reference genomes for both host and symbiont will be important to test the hypotheses we present here, as are in silico restriction digests that test the effectiveness of different enzyme combinations.
From a practical standpoint, our study demonstrates that Symbiodineaceae DNA is not a consistent source of contamination in reduced representation libraries, and thus, those that feel strongly about employing a reference genome should weigh that decision against other important factors- particularly the final dataset size. Our holobiont dataset contained >2x the number of loci as our aposymbiotic dataset. While this did not significantly impact our overall results, many population and phylogeographic analyses such as FSC2 and other demographic simulation programs (e.g., Moments, dadi) show increased ability to differentiate between alternative models with increasing dataset size. Our holobiont FSC2 analyses showed a clear preference between the 2nd and 3rd best models, whereas our aposymbiotic FSC2 analyses did not (Table 3). If employing a conspecific genome, the impact on the resulting dataset size may be minimal, but using a genome from a more distant relative may result in a significant reduction in dataset size. Our study demonstrates that reference genomes within the same anthozoan family may serve as adequate genomic resources, but reference genomes that are simply within the same order are too distant to serve in the same capacity, at least for actiniarians: only five loci from B. annulata (suborder Anthemonae, superfamily Metridioidea, family Aiptasiidae) mapped to the genome of Nematostella vectensis (suborder Anenthemonae, superfamily Edwardsioidiea, family Edwardsiidae), and no more than 16 loci mapped to N. vectensis in our broader comparative dataset assembly analyses. For larger families or families known to be non-monophyletic (e.g., Actiniidae: Daly et al. 2017; Hormathiidae: Gusmão et al. 2020; Stichodactylidae: Titus et al. 2019b), a single family-level reference genome is likely to be insufficient.
For reduced representation approaches for symbiotic anthozoan groups without access to reference genomes or for researchers that decide against using them, we recommend that studies 1) employ extensive geographic sampling, or sample broadly across ecologically disjunct habitats (i.e., depth, temperature, nutrient concentration) to maximize the likelihood of sampling hosts that harbor diverse symbiodiniaceans, and 2) demonstrate empirically that multiple genera of Symbiodiniaceae are represented in the collected samples via PCR or sequencing (e.g., ITS, cp23s). In host species with highly specific endosymbiont associations, the approach to sampling and sequencing we describe here may be ineffective, as orthologous symbiodiniacean loci might be present in all samples and sample localities and de novo clustering programs may not filter them out. In these cases, employing approaches like those of Bongearts et al. (2017) or Leydet et al. (2018) may be required. We also recommend that studies using the assembly program pyRAD employ more conservative clustering and missing data thresholds, as these parameter settings produced the most consistent results. In Stacks, parameter settings had minimal effect on the inferred final population genetic structure, but the program also seemed to produce datasets with a weaker genetic structure overall.
Understanding the evolutionary and historical processes that have shaped the diversity of tropical anthozoans has been, and will continue to be, an important research priority for marine population geneticists (Bowen et al. 2013; Bowen et al. 2016). Our case study presents a promising framework and way forward for researchers wishing to employ these reduced representation sequencing approaches on symbiotic anthozoan species that do not have reference genomes readily available.
Data availability
All files and raw data are available on Dryad (https://doi.org/10.5061/dryad.kkwh70s6p).
