Abstract
Niche partitioning can lead to differences in the range dynamics of plant species through its impacts on habitat availability, dispersal, or selection for traits that affect colonization and persistence. We investigated whether niche partitioning into upland and riparian habitats differentiates the range dynamics of two closely related and sympatric eastern Australian trees: the mountain water gum (Tristaniopsis collina) and the water gum (T. laurina). Using genomic data from SNP genotyping of 480 samples, we assessed the impact of biogeographic barriers and tested for signals of range expansion. Circuit theory was used to model isolation-by-resistance across three palaeo-environment scenarios: the Last Glacial Maximum, the Holocene Climate Optimum and present-day (1950–2014). Both trees showed similar genetic structure across historically dry barriers, despite evidence of significant environmental niche differentiation and different post-glacial habitat shifts. Tristaniopsis collina exhibits the signature of serial founder effects consistent with recent or rapid range expansion, whilst T. laurina has genetic patterns consistent with long-term persistence in geographically isolated populations despite occupying a broader bioclimatic niche. We found the minor influence of isolation-by-resistance on both species, though other unknown factors appear to shape genetic variation. We postulate that specialized recruitment traits (adapted to flood-disturbance regimes) rather than habitat availability limited post-glacial range expansion in T. laurina. Our findings indicate that niche breadth does not always facilitate range expansion through colonization and migration across barriers, though it can promote long-term persistence in situ.
Similar content being viewed by others
Introduction
Range dynamics describe the changing geographic distributions of species in response to environmental shifts over time. The factors that govern climate-driven range shifts are key to understanding the evolutionary history of species and their projected vulnerability to future climate threats. Among plant species, range dynamics are broadly characterized by the ability to track suitable habitat through propagule dispersal and establishment (‘dispersal potential’) or to withstand environmental change in situ (‘persistence potential’). Habitat specialization is expected to differentiate these processes among species if they differ in eco-physiological tolerances (‘environmental niche breadth’; Ackerly 2003; Chase and Leibold 2003) or habitat connectivity across the landscape. However, these distinctions might be cryptic among sympatric species, as their present-day distributions could be the outcome of either recent range expansions or long-term local persistence. Furthermore, since the magnitude of climate change varies geographically, populations located in different parts of a species’ range can also be expected to respond differently (Sork et al. 2010). In this context, we seek to investigate whether habitat preference differentiates the range dynamics of closely related and sympatric species; and the extent to which this varies across their range.
The latitudinal climate gradient of mainland eastern Australia shifts from temperate to subtropical systems, making this an ideal region for investigating geographic patterns of climate-driven range shifts. The pollen fossil record indicates that substantial vegetation shifts occurred following the end of the Last Glacial Maximum (LGM, 21,000–17,000 years ago), yet there is a paucity of data on the widely distributed mesic flora of the region (although see Mellick et al. 2012). Glaciation during the LGM was less significant in eastern Australia than in the Northern Hemisphere, though conditions were much colder and drier than at present and the period was marked by an elevated fire frequency and intensity (Kershaw et al. 2007). In addition, several large, low-lying river systems formed dry habitat barriers for moisture-dependent species during glacial periods (Thom et al. 1994). Meanwhile, the onset of the Holocene about 12,000 years ago saw a dramatic rise in global temperatures and precipitation, which peaked around 6000 years ago during the Holocene Climactic Optimum (HCO). The impact of LGM habitat barriers and the rate of post-LGM range expansion appears to vary among ecologically-differentiated species (Mellick et al. 2013a, 2014; Rossetto et al. 2015a), though the roles of different factors remain unclear.
A major gap in our understanding of climate-driven range shifts is the relative importance of dispersal versus persistence in available habitat. A number of studies have demonstrated that dispersal mechanism and habitat preference affect restrictions across biogeographic barriers (Rossetto et al. 2007, 2009, 2015a, b; Milner et al. 2012; Worth et al. 2017), yet the ecological factors that determine survival in refugia are less clear. For instance, chloroplast phylogeographies of cool temperate trees indicate long-term persistence and local re-expansion from multiple glacial refugia across southeast Australia, regardless of dispersal mechanism (Clark and Carbone 2008; Worth et al. 2009, 2010; Nevill et al. 2010). On the other hand, species that are less cold-tolerant but wind-dispersed appear to have relied on long-distance dispersal from relatively few refugia (Worth et al. 2011), suggesting the role of bioclimatic tolerances. We explore these questions with tests of Holocene range expansion in two sympatric trees thought to differ in their potential for persistence and post-LGM dispersal.
Tristaniopsis collina Peter G. Wilson and J. T. Waterh (‘mountain water gum’) and Tristaniopsis laurina Peter G. Wilson and J. T. Waterh (‘water gum’) are medium-sized trees of the Myrtaceae family and are relatively widespread through the coast and ranges of eastern New South Wales (NSW; Fig. 1a). The species belong to sister clades in a monophyletic Australian lineage of Tristaniopsis but show marked ecological differences. While Tristaniopsis collina prefers habitats at elevation and typically occurs on rocky hillslopes or escarpment edges, T. laurina is restricted to rocky watercourses (Wilson 1991). The upslope and riparian habitats of the two species are thought to have been differently affected by climatic change over time and thus differentiate their persistence potential. Specifically, sites above 1000 m were too cold for woody species during the LGM (Sweller and Martin 2001; Hesse et al. 2003) and would have forced upland trees like T. collina into local extinction or to retreat to lower clines. Meanwhile, lower evapotranspiration rates at riparian sites within montane gullies are hypothesized to have provided microrefugia for lowland trees in periods of drought (Aide and Rivera 1998; de Lafontaine et al. 2014) and potentially enabled greater in situ persistence for T. laurina. To test this hypothesis, we first determine the degree of bioclimatic differentiation between the species and then assess their respective Holocene habitat availabilities with palaeo-environment models.
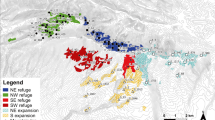
a The occurrence records of Tristaniopsis collina and T. laurina in eastern Australia, and their distributional extent indicated by the red box (inset map). Areas of significance are shown, including a number of known biogeographic barriers: the Clarence River Corridor (CRC), Manning River Valley (MRV), Hunter River Corridor (HRC), Cumberland Plain (CP), Shoalhaven River (SR), and Southern Transition Zone (STZ). b The study area in eastern New South Wales is indicated by the red box. Sample sites (n = 6) and their average population assignment proportions are represented as pie charts. Each colour represents a lineage identified by sNMF, assuming K = 4 for T. collina and c assuming K = 3 for T. laurina
Habitat connectivity is also an important factor in post-LGM dispersal (Sork et al. 2010) and is likely to have differentiated the study species. Riparian habitats are particularly distinct from other habitat types because their linear distribution patterns can function to rapidly disperse propagules across the landscape (Werth and Scheidegger 2014). However, hydrochory (propagule dispersal via water) can only transport genes downstream and is subject to the landscape terrain. This raises the question of whether or not hydrochory facilitates long-distance dispersal of T. laurina propagules within catchments (Nilsson et al. 2010). The impact of stream connectivity on population genetic structure has been demonstrated in a number of riparian plants (Wei et al. 2013; Werth and Scheidegger 2014). Yet there are few, if any studies that have evaluated the range dynamics of riparian versus upslope trees in a comparative framework.
Aside from their ecological differentiation, we chose to focus on Tristaniopsis species because, in addition to their geographically overlapping distributions (~1200 km), they frequently co-occur in the same community, sometimes only metres apart (M. Fahey, pers. obs). The two species also share numerous phenological traits, including similar flowering times and reproductive morphology, though there is no evidence that they hybridize (Wilson and Waterhouse 1982). Aside from minor differences in petal size and stamen abundance, an overall similarity in floral morphology suggests that the species receive similar floral visitors and pollen dispersers (Thomson et al. 2000; Castellanos et al. 2003; Rosas-Guerrero et al. 2011; Wilson et al. 2017). Both possess woody, capsular, 3-valved fruits that contain 30–40 small, terminally winged seeds thought to be mostly gravity-dispersed (Williams and Adam 2010). In this regard, the two species can be expected to encounter similar biogeographic influences and to have equal potential for pollen dispersal.
