Abstract
Evolutionary change is the change in trait values across generations, and usually occurs in multidimensional trait space rather than along isolated traits. Genetic covariation influences the magnitude and direction of evolutionary change and can be statistically summarized by the additive genetic (co)variance matrix, G. While G can affect the response to selection, it is exposed to evolutionary change by selection and genetic drift, but the magnitude and speed of these changes are poorly understood. We use comparative G matrix analyses to assess evolution of the shape and orientation of G over longer timescales in three species of Gomphocerine grasshoppers. We estimate 10 × 10 G matrices for five morphological traits expressed in both sexes. We find low-to-moderate heritabilities (average 0.36), mostly large cross-sex correlations (average 0.54) and moderate between-trait correlations (average 0.34). G matrices differ significantly among species with wing length contributing most to these differences. Wing length is the trait that is most divergent among species, suggesting it has been under selection during species divergence. The more distantly related species, Pseudochorthippus parallelus, was the most different in the shape of G. Projection of contemporary genetic variation into the divergence space D illustrates that the major axis of genetic variation in Gomphocerippus rufus is aligned with divergence from Chorthippus biguttulus, while the major axis of genetic variation in neither of the species is aligned with the divergence between Pseudochorthippus parallelus and the other two species. Our results demonstrate significant differences in G matrices with a phylogenetic signal in the differentiation.
Similar content being viewed by others
Introduction
The efficiency and direction of adaptive evolution are not only affected by the fitness landscape, but also by the amount and structure of heritable variation (Lande 1979; Teplitsky et al. 2014). While the univariate heritability acts mainly as an efficiency filter of inheritance among generations, the structure of genetic covariance among traits also affects the direction of the response (Lande 1979, 1982; Lande and Arnold 1983; Blows and McGuigan 2015). Genetic covariance arises either from pleiotropy or from linkage disequilibrium (Lynch and Walsh 1998). Since traits are often genetically coupled by such covariation, and therefore do not evolve in isolation, only a multidimensional view of trait evolution will capture the important intricacies of evolutionary dynamics that any unidimensional analysis will miss (Lande 1980b; Blows 2007; Walsh and Blows 2009). However, despite their relevance to the efficiency and direction of adaptive evolution, we know comparatively little about how genetic covariation evolves and how temporally stable genetic correlations are in natural populations.
The additive genetic variance–covariance matrix G offers a statistical summary of the amount and shape of genetic variation within populations and is integral to understanding multivariate evolution in quantitative traits (Lande 1979, 1980a). Among other things, G can be used to predict the evolutionary response to selection. The multivariate breeder’s equation Δz = Gβ represents the predicted change in mean trait values Δz in multivariate trait space as the matrix product of the additive genetic variance–covariance matrix, G with the vector of directional selection gradients, β (Lande 1979). It is apparent from this relationship that selection acting on a trait will usually produce an evolutionary response on traits that are genetically correlated, even though selection does not act directly on them. Similarly, the structure of G can constrain adaptive evolution by reducing the efficiency of adaptive responses or modifying the direction of response to selection (Teplitsky et al. 2014). While the mathematics of this relationship is simple and well worked out, empirical advance is hampered by the difficulty in estimating multivariate inheritance as well as selection in multivariate trait space (Turelli 1988; Arnold et al. 2008).
The genetic variance–covariance structure can change by natural selection, since selection in multivariate space is not solely directional at the level of individual traits, but also correlational on trait combinations (Lande and Arnold 1983; Phillips and Arnold 1989; Sinervo and Svensson 2002). Selection on G can be conceptualized by the analogy of the adaptive landscape (Arnold et al. 2001). The degree of the curvature at the hilltop of the multivariate adaptive landscape is defined by the strength of stabilizing selection and the orientation of any ridge by correlational selection. Stabilizing and correlational selection can be summarized in the γ matrix, the matrix of nonlinear selection gradients, where the diagonal elements contain the coefficients of stabilizing or disruptive selection γii and the off-diagonals of the coefficients of correlational selection γij (Phillips and Arnold 1989; Blows and Brooks 2003; Blows 2007). The effects of stabilizing and correlational selection on G can be particularly important when the covariances arise mostly from linkage disequilibrium rather than from pleiotropy (Arnold 1992). Adaptive changes in the shape of G can be caused by long-term weak selection or frequent periods of strong selection, but it is difficult to differentiate empirically between these processes (Roff 2000).
As well as being shaped by selection, the genetic variance–covariance matrix may change by random genetic drift (Roff 2000; Phillips et al. 2001; Steppan et al. 2002). Drift will mostly result in proportional changes in G matrix, but drift can also lead to changes in shape and orientation of G (Phillips et al. 2001). It is difficult to disentangle the effects of drift and selection as their signatures on the structure of G are very similar (Merilä 1999; Phillips et al. 2001). Furthermore, it is likely that over extended periods of time both selection and drift contribute to changes in G and it will thus be difficult to assign changes to one cause or the other. While it would be desirable to separate the influence of selection and drift on G matrices, their joint effect is of relevance to how contemporary populations may respond to selection and how persistent effects of genetic covariance are in affecting evolutionary change.
The key issue in the estimation of changes in the genetic covariance matrix is the small expected effect size and the substantial sampling variance in empirical quantifications of G. Comparative quantitative genetic studies offer a solution to this problem, because they capitalize on the cumulative change in G across many generations (Schluter 1996; Steppan et al. 2002). Comparative analyses can also shed some light on the contribution of selection to changes in G. Divergence among species is expected to be, at least partially, caused by past selection, hence any alignment between G and multivariate divergence matrix D (where D represents the matrix of phenotypic differentiation estimated as the (co)variance among mean trait values across species) will suggest an effect of selection (although the effect might go both ways). Misalignment, however, is suggestive for a role of genetic drift. Comparative studies thus help to give empirical answers to the persistent conundrums about the stability of genetic covariance structure in natural populations. Surprisingly few studies have used this comparative option, probably because of the difficulty in estimating G in multiple species (see below).
An intriguing special case of multivariate evolution is sexually dimorphic trait expression. Sexual dimorphism is very widespread and arguably represents the most conspicuous form of intraspecific phenotypic diversity. Sexual dimorphism ultimately arises from sex-specific selection, but is typically constrained by the shared genetic variation among females and males that results in intra-locus sexual conflict if selection is sex specific (Lande 1980b; Lande 1987). Unlike functionally unrelated traits expressed in the same sex, where we may expect low genetic correlations, the ancestral genetic correlation among homologous traits in the two sexes is expected to be large and only reduced by persistent sexually antagonistic selection (Poissant et al. 2010; Griffin et al. 2013). Such cross-sex genetic correlations cannot be shaped directly by correlational selection on the same trait expressed in males and in females when the traits (in gonochorous species) are never co-expressed in any single individual and correlational selection is thus absent.
The degree of stability of G matrix has been an enigmatic question in the evolution of quantitative characters (Steppan et al. 2002; Arnold et al. 2008). Theory and computer simulations predict that G can remain stable if the shape of the fitness surface is stable and the mutational covariance is neutral with respect to G (Turelli. 1988; Jones et al. 2003). However, since these conditions are likely to be violated over longer evolutionary time scale, G is bound to change (McGuigan. 2006). Among other things, the type of traits considered affects the stability of G. Fitness components, for example, are predicted to have unstable G matrices, while bilateral traits have a more stable structure of G due to high correlations between mutational effects and strong correlational selection on both sides (Jones et al. 2003). However, there is no general theoretical answer to the stability of G in natural populations, since factors and conditions that can stabilize or destabilize G are likely to coexist.
