Abstract
Microphthalmia with brain and digital anomalies (MCOPS6, MIM# 607932) is an autosomal dominant disorder caused by loss-of-function variants or large deletions involving BMP4, which encodes bone morphogenetic protein 4, a member of the TGF-β protein superfamily. BMP4 has a number of roles in embryonic development including neurogenesis, lens induction, development of cartilage and bone, urogenital development, limb and digit patterning, hair follicle regeneration, as well as tooth formation. In addition to syndromic microphthalmia, BMP4 variants have been implicated in non-syndromic cleft lip with or without cleft palate and congenital healed cleft lip indicating different allelic presentations. MCOPS6 subjects may also lack some of the major phenotypic hallmarks of the disorder, including microphthalmia, indicating variable expressivity. As only a handful of individuals with MCOPS6 have been described, we review the clinical findings in previously reported cases with either deletions or loss-of-function variants in BMP4. We describe three new cases, including two subjects with novel deletions and one subject with a likely pathogenic de novo nonsense variant [c.1052C>G, p.(S351*)] in BMP4. One of the subjects had dual molecular diagnoses including a co-occurring microdeletion at 17q21.31 associated with Koolen de Vries syndrome, which has a partially overlapping disease phenotype. None of these individuals had clinically apparent microphthalmia or anopthalmia, which have been reported in a majority of previously described cases. One subject had exophthalmia and strabismus, while another had bilateral Peters anomaly and sclerocornea, thus expanding the phenotype associated with BMP4 loss-of-function variants.
Similar content being viewed by others
Introduction
Microphthalmia, Anophthalmia, and Coloboma together make up the MAC spectrum of developmental eye defects. Coloboma, microphthalmia, and anophthalmia, in order of decreasing prevalence, are observed in ∼1/5000, ∼1/7000, ∼1/30,000 live births, respectively [1]. This spectrum of ocular phenotypes has been associated with a number of genes expressed during early ocular development and are observed clinically in the context of both syndromic and non-syndromic conditions. Studies of patient cohorts have demonstrated that a genetic cause can be identified in up to 80% of individuals with severe, bilateral anophthalmia, or microphthalmia and up to 20% of all individuals that fall in the MAC spectrum [1]. The most common single genes associated with MAC include SOX2 (syndromic microphthalmia-3 (MCOPS3), MIM# 206900) and OTX2 (MCOPS6, MIM# 610125) which account for ~15–20% and ~5% of all cases, respectively [2].
BMP4 is a secreted bone morphogenetic protein family member and is part of the TGFβ superfamily [3]. The full-length protein is 408 amino acids and is generated as an inactive precursor protein that homodimerizes or heterodimerizes with other BMPs and is subsequently cleaved to generate prodomain (292 AA) and mature domain (116 AA) fragments [3]. Both the mature ligand and the prodomain are secreted but only the mature domain fragment is able to bind to and activate type I and type II serine/threonine kinase receptors [4]. Upon ligand binding, the type II receptor phosphorylates a type I signaling receptor, leading to downstream signaling via both SMAD-dependent and SMAD-independent pathways [5].
BMP4 has several conserved roles in embryonic development including neurogenesis, eye lens induction, development of cartilage and bone, limb and digit patterning, and tooth formation. Loss-of-function variants in BMP4 are a rare cause of syndromic microphthalmia and likely make up <1% of MAC spectrum cases [6]. In addition, several families have been described with large deletions encompassing the BMP4 gene [7, 8]. The clinical phenotypes have been observed to be highly variable and individuals may present with or without associated ocular malformations in addition to syndromic features. One subject presented with features consistent with SHORT (short stature, hyperextensibility, hernia, ocular depression, Rieger anomaly, and teething delay; MIM# 269880) syndrome [9]. BMP4 haploinsufficiency has also been identified as the likely cause of Frías syndrome (MIM# 609640), which was initially described in a large multigenerational family with a 4.06 Mb interstitial deletion at 14q22.1q22.3 [8, 10]. Several large contiguous gene deletions have been described within 14q22, which include both BMP4 and OTX2, and are associated with a more severe eye phenotype with several additional syndromic features [1, 11].