In this study, we evaluate the population genetic structure of T. collina and T. laurina with single-nucleotide polymorphisms (SNPs) and construct palaeo-environment models to investigate their range dynamics in eastern NSW, Australia. Here we use a relatively novel demographic model to test for the signature of serial founder spatial expansions within species (Peter and Slatkin 2013, 2015). We also calculate resistance surfaces based on circuit theory (Shah and McRae 2008) to model the change in habitat connectivity for each species during the Holocene, which is poorly tested in tree species (although see Ortego et al. 2015). Specifically, we tested the following hypotheses:
(1) There is environmental differentiation between the upslope and riparian habitats of T. collina and T. laurina, and these were differently affected by Holocene climatic fluctuations.
(2) The influence of biogeographic barriers varies between species, due to differences in habitat availability and/or the relative efficacy of hydrochory versus gravity dispersal.
(3) Environmental niche partitioning differentiates the two study species in their potential for range expansion and persistence in refugia.
(4) Within each species, range dynamics vary across the study area, owing to spatial variation in landscape barriers and a latitudinal climate gradient.
Materials and methods
Study system and sampling strategy
We collected foliage samples from the core range of Tristaniopsis collina and T. laurina in NSW (Fig. 1). This included lowland coastal and adjacent upland locations, and sites separated by putative biogeographic barriers. The Clarence River Corridor in northern NSW is largest coastal catchment in the state, and vicariance across its broad floodplain has been demonstrated in various rainforest trees (Mellick et al. 2011; Heslewood et al. 2014; Rossetto et al. 2015b). The Hunter River Corridor in central NSW is the oldest known barrier in eastern Australia (Chapple et al. 2011) and disrupts latitudinal gene flow in several plant and animal species (Playford et al. 1993; Worth et al. 2009; Chapple et al. 2011; Milner et al. 2012; Heslewood et al. 2014).
Also influential are the climatic gradients of the ‘Southern Transition Zone’ (Milner et al. 2012), which extends southwards from the Shoalhaven River to Victoria. This region has had a complex history of genetic breaks along the coastal escarpment as lowland and upland populations were differently affected by glaciation (Chapple et al. 2011; Milner et al. 2012). The impact of the Southern Transition Zone on the genetic structure of different species is hypothesized to vary with their physiological tolerances (Milner et al. 2012).
We selected the most proximate sites possible between species, so as to have a uniform and comparable sampling design. We took genetic material from six individuals separated by a minimum distance of 30 m from each locality, which included a total of 34 populations of T. collina and 46 populations of T. laurina (Supplementary Table S1). All collections were made during April–May 2015 and February–May 2016. Based on our previous analyses of SNP data with DArTseq, we found that six samples per population is sufficient to capture site-level genetic variation (Rossetto et al. 2018).
Quantification of environmental niche breadth and differentiation
In order to assess whether habitat preference varies the range dynamics of T. collina and T. laurina, we first sought to determine the degree of environmental differentiation between them. We quantified environmental niche separation between the study species from environmental data recorded at their georeferenced occurrence records. The occurrence data were derived from our sample sites, in addition to records obtained from Australia’s Virtual Herbarium (http://avh.chah.org.au). The occurrence data were filtered to remove records older than 1970 or with spatial uncertainty >1 km. To limit spatial bias, we only included one record per 5 km radius. The final data set consisted of 112 occurrence points for T. collina and 118 occurrence points for T. laurina.
To reconstruct the environmental background of the study area, we obtained 23 bioclimatic and seven geomorphological variables at a spatial resolution of 36 s (c. 1 km) from eMAST (Xu and Hutchinson 2013). This included the 19 BioClim variables, Mean Monthly Evaporation, Minimum Monthly Evaporation, Maximum Monthly Evaporation, and Evaporation Seasonality. The geomorphological variables included Slope, Aspect, Topographic Wetness Index, Topographic Position Index, Clay %, Sand %, and Silt %. The spatial extent of the study area was bounded to include a 20 km buffer zone around the northern- and southernmost occurrences of T. laurina (−24.8° S, −37.8° S, 147° E, 154° E).We quantified niche overlap between T. collina and T. laurina and the niche breadth of each species in multidimensional environmental space, using a principal components analysis (PCA) method developed by Broennimann et al. (2012) and implemented in the R package Ecospat (Broennimann et al. 2012; Di Cola et al. 2017). This method estimates the niche breadth of each species from the first two axes of a PCA of the 30 predictor variables and visualizes niche space with boxplots for each axis. Niche overlap between species was estimated with Schoener’s D similarity index, ranging between 0 (no overlap) and 1 (complete overlap; Schoener 1970). We also conducted a niche equivalency test to determine whether the niches of two entities from distinct geographical ranges are equivalent (D).
One limitation with our models is that the environmental layers could not be downscaled to the resolution of hydrological datasets (25 m), and so we could not accurately model the riparian microclimate. The spatial resolution is also potentially too coarse to distinguish between the localized changes in Topographic Wetness Index and slope that are believed to differentiate the species. While this would lead to an over-estimated niche breadth for T. laurina, the bioclimatic variables are still useful for inferring broader biogeographic trends across the landscape and between species.
Genomic methods
DNA extraction from leaf samples and SNP genotyping using DArTseq technology (Sansaloni et al. 2011) was undertaken at Diversity Arrays Technology Pty Ltd (Canberra, Australia). We evaluated the impact of data quality on downstream diversity analyses through the comparison of SNP data sets filtered according to different quality thresholds: reproducibility average (proportion of technical replicates for which the marker score is consistent) and call rate (proportion of individuals with non-missing scores). We selected markers with a reproducibility average of 0.96 and call rate >0.95, which yielded a final data set of 8215 SNPs for 204 samples of T. collina and 10,883 SNPs for 276 samples of T. laurina.
Assessment of genotypic diversity and population structure
We sought to assess whether population structure within the study species is characterized by divergence across ancient habitat barriers. First, we used the individual ancestry and population clustering algorithm implemented by the R package sNMF (Frichot et al. 2014) to identify putative ancestral lineages. In a procedure similar to that implemented by the widely used program STRUCTURE (Pritchard et al. 2000), ancestry coefficients (K) were estimated by running 10 iterations each of K = 2 to K = 10, and the best-supported K was assessed with a cross-entropy criterion (see Frichot et al. 2014). The find.clusters and dapc function in adegenet was used to verify the results of sNMF. This procedure transforms data using a PCA and runs successive k-means with an increasing number of clusters (k), before performing a linear discriminant analysis on the principal components to assign individuals to clusters (Jombart et al. 2010). For each model, a statistical measure of goodness of fit is computed with the Bayesian information criterion to identify the optimal k. Clustering methods have been critiqued for assigning discrete clusters to species potentially characterized by continuous patterns of genetic differentiation (Lawson et al. 2018). Therefore, we ran a PCA of the individual pairwise genetic distance matrix using the R package adegenet 2.0.0 (Jombart 2008; Jombart and Ahmed 2011). Ordination of the first two principal components was used to visualize genetic similarity among samples and does not assume clusters.
To quantify genetic distance between sample sites, we calculated pairwise FST values in R. Then, to assess different sources of landscape structure within species, including biogeographic barriers, we used Poppr 2.8.1 (Kamvar et al. 2014, 2015) to quantify the degree of genetic variance across the population hierarchy with a three-way analysis of molecular variance (AMOVA) following Weir and Cockerham (1984). Variance was assessed across pairwise FST values between individuals within sites, values averaged by sample site, and values averaged by lineages identified by sNMF. To assess the influence of catchment connectivity on gene flow within T. laurina, we performed a four-way AMOVA that included catchment membership within lineages. Significance was tested with a randomization test as per Excoffier et al. (1992), using 999 permutations.
Genetic diversity statistics were calculated and compared across sample sites within species and averaged across sites to compare overall diversity between species. The following estimates were calculated with the R package diveRsity (Keenan et al. 2013): observed heterozygosity (HO), unbiased expected heterozygosity (uHE), and allelic richness (AR).
Palaeo-environment modelling
We used environmental niche modelling (ENM) to infer the geographic distribution of suitable palaeo-environments for each species, to test whether current and/or past habitat connectivity can explain observed patterns of genetic structure. There are a number of assumptions that reduce our confidence in niche projections, including the inability to account for the likelihood that some past environments had no modern analogues to train the models (Williams and Jackson 2007), or the possibility that the study species’ current distributions are in disequilibrium with the environment due to dispersal constraints (Svenning and Skov 2007) or that they underwent changes in existing realized niche since the LGM (Worth et al. 2014). Nevertheless, a number of studies have found that such models corroborate genetic evidence for glacial refugia and post-glacial shifts (Mellick et al. 2012, 2014, 2013b; Ortego et al. 2015).