Empirical studies allow insights into the evolutionary dynamics of G. Previous studies report stable G among populations of the individual species for life history (Spitze et al. 1991; Delahaie et al. 2017), morphology (Brodie 1993; Delahaie et al. 2017), and behavior (Brodie 1993). In other cases, G matrices for morphology, life history, and behavior seem to differ between populations (Shaw et al. 1995; Doroszuk et al. 2008; Careau et al. 2015; Karlsson Green et al. 2016; Sniegula et al. 2018) or are differentiated by habitat or ecotype (Calsbeek et al. 2011; Eroukhmanoff and Svensson 2011). Similarly, longitudinal studies on single populations of passerine birds have reported either temporal stability for reproductive traits (Garant et al. 2008) or remarkable changes as for morphological traits (Björklund et al. 2013). Furthermore, experimental populations of inbred lines in Drosophila show divergence for morphological G matrices (Phillips et al. 2001). Between-species comparisons that build on longer divergence times are rather few. As for within-species comparisons, there is evidence for stability of G matrices (Bégin and Roff 2003) as well as evidence for G matrix divergence for morphology (Paulsen 1996; Roff and Mousseau 1999; McGlothlin et al. 2018) in among-species comparisons. Differences among studies might reflect differences in traits, ecological context, population history, etc. Overall, there seems to be substantial variation in outcomes, a little more evidence for G matrix divergence on longer timescales, but also some cases of rather rapid changes.
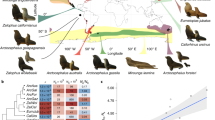
We measured five morphological traits and performed an among-species comparison to test empirically for G matrix stability in a group of grasshoppers. The traits measured are femur length, wing length, antenna length, eye height, and lobe height, separately in both the sexes. Specifically, we studied three species of grasshoppers from the subfamily Gomphocerinae, a clade of grasshoppers with about 230 species, world-wide distribution (Cigliano et al. 2017) and with rather conserved general morphology. Though exact divergence times are unknown, the Gomphocerinae have started to diversify about 30 Mya (Contreras and Chapco 2006). Within this clade we selected two closely related species, Chorthippus biguttulus and Gomphocerippus rufus, that show about 4.8% mitochondrial sequence divergence (Contreras and Chapco 2006). For comparison, we chose a more distantly related species, Pseudochorthippus parallelus (Nattier et al. 2011; Vedenina and Mugue 2011). We predict the genetic (co)variance structure to be more similar between the two more closely related species as compared with the more diverged species. Furthermore, we predict an alignment between G and the divergence matrix D if either selection has acted to align G with the vector of selection or if G has constrained the response to selection to be aligned with the main axis of genetic variation. We expect less alignment if drift has contributed substantially to G matrix divergence.
We performed a full matrix comparison among the three species using various matrix comparison tools (Roff et al. 2012). Comparisons advanced in five steps of increasing complexity. (1) We estimated heritabilities, shared environmental effects, and permanent environment for all five morphological traits in both males and females, and compared the contribution of sources of variances to phenotypic variation among species and sexes. (2) We compared the multivariate trait covariation among five morphological traits and compared them among the sexes and among species. (3) We compared between sex correlations within species to address the constraint to evolution of sexual dimorphism. (4) We used four different matrix comparison methods to assess the overall (dis)similarity in the structure of genetic (co)variation and how this similarity depends on phylogenetic relatedness. (5) We assessed whether the overall genetic variance–covariance structure is aligned with the phenotypic divergence among the three species. Our work contributes to the pool of empirical evidence of G matrix divergence over longer time scale through among-species comparison of morphological quantitative traits.
Materials and methods
Study organisms
We studied three species of Acridid grasshoppers, Chorthippus biguttulus, Pseudochorthippus parallelus, and Gomphocerippus rufus (for brevity we refer to them without the genus name in the following). All of the three species are sexually dimorphic in morphology: females are generally bigger than males, but wings are relatively shorter in females than in males and antennae show significant sexual dimorphism (Table S1). We collected biguttulus and parallelus from in and around Bielefeld, Germany (52°01′ N; 08°28′ E). These two species prefer different habitats: biguttulus inhabits dry grasslands, whereas parallelus is found in lush green meadows. However, both species have wide ecological amplitude and co-occur in many places. The third species rufus was collected from Tübingen, Germany (48°30′ N; 09°04′ E), where it occurs on semi-open slopes with tall grasses and herbs. Only final (fourth) instar nymphae were caught from the field to ensure virginity. Nymphae were kept in netted plastic cages and provided with grass as a food source. Upon emergence as adults, virgin males and females were kept separately in large mesh cages (47.5 × 47.5 × 90 cm3).
Breeding design
We set up a paternal half-sib breeding design in the laboratory at Bielefeld University. Each male was mated to two virgin females to form paternal half-sib families. Females were housed and paired separately in mesh cages (22 × 16 × 16 cm3). Males were swapped between their assigned female cages every 2–3 days until the male died. Females that died were replaced by new virgin females if the male was still alive. Each cage was provided with sand pots as egg-laying substrate. The sand was sieved once per week for egg pods (ca. 1 cm long solid structures containing typically 6–12 eggs, and only occasionally more (Chakrabarty et al. 2019)). Egg pods were then collected and kept in petri dishes lined with filter paper. Each pod was kept on a separate dish and dishes were sprayed regularly to keep the eggs moist. Egg dishes were kept in refrigerators at 0–10 °C for a period of at least 3 months starting from October.
F1 animals
Petri dishes with egg pods were taken out of the refrigerators in five cohorts between early January and early June. Upon hatching, individuals from the same egg pod were kept in the same mesh cage (dimensions: 22 × 16 × 16 cm3) and were provided with ad libitum food (freshly cut grass provided in a vial with water). Egg pods produced on average 6 hatchlings per pod and the mean and SD of surviving hatchlings per egg pod were: 3.21 ± 2.28 in biguttulus, 4.13 ± 2.35 in rufus, and 3.13 ± 2.10 in parallelus. Hatchlings were kept at a temperature of 25–30 °C and 35–55% relative humidity. Newly emerged adults were individually marked with numbered bee tags and transferred to larger communal cages (dimensions 43.5 × 43.5 × 93 cm3) in groups of ~25 individuals. The number of F1 animals that survived to adulthood were 1237 biguttulus, 897 rufus, and 390 parallelus that hatched from 383 egg pods in biguttulus, 217 in rufus, and 120 in parallelus. The total number of full-sib families was 112 in biguttulus, 70 in rufus, and 66 parallelus. The numbers of half-sib families were: 73 for biguttulus, 48 rufus, and 51 in parallelus. The number of males in biguttulus is 656 and number of females is 581, in rufus is 463 and 434, and in parallelus is 195 and 195.
Morphometrics
Standardized photographs of both males and females (adults only) from F1 generation were taken after collection or natural death. Hind legs, forewings (tegmina), and antennae were first detached from the main body. Photographs of the antennae, forewings, and hind legs were taken on a white background with a scale next to the body parts. For pictures of pronotal lobes and eyes, the animal was placed on a dish full of fine-grained sand with a scale next to it. The sand allowed adjusting the body for plain dorsal and lateral views. An artificial light source was used for the photographs, which were all taken by a Fuji camera (FinePix HS35 EXR). Pictures were analyzed using the software ImageJ 1.46r (Schneider et al. 2012). We measured postfemur length, forewing length, length of the antennae, height of the pronotal lobes, and the vertical diameter of the eye. We refer to those as femur length, wing length, antenna length, lobe height, and eye height in the following. All traits were sexually dimorphic with weakest sexual dimorphism in eye height (Table S1). Each measurement was calibrated using a 20 mm scale.
