Abstract
Many Odonata species exhibit female-limited polymorphisms, where one morph is similar to the conspecific male in body color and other traits (andromorph), whereas one or more other morphs differ from the male (gynomorphs). Here we investigated the differentially expressed transcripts (DETs) among males and two female morph groups (gynomorphs and andromorphs) using RNA-seq to identify candidate transcripts encoding female-limited polymorphisms in the damselfly Ischnura senegalensis. Seven DETs that had significantly different expression levels between males and gynomorphs, but not between males and andromorphs, were identified. The expression levels of four of these candidate genes, doublesex (dsx), black, ebony, and chaoptin (chp), were selected for further analysis using qRT-PCR. Sequence analysis of the dsx amplicons revealed that this gene produced at least three transcripts. Two short transcripts were mainly expressed in males and andromorphs, whereas the long transcript was specifically expressed in both morph female groups; that is, the expression pattern of the dsx splice variants in andromorphs was an intermediate between that of males and gynomorphs. Because the dsx gene functions as a transcription factor that regulates the sex-specific expression of multiple genes, its splice variants in I. senegalensis may explain why the andromorph is female but exhibits some masculinized traits. Because we did not detect different coding sequences of the candidate genes among the different morphs, a diallelic genomic region controlling alternative splicing of dsx, thus determining female-limited polymorphism in I. senegalensis most likely lies in a non-coding region of the dsx gene or in a gene upstream of it.
Similar content being viewed by others
Introduction
Color polymorphism (the co-occurrence of two or more discrete color morphs within populations) contributes to sexual and deceptive signaling, crypsis, thermoregulation, and aposematism. Numerous studies have explored the maintenance and consequences of color polymorphism (Gray and McKinnon 2007; White and Kemp 2016). Although color polymorphisms occur in numerous taxa, sex-limited color polymorphisms are not common in animals (van Gossum et al. 2008). For example, only a single species of copepods (Fava 1986) and 25 bird species have been reported to exhibit sex-limited polymorphisms (Galeotti et al. 2003). In contrast, sex-limited polymorphisms are common among Odonata and occur in 43% of European Coenagrionids (Cordero-Rivera and Andres 1996). The evolution of female-limited color polymorphisms in Odonata appears to be associated with polyandrous mating systems (Robinson and Allgeyer 1996), and the evolution of female-limited color polymorphism occurred multiple times in Odonata through convergent evolution (Fincke et al. 2005).
Among the female-limited color polymorphisms observed in many species of Odonata, one morph is similar to the conspecific male in body or wing color (andromorph), whereas one or two morphs (gynomorphs) differ from males in these two traits (Fincke et al. 2005). The convergent evolution of female-limited polymorphism in Odonata suggests that they contain a genetic mechanism and architecture that allow the repeated evolution of such polymorphism (Fincke et al. 2005). Studies on female-limited color polymorphisms of damselflies suggest that balancing selection maintains color polymorphisms in nature (Andres et al. 2000, 2002; Wong et al. 2003; Takahashi et al. 2013, 2014). In Ischnura spp., multiple female morphs are maintained by negative frequency-dependent selection caused by sexual conflict (Takahashi et al. 2010). Although in some species or populations andromorphs are not preferred by males, even if they are the most common morph (Cordero-Rivera and Sánchez-Guillén 2007), males will at least attempt to mate with andromorphs during oviposition (Takahasi and Watanabe 2009). If females are attacked by males while they are ovipositing, they are forced to cease ovipositing and move away from suitable oviposition sites. Therefore, the reproductive success of common female morphs is lower than that of rare morphs, leading to the coexistence of female color polymorphisms under native frequency-dependent selection (Svensson et al. 2005; Takahashi and Watanabe 2009; Takahashi et al. 2010).
Interestingly, female color morphs correlate with phenotypic differences other than colors, such that andromorphs morphologically and behaviorally resemble the males. For example, andromorphs tend to perch in less vegetated areas, like males do, while gynomorphs spend more time hiding in vegetation and they fly greater distances (van Gossum et al. 2001; Sánchez-Guillén et al. 2017). Moreover, andromorphs engage in male-like face-off behavior when approached by conspecific males, while gynomorphs tend to escape (Robertson 1985). In addition, the wing lengths of andromorphs are shorter than those of gynomorphs, and are therefore similar to those of males (Abbott and Gosden 2009; M. Takahashi et al. unpublished). These findings suggest that andromorphs are either defeminized or masculinized females. Furthermore, the size and number of mature eggs and the number of ovarioles in an ovary are known to differ between color morphs (Takahashi and Watanabe 2010; Takahashi et al. 2011; Sánchez-Guillén et al. 2017). Abbott and Gosden (2009) suggested that the integration of multiple traits in andromorphs may be attributable to pleiotropic effects of the locus controlling the color polymorphism.