Missense variants in BMP4 have also been implicated in congenital renal malformations, subepithelial microform cleft lip and palate, and colorectal cancer [12,13,14]. Previous studies in large multigenerational families with syndromic microphthalmia suggest that the disorder may vary widely in its phenotypic expression and some confirmed carriers appear to be unaffected, indicating reduced penetrance. Overall, only a few studies have been published examining the phenotypic spectrum associated with BMP4 deletions and loss-of-function variants. Recently, a large family with dopa-responsive dystonia, eye, and skeletal anomalies was reported with a complex chromosomal rearrangement involving at 14q21q22, which resulted in the deletion of BMP4 and GTP cyclohydrolase 1 (GCH1, MIM# 600225); deletion of these genes could independently explain the complex phenotype and inheritance pattern observed in this family [15]. Several new studies have also emphasized the importance of considering multiple, independent molecular diagnoses in subjects with atypical or ‘expanded’ phenotypes [16]. In order to clarify the phenotypic spectrum of MCOPS6, we review previously reported cases and describe three new subjects including one with a novel nonsense variant in BMP4 and two subjects with microdeletions involving BMP4 that do not include OTX2.
Subjects and methods
Subjects
Three unrelated subjects were identified and referred to genetics practice for developmental concerns. Subject 1 was seen at the Mayo Clinic, Rochester Minnesota, where he underwent a comprehensive evaluation in the Department of Clinical Genomics. His family history was negative for similar findings including developmental delay, hypotonia, macrocephaly, and polydactyly. Subject 2 was evaluated by the medical genetics department at Centre Hospitalier Universitaire (CHU) in Nantes, France. Subject 3 was evaluated at the Hospital Universitario La Paz in Madrid, Spain. The subjects or their parents consented for sample collection and subsequent analysis under a protocol approved by an institutional review board in accordance with the Declaration of Helsinki. In addition, written informed consent was obtained to publish subject photographs.
Whole exome sequencing (WES) and analysis
Subject 1
Blood samples were collected from subject 1 and his parents and genomic DNA was extracted. Clinical WES was performed at the Baylor Miraca Genetics Laboratories at Baylor College of Medicine (BCM) as previously described [17, 18]. Briefly, exome capture and hybridization were performed using the VCRome 2.1 exome capture reagent (Roche NimbleGen, Madison, WI) and 100-bp paired end reads were generated using the Illumina HiSeq platform. Data was converted to FastQ format using CASAVA 1.8 software (Illumina) and mapped to GRCh37/hg19 using the Burrows-Wheeler Aligner (BWA) [19]. Variant calling was performed using Atlas-SNP and Atlas-indel (Human Genome Sequencing Center (HGSC), BCM) [20]. Synonymous and intronic variants as well as variants with a minor allele frequency (MAF) >1% in the exome sequencing project (ESP) 5400 exomes or the 1000 Genomes Project control databases were filtered. The remaining variants were interpreted according to ACMG guidelines and informed by the subject’s reported clinical phenotype [21]. Confirmatory Sanger sequencing was performed on the subject and his parents in order to determine the mode of inheritance for variants of interest. An Illumina HumanExome-12v1 quality control (QC) array was run in parallel with testing in the proband.
Microarray
Subject 2
Array comparative genomic hybridization (aCGH) experiments were performed using Agilent Human Genome CGH 60K oligonucleotide arrays (Agilent, Santa Clara, CA, USA; www.agilent.com) with the ISCA design (www.iscaconsortium.org). The arrays were analyzed with the Agilent scanner and the Feature Extraction software (v.9.1.3). Graphical overview was obtained using the CGH analytics software (v.3.5.14).
Subject 3
High-density aCGH were performed using a custom Agilent-based SurePrint G3 Human CGH Microarray 8 × 60K (KaryoArray® v3.014), an oligonucleotide chip that contains around 60,000 distinct biological features with a probe spacing 9 kB overall in interest regions (mainly microdeletion/microduplication syndromes) with genomic data sourced from—UCSC hg19 (NCBI Build 37) [22]. Array experiments were performed as recommended by the manufacturer (Agilent Technologies, Santa Clara, CA, USA). The array was analyzed with the Agilent scanner using the Feature Extraction software (v9.1 Agilent Technologies).