Individuals of two species rufus and biguttulus are all long-winged, while the wings are short in parallelus. Specifically, female parallelus shows forewings about half the length of the abdomen and hindwings that are reduced to stubs. Male parallelus also shows very narrow, short hindwings, but slightly longer forewings that cover most of the abdomen. Both sexes of parallelus are incapable of flight, which would require developed hindwings. However, most natural populations harbor individuals with fully developed fore- and hindwings at low frequencies, and this wing length polymorphism is apparently environmentally induced (Harrison 1980; Ritchie et al. 1987). These individuals with fully developed wings are called macropterous. While all our founding individuals were short-winged, we found 13 macropterous individuals (1.8%) from seven different half-sib families among the offspring. Because these individuals represented outliers with respect to the normal distribution of wing length in the population (Fig. S1), we excluded the wing lengths of these individuals from the analysis of this trait. We also performed an analysis that excluded macropterous individuals entirely, but this hardly affected the results (Table S5).
Statistical analyses
All models were fitted in R 3.5.1 (R Core Team 2018) under a Bayesian framework as implemented in the MCMCglmm package (Hadfield 2010). We fitted multivariate animal models and extracted the posterior distribution of G as well as other (co)variances. For each individual trait the key components to the model were
where y is a vector of trait values, β is the vector of fixed effects, a is the vector of additive genetic effects, s is the vector of shared environmental effects, i is the vector of individual identity effects (individuals have two measurements for each bilateral traits), and e is the vector of residual errors. The shared environmental effect is the part of the phenotypic variance originating from individuals sharing the same egg pod (hence also the same mother) and shared rearing conditions in the same nymphal cage. The individual identity effect is the part of the total variance that is reproducible between sides within individuals (hence excluding most measurement error and fluctuating asymmetry) beyond similarity among siblings. The residual component consists of measurement error and fluctuating asymmetry between the two sides. Z1, Z2, and Z3 are the respective incidence matrices for three vectors of random effects (based on pedigree relationships, egg pod identities, and individual identities, respectively) and X is the design matrix for the fixed effects. Heritabilities were calculated with the residual component removed, since we were not interested in fluctuating asymmetry and the individual repeatable component is deprived of measurement error and thus closer to the genetic signal.
We fitted ten-trait models (five morphological traits expressed in both sexes) for all species in order to estimate 10 × 10 G matrices (and we will refer to the dimension of G as n in the following). Side was fitted as a fixed effect with two levels (coded as numeric, left = −0.5 and right = 0.5). For the estimation of individual identity effects, we split the 10 × 10 matrix into two 5 × 5 matrices for males and females, because within-individual covariation cannot exist between the sexes. By doing so, individual-specific covariances between the sexes were effectively constrained to zero. We estimated the between sex correlations as well as the within sex between trait correlations from the above models.
We used parameter expanded (half Cauchy) priors for our model as they are less informative than regularly used inverse-Wishart priors (Gelman 2006). The reason for using a weakly informative prior is to ensure that the information comes chiefly from the data. The degree of belief parameter ν was set to ν = 11 for all the random effects except for the individual identity. We also fitted models with ν = 9, ν = 10, and ν = 12 for sensitivity assessment, but the choice of ν had rather little influence on critical measurements (see Figs. S5–S7). For individual identity, the 10 × 10 matrix was split into two 5 × 5 matrices, one for males and one for females with ν = 6 for each matrix. The ν for residual effects was set to 0.002. The posterior distribution of each model was estimated from 1,100,000 MCMC iterations with a thinning interval of 1000 and a burn-in period of 100,000. We ran two independent chains per model yielding a total of 2 × 1000 samples form the posterior distribution for each parameter. Model convergence was visually inspected using the trace plots and using Gelman and Rubin diagnostics (Gelman and Rubin 1992).
Matrix comparisons
We used four established matrix comparison methods that allow exploring different aspects of variance–covariance matrices with subtly different inferences. The Krzanowski’s subspace analysis determines whether the subspaces containing maximum genetic variation are similar across species. The random skewer analysis allows evaluating differences in orientation of G matrices. The Flury hierarchy analysis implements a formal assessment of the number of shared eigenvectors. The tensor analysis quantifies specifically the differences among variances and covariances across G matrices.
We compared raw matrices on the original scale, because trait units were identical and size difference were not excessive. However, in order to remove size effects, we also compared matrices after division by the square of trait means (Houle 1992) to account for allometric scaling with size. Results for matrix comparisons on mean-standardized variance–covariances matrices were qualitatively similar to unstandardized matrices.
Krzanowski’s common subspace analysis
One approach to compare genetic architecture is to identify the part of G containing most of the genetic variance and to test whether this part overlaps among species. Krzanowski’s subspace analysis is a method to evaluate which part of G contains the maximum variance and whether the eigenvectors explaining most of the genetic variance is similar across species/populations (Krzanowski 1979; Aguirre et al. 2014; Gosden and Chenoweth 2014). The similarity among subspaces of G matrices that captures the greatest amount of genetic variance can be tested using this analysis. The common subspace H among the p = 3 species is given by (Krzanowski 1979)
where t = 1, …., p indexes species and At contains the subset of kt eigenvectors of Gt as columns and k is an integer smaller than n (the dimensions of G) that is chosen a priori (see below). These k vectors define the dominant subspace of the three G matrices and only a k dimensional comparison is interpretable as adding any further dimension will make that orthogonal to one of the species subspaces (k = min (ki), = 1, …, p) (Krzanowski 1979). We chose to analyze the common subspace using a fixed number of k = 5 eigenvectors of G following previous studies that used k equal to half the size of the original G matrix (Aguirre et al. 2014; Gosden and Chenoweth 2014).
We conducted an eigendecomposition of the H matrix, where the five eigenvectors hi of H contain the genetic variation of the linear trait combination that is shared across the three species. The eigenvalues of H can reach a maximum value of p = 3. At the limit, an eigenvector with an associated eigenvalue of 3 shows that a linear combination of traits completely explains the genetic variance that is shared across the three species (Gosden and Chenoweth 2014). A departure of an eigenvalue from this maximum value of p = 3 shows that the trait combination of the corresponding eigenvector of H cannot be rebuilt from the k = 5 eigenvectors of G in at least one of the species. This indicates that the dominant part of the G matrix of at least one species is not perfectly aligned with the eigenvector of H (Aguirre et al. 2014; Gosden and Chenoweth 2014).
The difference in alignment can be measured by the angle δt between each eigenvector of H and the subspaces of species t (Krzanowski 1979; Gosden and Chenoweth 2014)
This comparison when done under a Bayesian framework can use the samples form the posterior distributions of G matrices, and thus gives a measure of uncertainty in estimates (Aguirre et al. 2014). In order to test statistically whether the observed differences among the G matrices of the three species are caused by sampling variance, we compared the observed data from the subspace analysis with a null model where we expect the G matrix differences are solely due to random sampling. Randomized G matrices were created from the posterior predictive breeding values of the observed set of G matrices followed by an estimation of the covariance of breeding values among traits. p values were calculated as the proportion of randomized samples that show equal or smaller values than the original MCMC samples. The R code for the analysis was adapted from Aguirre et al. (2014).
Random skewers analysis
Random skewer analysis is a method for comparing the population-level consequences of matrix differences (mostly in shape) when a population is exposed to random linear selection gradients (Cheverud 1996; Cheverud and Marroig 2007; Roff et al. 2012; Aguirre et al. 2014). This method primarily compares differences in matrix orientations. Random skewer analysis builds on the multivariate breeder’s equation, Δz = Gβ, where Δz is the vector of trait changes and β is a vector of selection gradients, and makes use of its biological interpretation. Randomly generated selection vectors β are projected through the G matrices, and the predicted response to selection, Δz, is calculated. The angle θ between Δz with β quantifies the amount of deflection and the angle between two Δz from different matrices measures the difference in the direction of deflection when exposed to the same vector of random selection gradients.