It is important to identify the genetic and molecular mechanisms determining female-limited polymorphisms in coenagrionids in order to understand how one locus controls the production of different color morphs, including male-resembling andromorphs. For example, the Papilio butterfly shows female-limited mimetic polymorphism, that is, only females resemble different model species. Recent studies have shown that a single gene, doublesex (dsx), which controls mimicry, can switch the entire pattern of wing phenotypes (Kunte et al. 2014). Furthermore, Nishikawa et al. (2015) found that mimetic coloration patterns are induced by female-specific Dsx isoforms expressed by a single autosomal inversion region that includes dsx, suggesting that this tightly linked inversion region contributes to the maintenance of female-limited polymorphisms. Besides, in many insects, sex-limited splice variants of dsx, which are often related to sex determination, have been suggested to also control multiple sex-limited traits such as male courtship behavior, female responsiveness to male courtship behavior, and sexual dimorphic abdominal pigmentation in Drosophila (Kopp 2012; Matson and Zarkower 2012), as well as sex-limited horns and mandibles in beetles (Rideout et al. 2007, 2010; Ito et al. 2013; Gotoh et al. 2014; Zhou et al. 2014). Pleiotropic genes such as dsx may therefore control the female-limited color polymorphisms and trait integration that occur in Odonata.
The purpose of the present study was to detect candidate transcripts responsible for producing female-limited color polymorphisms in the damselfly Ischnura senegalensis. Here we used RNA-seq to detect differentially expressed transcripts (DETs) among phenotypes. The expression levels of selected candidate transcripts were confirmed with quantitative reverse transcriptase-PCR (qRT-PCR). We discuss here the results in relation to the genetic control of female-limited polymorphisms in the damselfly.
Methods
Species
In Japan, I. senegalensis (Fig. 1) generally inhabits fens and stagnant pond water along the coast from Okinawa to the southern Tohoku district. The larval stage lasts for approximately 3 months. This species exhibits female-limited dimorphism, comprising andromorphs and gynomorphs. The gynomorphic allele (D) and andromorphic allele (d) at a putative autosomal locus determine the color morph (Takahashi et al. 2014). The gynomorphic allele is dominant over the andromorphic allele; thus, the genotypes DD, Dd, and dd express gynomorphic, gynomorphic, and andromorphic phenotypes, respectively.
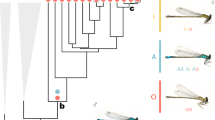
Female-limited polymorphism in Ischnura senegalensis. Male (top) and andromorph (middle) have similar body color patterns, whereas gynomorphs (bottom) differ from males and andromorphs. The thoracic melanic (black) regions of males and andromorphs are larger than those of gynomorphs. The total body length of both males and females is approximately 30 mm
Sampling and RNA extraction
Samples used for RNA-seq, PCR amplification, and qRT-PCR are shown in Table 1 and Fig. S1. For RNA-seq, males and females with the three different genotypes (DD, Dd, and dd) were used. They were born from females with a similar genetic background captured from two adjacent populations with different morph frequencies (Heiwasouzou and Kakinohana) on Okinawa Island in November 2014 (Table 1). In addition, for PCR amplification and qRT-PCR, parental females were collected from five locations on Okinawa Island (Heiwasouzou, Kakinohana, Uranishi, Shikiya, and Makabe) in May 2016 (Table 1). The frequencies of color morphs differed greatly among the populations, although there is sufficient gene flow among them (Takahashi et al. 2011; Inomata et al. 2015). In Heiwasouzou, Uranishi, and Makabe, most of the females were gynomorphs, whereas andromorphs were common in Kakinohana and Shikiya.
The offspring of wild females were collected to obtain teneral individuals (adults immediately after emergence). The wild females were kept in plastic cases (diameter 11 cm, height 4 cm), in which a wet paper filter was placed to promote prompt oviposition. The plastic cases with the females were placed under fluorescent light for 1 or 2 days to allow the females to lay eggs onto the paper filter. Eggs laid by a female were retained for 10–12 days in the plastic case in water until hatching, and the larvae were reared for approximately 1 month. Thereafter, the larvae were individually kept in small plastic cases (diameter 3 cm, height 5 cm) and reared with Artemia and Tubifex until emergence. The thoraxes of adults were left for a few hours after emergence, and those of wild mature adults were dissected immediately to extract RNA or fixed in RNA stabilization solution (RNAlater) for later RNA extraction. The fresh or fixed thoraxes were homogenized, and total mRNA was extracted using a Maxwell 16 LEV simplyRNA Tissue Kit (Promega).
For samples analyzed with RNA-seq, the genotype of gynomorphic offspring born from gynomorphs of the Heiwasouzou population was assumed to be DD when all the sisters showed the gynomorph phenotype, because the andromorphic allele d is likely quite rare in such a group, although this cannot be confirmed without knowing the phenotype of the next generation. The genotype of andromorphic offspring was dd and that of the gynomorphic offspring born from andromorphs was Dd. However, there were no andromorphs among the 14 offspring of the gynomorphs from Heiwasouzou. For gynomorph samples used in PCR amplification and qRT-PCR, offspring from gynomorph females were used. The genotype of gynomorph used was either DD or Dd. Mature individuals used for qRT-PCR were collected in June 2016 from Tsukuba, Ibaraki, Japan (36 °15′78′′N, 140 °06′38′′E).
mRNA sequencing and de novo transcriptome assembly
Three males, three homozygous recessives (andromorphs), three heterozygotes (gynomorphs born from andromorphs), and three putative homozygous dominants (gynomorphs whose sisters were gynomorphs, born from gynomorphs of Heiwasouzou) were used for transcriptome analyses (RNA-seq). Library preparations and RNA-seq were performed at the Beijing Genomics Institute (BGI, Shenzhen, China). The libraries were sequenced with an Illumina HiSeq 4000 (paired-end reads). The raw sequencing reads were trimmed to remove adapter sequences and filtered to eliminate low-quality reads, as well as contaminating sequences. The reads derived from 12 individuals were assembled de novo using Trinity (ver. 2.0.6) (Grabherr et al. 2011), yielding contigs from all pooled individuals.