We modelled the current distribution of habitat suitability for each species using the maximum entropy algorithm implemented by MaxEnt 3.3.3 (Phillips et al. 2006; Phillips and Dudík 2008). Because widely distributed species are often composed of locally adapted populations with distinct environmental tolerances and potential ranges (Banta et al. 2012), intraspecific genotypes have been identified as useful units for climate envelope modelling (Eckert et al. 2009; Fournier-Level et al. 2011; Hancock et al. 2011). We produced separate models for the within-species lineages identified by sNMF (sensu Jezkova et al. 2009; Mellick et al. 2011; Hornsby and Matocq 2012; Banta et al. 2012), as well as models based on the full distribution of each species.
The full-species models were constructed using the same occurrence records, extent, and resolution as described for the niche models implemented in Ecospat. The lineage-based models were trained on a background area constrained to a 200 km radius around each occurrence point, to avoid very different climates inflating the strength of predictions. We qualitatively assessed the accuracy of predicted occurrences and used model-evaluation statistics to select sets of non-correlated bioclimatic and geomorphological variables for each lineage. Only sets of variables with a variance inflation factor <12 were used for each model (Powney et al. 2011) which was iteratively calculated using the R package usdm (Naimi 2013).
To produce palaeo-models for the LGM and HCO climate scenarios, we projected contemporary species-environment relationships onto General Circulation Models obtained from the Inter-comparison Project 5 data repository maintained by the Earth Survey Grid Federation, accessed via the Lawrence Livermore National Laboratory node (https://esgf-node.llnl.gov/projects/esgf-llnl; Supplementary Table S2). For each General Circulation Model, we ran 10 replicates under the cross-validation method with linear, product, and quadratic features. Current and palaeo-habitat suitability maps based on the logistic output of MaxEnt were generated to visualize habitat availability through time. We compared the output from each model, since the climate scenarios simulated by different General Circulation Models vary considerably (Whetton et al. 2015). We also calculated the mean logistic output scores across models to use as a consensus habitat suitability model in downstream analyses. Finally, we used the sum of the logistic output of each climatic period to estimate Holocene habitat suitability stability (sensu Devitt et al. 2013). Since the ENMs do not account for the reduced atmospheric CO2 concentrations which would have exacerbated the physiological effects of aridity during the LGM, we take a conservative estimate of habitat suitability as >0.7.
Landscape genetic analyses
We sought to test whether niche specialization differentiates the dispersal potential of the study species, and whether dispersal varies across the study area. First, we applied circuit theory to evaluate whether gene flow within lineages reflects isolation-by-distance (IBD) or different isolation-by-resistance (IBR) scenarios. IBR assumes that habitat suitability constrains dispersal across spatially heterogeneous landscapes more than can be expected of geographic distance alone (McRae and Beier 2007). IBR is typically used to estimate animal movement across habitat networks, though it can be applied at a broader scale to evaluate dispersal as the number of migrants exchanged between subpopulations per generation (Shah and McRae 2008; Ortego et al. 2015).
We used CIRCUITSCAPE 4.0.5 to calculate resistance distance matrices between all pairs of population sample sites, considering an eight-neighbour cell connection scheme (Shah and McRae 2008). The resistance distance is calculated according to the minimum movement cost and weighted by the availability of alternative pathways. In summary, the resistance distance is small when two cells are connected by many paths with low resistance values and large when there are few paths with high resistance.
We used habitat suitability data (i.e. the mean logistic output of the ENMs) as conductance grids to calculate resistance values that reflect four IBR scenarios: (1) current habitat suitability (IBRCURRENT); (2) HCO habitat suitability (IBRHCO); (3) LGM habitat suitability (IBRLGM); and (4) Holocene habitat suitability stability (IBRSTABILITY; see above). To model IBD, we calculated a matrix of log transformed Euclidean geographic distances between sampled populations. We modelled IBR and IBD separately for each lineage as well as across the full range of the study species. This enabled us to assess whether there is regional variation in landscape influences on the range dynamics of the species, and control for the confounding effect of biogeographic barriers on the pattern of IBD (Slatkin 1993; Rousset 1997; Garnier et al. 2004).
IBD and IBR matrices were tested against genetic distance matrices using multiple matrix regression with randomization (Wang 2013) implemented in the R package PopGenReport 3.0.0 (Adamack and Gruber 2014; Gruber and Adamack 2015) and partial Mantel tests in the package ncf 1.2–6 (Bjornstad and Cai 2018). We constructed and evaluated models with each possible combination of explanatory terms fitted. However, final models do not include >2 independent variables because multi-collinearity hampered interpretation of results. The coefficient of determination (r2) indicates the overall model fit whilst the regression coefficient (βn) indicates the strength of the association between linearized FST and each individual distance variable after other variables have been accounted for. Finally, for each lineage in each species, we performed a Pearson’s correlation on population genetic diversity indices (AR, HO, and uHE) with their latitude and IBRSTABILITY scores.
Tests for range expansion
We sought to compare the study species in their potential for range expansion versus persistence in refugia. We used an approach developed by Peter and Slatkin (2013) that tests whether drift under population expansion can explain the observed genetic structure. This approach infers the strength of founder effects associated with spatial expansion and the most likely expansion origin, and tests significance against a null model of equilibrium IBD (Peter and Slatkin 2013, 2015).
The model assumes that present-day populations have expanded on a stepping-stone model from a single deme (i.e., refugium), assuming a founder effect (ke/Ne) for each colonization event and one generation of drift between events. In an island population model, Ne corresponds to the effective population size and ke corresponds to the effective number of founders on a given island. Given that we do not know the actual deme size of subpopulations for most species, the founder effect is arbitrarily re-scaled to 0.99. This enables us to compare expansions using the distance over which that founder effect occurs, where a low founder distance indicates a strong founder effect. Assuming ke/Ne = 0.99, the effective founder distance (d) is calculated as the deme size (in km) for which Ne is reduced by 1% in a founder event. As per the assumption of a single source population, we constructed separate models for the lineages identified by sNMF. which would have had distinct expansion origins (Peter and Slatkin 2013).
The model will detect weak or non-significant founder effects if range expansion was gradual, the species underwent long periods of post-expansion drift, or the species had multiple reticulate expansions per lineage. Based on results from an Arabadopsis thaliana dataset (Peter and Slatkin, 2015), we consider d < 5 km to indicate strong founder effects and d > 25 km to indicate weak founder effects.
Results
Environmental niche differentiation between species
Niche overlap is low between Tristaniopsis collina and T. laurina (D = 0.311), supporting the hypothesis that environmental preferences differ between the species. Niche equivalency was rejected (P = 0.0198) and niche similarity was greater than expected by chance (P = 0.099), indicating that the observed niche differentiation between species is a function of habitat selection rather than an artefact of underlying differences in available environments between the two species.
The PCA separates the occurrence of T. collina and T. laurina in environmental space and the first two principal components explain 68.86% of the total environmental variation between the species (Fig. 2). The PC loadings of predictor variables indicate that T. collina occupies a cooler, wetter, and more aseasonal range of climates than does T. laurina (Supplementary Table S4). We found that T. laurina has a greater niche breadth (PC1 variance = 4.98, PC2 variance = 7.66) than does T. collina (PC1 variance = 3.311, PC2 variance = 4.64), indicative of broader bioclimatic tolerances.
A model of environmental niche occupancy (grey gradient) and availability across the distribution of a Tristaniopsis collina, and b T. laurina. The axes define environmental space in the model, according to the first two principal components that describe variance across twenty-three bioclimatic and seven geomorphological predictor variables. The solid line indicates 100% of available environments and the dashed line shows the 50% most frequent available environments. The environmental niches of the two species differ according to tests of niche equivalency (P = 0.020) and niche similarity (P = 0.099). Tristaniopsis laurina shows greater niche breadth but a smaller core density of occurrences
Genotypic diversity and population structure
Genetic subdivision was detected between all populations of T. collina and T. laurina, based on pairwise population FST estimates (P < 0.001; Supplementary Tables S4–S5). Subdivision was slightly lower among populations of T. collina (mean FST = 0.170; Supplementary Table S5) than among those of T. laurina (mean FST = 0.194; Supplementary Table S6). This excludes populations north of the Hunter River Corridor, where T. collina has greater FST values than does T. laurina.