We generated 1000 random selection vectors sampled from a multivariate normal distribution with uncorrelated axes. All 1000 random skewers were projected through MCMC samples of each G matrix generating a posterior distribution of response vectors (applying the same 1000 skewers to the two chains). The angle between response vectors was calculated as
where v1 and v2 are the two vectors to be compared. Since angle calculations were done on each MCMC sample, we extracted 2000 angles that together represent the posterior distribution of the estimate.
Flury hierarchy analysis
Flury hierarchy is an approach of matrix comparison where a series of models are built and ranked, starting from matrix inequality to equality (Flury 1988; Arnold and Phillips 1999; Phillips and Arnold 1999; Steppan et al. 2002; Roff et al. 2012). It is a test of overall similarity of matrices through a series of hierarchical tests. One of the most important contributions of Flury hierarchy in comparing G matrices is the stepwise analysis of matrix similarities and differences. As matrix comparison is a multivariate exercise, there are several possible states between the extreme conditions of matrix equality and inequality (Arnold and Phillips 1999). Besides this broad classification of equal or unequal, matrices can differ in other characters like difference in eigenvalue or eigenvector. If the matrices have similar eigenvectors but their eigenvalues differ by a constant, then the matrices are said to be proportional. Matrices can also differ in having different eigenvalues but having all eigenvectors in common. This is tested by the common principal component model (CPC) that assumes eigenvectors to be identical. Sharing of eigenvectors can also be partial, which is tested by partial CPC models (Phillips and Arnold 1999) that allow for 1 to n−2 eigenvectors to be identical between matrices, where n is the dimension of the G matrices.
An approach for comparing the model fit by Akaike information criteria (AIC) was described by Flury (Flury 1988; Phillips and Arnold 1999). AIC adjusts the log-likelihood of a particular model for the number of parameters used to fit a particular model. Models with smaller AIC values are considered better fits. We used the R package ‘cpc’ (Pepler 2015) to perform the Flury hierarchical tests using the model building approach, on the three species-specific G matrices. We reiterated tests for our 2000 MCMC samples to get a posterior distribution of AIC values and ranked models by average AIC.
Genetic covariance tensor analysis
Genetic covariance tensor analysis is a method to explore and determine directions in which divergence in G matrices occur among populations or species (Hine et al. 2009; Aguirre et al. 2014). It quantifies the variance in (co)variances across G matrices and thus offers a quantitative summary of matrix differences. The structure of those variances in (co)variances can then be analyzed by eigenanalyses.
In multilinear algebra, covariance tensors are fourth order arrays that are used to define variation in lower order variables like vectors (first order tensors) and matrices (second order tensors). The covariance structure of a set of traits of a single species as summarized by a two-dimensional G matrix thus represents a second order tensor with elements indexed by i and j for the 1 to n traits. If more than one species are concerned, then the covariance elements of G can be characterized by a four-dimensional genetic covariance tensor Σ, which is indexed by i, j, k, and l each varying from 1 to n traits in two or more G matrices (traits indexed i and j in one matrix and k and l in the other). Elements of Σ are thus defined by the covariance in (co)variances of multiple G matrices as (Hine et al. 2009)
The genetic covariance tensor Σ can also be summarized by a symmetric matrix S with dimensions m × m, where \(m = \frac{{n(n + 1)}}{2}\). The S matrix summarizes Σ in 2D format (see Hine et al. 2009 for details).
An eigenanalysis of Σ can be done in a similar fashion like an eigenanalysis of a G matrix, except that what represents an eigenvector in the case of an eigendecomposition of a two-dimensional G is represented by a two-dimensional eigentensor (a matrix, symbolized by E) in the case of eigendecomposition of a four-dimensional Σ. As in eigenanalysis of G, each eigentensor E is associated with an eigenvalue that quantifies how much variation among G matrices is captured by each E (Hine et al. 2009; Careau et al. 2015). The maximum number of nonzero eigenvalues of Σ is \(\frac{{n(n + 1)}}{2}\) or (p−1), whichever is smaller, where n is the dimensions of the G matrices and p the number of G matrices to be compared (Hine et al. 2009; Aguirre et al. 2014). In our study, this number is p − 1 = 2. Further exploration of the variation among G matrices can be done using orthogonal linear combination of traits, which portray the independent changes among G matrices, by eigenanalysis of the eigentensor E where the eigenvectors of E are denoted as e. If the largest eigenvalue of an E is close to 1, then the change in covariance pattern as defined by the eigentensor can be attributed to the change in VA for a particular trait combination.
We used a Bayesian framework, as outlined in Aguirre et al. (2014), to determine which independent facets of the genetic covariance structure as described by the tensor show significant variation among three species of grasshoppers. We determined Si, the matrix representation of a tensor for the ith MCMC sample of the set of three G matrices and extracted the posterior mean of S based on all 2000 MCMC samples and estimated the variance among G matrices αij, explained by each eigenvector. This enabled us to calculate the amount of additive genetic variation VA in the direction of greatest genetic variation among the three species-specific G matrices (Careau et al. 2015). The posterior distribution of αj contains the uncertainty in the variance of the covariance structure as described by each eigentensor E. This posterior distribution of αj is then compared with the posterior distribution extracted from the null model, where the variation among matrices is solely caused by sampling variation after randomizing breeding values.
Alignment of G with D
We quantified phenotypic divergence in morphospace among the species as a proxy for the long-term direction of evolutionary change in the past (Schluter 1996). We constructed a species mean trait value variance–covariance matrix D across all three species and the ten sex-specific traits. Since subjects were raised in the same environment, phenotypic means are expected to be representative for genetic divergences among species. We used eigenanalysis of the D matrix to quantify the first and second principal axes of species divergence. This can be done separately by sex (with five traits in each sex) and for sexes pooled (ten traits, five in each sex). Both are of interest because the first one captures sex-specific divergence and the second captures species-specific divergence. Breeding values estimated from our MCMCglmm animal model were then projected into divergence space for display.
Results
Heritabilities
We estimated heritabilities and other variance components (Fig. 1) for ten sex-specific traits from multivariate animal models. Heritabilities averaged h2 = 0.36 (Table 1). Females tended to show lower heritabilities than males (0.32 in females vs. 0.39 in males). Though the heritabilities of the sexes overlapped on a trait-by-trait basis, we find the same tendency replicated over most traits and species (Table 1).
Genetic correlations
Genetic correlations between traits within sexes were moderate (average rG = 0.34) with strongest correlations in parallelus (average rG = 0.45) and lowest in rufus (average rG = 0.23, Table 2). Correlations tended to be higher in males (average rG = 0.38) than in females (average rG = 0.30) and in traits that involved femurs, lobes or eyes (rG = 0.35–0.43) than correlations that involved wings or antennae (rG = 0.24).
We estimated cross-sex genetic correlations rMF between males and females in all three species. Cross-sex correlations were moderate and positive (average rMF = 0.54, Table 2), were similar across species (lowest in parallelus, average rMF = 0.48, highest in biguttulus, average rMF = 0.58) and higher for femur, lobes, and eyes (rMF = 0.58–0.66) than for wings and antennae (rMF = 0.40–0.44). Among-trait correlations across sexes were weaker (average rG = 0.27) than correlations within traits across sexes and across-traits within sexes, with correlations involving femur, lobes, or eyes being higher (rG = 0.28–0.36) than correlations involving wings or antennae (rG = 0.16–0.18) (Table S3).