Detecting DETs
Count data and fragments per kilobase of transcript per million mapped reads (FPKM) were used as an index of the expression levels of all contigs, as estimated using align_and_estimate_abundance.pl of Trinity for each individual (estimation method, RSEM; alignment method, bowtie). Transcripts expressed at very low levels (average FPKM of 12 individuals < 0.2) were excluded from subsequent analyses to remove background data. DETs between phenotypes were detected using the TCC package (false discovery rate < 0.05) in edgeR (Sun et al. 2013).
To identify transcripts contributing to the expression of the male-like phenotype in females, we conducted two comparisons as follows: First, we compared the expression levels between phenotypes featuring different colors, that is, males and heterozygous gynomorphic females (Dd), males and homozygous gynomorphic females (DD), andromorphic females (dd) and heterozygous gynomorphic females, and andromorphic females and homozygous gynomorphic females; then, we analyzed the transcripts with significantly different expression levels between the four comparisons listed above. Second, we compared the expression levels between the male and three female genotypes, that is, between males and andromorphs (dd), males and heterozygous gynomorphic females (Dd), and males and homozygous gynomorphic females (DD). In this second group of comparisons, we defined DETs as those with expression levels that were significantly different between those morphs with a false discovery rate cut-off of <0.05. DETs of males and heterozygous or homozygous gynomorphs but not those of males and andromorphs were designated as candidate DETs, as these possibly contribute to the differences between the male/male-like and female phenotypes. We annotated the candidate transcripts as queries for BLASTx analysis with the NCBI non-redundant database (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and identified functions using FlyBase (http://flybase.org). Of the candidate transcripts, doublesex (dsx), black, ebony, and chaoptin (chp), were designated as strong candidate genes for the determination of female-limited color polymorphisms.
Among the four candidate transcripts, dsx is known to have multiple splice variants. Thus, to identify the sequences in our dataset specific to each transcript, we generated contigs for each phenotype (males, andromorphs, heterozygous gynomorphs, and putative homozygous gynomorphs) using Trinity and used the dsx transcripts as queries for BLAST analyses of these contigs. Hit contigs were aligned by ClustalW and the alignments were manually corrected using Integrative Genomics Viewer version 2.3.67 (Thorvaldsdóttir et al. 2013). The contigs were unfortunately short, particularly those from females, and only covered fragments of the reference transcripts, therefore we were unable to determine the complete sequence of each transcript variant.
To obtain the full dsx transcript variant sequences, we next cloned and sequenced the complementary DNA (cDNA) of dsx. cDNA was prepared from the total RNA of each individual using SuperScript III Reverse Transcriptase (Thermo Fisher), in accordance with the manufacturer’s instructions. For the tenerals, 8 males, 3 andromorphs, and 11 gynomorphs were prepared for both PCR amplification and qRT-PCR, whereas for the adults, 4 males, 4 andromorphs, and 4 gynomorphs were prepared for qRT-PCR (Table 1). PCR was conducted using the forward primer (dsx_F1: 5′-GAGACAAGTGCTCCCAGTCG-3′) and reverse primer (dsx_R1: 3′-CATGGCGCGGATATTCTCCT-5′) specific to the flanking sequences of dsx (see Fig. 2a, red arrows on regions 1 and 5). Reactions were catalyzed by Tks Gflex DNA Polymerase (Clontech), in accordance with the manufacturer’s instructions. The short dsx amplicons from the males and the long dsx amplicons of the gynomorphs were inserted into the pGEM-T Easy Vector (Promega) and sequenced using an Applied Biosystems 3130 Genetic Analyzer with seven primers (see Table S1).
Splice variants (a) and PCR products of dsx (b) using RNA extracted from the thorax of teneral individuals. a The long transcript that includes region 4 and the short transcripts that lack this region were detected using 25 teneral individuals. The red box indicates the conserved functional domain (DM1). The blue boxes indicate the open reading frames. The red arrows indicate primers used in PCR amplification and sequencing. The numbers above the boxes are region identifiers. b PCR products of dsx of a teneral male (M), an andromorph (A), and a gynomorph (G) are shown. The long transcript was specifically expressed by andromorphic (n = 3) and gynomorphic females (n = 11), and the short transcripts were expressed by males (n = 8) and andromorphic females (n = 3). Short transcripts were detected in some of the gynomorphs when 40 cycles of PCR amplification were performed. The numbers on the right-hand side of the figure show the combination of putative exons corresponding to each band
Quantitative RT-PCR
qRT-PCR analyses of dsx, black, ebony, and chp transcription were conducted for teneral and mature individuals (Table 1). Sample sizes were as follows: teneral males, N = 8; teneral andromorphs, N = 3; teneral gynomorphs, N = 11; mature males, N = 4; mature andromorphs, N = 4; and mature gynomorphs, N = 4. The expression levels of each differentially spliced transcript of dsx and those of black, ebony, and chp were quantified using the StepOne Real-Time PCR System (Thermo Fisher) with Power SYBR Green PCR Master Mix (Applied Biosystems). As dsx encodes many different isoforms including several minor ones (Gotoh et al. 2016), we facilitated data analysis by using primer pairs complementary to putative exon junctions for the long and short dsx transcripts (see Table S1). Normalized gene expression values were estimated using ribosomal protein L13 (RpL13) levels as an internal control. The expression levels were visualized relative to maximal expression, which was defined as a level of 1 for each transcript. Error bars represent the standard error of the mean. The significance of the differences in the expression levels among the phenotypes was evaluated using the generalized linear model (GLM) assuming a gamma distribution, followed by post-hoc pairwise comparisons with Holm corrections.