References
Abalaka J, Hudin NS, Ottosson U, Bloomer P, Hansson B (2015) Genetic diversity and population structure of the range restricted rock firefinch Lagonosticta sanguinodorsalis. Conserv Genet 16:411–418
Allio R, Donega S, Galtier N, Nabholz B (2017) Large variation in the ratio of mitochondrial to nuclear mutation rate across animals: implications for genetic diversity and the use of mitochondrial DNA as a molecular marker. Mol Biol Evol 34:2762–2772
Andras JP, Rypien KL, Harvell CD (2013) Range-wide population genetic structure of the Caribbean sea fan coral, Gorgonia ventalina. Mol Ecol 22:56–73
Aranda M, Li Y, Liew YJ, Baumgarten S, Simakov O, Wilson MC et al. (2016) Genomes of coral dinoflagellate symbionts highlight evolutionary adaptations conducive to a symbiotic lifestyle. Sci Rep 6:39734
Arbogast BS (2001) Phylogeography: the history and formation of species. Am Zool 41:134–135
Avise JC (2009) Phylogeography: retrospect and prospect. J Biogeogr 36:3–15
Avise JC, Bowen BW, Ayala FJ (2016) In the light of evolution X: comparative phylogeography. Proc Natl Acad Sci 113:7957–7961
Baker AC (2003) Flexibility and specificity in coral-algal symbiosis: diversity, ecology, and biogeography of Symbiodinium. Annu Rev Ecol Evol Syst 34:661–689
Baumgarten S, Simakov O, Esherick LY, Liew YJ, Lehnert EM, Michell CT et al. (2015) The genome of Aiptasia, a sea anemone model for coral symbiosis. Proc Natl Acad Sci 112:11893–11898
Baums IB, Miller MW, Hellberg ME (2005) Regionally isolated populations of an imperiled Caribbean coral, Acropora palmata. Mol Ecol 14:1377–1390
Bongearts P, Riginos C, Brunner R, Englebert N, Smith SR, Hoegh-Guldberg O (2017) Deep reefs are not universal refuges: reseeding potential varies among coral species. Sci Adv 3:e1602373
Bowen BW, Meylan AB, Ross JP, Limpus CJ, Balazs GH, Avise JC (1992) Global population structure and natural history of the green turtle (Chelonia mydas) in terms of matriarchal phylogeny. Evolution 46:865–881
Bowen BW, Kamezaki N, Limpus CJ, Hughes GR, Meylan AB, Avise JC (1994) Global phylogeography of the loggerhead turtle (Caretta caretta) as indicated by mitochondrial DNA haplotypes. Evolution 48:1820–1828
Bowen BW, Rocha LA, Toonen RJ, Karl SA, ToBo Lab (2013) The origins of tropical marine biodiversity. Trends Ecol Evol 28:359–366
Bowen BW, Gaither MR, DiBattista JD, Iacchei M, Andrews KR, Grant WS et al. (2016) Comparative phylogeography of the ocean planet. Proc Natl Acad Sci 113:7962–7969
Brugler MR, González‐Muñoz RE, Tessler M, Rodríguez E (2018) An EPIC journey to locate single‐copy nuclear markers in sea anemones. Zoologica Scr 47:756–776
Burnham KP, Anderson DR (2002) Model selection and multimodal inference: a practical information theoretic approach, 2nd ed. Springer-Verlag, New York, NY
Carstens B, Lemmon AR, Lemmon EM (2012) The promises and pitfalls of next-generation sequencing data in phylogeography. Syst Biol 61:713–715
Carstens BC, Pelletier TA, Reid NM, Satler JD (2013) How to fail at species delimitation. Mol Ecol 22:4369–4383
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22:3124–3140
Combosh DJ, Vollmer SV (2015) Trans-Pacific RAD-seq population genomics confirms introgressive hybridization in Eastern Pacific Pocillopora corals. Mol Phylogenetics Evol 88:154–162
Cowman PF, Quattrini AM, Bridge TC, Watkins-Colwell GJ, Fadli N, Grinblat M et al. (2020) An enhanced target-enrichment bait set for Hexacorallia provides phylogenomic resolution of the staghorn corals (Acroporidae) and close relatives. Mol Phylogenetics Evol 153:106944
Cunning R, Bay RA, Gillette P, Baker AC, Traylor-Knowles N (2018) Comparative analysis of the Pocillopora damicornis genome highlights role of immune system in coral evolution. Sci Rep 8(1):1–10
Daly M, Crowley LM, Larson P, Rodríguez E, Saucier EH, Fautin DG (2017) Anthopleura and the phylogeny of Actinioidea (Cnidaria: Anthozoa: Actiniaria). Org Diversity Evol 17(3):545–564
Daly M, Gusmão LC, Reft AJ, Rodríguez E (2010) Phylogenetic signal in mitochondrial and nuclear markers in sea anemones (Cnidaria, Actiniaria). Integr Comp Biol 50:371–388