Subspace analysis
We used the first five eigenvectors of G (together explaining 97% of the variance in G in biguttulus and parallelus and 96% in rufus) to test whether the dominant subspaces A of G were shared among the three different species. Eigenvalues were compared with a value of 3 that would indicate identical subspaces. In order to accommodate sampling variance, we compared estimated with randomized eigenvalues. The first four eigenvalues of H were significantly lower than 3, which indicate significant differences in orientation among the G matrices of the three species (Fig. 2). The angle between each eigenvector of H and each of the k subspaces of G shows only minor differences among species (Table S8). If the angles are close to zero, the specific eigenvector of h is closer to the species subspace and explains the genetic variance better for that subspace. The overlap of the credibility intervals for the angles shows that there is little difference in degree of divergence among the subspaces, though the subspaces themselves have diverged in orientation.
Krzanowski’s subspace H for the comparison of G matrices among three species of grasshoppers. The x-axis denotes the five eigenvectors h1–h5 of the H matrix and the y-axis denotes the eigenvalues of H. Filled symbols show empirical estimates with 95% CI and open symbols show randomized values. P values denote the proportion of randomized values that show equal or lower values than empirical estimates (incorporating variability in both empirical and randomized values)
Random skewer analysis
The random skewer analysis showed marked difference of genetic covariance matrices across three species. Based on 2000 random skewer projections, the angle of deflection in biguttulus was 56.9° ± 10 (36.9–75), in rufus it was 57.3° ± 10 (35.5–74.8), and in parallelus the angle was 60.2° ± 9.9 (39–76.4) with no significant differences among them (Fig. S13). Hence, response vectors were deflected to a similar magnitude in all species. We also calculated angles between predicted response vectors of different species to evaluate if they were deflected in a similar direction. The mean angles between the response vectors of biguttulus and rufus was 51.5° ± 19.7 (19.2–93.3), between biguttulus and parallelus was 53.9° ± 21.2 (19.1–102.6) and that between rufus and parallelus was 49.6° ± 20.2 (17.6–94.3). The direction of deflection was thus markedly different in all pairwise comparisons (all > 49°). However, the random skewer analysis was associated with large uncertainties. Even randomized samples from the posterior distributions of response vectors produced average angles of 38.3° ± 18.1 (12.2–80.8) for biguttulus, 38.1° ± 18.7 (12.4–83.6) for rufus, and 37.1° ± 21.6 (10.4–95) for parallelus.
Flury hierarchy analysis
The Flury hierarchy analysis for the three matrices reveal that the G matrices are not equal or proportional, or share any CPCs, as the best fitting model is the one of unequality or heterogeneity. Models with CPCs yielded significantly worse fits with AIC increasing almost steadily with the number of CPC being added. Hence, the matrices do not show stability in terms of the stability of eigenvectors.
Genetic covariance tensor analysis
Genetic covariance tensor analysis estimates (co)variation among elements of G across species. In our study with three species-specific G matrices, the maximum possible number of nonzero eigenvalues of the genetic covariance tensor is 2. The first eigentensor E1 explains 71% of the variation among G matrices (Fig. 3, Table S9). The 95% CI of the eigenvalues did not overlap between observed and randomized G matrices for both the eigentensors E1 and E2, illustrating significant variation among matrices. Further, the eigenanalysis of the eigentensor showed that the first eigenvector e11 of E1 explains 74% and first eigenvector e21 of E2 explains 40% of variation in eigentensors. Wing length loads on both eigenvectors in particular in the mean-standardized analysis (Table S9). This suggests that wing length contributes most to the major axis of variation among G matrices compared with other traits, and that the matrices must have diverged along wing length. The second eigenvector e22 of the second eigentensor E2 captures 39% of the variation in E2. Hence the two eigenvalues of substantial size suggests that the independent genetic change represented by E2 occurs mainly in two genetically independent trait combinations (Table S9) (Hine et al. 2009). The projection of the eigenvectors onto the observed G matrices showed that the change in genetic variance represented by e11 and e21 is mostly attributable to parallelus. Though the credibility intervals overlap, there is a trend especially for e11 (Fig. 3). Hence, along e11, there is differentiation among parallelus on the one hand and the species pair biguttulus/rufus on the other, and this change in VA probably bears the signature of divergence of parallelus from the latter two.
Genetic covariance tensor analysis. a Variance in variation among G matrices as explained by the two eigentensors E1 and E2 of the covariance tensor Σ along with credibility intervals. Observed values (filled symbols) were compared with values after randomization (open symbols). Eigenanalysis of each E identifies the major axis of genetic variation among G matrices. Figures show the amount of additive genetic variance in each species along b the major axes e11 of E1 and c the major axis e21 of E2
Divergence analysis
Species divergence can be summarized by the divergence matrix D in mean trait values (Table S6). We used eigenanalysis of D to summarize the main axes of divergence. The first two eigenvalues of D explained the majority of species divergence (93% and 7%, respectively). Wing length loaded heavily on the divergence axis 1 and antenna and femur on the axis 2 (Table S7). We projected contemporary genetic variation as summarized by G into the main axes of historical divergence space (Fig. 4). Rufus is most strongly aligned with the second principle component axis that captures mostly the differences rufus/biguttulus, whereas biguttulus has a less pronounced alignment with least structure, which is indicative of the weak genetic correlations. Contemporary genetic variation in parallelus is oriented away from divergence axis 1, but the breeding value ellipse illustrates high genetic correlations.
Contemporary genetic variation projected into divergence space among the three species of grasshoppers. The axes show the first two principal components of the species divergence matrix D (Table S7). Breeding values of the three species are plotted onto the same plane. Ellipses around breeding values show 95% confidence level. Lines indicated the main direction of divergence as they connect the center of the more closely related Chorthippus biguttulus and Gomphocerippus rufus as well as the midpoint of these two species to the center of Pseudochorthippus parallelus
Discussion
We estimated and compared the genetic (co)variance structure of 10 × 10 G matrices consisting of five morphological traits expressed in both sexes in three grasshopper species. The subspace analysis shows significant difference in dominant subspaces among G matrices. The random skewer analysis also suggests marked differences in deflection angles, although this is accompanied by large uncertainties. In line with this, the Flury analysis identifies no shared principle components. The tensor analysis indicates difference in shape and orientation of G matrices with most standing genetic variation in the direction of G matrix divergence in Pseudochorthippus parallelus. It also identifies wing length as the most influential trait. The same pattern is also seen in the divergence analysis. Furthermore, the divergence analysis illustrates that the main axis of species divergence is only partly aligned with genetic variation within species. It is mostly Gomphocerippus rufus that clearly shows most genetic variation in the direction of divergence from Chorthippus biguttulus, while genetic variation in parallelus is oriented away from the direction of divergences from the other two species. Overall, these matrix comparisons illustrate rather substantial differences in G matrices. The comparisons reveal a phylogenetic signal that related species are more similar in their G matrices, that alignment with the main axis of divergence is prominent only in the youngest species pair, and finally they identify wing length as a key trait that contributes substantially to matrix differences.
The motivation for the comparative analysis builds on the assumption that the divergence axis is an indicator of past selection (Schluter 1996). Under these conditions, the comparison of species divergence and standing genetic (co)variation allow insights into the alignment of the main axis of G with the direction of selection. There are three reasons why the main axis of genetic variation, gmax, might be aligned with the main axis of selection. First, divergence might be faster and more efficient in the direction of gmax and that the divergence axis therefore represents a compromise between the forces of selection and the influence of genetic covariance (axis of least genetic resistance hypothesis, (Schluter 1996)). Second, the genetic covariance structure might be shaped by correlational selection to align with the dominant direction of selection (Lande and Arnold 1983; Phillips and Arnold 1989; Sinervo and Svensson 2002). Finally, the axis might be aligned by chance.