Results
Differentially expressed transcripts
A summary of the assembly statistics is shown in Table 2. In the comparison of expression levels between phenotypes with different body colors, common transcripts with different expression levels were not detected. On the other hand, comparison of the expression levels between the males and the three female genotypes found several candidate transcripts. The numbers of DETs between males and heterozygous gynomorphs, between males and andromorphs, and between males and putative homozygous gynomorphs were 202, 93, and 70, respectively (Fig. 3). Analysis of the count data using the TCC package (false discovery rate < 0.05) in edgeR revealed seven candidate transcripts that were expressed at significantly different levels between males and heterozygous or homozygous gynomorphs, but not at significantly different levels between males and andromorphs. The annotated transcripts were homologs of dsx, black, ebony, chp, easily shocked (eas), rho guanine nucleotide exchange factor (RhoGEF), and nuclear factor I (NfI). The expression levels revealed by FPKM analysis of these transcripts are shown in Fig. 4. The expression levels of dsx, eas, RhoGEF, and Nf1 in males were lower than those in all females. The expression levels of black, ebony, and chp in males and andromorphs were lower than those in gynomorphs, and the expression levels of these genes in heterozygous gynomorphs were lower than those in homozygous gynomorphs (Fig. 4). There were no fixed nucleotide differences between transcripts from the different color morphs. dsx affects sex-limited polymorphism in Papilio polytes, whereas black, ebony, and chp affect body color in several insect species (Burtis and Baker 1989; Raymond et al. 1998; Ohbayashi et al. 2001; Suzuki et al. 2003, 2005; Phillips et al. 2005; Mummery-Widmer et al. 2009; Kunte et al. 2014; Nishikawa et al. 2015). Therefore, we focused on these genes for further analysis of their expression levels using qRT-PCR.
The numbers of differentially expressed transcripts (DETs) between males and andromorphs (blue), males and heterozygous gynomorphs (orange), and males and homozygous gynomorphs (red). Seven candidate transcripts (white circle) were detected with expression levels that differed significantly between males and heterozygous and homozygous gynomorphs, but not between males and andromorphs
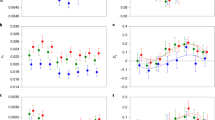
Fragments per kilobase of transcript per million mapped reads (FPKM) of seven DETs of each genotype (N = 3 for each genotype). a dsx, b black, c ebony, d chp, e eas, f RhoGEF, and g NfI. The FPKM of each DET did not generally differ significantly between males and andromorphs, although there were significant differences between males and heterozygous or homozygous gynomorphs (false discovery rate < 0.05). Data represent the mean ± S.E.
PCR and sequence analyses of dsx
The relative abundance of transcripts revealed by FPKM showed that the expression patterns of dsx differed among males, andromorphs, and gynomorphs (Fig. 4a). When dsx cDNA was amplified using a PCR primer pair complementary to the flanking regions of the female-limited expression region (dsx_F1 and dsx_R1, Table S1), dsx transcripts mainly comprised long and short transcripts. The sequences of the dsx transcripts indicated that the mRNA harbored a region specifically expressed in all females (region 4 in Fig. 2) and some common regions expressed irrespective of sex (regions 1, 3, and 5 in Fig. 2). The sequences flanking the primers (see red arrows in Fig. 2a) were estimated from dsx contigs produced by RNA-seq. The long transcript was expressed specifically in all females, whereas the short transcripts were expressed in males and andromorphs (Fig. 2b). Cloning and sequencing of the dsx transcripts revealed that the short transcripts represented regions 1, 3, and 5 or regions 1, 2, 3, and 5, and the long transcript comprised regions 1, 3, 4, and 5 (Fig. 2b).
Confirmation of the expression levels of dsx, black, ebony, and chp
We confirmed the expression levels of dsx, black, ebony, and chp using qRT-PCR. qRT-PCR experiments revealed that the expression levels of the dsx long and short transcripts and black transcripts significantly varied among the males, andromorphs, and gynomorphs irrespective of age. For dsx, the expression levels of the long transcript in males and andromorphs were significantly lower than in gynomorphs. Mature andromorphs exhibited slightly higher expression levels of the long transcript than that observed in males (Figs. 5a, b). For the short transcripts, although the expression levels in males were significantly higher than those in the two female morphs at the mature stage, at the teneral stage the expression levels in males tended to be only slightly higher than that in the two female morphs (Figs. 5c, d). For black, the expression level in gynomorphs was significantly higher than in males and andromorphs, independent of age, although the level in teneral males was slightly higher than in andromorphs (Figs. 6a, b). Although there were no significant differences in the expression levels of ebony and chp among the mature males, andromorphs, and gynomorphs, the expression level of chp in teneral males was significantly higher than in the two teneral female morphs (Figs. 6c–f).