Davies SW, Marchetti A, Ries JB, Castillo KD (2016) Thermal and pCO2 stress elicit divergent transcriptomic responses in a resilient coral. Front Mar Sci 3:112
DeBiasse MB, Richards VP, Shivji MS, Hellberg ME (2016) Shared phylogeographical breaks in a Caribbean coral reef sponge and its invertebrate commensals. J Biogeogr 43:2136–2146
Devlin-Durante MK, Baums IB (2017) Genome-wide survey of single-nucleotide polymorphisms reveals fine-scale population structure and signs of selection in the threatened Caribbean elkhorn coral, Acropora palmata. PeerJ 5:e4077
Drury C, Schopmeyer S, Goergen E, Bartels E, Nedimyer K, Johnson M et al. (2017) Genomic patterns in Acropera cervicornis show extensive population structure and variable genetic diversity. Ecol Evol 7:6188–6200
Eaton DA (2014) PyRAD: assembly of de novo RADseq loci for phylogenetic analyses. Bioinformatics 30:1844–1849
Erickson KL, Pentico A, Quattrini AM, McFadden CS (2020) New approaches to species delimitation and population structure of anthozoans: Two case studies of octocorals using ultraconserved elements and exons. Mol Ecol Resour 21:78–92
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Dupanloup I, Huerta-Sánchez E, Sousa VC, Foll M (2013) Robust demographic inference from genomic and SNP data. PLOS Genet 9:e1003905
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Forsman ZH (2017) Coral hybridization or phenotypic variation? Genomic data reveal gene flow between Porites lobata and P. compressa. Mol Phylogenetics Evol 111:132–148
Foster NL, Paris CB, Kool JT, Baums IB, Stevens JR, Sanchez JA et al. (2012) Connectivity of Caribbean coral populations: complementary insights from empirical and modelled gene flow. Mol Ecol 21:1143–1157
Gates RD, Edmunds PJ (1999) The physiological mechanisms of acclimatization in tropical reef corals. Am Zool 39:30–43
Glon H, Quattrini A, Rodríguez E, Titus BM, Daly M (2021) Comparison of sequence-capture and ddRAD approaches in resolving species and populations in hexacorallian anthozoans. Mol Phylogenetics Evol 163:107233
Grajales A, Rodriguez E (2014) Morphological revision of the genus Aiptasia and the family Aiptasiidae (Cnidaria, Actiniaria, Metridioidea). Zootaxa 3826:55–100
Grajales A, Rodríguez E (2016) Elucidating the evolutionary relationships of the Aiptasiidae, a widespread cnidarian–dinoflagellate model system (Cnidaria: Anthozoa: Actiniaria: Metridioidea). Mol Phylogenetics Evol 94:252–263
Grajales A, Rodríguez E, Thornhill DJ (2015) Patterns of Symbiodinium spp. associations within the family Aiptasiidae, a monophyletic lineage of symbiotic sea anemones (Cnidaria, Actiniaria). Coral Reefs 35:345–355
Grinblat M, Cooke I, Shlesinger T, Ben-Zvi O, Loya Y, Miller DJ et al. (2021) Biogeography, reproductive biology and phylogenetic divergence within the Fungiidae (mushroom corals). Mol Phylogenetics Evol 164:107265
Gusmão LC, Van Deusen V, Daly M, Rodríguez E (2020) Origin and evolution of the symbiosis between sea anemones (Cnidaria, Anthozoa, Actiniaria) and hermit crabs, with additional notes on anemone-gastropod associations. Mol Phylogenetics Evol 148:106805
Harvey MG, Judy CD, Seeholzer GF, Maley JM, Graves GR, Brumfield RT (2015) Similarity thresholds used in DNA sequence assembly from short reads can reduce the comparability of population histories across species. PeerJ 3:p.e895
Huebner LK, Chadwick NE (2012) Reef fishes use sea anemones as visual cues for cleaning interactions with shrimp. J Exp Mar Biol Ecol 416:237–242
ICZN (2017) Opinion 2404 (Case 3633)–Dysactis pallida Agassiz in Verrill, 1864 (currently Aiptasia pallida; Cnidaria: Anthozoa: Hexacorallia: Actiniaria): precedence over Aiptasia diaphana (Rapp, 1829), Aiptasia tagetes (Duchassaing de Fombressin & Michelotti, 1864), Aiptasia mimosa (Duchassaing de Fombressin & Michelotti, 1864) and Aiptasia inula (Duchassaing de Fombressin & Michelotti, 1864) not approved. Bull Zool Nomenclature 72:130–131