For T. collina, the cross-entropy evaluation implemented in sNMF suggests K = 4–10 and the k-means analysis implemented in adegenet identified 3–5 clusters (Supplementary Figs. S1–S2). We assumed K = 4 in subsequent analyses of T. collina because this is the simplest most parsimonious model. The four ancestral lineages (Shield Volcano, North, Central, and South) are diverged across three biogeographic barriers: Clarence River Corridor, Hunter River Corridor, and Cumberland Plain (Fig. 1b). However, a fifth cluster was identified in the Barrington region and also makes biogeographic sense. Both analyses identified three ancestral lineages in T. laurina (North, Central, and South; Supplementary Figs. S3–S4) bounded by two barriers (Hunter River Corridor and Cumberland Plain; Fig. 1c).
The mean population assignment proportions are generally high (0.724 in T. collina and 0.861 in T. laurina). If the clusters represent ancestral lineages as per the assumption of sNMF, both species show admixture indicative of secondary contact zones south of Central in the Illawarra plateau and to the north in the Barrington region (Fig. 1). This region either admixed in T. collina assuming K = 4 or differentiated due to drift under geographic isolation assuming K = 5.
The PCA ordinations identify the same major groupings as sNMF (albeit with hierarchical structure) and reveal opposite trends between the study species. Within T. collina, strong geographic structure is evident in North, which separates populations across the Manning River Valley and the Hunter River Corridor (PC1 = 16.98; Fig. 3a). Along PC2 (2.13), Central lacks structure and South shows moderate geographic structure. The primary source of variation in T. laurina is between populations separated by the Hunter River Corridor (PC1 = 15.28; Fig. 3b). Populations in the Hunter catchment also separate from other populations in Central along PC1. There is little geographic structure in North, and continuous variation within South along PC2 (2.74).
The AMOVA revealed that the two study species have similar levels of structure across barriers, although greater within-lineage differentiation is apparent for T. laurina populations (Table 1). Among-catchment differentiation is an equivalent source of variation to lineage divergence in T. laurina when included as a separate variable in the three-way AMOVA. The four-way AMOVA indicates there is greater variation within catchments than between catchments, discounting the hypothesis that hydrochory facilitates long-distance dispersal within catchments. Finally, using Welch’s t-test we found greater genetic diversity in T. laurina across all estimates and all lineages (P < 0.001; Supplementary Table S1), suggesting longer population persistence in this species.
Inference of post-LGM habitat shifts
The modelled present-day distributions of Tristaniopsis are consistent with their observed occurrences (excluding the full model of T. laurina), and the average area-under-the-curve scores (all models >0.929) indicate a good model fit (Supplementary Table S3). The full distribution models under-predicted habitat availability relative to the lineage trained models, though both methods generally agreed. Likewise, there was overall concordance between models derived from the different General Circulation Models. Excluding the LGM models for T. laurina North and South and the HCO models for T. collina South, visual inspection showed at least 75% of General Circulation Models predicted the same habitat suitability distributions for each lineage. However, some General Circulation Models were spatially inconsistent in the amount of habitat suitability predicted across the study region when compared to the lineage means.
Overall, the full-distribution and lineage-based ENMs indicate a larger and more stable environmental envelope for T. laurina than for T. collina, with contrasting habitat suitability shifts since the LGM (Fig. 4; Supplementary Fig. S5). The models show long-term habitat barriers across the Clarence River Corridor, Hunter River Corridor, and Cumberland Plain, which is consistent with the lineage boundaries identified in T. collina assuming K = 4. Interestingly, T. laurina shows greater connectivity across these barriers, despite the greater genetic differentiation within this species.
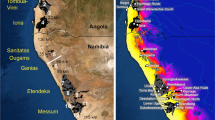
Environmental niche modelling for three lineages of Tristaniopsis collina (a–c) and T. laurina (d–f) in eastern New South Wales. For each lineage, three Holocene climate scenarios are shown: the Last Glacial Maximum (LGM; c. 21,000 years ago), the Holocene Climatic Optimum (HCO; c. 6000 years ago), and the present day. The predictor variables and climatic models used for the LGM and HCO distributions are listed in Supplementary Tables S2 and S3. The colour scale refers to the logistic probability (range: 0–1) of habitat suitability in the study area
For T. collina, we produced a combined Shield Volcano and North model because there were insufficient data points to train a separate model for Shield Volcano. The models indicate putative refugia (habitat suitability >0.7) at three upland regions for Shield Volcano–North (Fig. 4a). However, significant local and regional habitat contractions were projected for the HCO (Fig. 4a). The ENMs for Central projected low LGM and HCO habitat suitability (Fig. 4b), though identified microrefugia on either side of the Hunter River Corridor. The models indicate that the lineage underwent a large-scale habitat expansion between the HCO and the present day. The models for South suggest fairly stable fragments of suitable habitat across each period, though a rapid expansion of habitat availability was inferred between the LGM and HCO (Fig. 4c). All ENMs (including trial models with different predictor variables) identified suitable habitat in Victoria. This is outside the current range of T. collina and may indicate range disequilibrium with the environment.
From the North model for T. laurina, we identified three large areas of suitable habitat that expanded between the LGM and HCO; and contracted again in the present day (Fig. 4d). We did not find suitable LGM habitat in Central (Fig. 4e), and the models suggest that the species underwent significant expansion from coastal microrefugia to inland regions, particularly during the HCO. Finally, the models for South identify fragments of refugia in southeast Victoria and along the extent of the southern NSW coastline (Fig. 4f). The models also suggest significant habitat expansion from refugia between the LGM and HCO, and a subsequent contraction under the current climate.
Landscape genetic analyses
Due to multicollinearity between the distance matrices, we were not able to reliably assess models with more than two independent variables. We found an overall weak influence of IBD and IBR variables on FST in both species. Support from the single GCM models was mixed. While the regression coefficient of IBR variables was often congruent with the mean model, many lost significance with the inclusion of IBD. While this reduced our ability to assess the relative contribution of each variable to genetic isolation, we were able to infer that dispersal constraints slightly differ among species, and among lineages within species (Table 2).
The effect of IBD on genetic differentiation within T. collina is small and its significance was usually lost when IBR variables were included in the models. IBRHCO and IBRSTABILITY best explained pairwise FST across the full range of T. collina as did IBRLGM in Shield Volcano–North; though all IBR variables were significant. For the other lineages, IBRLGM in South was the only significant variable when other variables were controlled for. The IBR variables had a greater influence on the full range model of T. laurina compared with T. collina, and IBRSTABILITY was the most influential. There was no support for the influence of IBR variables in the individual lineages. We found a moderate influence of IBD within Sydney and South in the partial mantels, though not in the multiple matrix regression models when IBR variables were included.
For both species, we found a significant negative correlation between AR and uHE with latitude in North and a significant positive correlation in South (Table 3). The same trends were also significant for HO in T. laurina. This suggests that genetic diversity increases at lineage boundaries with Central, potentially due to admixture between lineages. We did not find significant correlations between diversity values and IBRSTABILITY (all P > 0.1).
Range expansion models
For T. collina, we produced a single Shield Volcano–North model because there were insufficient data points to produce separate models for each of the northern groupings identified in the PCA. We also included models of the Central and Shield Volcano–North groups both with and without the Barrington region, owing to the uncertain boundary between the two lineages in this species. Overall, there was strong support (P < 0.0001) for the serial-founder expansion model among T. collina lineages, indicating rapid or recent range expansion (Table 4). Models for Shield Volcano–North inferred a primarily southward expansion originating west of the Border Ranges; though this is not consistent with the multiple refugia identified from by the palaeo-environment models. The strongest founder effect was detected in Central (without Barrington), and we inferred the expansion origin from a mid-altitude site in the Sydney Basin to montane areas in the Greater Blue Mountains. This is consistent with the habitat expansion inferred from the palaeo-environment models. For South, we inferred a range expansion from the southernmost range limit of T. collina. However, the low R2 (0.4113) reduces our confidence in the predicted site of origin. The weak founder effects in this lineage are consistent with the greater habitat connectivity and stability inferred from the ENMs.