The expression levels of dsx in male, andromorphic, and gynomorphic individuals. Long transcripts of teneral (a) and mature (b) individuals. Short transcripts of teneral (c) and mature (d) individuals. Teneral individuals were born to females captured on Okinawa Island, whereas mature individuals were captured in Tsukuba. Data represent the mean ± S.E. Different letters in the figure indicate significant differences evaluated using the generalized linear model (GLM) assuming a gamma distribution, followed by post-hoc pairwise comparison with Holm corrections
The expression levels of black and ebony in males, andromorphs, and gynomorphic individuals. Expression of black in teneral (a) and mature (b) individuals. Expression of ebony in teneral (c) and mature (d) individuals. Expression of chp in teneral (e) and mature (f) individuals. Teneral individuals were born to females captured on Okinawa Island, whereas mature individuals were captured in Tsukuba. Data represent the mean ± S.E. Different letters in the figure indicate significant differences evaluated using the generalized linear model (GLM) assuming a gamma distribution, followed by post-hoc pairwise comparison with Holm corrections
Discussion
In the present study, we attempted to detect DETs among males and two female morphs (gynomorphs and andromorphs) using RNA-seq. We identified seven candidate transcripts, dsx, black, ebony, chp, eas, RhoGEF, and NfI, which exhibited significantly different expression between the three groups. We focused on four of these candidates, dsx, black, ebony, and chp, because studies of other insect species have indicated that dsx controls sexual and sex-limited dimorphic traits and that black, ebony, and chp are associated with body color (Jacobs 1960; Phillips et al. 2005; Mummery-Widmer et al. 2009). The differential expression of these genes detected using RNA-seq was confirmed using qRT-PCR. Thus, the transcripts that were identified as differentially expressed by qRT-PCR are likely associated with differences in body color and other traits among males, gynomorphs, and andromorphs.
The dsx gene and female-limited polymorphisms
The dsx transcript is alternatively spliced to yield different sex-specific proteins that control sex-specific traits in many insect species (e.g., fly, bee, beetle, and moth) (Verhulst et al. 2010; Kopp 2012; Matson and Zarkower 2012). Among Drosophila species, evolutionary modification of DSX binding sites in cis-regulatory elements causes gain or loss of transcriptional regulation of the downstream genes, leading to sex-specific gene expression (Matson and Zarkower 2012). In Papilio polytes, dsx is a single autosomal locus with two alleles that control female-limited mimetic polymorphism (Kunte et al. 2014; Nishikawa et al. 2015).
PCR amplification and sequencing of the dsx transcripts of I. senegalensis revealed that dsx mainly produced two transcripts, which were classified as one long and two short transcripts. The latter were mainly expressed in males and andromorphs, and the former was specifically expressed in females (both andromorphs and gynomorphs). Thus, andromorphs may have both male and female elements expressed by dsx, and this may explain why these morphs are female but with masculinized color or behavioral traits.
qRT-PCR analysis revealed that the expression level of the long transcript in teneral gynomorphs was significantly higher than those of teneral males and andromorphs, and the expression level of the short transcripts of teneral males was higher than that of teneral gynomorphs. The inconsistency between the results of PCR amplification (strong expression of the long transcript in andromorphs) and qRT-PCR (low expression level of the long transcript in andromorphs) may be explained by the effect of minor dsx transcripts (Gotoh et al. 2016). If this effect occurred, the PCR results would be more credible than the qRT-PCR results. The female-limited polymorphism in I. senegalensis is thought to be controlled by a diallele at an autosomal locus (Takahashi et al. 2014). Because we did not detect different coding sequences among different morphs, the diallelic region determining female-limited polymorphism in I. senegalensis must therefore lie in a non-coding region of dsx or in a gene upstream of it.
The functions of black and ebony
The qRT-PCR results showed that the expression levels of black in gynomorphs were higher than those in males and andromorphs, whereas those of ebony were not significantly different among males, andromorphs, and gynomorphs. In Drosophila, black and ebony contribute to the production of N-β-alanyl-dopamine (NBAD), leading to cuticle sclerotization from dopamine (Phillips et al. 2005), which prompts cuticular melanization (Walter et al. 1996; Gibert et al. 2007). In ebony or black mutants, the overabundance of dopamine causes cuticular melanization (Jacobs 1960; Phillips et al. 2005). In I. senegalensis, males and andromorphs exhibit broad black stripes and a blue–green thorax and other body parts (Fig. 1). Prum et al. (2004) suggested that non-iridescent structural colors derived from epidermal structures may produce the blue–green thoracic color of members of Odonata.
Epidermal melanin affects the blue–green color that is produced by selective scattering from nanostructures (non-iridescent structural color) (Dyck 1971; Shawkey and Hill 2006). Thus, interactions between cuticular melanization and sclerotization may be involved in producing differences between males/andromorphs (blue–green and black stripes) and gynomorphs (orange thorax and fewer black stripes). Black functions as an aspartate decarboxylase that converts aspartate to β-alanine (Phillips et al. 2005). β-Alanine is bound to dopamine by ebony, which functions as a β-alanyl-dopamine synthetase that produces NBAD (Richardt et al. 2003). Lower expression levels of black in males and andromorphs may inhibit β-alanine production and promote cuticle melanization and suppression of sclerotization, which produces the coloration observed for males and andromorphs.