Iguchi A, Yoshioka Y, Forsman ZH, Knapp IS, Toonen RJ, Hongo Y et al. (2019) RADseq population genomics confirms divergence across closely related species in blue coral (Heliopora coerulea). BMC Evolut Biol 19(1):187
Jennison BL (1981) Reproduction in three species of sea anemones from Key West, Florida. Can J Zool 59:1708–1719
Johnston EC, Forsman ZH, Flot JF, Schmidt-Roach S, Pinzon JH, Knapp ISS et al. (2017) A genomic glace through the fog of plasticity and diversification in Pocillopora. Sci Rep 7:5991
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 11:1–15
Jombart T, Ahmed I (2011) adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:3070–3071
Kenkel CD, Matz MV (2017) Gene expression plasticity as a mechanism of coral adaptation to a variable environment. Nat Ecol Evol 1:0014
Kenkel CD, Moya A, Strahl J, Humphrey C, Bay LK (2018) Functional genomic analysis of corals from natural CO2‐seeps reveals core molecular responses involved in acclimatization to ocean acidification. Glob Change Biol 24:158–171
Knowles LL (2009) Statistical phylogeography. Annu Rev Ecol Evol Syst 40:593–612
LaJeunesse TC, Parkinson JE, Gabrielson PW, Jeong HJ, Reimer JD, Voolstra CR et al. (2018) Systematic revision of Symbiodiniaceae highlights the antiquity and diversity of coral endosymbionts. Curr Biol 28:2570–2580
Leydet KP, Grupstra CG, Coma R, Ribes M, Hellberg ME (2018) Host‐targeted RAD‐Seq reveals genetic changes in the coral Oculina patagonica associated with range expansion along the Spanish Mediterranean coast. Mol Ecol 27:2529–2543
Liu H, Stephens TG, Gonzalez-Pech RA, Beltran VH, Lapeyre B, Bongaerts P et al. (2018) Symbiodinium genomes reveal adaptive evolution of functions related to coral-dinoflagellate symbiosis. Nat Commun Biol 1:95
McCormack JE, Hird SM, Zellmer AJ, Carstens BC, Brumfield RT (2013) Applications of next-generation sequencing to phylogeography and phylogenetics. Mol Phylogenetics Evol 66:526–538
McFadden CS, Quattrini AM, Brugler MR, Cowman PF, Dueñas LF, Kitahara MV et al. (2021) Phylogenomics, origin, and diversification of anthozoans (Phylum Cnidaria). Syst Biol 70:635–647
Mieog JC, van Oppen MJH, Cantin NE, Stam WT, Olsen JL (2007) Real-time PCR reveals a high incidence of Symbiodinium clade D at low levels in four scleractinian corals across the Great Barrier Reef: implications for symbiont shuffling. Coral Reefs 26:449–457
Muscatine LR, McCloskey LE, Marian R (1981) Estimating the daily contribution of carbon from zooxanthellae to coral animal respiration. Limnol Oceanogr 26:601–611
Muscatine L, Ferrier-Pages C, Blackburn A, Gates RD, Baghdasarian G, Allemand D (1998) Cell-specific density of symbiotic dinoflagellates in tropical anthozoans. Coral Reefs 17:329–337
Nakabayashi A, Yamakita T, Nakamura T, Aizawa H, Kitano YF, Iguchi A et al. (2019) The potential role of temperate Japanese regions as refugia for the coral Acropora hyacinthus in the face of climate change. Sci Rep 9:1–12
O’Leary SJ, Puritz JB, Willis SC, Hollenbeck CM, Portnoy DS (2018) These aren’t the loci you’re looking for: Principles of effective SNP filtering for molecular ecologists. Mol Ecol 27:3193–3206
O’Reilly EE, Chadwick NE (2017) Population dynamics of corkscrew sea anemones Bartholomea annulata in the Florida Keys. Mar Ecol Prog Ser 567:109–123
O’Reilly EE, Titus BM, Nelsen MW, Ratchford S, Chadwick NE (2018) Giant ephemeral anemones? Rapid growth and high mortality of corkscrew sea anemones Bartholomea annulata (Le Sueur, 1817) under variable conditions. J Exp Mar Biol Ecol 509:44–53
Pante E, Puillandre N, Virice A, Arnaud‐Haond S, Aurelle D, Castelin M et al. (2015) Species are hypotheses: avoid connectivity assessments based on pillars of sand. Mol Ecol 24:525–544
Pelletier TA, Carstens BC (2014) Model choice for phylogeographic inference using a large set of models. Mol Ecol 23:3028–3043
Porro B, Mallien C, Hume BC, Pey A, Aubin E, Christen R et al. (2020) The many faced symbiotic snakelocks anemone (Anemonia viridis, Anthozoa): host and symbiont genetic differentiation among colour morphs. Heredity 124:351–366