By comparison, founder effects in T. laurina North and South are one order of magnitude weaker (Table 4); suggesting older, slower, or more reticulate range expansion scenarios in this species. For North, we inferred a predominantly north-to-south range expansion from a lowland coastal site. The weak founder effects are consistent with the stable habitat connectivity inferred from the ENMs. For South, we inferred a multidirectional expansion from the far south coast of NSW. The moderate founder effects are not consistent with the high habitat availability inferred for this region. Model support was not significant for Central (P < 0.01) which indicates that the observed structure in this lineage cannot be differentiated from equilibrium isolation by distance.
Discussion
A broader aim of this study was to investigate the relative importance of dispersal versus persistence on the range dynamics of long-lived trees, and whether this relates to habitat preference. We found that the tree species Tristaniopsis collina and T. laurina have similar restrictions across major biogeographic barriers and overall genetic structure, despite evidence of significant environmental niche differentiation and contrasting habitat shifts under Holocene climate change. However, our range expansion models and landscape genetic analyses do reveal cryptic differences in the dispersal potential of the study species and their persistence in refugia. We discuss these findings in reference to the bioclimatic preferences, habitat connectivity and recruitment of our two study species.
Niche divergence in Tristaniopsis
As a first step towards demonstrating the impact of niche preference on range dynamics, we verified that environmental niche separation between the study species is a function of habitat selection. The species are significantly differentiated across environmental space, and we confirmed a smaller bioclimatic niche for T. collina. This can be attributed to the tendency of T. collina to occupy higher altitudes across its range, where it encounters lower temperatures and greater precipitation than does T. laurina. However, the coarse resolution of the bioclimatic variables must be interpreted with caution. Considering that T. laurina is known to be drought sensitive (Melick 1990a), its greater distribution in slightly warmer-drier climates than T. collina should not be taken to indicate that the species is better adapted to those conditions. A more likely explanation is that the riparian habitats of T. laurina are sometimes nested within areas that receive less precipitation than would otherwise be suitable. So, although T. laurina is spatially restricted at a local scale, the riparian microclimate enables the species to occupy a broader geographic range.
Equivalent dispersal across ancient habitat barriers
We used multiple clustering methods to investigate the influence of ancient habitat barriers on within-species divergence. Congruence with the PCA ordination suggests that the ancestral lineages presented in Fig. 1 are a good approximation of the overarching genetic structure of the study species, though hierarchical structure north of the Hunter may have led to an under-estimation of K in T. collina. The continuous genetic variation observed within T. laurina lineages suggests K = 3 is most parsimonious. However, the algorithm implemented by sNMF cannot differentiate between recent admixture and other demographic histories. Consequently, admixed populations might be the outcome of admixture with an extinct lineage or a recent bottleneck (Lawson et al. 2018) rather than secondary contact. Given our thorough sample scheme, it is unlikely the results have been confounded by un-sampled genotypes.
We anticipated that environmental niche differentiation and contrasting range shifts would have varied the influence of biogeographic barriers on the study species. While our palaeo-environment models suggest that the two species had differing degrees of habitat availability over the last 21,000 years, both species show divergence across dry barriers north and south of Sydney that likely pre-dates the LGM. This includes the Cumberland Plain, a low altitude 2750 km² basin located in the rain shadow between the Greater Blue Mountains and coastal Sydney (Fig. 1). To our knowledge, this study is the first to highlight the region as an ancient-dry genetic barrier, and it is interesting that such restrictions are not evident in rainforest trees genotyped from the region (e.g. Mellick et al. 2011; Heslewood et al. 2014).
Our expectation that dispersal mechanism or environmental niche would differentiate the impact of biogeographic barriers on the study species was not supported. Although we found that catchment connectivity facilitates gene flow within T. laurina, the overall equivalent partitioning of genetic diversity within each of the study species suggests that they are characterized by similar dispersal potential (Duminil et al., 2007). This either discounts hydrochory as a frequent mechanism of long-distance dispersal in T. laurina or suggests other factors regulate gene flow (see section below). Other studies have demonstrated that dispersal traits such as fruit size (Rossetto et al. 2009; Rossettoet et al. 2015a) and fruit fleshiness (Rossetto, Mcpherson, et al. 2015; Worth et al. 2017) have a greater influence on migration across barriers. Likewise, compared with co-distributed trees from wet versus dry forests (Milner et al. 2012; Worth et al. 2017) or nutrient-rich versus nutrient-poor soils (Rossetto et al. 2009), niche differentiation did not significantly vary barrier effects between the study species. Nevertheless, higher rainfall requirements can account for the greater structure of T. collina across the dry valley systems of northern NSW. In contrast, the palaeo-models suggest that these barriers were less important for T. laurina after the LGM.
Dispersal or persistence under Holocene climate change
The palaeo-environment models and genetic data indicate that range shifts through the Holocene can partially account for genetic variation within T. collina and we deduce that greater climate sensitivity has reduced its persistence potential compared with T. laurina. An increasing number of studies have identified high-altitude LGM refugia for tree species in areas outside their current bioclimatic limits (Nevill et al. 2014; Zeng et al. 2015). Yet for T. collina, we inferred a serial-founder range expansion from midland refugia to high-altitude localities in Central. This is consistent with palynological surveys in the region that have identified the LGM treeline to be as low as 1000 m (Hesse et al. 2003) (Sweller and Martin 2001). As a gravity-dispersed tree, range expansion in T. collina is likely to have occurred through a series of stepping-stone colonization events. This demonstrates that tree species are capable of rapid climate-driven range shifts even in the absence of long-distance dispersal mechanisms.
In contrast, T. laurina exhibits slightly higher genetic diversity and stronger population structure more consistent with spatial conservatism and periodic isolation (Hewitt 1996) than with rapid range expansion. Combined with the palaeo-environment models, this suggests that persistence in multiple refugia has maintained a relatively stable distribution of T. laurina through the Holocene. Meanwhile, weak founder effects suggest that as conditions improved during the HCO, the species underwent only gradual and/or localized expansions. Localized elevational shifts have been inferred for montane trees across a range of environments in response to past climatic change (Gugger et al. 2013; Mellick et al. 2013b, 2014; Nevill et al. 2014). Here we find evidence for in situ persistence during the last interglacial, which supports the hypothesis that riparian sites can provide important microrefugia (de Lafontaine et al. 2014) particularly for lowland species (Aide and Rivera 1998).
Following observations of co-distributed trees in other ecosystems, we anticipated that genetic structure within Tristaniopsis would reflect their differing sensitivities to Pleistocene climatic change (Collevatti et al. 2001, 2013, 2014; Ramos et al. 2007; Novaes et al. 2010, 2013; Souza et al. 2017). The greater structure in T. laurina was therefore surprising given its broader environmental niche and potential for connectivity along riparian corridors. This disparity implies that species differences are not fully explained by their past or current habitat availability. We hypothesize that specialized recruitment traits limit the colonization success of T. laurina and reduce genetic connectivity relative to T. collina. For instance, T. laurina seedlings are adapted to the nutrient and light pulses that follow flood-disturbance events (Melick 1990a, b) and so colonization is rare in the absence of high rainfall and disturbance (Melick and Ashton 1991). To explore this hypothesis, it is worth investigating whether T. collina seedlings germinate and colonize new habitats more readily than T. laurina.
Like other species that have had a relatively continuous (albeit fragmented) distribution of suitable habitat since the LGM (e.g. Ortego et al. 2015), we did not find a relationship between genetic diversity and habitat stability. This contrasts with tree species that underwent more dramatic range shifts from a large central refugium, in which diversity is either associated with habitat stability or distance from a refugium core (Collevatti et al. 2013; Tsuda et al. 2015; Zeng et al. 2015; Souza et al. 2017; Kim et al. 2018). However, in both Tristaniopsis species, we did find a latitudinal cline in diversity that increased at sites of admixture between the putative lineages, supporting a scenario of secondary contact following Holocene warming. Similar patterns of elevated diversity in admixed zones has been widely reported in trees from the Northern Hemisphere (Comps et al. 2001; Walter and Epperson 2001; Petit et al. 2003; Heuertz et al. 2004; Sakaguchi et al. 2011; Lee et al. 2014; Havrdová et al. 2015).