Despite these reasons for alignment, our observation does not support this. Instead, the results suggest that any alignment is a matter of temporal scale. There is some indication for alignment between rufus and biguttulus, in particular in the genetic covariance structure of rufus. On a larger temporal scale, the more distantly related G matrix of parallelus is poorly aligned. This is expected for example if the direction of selection might have fluctuated since the split between species and we do not know if it is the more recent history of selection that dominates the shape of G or the average long-term pattern of selection (Steppan et al. 2002; McGuigan 2006; Careau et al. 2015). Previous studies on within-species comparisons that illustrate differences in G (see ‘Introduction' section) suggest that changes in G can be strongly affected by the recent past. On the other hand, there are also studies on congeneric species that suggest alignment over longer timescales (Schluter 1996). Only additional empirical results on different species will allow a better understanding of which trends predominate and how the peculiarities of every individual system affect the outcome.
We found evidence for a phylogenetic signal in the G matrix divergence among species, such that the G matrix for the most distantly related species is the least similar. Such a phylogenetic signal is not always observed in comparative G matrix analyses. The G matrices of crickets, for example, do not diverge according to phylogenetic relatedness and are overall rather similar (Bégin and Roff 2004). Specifically, we found alignment with the main axis of divergence only in the youngest species pair, but not with respect to a more distantly related species. Previous studies have reported an alignment of the divergence with the genetic variation in deeply diverged sets of Anolis lizards (McGlothlin et al. 2018) as well as in a much younger radiation of ecotypes in plants (Walter et al. 2018). Overall, there is still little data on whether G matrix similarity reflects phylogenetic relatedness, but there is some evidence for alignment of divergence with genetic covariation. Our data suggests that both G matrix shape and alignment with the axis of divergence are a matter of temporal scale.
Wing length turned out to be a particularly influential trait in our analyses. It is one of the longest structures that we have included in our study (Table S1) and it might seem intuitive to assume that the size itself is causing the dominance on G matrix divergence. However, the same patterns are also present in the mean-standardized analysis, illustrating that its influence is not only due to the scaling of the variance with the mean (Houle 1992), but that variation in wing length is large even when accounted for its size. The length of the wings is among the most variable characters of grasshoppers, with long wings in some species (such as biguttulus, but also others with even longer wings) and wings significantly reduced in others (such as parallelus). Wing length is also highly variable within species, and tends to show substantial sexual dimorphism. Males typically have longer wings and they use their forewings for stridulation (Uvarov 1966). Females do not produce advertisement songs and their wing length is often reduced compared with males. Finally, wing length is sometimes polymorphic within populations independent of sex (Harrison 1980; Roff 1986a; Roff and Fairbairn 2007), further illustrating that it varies independent of other morphological traits. Wing length thus seems to respond quickly to selection and is genetically decoupled from other morphological traits.
We used four formal matrix comparison methods and projection into divergence space to compare matrices. Although other methods exist (Roff et al. 2012; Teplitsky et al. 2014), our analysis represents a very comprehensive exploitation of matrix comparison methods. Overall, results were consistent across methods, but there are also relevant differences. The random skewer analysis was indicative of differences among G matrices, but was accompanied by large sampling variation suggesting relatively low power. Low power of the random skewer analysis has also been reported in simulation studies (Teplitsky et al. 2014). On the other hand, the Flury analysis seems to indicate rather substantial differences in the covariance structure of the matrices, with none of the matrices having any CPCs. It seems possible that the Flury analysis overemphasizes differences between matrices. The subspace and the tensor analysis seem to be most nuanced, illustrating significant differences without dismissing the similarities that do exist among matrices. Both methods also suggest that parallelus contributes most to these differences. The tensor analysis furthermore identifies the traits contributing most to differences among G matrices.
Our analysis is one of the first of its kind to show the distribution of breeding values in divergence space (but see (McGlothlin et al. 2018)). Phylogenetically, biguttulus is more closely related to rufus than parallelus (Vedenina and Mugue 2011). Using average trait values of the more closely related species pairs implicitly applies an ancestral state reconstruction that may be very simple, but offers a point of comparison for the more distantly related species. The divergence analysis shows that the two ellipses for rufus and biguttulus are aligned in the same direction, compared with parallelus, which is oriented away from the divergence axis. Phenotypic divergence is used as a surrogate for past selection (Schluter 1996), although it is evidently the consequence of selection and drift and also influenced by the (unknown) ancestral shape of G.
Conclusions about G matrix similarities are also influenced by the choice of traits. In our analysis, we treated traits in females and males as sex-specific traits. As expected, correlations among traits expressed in the two sexes produce rather strong cross-sex genetic correlations and hence structure in G that is similar across species. The decision to treat traits as sex specific thus influences the amount of structure in G and the similarity in comparisons. Furthermore, our analysis is focused on bilateral traits. We treated the two sides as replicate observations that were effectively averaged for the same trait as a tool to reduce measurement error. However, we could have treated left and right sides as distinct traits which would have created strong structure in G, since genetic integration almost certainly produces strong genetic correlations among sides and again this structure would be shared among species. The choice to either treat sides as distinct or the same trait most likely influences the outcome of G matrix comparisons.
Statistical power is always a concern when estimating genetic covariation. We here report G matrices based on decent sample sizes of around 900–1100 for two of the species, but clearly less for the third species. However, our results also illustrate that it is not only the sample size that determines the outcome. We describe biologically expected patterns in the structure of the G matrix that are unlikely to be produced with insufficient data. In particular, we find a ranking of genetic correlations being highest for the same traits expressed in the two sexes, lower for genetic correlations among traits within sexes and lowest for correlations among traits among sexes. Furthermore, we show that the magnitude and precision of estimates depend on the magnitude of both heritabilities and of genetic covariation (Figs. S8 and S9, Table S11). Hence the structure of genetic variation partly determines the precision of estimates independent of sample size. The structure of genetic variation is not under the experimenter’s control, making it difficult to perform power analysis if the magnitude of genetic (co)variation is unknown.
Of special interest in G matrix analyses are the cross-sex genetic correlations, since they are relevant to the evolution of sexual dimorphism. Low cross-sex correlation in homologous traits suggests that sex-specific selection is at work to a certain degree (Chenoweth and Blows 2004; Day and Bonduriansky 2004; Bonduriansky and Rowe 2005). This helps the sexes to achieve their sex-specific optima (Bonduriansky and Chenoweth 2009). A review of cross-sex correlations reported a mean rMF of 0.80 for morphological traits predicted when there is no sexual dimorphism (Poissant et al. 2010). Our study shows that the evolution of sexual dimorphism might be somewhat impeded by covariation, although the constraint is not absolute as is illustrated by the non-perfect correlations (average 0.54) and the existence of sexual dimorphism in all traits. Wing length and antenna length are the two traits with lowest rMF (average 0.44 and 0.40, respectively). Wings are involved in sound production in grasshoppers and show marked sexual dimorphism in many species (Roff 1986a; Gäde 2002; Rosetti and Remis 2018) including some crickets and bush crickets (Roff 1986b; Roff and Fairbairn 1993; Heidinger et al. 2018). The antennae are also involved in courtship display in particular in rufus (Riede 1986). Both traits are thus likely to be the target of sex-specific selection and even tend to show reverse sexual dimorphism by being larger in otherwise smaller males (Table S1).