Other phenotype-associated DETs
RNA-seq also identified chp, eas, RhoGEF, and NfI as DETs. Because chp mutants display body color defects in Drosophila (Mummery-Widmer et al. 2009), we conducted qRT-PCR analysis of chp. However, the expression levels of chp did not differ significantly among males, andromorphs, and gynomorphs. The reason why the FPKM analysis of the RNA-seq data for chp was inconsistent with the qRT-PCR results may be because RNA-seq is an exhaustive method that also detects splice variants of chp.
NfI comprises a family that encodes sequence-specific DNA binding proteins, which function as transcription factors and mediate DNA replication (Gronostajski 2000). Consequently, NfI may regulate the expression levels of other DETs.
In damselflies, flying distances (van Gossum et al. 2001) and hindwing lengths (Abbott and Gosden 2009; M. Takahashi et al., unpublished) differ between andromorphs and gynomorphs. In Drosophila, mutants of eas and RhoGEF cause flight defects and affect wing morphology, respectively (Widmann and Dahmann 2009; Iyengar and Wu 2014). Therefore, differing expression levels of eas and RhoGEF may result in differences in wing morphology and flight ability between damselfly morphs.
Integration of multiple traits and pleiotropic effects
In Ischnura spp., several traits and behaviors are associated with color morphs of females (Robertson 1985; van Gossum et al. 2001; Takahashi and Watanabe 2010; Takahashi et al. 2011; Takahashi and Kawata 2013; Sánchez-Guillén et al. 2017). The integration of several traits could be explained by the fact that dsx encodes a transcription factor with pleiotropic functions (Chatterjee et al. 2011). For example, in Drosophila, dsx controls the expression of downstream genes involved in sexual dimorphism (Matson and Zarkower 2012) including bric à brac, which produces a sexually dimorphic pigmentation pattern on abdominal segments (Kopp et al. 2000). In the present study, different splice variants of dsx were expressed between males and gynomorphs, and both of these variants were expressed in andromorphs. Thus, it is possible that dsx controls sexually dimorphic coloration.
Among female-limited color polymorphisms observed in Ischnura, the andromorph is similar to the conspecific male in body color and other related traits, and dsx may regulate genes such as black to confer sexual mimicry. Therefore, our present results are consistent with a previous hypothesis that pleiotropism affects multiple traits in the damselfly (Abbott and Gosden 2009), and indicate that dsx produces sex-specific and morph-specific splice variants that may control the expression of multiple traits such as color pattern, behavior, habitat choice, and morphology. However, it remains unknown whether dsx forms part of the diallelic autosomal locus that Takahashi et al. (2014) inferred to determine female-limited color polymorphisms, because we were unable to detect differences among the coding region sequences of different morphs. The locus responsible for female-limited polymorphism in I. senegalensis is expected to be a non-coding region of dsx or a gene upstream of it that regulates its splice variants.
Conclusion
It follows from the function of dsx in other invertebrates that the products of dsx can control multiple genes such as black to produce morphs with different body color, flight ability, and wing morphology. However, the coding region of dsx does not appear to determine female-limited color polymorphisms in a diallelic manner. Thus, further studies of transcriptional control and the upstream and non-coding regions of dsx should be conducted to identify the diallelic region that determines the differences in body color of females and to reveal the evolution of female-limited polymorphisms and their underlying mechanisms of inheritance.
Data archiving
RNA-seq raw sequence reads are available through the DDBJ Sequence Read Archive http://ddbj.nig.ac.jp/ under accession no. DRA005915. Count data and FPKM estimated by using align_and_estimate_abundance.pl of Trinity are listed in Supplementary Tables S2 and S3, respectively. The expression levels of dsx from qRT-PCR are listed in Supplementary Table S4. The expression levels of black and ebony from qRT-PCR are listed in Supplementary Table S5.