As we found a weak signal IBD and mixed support for IBR in both our study species, we must consider potential shortcomings in their palaeo-environment models. This could include overfitting or inappropriate variable selection, as we only calculated resistance matrices from one set of variables per lineage. It is also possible that the genetic structure in the species is more closely associated with the climate of other periods than those explored in the present study. Finally, we acknowledge that other population models might be more appropriate for our species. For instance, isolation-by-environment is widely reported in plant species (Sexton et al. 2014; Wang and Bradburd 2014). Alternatively, selection against migrants can reinforce founder effects created by range expansions, and drive a pattern of isolation-by-colonization that overrides the influence of environmental gradients (Orsini et al. 2013).
Range dynamics vary between regions
Finally, we have demonstrated that climate-driven range shifts can vary significantly across the distribution of widespread species. This was particularly evident among the different lineages of T. collina. Regional differences can be explained in terms of the latitudinal climatic gradient along eastern Australia that would have been differently affected by Holocene warming, as well as the spatial variation in topographic relief.
Tristaniopsis laurina also showed regional differentiation. As has been demonstrated in other riparian plants, mountain ridges between rivers act as genetic barriers whilst river valleys act as corridors for gene flow (Tero et al. 2003; Fer and Hroudava 2008; Wei et al. 2013; Werth and Scheidegger 2014). Following this, the greater genetic structure within South might be attributed to the more dissected landscape in which the lineage is located, compared with North where low topographic relief across extensive coastal plains is likely to facilitate greater dispersal within and between catchments.
Conclusions
This study investigated environmental niche partitioning and its impact on the range dynamics of two widely distributed, closely related sympatric trees in response to Holocene climate change. We conclude that riparian specialization has enabled Tristaniopsis laurina to occupy a broader climatic envelope and persist in multiple refugia across its range, while specialized recruitment traits potentially limit colonization of new habitat. In contrast, a narrow climatic niche has driven a history of habitat instability and larger-scale range shifts for T. collina throughout the Holocene. However as both species had refugia across their respective ranges, we can expect more dramatic differences in the range dynamics of species that occupy different soil types or biomes.
The patterns observed in Tristaniopsis have several implications for the study of range shifts and evolution in response to climate change. First, we have demonstrated cryptic differences in the historical ranges of species with present-day sympatry and similar biogeographic restrictions. This reveals how divergent processes can produce similar distributional patterns and highlights the utility of SNP-based demographic models to differentiate between alternative drivers of population structure, such as drift via founder effects versus long-term geographic isolation. We also found that a greater niche breadth and capacity for long-distance dispersal does not necessarily enhance species’ migration across habitat barriers or promote post-glacial range expansion. Finally, our results demonstrate that spatial variation in landscape barriers and climatic gradients can create regional differences in the range dynamics of widely distributed species.
Further research is needed to address some of the uncertainties in our study. In particular, the history of lineage divergence and admixture requires clarification. The mixed support for IBR and weak influence of IBD also suggests that other potential drivers of genetic variation need to be explored. Adaptive divergence could be frequent within species like T. laurina that have experienced local climate-driven range shifts but maintained stable ranges over time (Gugger et al. 2013; Ortego et al. 2015). Coalescent analyses can help verify the within-species lineages we identified and test whether the weak founder effects in T. laurina reflect a history of gradual, localized and/or ancient expansions. This could be combined with gene-environment association analyses to investigate climate adaptation as an alternative driver of population structure, and whether this differs between the two species. Finally, germination trials are required to test whether recruitment traits differentiate the colonization and range dynamics of T. collina and T. laurina.
Data archiving statement
Genomic data in the form of binary-coded SNPs (DArTseq) and geographic coordinates for all samples are available from Dryad: https://doi.org/10.5061/dryad.0p6bj53
References
Ackerly DD (2003) Community assembly, niche conservatism, and adaptive evolution in changing environments. Int J Plant Sci 164:S165–S184
Adamack AT, Gruber B (2014) PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol 5:384–387
Aide TM, Rivera E (1998) Geographic patterns of genetic diversity in Poulsenia armata (Moraceae): implications for the theory of Pleistocene refugia and the importance of riparian forest. J Biogeogr 25:695–705
Banta JA, Ehrenreich IM, Gerard S, Chou L, Wilczek A, Schmitt J et al. (2012) Climate envelope modelling reveals intraspecific relationships among flowering phenology, niche breadth and potential range size in Arabidopsis thaliana. Ecol Lett 15:769–777
Bjornstad ON, Cai J (2018) Package ‘ncf’: Spatial Covariance Functions. pp. 1–44
Broennimann O, Fitzpatrick MC, Pearman PB, Petitpierre B, Pellissier L, Yoccoz NG et al. (2012) Measuring ecological niche overlap from occurrence and spatial environmental data. Glob Ecol Biogeogr 21:481–497
Castellanos MC, Wilson P, Thomson JD (2003) Pollen transfer by hummingbirds and bumblebees, and the divergence of pollination modes in Penstemon. Evolution 57:2742–2752
Chapple DG, Chapple SNJ, Thompson MB (2011) Biogeographic barriers in south-eastern Australia drive phylogeographic divergence in the garden skink, Lampropholis guichenoti. J Biogeogr 38:1761–1775
Chase JM, Leibold MA (2003) Comparing Classical and Contemporary Niche Theory. Ecological niches: linking classical and contemporary approaches. University of Chicago Press, Chicago, pp 51–59
Clark CM, Carbone I (2008) Chloroplast DNA phylogeography in long-lived Huon pine, a Tasmanian rain forest conifer. Can J Res 38:1576–1589
Di Cola V, Broennimann O, Petitpierre B, Breiner FT, D’Amen M, Randin C et al. (2017) ecospat: an R package to support spatial analyses and modeling of species niches and distributions. Ecography 40:774–787
Collevatti RG, Grattapaglia D, Hay JD (2001) Population genetic structure of the endangered tropical tree species Caryocar brasiliense, based on variability at microsatellite loci. Mol Ecol 10:349–356
Collevatti RG, Lima-Ribeiro MS, Terribile LC, Guedes LBS, Rosa FF, Telles MPC (2014) Recovering species demographic history from multi-model inference: the case of a Neotropical savanna tree species. BMC Evol Biol 14:1–13
Collevatti RG, Telles MPC, Nabout JC, Chaves LJ, Soares TN (2013) Demographic history and the low genetic diversity in Dipteryx alata (Fabaceae) from Brazilian Neotropical savannas. Heredity 111(2):97–105
Comps B, Gömöry D, Letouzey J, Thiébaut B, Petit RJ (2001) Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 157:389–397
Devitt TJ, Devitt SEC, Hollingsworth BD, McGuire JA, Moritz C (2013) Montane refugia predict population genetic structure in the Large-blotched Ensatina salamander. Mol Ecol 22:1650–1665
Duminil J, Fineschi S, Hampe A, Jordano P, Salvini D, Vendramin GG et al. (2007) Can population genetic structure be predicted from life‐history traits? Am Nat 169:662–672
Eckert AJ, Bower AD, Wegrzyn JL, Pande B, Jermstad KD, Krutovsky KV et al. (2009) Association genetics of coastal Douglas fir (Pseudotsuga menziesii var. menziesii, Pinaceae). I. Cold-hardiness related traits. Genetics 182:1289–1302