Besides genetic variation, we find substantial amounts of individual identity effects illustrating a large amount of phenotypic flexibility. This is not too surprising for grasshoppers that often show strong developmental plasticity, for example in temperature-dependent variation in overall body size and life-history traits (Hinks and Erlandson 1994; Willott and Hassall 1998). Despite the common garden situation, there is always some heterogeneity in conditions such as local temperature, food quality, and competition that might affect the development. Idiosyncratic effects might arise in particular if particular environmental stages are sensitive to have a disproportional effect on final body size. The first nymphal stages in particular seem to be quite variable in molting times, activities, and growth, possibly representing a critical stage during development.
An auxiliary finding of our study is the spontaneous occurrence of macropterous parallelus in a population of exclusively short-winged adults. Phenotypic plasticity in wing length is well documented in grasshoppers (Roff and Fairbairn 2007; Forsman 2015). The fact that macropterous individuals were neither clustered in genetic families nor in rearing cages suggests that neither a simple genetic mechanism nor any strong environmental trigger produces macropterous individuals. Crowding has been implicated in macropterism in a number of Orthopterans (Harrison 1980). Although not designed for this purpose, our data does not suggest a role of crowding in wing dimorphism of parallelus.
Overall, the structure of genetic variation is remarkably variable among the three species. Notably, however, the G matrices of biguttulus and rufus, the more recently diverged species pair, are better aligned to the direction of divergence in morphology, while the more distantly related parallelus is not. Wing length is contributing most to these differences and wing length is also the trait that is most variable between and within species, including variability among the sexes. It is thus the trait that is most affected by both population-specific and sex-specific selection. This might indicate a role of selection shaping G matrix differences. Our analysis thus illustrates that the structure of the G matrix can be variable when assessed over longer evolutionary timescales even for largely conserved morphological traits.
Data archiving
Data available from the Dryad Digital Repository: https://doi.org/10.5061/dryad.rjdfn2z5x.
References
Aguirre J, Hine E, McGuigan K, Blows M (2014) Comparing G: multivariate analysis of genetic variation in multiple populations. Heredity 112:21–29
Arnold SJ (1992) Constraints on phenotypic evolution. Am Nat 140:S85–S107
Arnold SJ, Bürger R, Hohenlohe PA, Ajie BC, Jones AG (2008) Understanding the evolution and stability of the G‐matrix. Evolution 62:2451–2461
Arnold SJ, Pfrender ME, Jones AG (2001) The adaptive landscape as a conceptual bridge between micro- and macroevolution. Genetica 112:9–32
Arnold SJ, Phillips PC (1999) Hierarchical comparison of genetic variance-covariance matrices. II. Coast-inland divergence garter snake, Thamnophis elegans. Evolution 53:1516–1527
Bégin M, Roff DA (2003) The constancy of the G matrix through species divergence and the effects of quantitative genetic constraints on phenotypic evolution: a case study in crickets. Evolution 57:1107–1120
Bégin M, Roff DA (2004) From micro to macroevolution through quantitative genetic variation: positive evidence from field crickets. Evolution 58:2287–2304
Björklund M, Husby A, Gustafsson L (2013) Rapid and unpredictable changes of the G‐matrix in a natural bird population over 25 years. J Evol Biol 26:1–13
Blows MW (2007) A tale of two matrices: multivariate approaches in evolutionary biology. J Evol Biol 20:1–8
Blows MW, Brooks R (2003) Measuring nonlinear selection. Am Nat 162:815–820
Blows MW, McGuigan K (2015) The distribution of genetic variance across phenotypic space and the response to selection. Mol Ecol 24:2056–2072
Bonduriansky R, Chenoweth SF (2009) Intralocus sexual conflict. Trends Ecol Evol 24:280–288
Bonduriansky R, Rowe L (2005) Intralocus sexual conflict and the genetic architecture of sexually dimorphic traits in Prochyliza xanthostoma (Diptera: Piophilidae). Evolution 59:1965–1975
Brodie ED, III (1993) Homogeneity of the genetic variance‐covariance matrix for antipredator traits in two natural populations of the garter snake Thamnophis ordinoides. Evolution 47:844–854
Calsbeek B, Lavergne S, Patel M, Molofsky J (2011) Comparing the genetic architecture and potential response to selection of invasive and native populations of reed canary grass. Evolut Appl 4:726–735
Careau V, Wolak ME, Carter PA, Garland T, Jr (2015) Evolution of the additive genetic variance-covariance matrix under continuous directional selection on a complex behavioural phenotype Proc R Soc B 282:20151119
Chakrabarty A, van Kronenberg P, Toliopoulos N, Schielzeth H (2019) Direct and indirect genetic effects on reproductive investment in a grasshopper. J Evol Biol 32:331–342
Chenoweth SF, Blows MW (2004) Contrasting mutual sexual selection on homologous signal traits in Drosophila serrata. Am Nat 165:281–289
Cheverud J (1996) Quantitative genetic analysis of cranial morphology in the cotton‐top (Saguinus oedipus) and saddle‐back (S. fuscicollis) tamarins. J Evol Biol 9:5–42
Cheverud JM, Marroig G (2007) Comparing covariance matrices: random skewers method compared to the common principal components model. Genet Mol Biol 30:461–469
Cigliano MM, Braun H, Eades DC, Otte D (2017) Orthoptera Species File (Version 5.0/5.0). http://Orthoptera.SpeciesFile.org
Contreras D, Chapco W (2006) Molecular phylogenetic evidence for multiple dispersal events in gomphocerine grasshoppers. J Orthoptera Res 15:91–98
Day T, Bonduriansky R (2004) Intralocus sexual conflict can drive the evolution of genomic imprinting. Genetics 167:1537–1546
Delahaie B, Charmantier A, Chantepie S, Garant D, Porlier M, Teplitsky C (2017) Conserved G-matrices of morphological and life-history traits among continental and island blue tit populations. Heredity 119:76–87
Doroszuk A, Wojewodzic MW, Gort G, Kammenga JE (2008) Rapid divergence of genetic variance-covariance matrix within a natural population. Am Nat 171:291–304
Eroukhmanoff F, Svensson E (2011) Evolution and stability of the G‐matrix during the colonization of a novel environment. J Evol Biol 24:1363–1373
Flury B (1988) Common principal components & related multivariate models. John Wiley & Sons, Inc.