References
Abbott JK, Gosden TP (2009) Correlated morphological and colour differences among females of the damselfly Ischnura elegans. Ecol Entomol 34:378–386
Andres JA, Sanchez-Guillen RA, Cordero RA (2002) Evolution of female colour polymorphism in damselflies: testing the hypotheses. Anim Behav 63:677–685
Andres JA, Sanchez RA, Cordero RA (2000) Molecular evidence for selection on female color polymorphism in the damselfly Ischnura graellsii. Evolution 54:2156–2161
Burtis KC, Baker BS (1989) Drosophila doublesex gene controls somatic sexual differentiation by producing alternatively spliced mRNAs encoding related sex-specific polypeptides. Cell 56:997–1010
Chatterjee SS, Uppendahl LD, Chowdhury MA, Ip P, Siegal ML (2011) The female-specific Doublesex isoform regulates pleiotropic transcription factors to pattern genital development in Drosophila. Development 138:1099–1109
Cordero-Rivera, Andres JA (1996) Colour polymorphism in odonates: females that mimic males? J Br Dragonfly Soc 12:50–60
Cordero-Rivera A, Sánchez-Guillén RA (2007) Male-like females of a damselfly are not preferred by males even if they are the majority morph. Anim Behav 74:247–252
Dyck JA (1971) Structure and colour-production of the blue barbs of Agapornis roseicollis and Cotinga maynana. Z fur Zellforsch und Mikrosk Anat 115:17–29
Fava G (1986) Genetic differentiation in experimental populations of the copepod Tisbe clodiensis from two aquatic lagoons. J Exp Mar Bio Ecol 97:51–61
Fincke OM, Jödicke R, Paulson DR, Schultz TD (2005) The evolution and frequency of female color morphs in Holarctic Odonata: why are male-like females typically the minority? Int J Odonatol 8:183–212
Galeotti P, Rubolini D, Dunn PO, Fasola M (2003) Colour polymorphism in birds: causes and functions. J Evol Biol 16:635–646
Gibert JM, Peronnet F, Schlötterer C (2007) Phenotypic plasticity in Drosophila pigmentation caused by temperature sensitivity of a chromatin regulator network. PLoS Genet 3:266–280
Gotoh H, Miyakawa H, Ishikawa A, Ishikawa Y, Sugime Y, Emlen DJ et al. (2014) Developmental link between sex and nutrition; doublesex regulates sex-specific mandible growth via juvenile hormone signaling in stag beetles. PLoS Genet 10:1–9
Gotoh H, Zinna RA, Warren I, Denieu M, Niimi T, Dworkin I et al. (2016) Identification and functional analyses of sex determination genes in the sexually dimorphic stag beetle Cyclommatus metallifer. BMC Genom 17:250
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I et al. (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Gray SM, McKinnon JS (2007) Linking color polymorphism maintenance and speciation. Trends Ecol Evol 22:71–79
Gronostajski RM (2000) Roles of the NFI/CTF gene family in transcription and development. Gene 249:31–45
Inomata N, Hironaka K, Sawada K, Kuriwada T, Yama K (2015) Discrepancy in the degree of population differentiation between color-morph frequencies and neutral genetic loci in the damselfly Ischnura senegalensis in Okinawa Island, Japan. Genetica 143:271–277
Ito Y, Harigai A, Nakata M, Hosoya T, Araya K, Oba Y et al. (2013) The role of doublesex in the evolution of exaggerated horns in the Japanese rhinoceros beetle. EMBO Rep 14:561–567
Iyengar A, Wu C (2014) Flight and seizure motor patterns in Drosophila mutants: simultaneous acoustic and electrophysiological recordings of wing beats and flight muscle activity. J Neurogenet 28:316–328
Jacobs ME (1960) Influence of light on mating of Drosophila melanogaster. Ecology 41:182–188
Kopp A (2012) Dmrt genes in the development and evolution of sexual dimorphism. Trends Genet 28:175–184
Kopp A, Duncan I, Carroll SB (2000) Genetic control and evolution of sexually dimorphic characters in Drosophila. Nature 408:553–559
Kunte K, Zhang W, Tenger-Trolander A, Palmer DH, Martin A, Reed RD et al. (2014) doublesex Is a mimicry supergene. Nature 507:229–232
Matson CK, Zarkower D (2012) Sex and the singular DM domain: insights into sexual regulation, evolution and plasticity. Nat Rev Genet 13:163–174
Mummery-Widmer JL, Yamazaki M, Stoeger T, Novatchkova M, Bhalerao S, Chen D et al. (2009) Genome-wide analysis of Notch signalling in Drosophila by transgenic RNAi. Nature 458:987–992
Nishikawa H, Iijima T, Kajitani R, Yamaguchi J, Ando T, Suzuki Y et al. (2015) A genetic mechanism for female-limited Batesian mimicry in Papilio butterfly. Nat Genet 47:405–409
Ohbayashi F, Suzuki MG, Mita K, Okano K (2001) A homologue of the Drosophila doublesex gene is transcribed into sex-specific mRNA isoforms in the silkworm, Bombyx mori. Comp Biochem Physiol Part B 128:145–158
Phillips AM, Smart R, Strauss R, Kelly LE (2005) The Drosophila black enigma: the molecular and behavioural characterization of theblack1 mutant allele. Gene 351:131–142
Prum RO, Cole JA, Torres RH (2004) Blue integumentary structural colours in dragonflies (Odonata) are not produced by incoherent Tyndall scattering. J Exp Biol 207:3999–4009
Raymond CS, Shamu CE, Shen MM, Seifert KJ, Hirsch B, Hodgkin J et al. (1998) Evidence for evolutionary conservation of sex-determining genes. Nature 391:691–695
Richardt A, Kemme T, Wagner S, Schwarzer D, Marahiel MA, Hovemann BT (2003) Ebony, a novel nonribosomal peptide synthetase for beta-Alanine conjugation with biogenic amines in Drosophila. J Biol Chem 278:41160–41166
Rideout EJ, Billeter JC, Goodwin SF (2007) The sex-determination genes fruitless and doublesex specify a neural substrate required for courtship song. Curr Biol 17:1473–1478
Rideout EJ, Dornan AJ, Neville MC, Eadie S, Goodwin SF (2010) Control of sexual differentiation and behavior by the doublesex gene in Drosophila melanogaster. Nat Neurosci 13:458–466
Robertson HM (1985) Female dimorphism and mating behaviour in a damselfly, Ischnura ramburi: females mimicking males. Anim Behav 33:805–809
Robinson JV, Allgeyer R (1996) Covariation in life-history traits, demographics and behaviour in ischnuran damselflies: the evolution of monandry. Biol J Linn Soc 58:85–98
Sánchez-Guillén RA, Wellenreuther M, Chávez-Ríos JR, Beatty CD, Rivas-Torres A, Velasquez-Velez M, et al. (2017) Alternative reproductive strategies and the maintenance of female color polymorphism in damselflies. Ecol Evol 7:5592–5602.