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Prada C, Hanna B, Budd AF, Woodley CM, Schmutz J et al. (2016) Empty niches after extinctions increase population sizes of modern corals. Curr Biol 26:3190–3194
Putnam NH, Srivastava M, Hellsten U, Dirks B, Chapman J, Salamov A et al. (2007) Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317:86–94
Quattrini AM, Faircloth BC, Dueñas LF, Bridge TC, Brugler MR, Calixto‐Botía IF et al. (2018) Universal target‐enrichment baits for anthozoan (Cnidaria) phylogenomics: New approaches to long‐standing problems. Mol Ecol Resour 18:281–295
Quattrini AM, Wu T, Soong K, Jeng MS, Benayahu Y, McFadden CS (2019) A next generation approach to species delimitation reveals the role of hybridization in a cryptic species complex of corals. BMC Evolut Biol 19(1):1–19
R Core Team (2015) R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. https://wwwR-project.org/
Reeb CA, Avise JC (1990) A genetic discontinuity in a continuously distributed species: mitochondrial DNA in the American oyster, Crassostrea virginica. Genetics 124:397–406
Reitzel AM, Herrera S, Layden MJ, Martindale MQ, Shank TM (2013) Going where traditional markers have not gone before: utility of and promise for RAD sequencing in marine invertebrate phylogeography and population genomics. Mol. Ecol. 22:2953–2970
Rippe JP, Matz MV, Green EA, Medina M, Khawaja NZ, Pongwarin T et al. (2017) Population structure and connectivity of the mountainous star coral, Orbicella faveolata, throughout the wider Caribbean region. Ecol Evol 7:9234–9246
Rodríguez E, Barbeitos MS, Brugler MR, Crowley LM, Grajales A et al. (2014) Hidden among sea anemones: the first comprehensive phylogenetic reconstruction of the order Actiniaria (Cnidaria, Anthozoa, Hexacorallia) reveals a novel group of hexacorals. PLOS ONE 9:e96998
Rosser NL, Thomas L, Stankowski S, Richards ZT, Kennington WJ, Johnson MS (2017) Phylogenomics provides new insight into evolutionary relationships and genealogical discordance in the reef-building coral genus Acropora. Proc R Soc B-Biol Sci 284:20162182
Rowan R, Powers DA (1991) A molecular genetic classification of zooxanthellae and the evolution of animal-algal symbioses. Science 251:1348–1351
Rowan R, Powers DA (1992) Ribosomal RNA sequences and the diversity of symbiotic dinoflagellates (zooxanthellae). Proc Natl Acad Sci 89:3639–3643
Rubin BER, Ree RH, Moreau CS (2012) Inferring phylogenies from RAD sequence data. PLoS ONE 7:e33394
Santos SR (2016) From one to many: the population genetics of cnidarian-Symbiodinium symbioses. In The Cnidaria, past, present and future (pp. 359–373). Springer, Cham.
Santos SR, Coffroth MA (2003) Molecular genetic evidence that dinoflagellates belonging to the Genus Symbiodinium Freudenthal are haploid. Biol Bull 1:10–20
Satler JD, Carstens BC (2017) Do ecological communities disperse across biogeographic barriers as a unit? Mol Ecol 26:3533–3545
Shearer TL, Van Oppen MJH, Romano SL, Wörheide G (2002) Slow mitochondrial DNA sequence evolution in the Anthozoa (Cnidaria). Mol Ecol 11:2475–2487
Shearer TL, Gutiérrez-Rodríguez C, Coffroth MA (2005) Generating molecular markers from zooxanthellate cnidarians. Coral Reefs 24:57–66
Shinzato C, Shoguchi E, Kawashima T, Hamada M, Hisata K, Tanaka M et al. (2011) Using the Acropora digitifera genome to understand coral responses to environmental change. Nature 476(7360):320–323
Shinzato C, Mungpakdee S, Arakaki N, Satoh N (2015) Genome-wide SNP analysis explains coral diversity and recovery in the Ryukyu Archipelago. Sci Rep 5:18211
Shoguchi E, Shinzato C, Kawashima T, Gyoja F, Mungpakdee S, Koyanagi R et al. (2013) Draft assembly of the Symbiodinium minutum nuclear genome reveals dinoflagellate gene structure. Curr Biol 23:1399–1408
Silverstein RN, Correa AM, Baker AC (2012) Specificity is rarely absolute in coral–algal symbiosis: implications for coral response to climate change. Proc R Soc Lond B: Biol Sci 279:2609–2618
Smith ML, Ruffley M, Espíndola A, Tank DC, Sullivan J, Carstens BC (2017) Demographic model selection using random forests and the site frequency spectrum. Mol Ecol 26:4562–4573