Forsman A (2015) Rethinking phenotypic plasticity and its consequences for individuals, populations and species. Heredity 115:276
Gäde G (2002) Sexual dimorphism in the pyrgomorphid grasshopper Phymateus morbillosus: from wing morphometry and flight behaviour to flight physiology and endocrinology. Physiol Entomol 27:51–57
Garant D, Hadfield JD, Kruuk LE, Sheldon BC (2008) Stability of genetic variance and covariance for reproductive characters in the face of climate change in a wild bird population. Mol Ecol 17:179–188
Gelman A (2006) Prior distributions for variance parameters in hierarchical models (Comment on an Article by Browne and Draper). Bayesian Anal 1:515–533
Gelman A, Rubin DB (1992) Inference from iterative simulation using multiple sequences. Stat Sci 7:457–472
Gosden TP, Chenoweth SF (2014) The evolutionary stability of cross‐sex, cross‐trait genetic covariances. Evolution 68:1687–1697
Griffin RM, Dean R, Grace JL, Ryden P, Friberg U (2013) The shared genome is a pervasive constraint on the evolution of sex-biased gene expression. Mol Biol Evol 30:2168–2176
Hadfield JD (2010) MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J Stat Softw 33:1–22
Harrison RG (1980) Dispersal polymorphisms in insects. Annu Rev Ecol Syst 11:95–118
Heidinger IMM, Hein S, Feldhaar H, Poethke H-J (2018) Biased dispersal of Metrioptera bicolor, a wing dimorphic bush-cricket. Insect Sci 25:297–308
Hine E, Chenoweth SF, Rundle HD, Blows MW (2009) Characterizing the evolution of genetic variance using genetic covariance tensors. Philos Trans R Soc B 364:1567–1578
Hinks C, Erlandson M (1994) Rearing grasshoppers and locusts: review, rationale and update. J Orthoptera Res 3:1–10
Houle D (1992) Comparing evolvability and variability of quantitative traits. Genetics 130:195–204
Jones AG, Arnold SJ, Bürger R, Houle D (2003) Stability of the G-matrix in a population experiencing pleiotropic mutation, stabilizing selection, and genetic drift. Evolution 57:1747–1760
Karlsson Green K, Eroukhmanoff F, Harris S, Pettersson LB, Svensson EI (2016) Rapid changes in genetic architecture of behavioural syndromes following colonization of a novel environment. J Evol Biol 29:144–152
Krzanowski W (1979) Between-groups comparison of principal components. J Am Stat Assoc 74:703–707
Lande R (1979) Quantitative genetic analysis of multivariate evolution, applied to brain: body size allometry. Evolution 33:402–416
Lande R (1980a) The genetic covariance between characters maintained by pleiotropic mutations. Genetics 94:203–215
Lande R (1980b) Sexual dimorphism, sexual selection, and adaptation in polygenic characters. Evolution 34:292–305
Lande R (1982) A quantitative genetic theory of life history evolution. Ecology 63:607–615
Lande R (1987) Genetic correlations between the sexes in the evolution of sexual dimorphism and mating preferences. In: Bradbury JW, Andersson MB, Heisler L (ed) Sexual selection: testing the alternatives. Wiley, Chichester, p 83–94
Lande R, Arnold SJ (1983) The measurement of selection on correlated characters. Evolution 37:1210–1226
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer, Sunderland, MA
McGlothlin JW, Kobiela ME, Wright HV, Mahler DL, Kolbe JJ, Losos JB et al. (2018) Adaptive radiation along a deeply conserved genetic line of least resistance in Anolis lizards. Evolution Lett 2:310–322
McGuigan K (2006) Studying phenotypic evolution using multivariate quantitative genetics. Mol Ecol 15:883–896
Merilä Björklund (1999) Population divergence and morphometric integration in the greenfinch (Carduelis chloris) – evolution against the trajectory of least resistance? J Evol Biol 12:103–112
Nattier R, Robillard T, Amedegnato C, Couloux A, Cruaud C, Desutter‐Grandcolas L (2011) Evolution of acoustic communication in the Gomphocerinae (Orthoptera: Caelifera: Acrididae). Zool Scr 40:479–497
Paulsen SM (1996) Quantitative genetics of the wing color pattern in the buckeye butterfly (Precis coenia and Precis evarete): evidence against the constancy of G. Evolution 50:1585–1597
Pepler T (2015) cpc: common principal component (CPC) Analysis and applications. R package version 0:1–5
Phillips PC, Arnold SJ (1989) Visualizing multivariate selection. Evolution 43:1209–1222
Phillips PC, Arnold SJ (1999) Hierarchical comparison of genetic variance‐covariance matrices. I. Using the Flury hierarchy. Evolution 53:1506–1515
Phillips PC, Whitlock MC, Fowler K (2001) Inbreeding changes the shape of the genetic covariance matrix in Drosophila melanogaster. Genetics 158:1137–1145
Poissant J, Wilson AJ, Coltman DW (2010) Sex‐specific genetic variance and the evolution of sexual dimorphism: a systematic review of cross‐sex genetic correlations. Evolution 64:97–107
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Riede K (1986) Modification of the courtship song by visual stimuli in the grasshopper Gomphocerus rufus (Acrididae). Physiol Entomol 11:61–74
Ritchie M, Butlin R, Hewitt G (1987) Causation, fitness effects and morphology of macropterism in Chorthippus parallelus (Orthoptera: Acrididae). Ecol Entomol 12:209–218
Roff D (2000) The evolution of the G matrix: selection or drift? Heredity 84:135–142
Roff D, Mousseau T (1999) Does natural selection alter genetic architecture? An evaluation of quantitative genetic variation among populations of Allenomobius socius and A. fasciatus. J Evol Biol 12:361–369
Roff DA (1986a) The evolution of wing dimorphism. Evolution 40:1009–1020
Roff DA (1986b) The genetic basis of wing dimorphism in the sand cricket, Gryllus firmus and its relevance to the evolution of wing dimorphisms in insects. Heredity 57:221–231
Roff DA, Fairbairn DJ (1993) The evolution of alternate morphologies: fitness and wing morphology in male sand crickets. Evolution 47:1572–1584
Roff DA, Fairbairn DJ (2007) The evolution and genetics of migration in insects. Bioscience 57:155–164
Roff DA, Prokkola JM, Krams I, Rantala MJ (2012) There is more than one way to skin a G matrix. J Evol Biol 25:1113–1126
Rosetti N, Remis MI (2018) Spatial variation in body size and wing dimorphism correlates with environmental conditions in the grasshopper Dichroplus vittatus (Orthoptera: Acrididae). Environ Entomol 47:519–526
Schluter D (1996) Adaptive radiation along genetic lines of least resistance. Evolution 50:1766–1774
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Shaw FH, Shaw RG, Wilkinson GS, Turelli M (1995) Changes in genetic variances and covariances: G whiz! Evolution 49:1260–1267
Sinervo B, Svensson E (2002) Correlational selection and the evolution of genomic architecture. Heredity 89:329–338
Sniegula S, Golab MJ, Drobniak SM, Johansson F (2018) The genetic variance but not the genetic covariance of life‐history traits changes towards the north in a time‐constrained insect. J Evol Biol 31:853–865
Spitze K, Burnson J, Lynch M (1991) The covariance structure of life‐history characters in Daphnia dulex. Evolution 45:1081–1090
Steppan SJ, Phillips PC, Houle D (2002) Comparative quantitative genetics: evolution of the G matrix. Trends Ecol Evol 17:320–327
Teplitsky C, Tarka M, Møller AP, Nakagawa S, Balbontín J, Burke TA et al. (2014) Assessing multivariate constraints to evolution across ten long-term avian studies. PLoS One 9:e90444
Turelli M (1988) Phenotypic evolution, constant covariances, and the maintenance of additive variance. Evolution 42:1342–1347
Uvarov BP (1966) Grasshoppers and locusts: anatomy, physiology, development, phase polymorphism, introduction to taxonomy, Vol 1. Cambridge UP. Published for the Anti-Locust Research Centre
Vedenina V, Mugue N (2011) Speciation in gomphocerine grasshoppers: molecular phylogeny versus bioacoustics and courtship behavior. J Orthoptera Res 20:109–125
Walsh B, Blows MW (2009) Abundant genetic variation+ strong selection= multivariate genetic constraints: a geometric view of adaptation. Annu Rev Ecol Evol Syst 40:41–59
Walter GM, Aguirre JD, Blows MW, Ortiz-Barrientos D (2018) Evolution of genetic variance during adaptive radiation. Am Nat 191:E108–E128
Willott S, Hassall M (1998) Life‐history responses of British grasshoppers (Orthoptera: Acrididae) to temperature change. Funct Ecol 12:232–241
Acknowledgements
We are grateful to Amy Backhouse, Julia Teckentrup, Pablo Valverde, Philipp van Kronenberg, and Niko Toliopoulos for their enormous help with field work, setting up of breeding experiments, and taking photographs. The project was funded by the Emmy Noether grant from DFG (SCHI 1188/1-1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Chakrabarty, A., Schielzeth, H. Comparative analysis of the multivariate genetic architecture of morphological traits in three species of Gomphocerine grasshoppers. Heredity 124, 367–382 (2020). https://doi.org/10.1038/s41437-019-0276-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-019-0276-1