Shawkey MD, Hill GE (2006) Significance of a basal melanin layer to production of non-iridescent structural plumage color: evidence from an amelanotic Steller’s jay (Cyanocitta stelleri). J Exp Biol 209:1245–1250
Sun J, Nishiyama T, Shimizu K, Kadota K (2013) TCC: an R package for comparing tag count data with robust normalization strategies. BMC Bioinform 14:219
Suzuki MG, Funaguma S, Kanda T, Tamura T, Shimada T (2003) Analysis of the biological functions of a doublesex homologue in Bombyx mori. Dev Genes Evol 213:345–354
Suzuki MG, Funaguma S, Kanda T, Tamura T, Shimada T (2005) Role of the male BmDSX protein in the sexual differentiation of Bombyx mori. Evol Dev 7:58–68
Svensson EI, Abbott J, Hardling R (2005) Female polymorphism, frequency dependence, and rapid evolutionary dynamics in natural populations. Am Nat 165:567–576
Takahashi Y, Kagawa K, Svensson EI, Kawata M (2014) Evolution of increased phenotypic diversity enhances population performance by reducing sexual harassment in damselflies. Nat Commun 5:1–7
Takahashi Y, Kawata M (2013) Alternative trait combinations and secondary resource partitioning in sexually selected color polymorphism. Ecol Evol 3:2038–2046
Takahashi Y, Morita S, Yoshimura J, Watanabe M (2011) A geographic cline induced by negative frequency-dependent selection. BMC Evol Biol 11:256
Takahashi Y, Nagata N, Kawata M (2013) Antagonistic selection factors induce a continuous population divergence in a polymorphism. Heredity 112:391–398
Takahashi Y, Watanabe M (2009) Diurnal changes and frequency dependence in male mating preference for female morphs in the damselfly Ischnura senegalensis (Rambur) (Odonata: Coenagrionidae). Entomol Sci 12:219–226
Takahashi Y, Watanabe M (2010) Morph-specific fecundity and egg size in the female-dimorphic damselfly Ischnura senegalensis. Zool Sci 27:325–329
Takahashi Y, Yoshimura J, Morita S, Watanabe M (2010) Negative frequency-dependent selection in female color polymorphism of a damselfly. Evolution 64:3620–3628
Thorvaldsdóttir H, Robinson JT, Mesirov JP (2013) Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14:178–192
van Gossum H, Sherratt TN, Cordero-rivera A (2008) The evolution of sex-limited colour polymorphism. In: Córdoba-Aguilar A (ed). Dragonflies and damselflies: model organisms for ecological and evolutionary research. Oxford University Press, Oxford, p 219–229
van Gossum H, Stoks R, Bruyn L, De (2001) Frequency-dependent male mate harassment and intra-specific variation in its avoidance by females of the damselfly Ischnura elegans. Behav Ecol Sociobiol 51:69–75
Verhulst EC, van de Zande L, Beukeboom LW (2010) Insect sex determination: it all evolves around transformer. Curr Opin Genet Dev 20:376–383
Walter MF, Zeineh LL, Black BC, McIvor WE, Wright TR, Biessmann H (1996) Catecholamine metabolism and in vitro induction of premature cuticle melanization in wild type and pigmentation mutants of Drosophila melanogaster. Arch Insect Biochem Physiol 31:219–233
White TE, Kemp DJ (2016) Colour polymorphism. Curr Biol 26:R517–R518
Widmann TJ, Dahmann C (2009) Dpp signaling promotes the cuboidal-to-columnar shape transition of Drosophila wing disc epithelia by regulating Rho1. J Cell Sci 122:1362–1373
Wong A, Smith ML, Forbes MR (2003) Differentiation between subpopulations of a polychromatic damselfly with respect to morph frequencies, but not neutral genetic markers. Mol Ecol 12:3505–3513
Zhou C, Pan Y, Robinett CC, Meissner GW, Baker BS (2014) Central brain neurons expressing doublesex regulate female receptivity in Drosophila. Neuron 83:149–163
Acknowledgements
We thank Takashi Makino for advices on analysis of bioinformatics, Shinichiro Maruyama and Yu Ishii for advices of experiments and Shoko Aoki for rearing individuals. Transcriptome analysis was comfortably carried out thanks to the supercomputer of the National Institute of Genetics.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
These authors contributed equally: Yuma Takahashi, Masakado Kawata.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Takahashi, M., Takahashi, Y. & Kawata, M. Candidate genes associated with color morphs of female-limited polymorphisms of the damselfly Ischnura senegalensis. Heredity 122, 81–92 (2019). https://doi.org/10.1038/s41437-018-0076-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-018-0076-z
This article is cited by
-
Sex-specific morphs: the genetics and evolution of intra-sexual variation
Nature Reviews Genetics (2023)
-
The genomics and evolution of inter-sexual mimicry and female-limited polymorphisms in damselflies
Nature Ecology & Evolution (2023)
-
Neues über Nävi und nävoide Hauterkrankungen
Die Dermatologie (2023)
-
A masculinizing supergene underlies an exaggerated male reproductive morph in a spider
Nature Ecology & Evolution (2021)
-
Does Heat Tolerance Explain Female Polymorphism in Damselflies?
Journal of Insect Behavior (2021)