Titus BM, Daly M (2017) Specialist and generalist symbionts show counterintuitive levels of genetic diversity and discordant demographic histories along the Florida Reef Tract. Coral Reefs 36:339–354
Titus BM, Daly M, Macrander J, Del Rio A, Santos SR, Chadwick NE (2017a) Contrasting abundance and contribution of clonal proliferation to the population structure of the corkscrew sea anemone Bartholomea annulata in the tropical Western Atlantic. Invertebr Biol 136:62–74
Titus BM, Vondriska C, Daly M (2017b) Comparative behavioral observations demonstrate the spotted “cleaner” shrimp Periclimenes yucatanicus engages in true symbiotic cleaning interactions. R Soc Open Sci 4:170078
Titus BM, Palombit S, Daly M (2017c) Endemic diversification in an isolated archipelago with few endemics: an example from a cleaner shrimp species complex in the Tropical Western Atlantic. Biol J Linnaean Soc 122:98–112
Titus BM, Blischak PD, Daly M (2019a) Genomic signatures of sympatric speciation with historical and contemporary gene flow in a tropical anthozoan (Hexacorallia: Actiniaria). Mol Ecol 28:3572–3586
Titus, BM, Benedict C, Laroche R, Gusmao LC, Van Deusen V, Chiodo T et al. (2019b) Phylogenetic relationships of the clownfish-hosting sea anemones. Mol Phylogenetics Evol https://doi.org/10.1016/j.ympev.2019.106526
van Oppen MJ, Bongaerts P, Frade P, Peplow LM, Boyd SE, Nim HT et al. (2018) Adaptation to reef habitats through selection on the coral animal and its associated microbiome. Mol Ecol 27:2956–2971
Wham DC, LaJeunesse TC (2016) Symbiodinium population genetics: testing for species boundaries and analysing samples with mixed genotypes. Mol Ecol 25:2699–2712
Acknowledgements
We are grateful to Erich Bartels, Annelise del Rio, Jose Diaz, Dan Exton, Lisle Gibbs, Natalie Hamilton, Alex Hunter, Anna Klompen, Jason Macrander, Spencer Palombit, Stephen Ratchford, Nancy Sheridan, Nuno Simoes, Jill Titus, Cory Walter, Eric Witt, Clay Vondriska, and the Operation Wallacea dive staff for assistance in the field and laboratory. We also thank Jordan Satler, Megan Smith, and Bryan Carstens for bioinformatic assistance and advice regarding fastsimcoal2 analyses and model selection. Paul Blischak and Theo Cavinto provided valuable bioinformatics assistance separating holobiont from aposymbiotic RADseq loci. Bellairs Research Station, the Bermuda Institute of Ocean Science, Cape Eleuthera Institute, CARMABI, Coral View Dive Center, Gerace Research Centre, the Honduran Coral Reef Foundation, Mote Marine Laboratory, Smithsonian Tropical Research Institute, and the University of the Virgin Islands provided valuable logistical support in the field. Specimens were collected from throughout the Tropical Western Atlantic under permits: SE/A-88- 15, PPF/DGOPA-127/14, CZ01/9/9, FKNMS-2012-155, SAL-12-1432A-SR, STT037-14, 140408, MAR/FIS/17, and 19985. This research was supported by funding from a National Science Foundation-Doctoral Dissertation Improvement Grant DEB-1601645 and Florida Fish and Wildlife Conservation Commission awards to B.M.T. & M.D. Operation Wallacea, American Philosophical Society, International Society for Reef Studies Graduate Fellowship, PADI Foundation Grant, and American Museum of Natural History Lerner Gray Funds supported field research for B.M.T. Additional funding was provided through the Trautman Fund of the OSU Museum of Biological Diversity, The Ohio State University, and National Science Foundation DEB-1257796 to MD.
Author information
Authors and Affiliations
Contributions
BMT and MD conceived the study and collected samples, BMT conducted laboratory work and analyzed the data. BMT and MD wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Jane Hughes.
Supplementary information
Rights and permissions
About this article
Cite this article
Titus, B.M., Daly, M. Population genomics for symbiotic anthozoans: can reduced representation approaches be used for taxa without reference genomes?. Heredity 128, 338–351 (2022). https://doi.org/10.1038/s41437-022-00531-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-022-00531-3