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fer T, Hroudava Z (2008) Detecting dispersal of Nuphar lutea in river corridors using microsatellite markers. Freshw Biol 53:1409–1422
Fournier-Level A, Korte A, Cooper MD, Nordborg M, Schmitt J, Wilczek AM (2011) A map of local adaptation in Arabidopsis thaliana. Science 334:86–89
Frichot E, Mathieu F, Trouillon T, Bouchard G, François O (2014) Fast and efficient estimation of individual ancestry coefficients. Genetics 196:973–983
Garnier S, Alibert P, Audiot P, Prieur B, Rasplus JY (2004) Isolation by distance and sharp discontinuities in gene frequencies: implications for the phylogeography of an alpine insect species, Carabus solieri. Mol Ecol 13:1883–1897
Gruber B, Adamack AT (2015) landgenreport: a new R function to simplify landscape genetic analysis using resistance surface layers. Mol Ecol Resour 15:1172–1178
Gugger P, Ikegami M, Sork V (2013) Influence of late Quaternary climate change on present patterns of genetic variation in valley oak, Née. Mol Ecol 22:3598–3612
Hancock AM, Brachi B, Faure N, Horton MW, Jarymowycz LB, Sperone FG et al. (2011) Adaptation to Climate Across the Arabidopsis thaliana genome. Science 334:83–86
Havrdová A, Douda J, Krak K, Vít P, Hadincová V, Zákravský P et al. (2015) Higher genetic diversity in recolonized areas than in refugia of Alnus glutinosa triggered by continent-wide lineage admixture. Mol Ecol 24:4759–4777
Heslewood MM, Lowe AJ, Crayn DM, Rossetto M (2014) Contrasting levels of connectivity and localised persistence characterise the latitudinal distribution of a wind-dispersed rainforest canopy tree. Genetica 142:251–264
Hesse PP, Humphreys GS, Selkirk PM, Adamson DA, Gore DB, Nobes DC et al. (2003) Late Quaternary aeolian dunes on the presently humid Blue Mountains, Eastern Australia. Quat Int 108:13–32
Heuertz M, Hausman J-F, Hardy OJ, Vendramin GG, Frascaria-Lacoste N, Vekemans X (2004) Nuclear microsatellites reveal contrasting patterns of genetic structure between Western and Southeastern European populations of the Common Ash (Fraxinus Eexcelsior L.). Evolution 58:976–988
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58:247–276
Hornsby AD, Matocq MD (2012) Differential regional response of the bushy-tailed woodrat (Neotoma cinerea) to late Quaternary climate change. J Biogeogr 39:289–305
Jezkova T, Jaeger JR, Marshall ZL, Riddle BR (2009) Pleistocene impacts on the phylogeography of the Desert Pocket Mouse (Chaetodipus penicillatus). J Mammal 90:306–320
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jombart T, Ahmed I (2011) adegenet 1.3: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:3070–3071
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Kamvar ZN, Brooks JC, Grünwald NJ (2015) Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Front Genet 6:208
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr : an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281
Keenan K, McGinnity P, Cross TF, Crozier WW, Prodöhl PA (2013) diveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol Evol 4:782–788
Kershaw AP, Bretherton SC, van der Kaars S (2007) A complete pollen record of the last 230 ka from Lynch’s Crater, north-eastern Australia. Palaeogeogr Palaeoclim Palaeoecol 251:23–45
Kim S-H, Cho M-S, Li P, Kim S-C (2018) Phylogeography and ecological niche modeling reveal reduced genetic diversity and colonization patterns of Skunk Cabbage (Symplocarpus foetidus; Araceae) from glacial refugia in eastern North America. Front Plant Sci 9:648
de Lafontaine G, Amasifuen Guerra CA, Ducousso A, Petit RJ (2014) Cryptic no more: soil macrofossils uncover Pleistocene forest microrefugia within a periglacial desert. New Phytol 204:715–729
Lawson DJ, van Dorp L, Falush D (2018) A tutorial on how not to over-interpret STRUCTURE and ADMIXTURE bar plots. Nat Commun 9:3258
Lee JH, Lee DH, Choi IS, Choi BH (2014) Genetic diversity and historical migration patterns of an endemic evergreen oak, Quercus acuta, across Korea and Japan, inferred from nuclear microsatellites. Plant Syst Evol 300:1913–1923
McRae BH, Beier P (2007) Circuit theory predicts gene flow in plant and animal populations. Proc Natl Acad Sci USA 104:19885–19890
Melick DR (1990a) Relative drought resistance of Tristaniopsis laurina and Acmena smithii from riparian warm temperate rainforest in Victoria. Aust J Bot 38:361–370
Melick DR (1990b) Regenerative succession of Tristaniopsis laurina and Acmena smithii in riparian warm temperate rainforest in Victoria, in relation to light and nutrient regimes. Aust J Bot 38:111–120
Melick DR, Ashton DH (1991) The effects of natural disturbances on warm temperate rain-forests in South-eastern Australia. Aust J Bot 39:1–30
Mellick R, Lowe A, Allen C, Hill RS, Rossetto M (2012) Palaeodistribution modelling and genetic evidence highlight differential post-glacial range shifts of a rain forest conifer distributed across a latitudinal gradient. J Biogeogr 39:2292–2302
Mellick R, Lowe A, Rossetto M (2011) Consequences of long- and short-term fragmentation on the genetic diversity and differentiation of a late successional rainforest conifer. Aust J Bot 59:351–362
Mellick R, Rossetto M, Allen C, Wilson P, Hill R, Lowe A (2013a) Intraspecific divergence associated with a biogeographic barrier and climatic models show future threats and long-term decline of a rainforest conifer. Open Conserv Biol J 7:1–10
Mellick R, Wilson PD, Rossetto M (2013b) Post-glacial spatial dynamics in a rainforest biodiversity hot spot. Diversity 5:124–138
Mellick R, Wilson PD, Rossetto M (2014) Demographic history and niche conservatism of tropical rainforest trees separated along an altitudinal gradient of a biogeographic barrier. Aust J Bot 62:438–450
Milner ML, Rossetto M, Crisp MD, Weston PH (2012) The impact of multiple biogeographic barriers and hybridization on species-level differentiation. Am J Bot 99:2045–2057
Naimi B (2013) usdm: Uncertainty analysis for species distribution models. R Packag version 1:1–12
Nevill PG, Bossinger G, Ades PK (2010) Phylogeography of the world’s tallest angiosperm, Eucalyptus regnans: Evidence for multiple isolated Quaternary refugia. J Biogeogr 37:179–192
Nevill PG, Després T, Bayly MJ, Bossinger G, Ades PK (2014) Shared phylogeographic patterns and widespread chloroplast haplotype sharing in Eucalyptus species with different ecological tolerances. Tree Genet Genomes 10:1079–1092
Nilsson C, Brown RL, Jansson R, Merritt DM (2010) The role of hydrochory in structuring riparian and Wetland vegetation. Biol Rev 85:837–858
Novaes RML, De Lemos Filho JP, Ribeiro RA, Lovato MB (2010) Phylogeography of Plathymenia reticulata (Leguminosae) reveals patterns of recent range expansion towards northeastern Brazil and southern Cerrados in Eastern Tropical South America. Mol Ecol 19:985–998
Novaes RML, Ribeiro RA, Lemos-Filho JP, Lovato MB (2013) Concordance between phylogeographical and biogeographical patterns in the Brazilian Cerrado: diversification of the endemic tree Dalbergia miscolobium (Fabaceae). PLoS ONE 8:e82198
Orsini L, Vanoverbeke J, Swillen I, Mergeay J, De Meester L (2013) Drivers of population genetic differentiation in the wild: isolation by dispersal limitation, isolation by adaptation and isolation by colonization. Mol Ecol 22:5983–5999
Ortego J, Gugger PF, Sork VL (2015) Climatically stable landscapes predict patterns of genetic structure and admixture in the Californian canyon live oak. J Biogeogr 42:328–338
Peter BM, Slatkin M (2013) Detecting range expansions from genetic data. Evolution 67:3274–3289
Peter BM, Slatkin M (2015) The effective founder effect in a spatially expanding population. Evolution 69:721–734
Petit RJ, Aguinagalde I, de Beaulieu J-L, Bittkau C, Brewer S, Cheddadi R et al. (2003) Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300:1563–1565
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Model 6:231–252
Phillips SJ, Dudík M (2008) Modeling of species distributions with Maxent: new extensions and a comprehensive evaluation. Ecography 31:161–175
Playford J, Bell J, Moran G (1993) A major disjunction in genetic diversity over the geographic range of Acacia melanoxylon R.Br. Aust J Bot 41:355–368
Powney GD, Roy DB, Chapman D, Brereton T, Oliver TH (2011) Measuring functional connectivity using long-term monitoring data. Methods Ecol Evol 2:527–533
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ramos ACS, Lemos-Filho JP, Ribeiro RA, Santos FR, Lovato MB (2007) Phylogeography of the tree Hymenaea stigonocarpa (Fabaceae: Caesalpinioideae) and the influence of quaternary climate changes in the Brazilian cerrado. Ann Bot 100:1219–1228
Rossetto M, Bragg J, Kilian A, McPherson H, van der Merwe M, Wilson P (2018) Restore and Renew: a genomics-era framework for species provenance delimitation. Restor Ecol 27:538–548
Rossetto M, Crayn D, Ford A, Mellick R, Sommerville K (2009) The influence of environment and life-history traits on the distribution of genes and individuals: a comparative study of 11 rainforest trees. Mol Ecol 18:1422–1438
Rossetto M, Crayn D, Ford A, Ridgeway P, Rymer P (2007) The comparative study of range-wide genetic structure across related, co-distributed rainforest trees reveals contrasting evolutionary histories. Aust J Bot 55:416–424
Rossetto M, Kooyman R, Yap JYS, Laffan SW (2015a) From ratites to rats: the size of fleshy fruits shapes species’ distributions and continental rainforest assembly Proc R Soc B Biol Sci 282:20151998
Rossetto M, Mcpherson H, Siow J, Kooyman R, van der Merwe M, Wilson PD (2015b) Where did all the trees come from? A novel multispecies approach reveals the impacts of biogeographical history and functional diversity on rain forest assembly. J Biogeogr 42:2172–2186
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rosas-Guerrero V, Quesada M, Armbruster WS, Pérez-Barrales R, Smith SDW (2011) Influence of pollination specialization and breeding system on floral integration and phenotypic variation in ipomoea. Evolution 65:350–364
Sakaguchi S, Takeuchi Y, Yamasaki M, Sakurai S, Isagi Y (2011) Lineage admixture during postglacial range expansion is responsible for the increased gene diversity of Kalopanax septemlobus in a recently colonised territory. Heredity 107(4):338–348
Sansaloni C, Petroli C, Jaccoud D, Carling J, Detering F, Grattapaglia D et al. (2011) Diversity Arrays Technology (DArT) and next-generation sequencing combined: genome-wide, high throughput, highly informative genotyping for molecular breeding of Eucalyptus. BMC Proc 5:54
Schoener TW (1970) Nonsynchronous spatial overlap of lizards in patchy habitats. Ecology 51:408–418
Sexton JP, Hangartner SB, Hoffmann AA (2014) Genetic isolation by environment or distance: which pattern of gene flow is most common? Evolution 68:1–15
Shah VB, McRae BH (2008) Circuitscape: a tool for landscape ecology. Proc 7th Python Sci Conf (SciPy 2008) 22:836–851
Slatkin M (1993) Isolation by distance in equilibrium and non-equilibrium populations. Evolution 47:264–279
Sork VL, Davis FW, Westfall R, Flint A, Ikegami M, Wang H et al. (2010) Gene movement and genetic association with regional climate gradients in California valley oak (Quercus lobata Née) in the face of climate change. Mol Ecol 19:3806–3823
Souza EHAV, Collevatti RG, Lima-Ribeiro MS, De Lemos-Filho JP, Lovato MB (2017) A large historical refugium explains spatial patterns of genetic diversity in a Neotropical savanna tree species. Ann Bot 119:239–252
Svenning JC, Skov F (2007) Could the tree diversity pattern in Europe be generated by postglacial dispersal limitation? Ecol Lett 10:866
Sweller S, Martin HA (2001) A 40,000 year vegetation history and climatic interpretations of Burraga Swamp, Barrington Tops, New South Wales. Quat Int 8385:233–244
Tero N, Aspi J, Siikamäki P, Jäkäläniemi A, Tuomi J (2003) Genetic structure and gene flow in a metapopulation of an endangered plant species, Silene antatarica. Mol Ecol 12:2073–2085
Thom B, Hesp P, Bryant E (1994) Last glacial ‘coastal’ dunes in Eastern Australia and implications for landscape stability during the Last Glacial Maximum. Palaeogeogr Palaeoclim Palaeoecol 111:229–248
Thomson JD, Wilson P, Valenzuela M, Malzone M (2000) Pollen presentation and pollination syndromes, with special reference to Penstemon. Plant Species Biol 15:11–29
Tsuda Y, Nakao K, Ide Y, Tsumura Y (2015) The population demography of Betula maximowicziana, a cool-temperate tree species in Japan, in relation to the last glacial period: its admixture-like genetic structure is the result of simple population splitting not admixing. Mol Ecol 24:1403–1418
Walter R, Epperson BK (2001) Geographic pattern of genetic variation in Pinus resinosa: area of greatest diversity is not the origin of postglacial populations. Mol Ecol 10:103–111
Wang IJ (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evolution 67:3403–3411
Wang IJ, Bradburd GS (2014) Isolation by environment. Mol Ecol 23:5649–5662
Wei X, Meng H, Jiang M (2013) Landscape genetic structure of a streamside tree species Euptelea pleiospermum (Eupteleaceae): contrasting roles of river valley and mountain ridge. PLoS ONE 8:e66928
Weir BS, Cockerham CC (1984) Estimating F-Statistics for the analysis of population structure. Evolution 38:1358
Werth S, Scheidegger C (2014) Gene flow within and between Catchments in the threatened riparian plant Myricaria germanica (GG Vendramin, Ed.). PLoS ONE 9:e99400
Whetton P, Ekström M, Gerbing C, Grose M, Bhend J, Webb L, Risbey J, Holper P, Clarke J, Hennessy K, Colman R, Moise A, Power S, Braganza K, Watterson I, Murphy B, Timbal B, Hope P, Dowdy AL (2015) Climate change in Australia information for Australia’s natural resource management regions: Technical report. Canberra
Williams, G, Adam P (2010) The Flowering of Australia’s Rainforests: A Plant and Pollination Miscellany, First. CSIRO Publishing: Canberra
Williams JW, Jackson ST (2007) Novel climates, no-analog communities, and ecological surprises. Front Ecol Environ 5:475–482
Wilson PG (1991) New South Wales Flora Online. PlantNET - Plant Inf Netw Syst R Bot Gard Domain Trust Sydney, Aust. http://plantnet.rbgsyd.nsw.gov.au/
Wilson PG, Waterhouse JT (1982) A review of the genus Tristania R. Br. (Myrtaceae): a heterogeneous assemblage of five genera. Aust J Bot 30:413–446
Wilson TC, Conn BJ, Henwood MJ (2017) Great Expectations: Correlations between Pollinator Assemblages and Floral Characters in Lamiaceae. Int J Plant Sci 178:170–187
Worth JRP, Holland BR, Beeton NJ, Schönfeld B, Rossetto M, Vaillancourt RE et al. (2017) Habitat type and dispersal mode underlie the capacity for plant migration across an intermittent seaway. Ann Bot 120:539–549
Worth JRP, Jordan GJ, Marthick JR, McKinnon GE, Vaillancourt RE (2010) Chloroplast evidence for geographic stasis of the Australian bird-dispersed shrub Tasmannia lanceolata (Winteraceae). Mol Ecol 19:2949–2963
Worth JRP, Jordan GJ, McKinnon GE, Vaillancourt RE (2009) The major Australian cool temperate rainforest tree Nothofagus cunninghamii withstood Pleistocene glacial aridity within multiple regions: Evidence from the chloroplast. New Phytol 182:519–532
Worth JRP, Marthick JR, Jordan GJ, Vaillancourt RE (2011) Low but structured chloroplast diversity in Atherosperma moschatum (Atherospermataceae) suggests bottlenecks in response to the Pleistocene glacials. Ann Bot 108:1247–1256
Worth JRP, Williamson GJ, Sakaguchi S, Nevill PG, Jordan GJ (2014) Environmental niche modelling fails to predict Last Glacial Maximum refugia: niche shifts, microrefugia or incorrect palaeoclimate estimates? Glob Ecol Biogeogr 23:1186–1197
Xu T, Hutchinson MF (2013) New developments and applications in the ANUCLIM spatial climatic and bioclimatic modelling package. Environ Model Softw 40:267–279
Zeng YF, Wang WT, Liao WJ, Wang HF, Zhang DY (2015) Multiple glacial refugia for cool-temperate deciduous trees in northern East Asia: The Mongolian oak as a case study. Mol Ecol 24:5676–5691
Acknowledgements
We thank P.G. Wilson for helpful discussions about the study species, and M.S. Fellows and C.M. Hoare for assisting with collections. Funding was provided by NSW Adaptation Research Hub (Office of Environment and Heritage) and the Restore & Renew flagship project (Botanic Gardens & Centennial Parklands).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fahey, M., Rossetto, M., Wilson, P.D. et al. Habitat preference differentiates the Holocene range dynamics but not barrier effects on two sympatric, congeneric trees (Tristaniopsis, Myrtaceae). Heredity 123, 532–548 (2019). https://doi.org/10.1038/s41437-019-0243-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-019-0243-x