Results
Clinical characterization
Subject 1
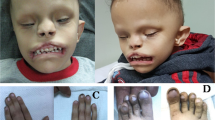
Subject 1 is a 4-year-old boy referred for evaluation regarding a history of developmental delay, macrocephaly with frontal bossing, and postaxial polydactyly (Table 1). The subject was born at term through a scheduled C-section to a 42-year-old G2P2 female after an unremarkable pregnancy (Fig. 1a). His birth weight was 3345 g and there were no neonatal complications aside from mild jaundice. The subject was noted to have postaxial polydactyly at birth. There were small hypoplastic phalanges in both hypoplastic sixth fingers (Fig. 1c). He also had macrocephaly and widely open fontanelles. His face was slightly flat with wide-set and prominent eyes, flat infraorbital ridges, long eyelashes, and mild down-slanting palpebral fissures (Fig. 1b). He was also noted to have micrognathia with a high arched palate. His external ears were over-folded and small.
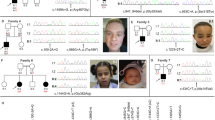
Family pedigrees and patient images. a Subject 1 family pedigree. The proband is indicated with an arrow in each pedigree. Patient variants are shown underneath symbols. Filled in symbols indicate affected status. b Photographs of subject 1 shows relative macrocephaly, prominent eyes, and mild dysmorphic features with midface hypoplasia and low-set ears. c Radiographs taken of subject 1 demonstrated bilateral postaxial polydactyly. d–g Head MRI from subject 1 at 3 years and 1 month of age demonstrated decreased volume of white matter in both cerebral hemispheres with the cortical margins nearly reaching the ventricular surface. While there was no convincing associated abnormal T2 signal the appearance is most suggestive of periventricular leukomalacia. The gyral patterns are unusual especially within the right occipital parietal region where there is a focal area of cortical thickness raising the possibility of polymicrogyria. The ventricular system is asymmetric with prominence of the left occipital pole. h Subject 2 family pedigree. i–l Photographs (i, k) and radiographs (j, l) taken of subject 2 at showing left hand postaxial polydactyly and left foot 4/5 toe syndactyly. Subject 2 ophthalmological examination at 8 years of age: (m) right sclerocornea at slit lamp examination, (n) right fundus, (o) left sclerocornea at slit lamp examination, (p) normal B-scan echography of the left eye; yellow line is axial length (26.56 mm). q Subject 3 family pedigree
At 2 years of age, he was noted to have mild myopia that was not progressive over a 2-year span. His external ocular movements and pupillary responses were normal, and he appropriately responded to stimuli in the peripheral fields. Fundus examination revealed normal optic discs. An early ultrasound of his kidneys and head were both unremarkable. He began pulling to stand at about 12 months and walking alone at 15 months with an uncoordinated gait. Mild delays in his fine motor abilities were also noted. The subject first spoke words at 18 months and by the age of 3 he spoke in full sentences, although a formal evaluation revealed mild delays in his receptive and expressive language skills (The Preschool Language Scales, Fifth Edition, Standard score = 81; percentile rank = 10; age equivalent = 2 years 6 months). He also had mild diffuse hypotonia with preserved muscle tone but with globally quiet reflexes and flexor plantar responses.
A head MRI obtained at age 3 showed reduced white matter volume in both cerebral hemispheres with the cortical margins nearly reaching the ventricular surface (Fig. 1d–g). While abnormal T2 signal was absent, the appearance was suggestive of periventricular leukomalacia. The gyral patterns were also unusual, particularly within the right occipital parietal region where there was a focal area of cortical thickness that raised the possibility of polymicrogyria. The ventricular system was also noted to be asymmetric with prominence of the left occipital pole.
The proband’s family history was negative for polydactyly, developmental delay, or similar problems in the extended pedigree.
Subject 2
Subject 2 is a 7-year and 7-month-old boy (Table 1, Fig. 1h). He presented with several congenital anomalies including postaxial polydactyly of the left hand (Fig. 1i–j), 4/5 toe syndactyly of the left foot (Fig. 1k–l), joint hyperextensibility, and bilateral Peters anomaly with sclerocornea. At birth the subject weighed 2520 g and was 48 cm in length. He had a normal head ultrasound. The subject was noted to have hypotonia and had some delays in gross motor milestones and only began ambulating at 28 months. On recent evaluation, he had dysmorphic facial features including a small nose and large forehead. His weight was 23.3 kg, his height was 122 cm, and his OFC was 54.3 cm (+1SD). He attends school and receives some supportive services. He has no language difficulties.
At ophthalmogic evaluation, the patient has severe visual impairment; he can see light, forms, colors, and large font letters. At 6 months old, he presented with poor vision with horizontal nystagmus and oculodigital sign. Slit-lamp examination showed bilateral sclerocornea, with a superficial punctate keratitis in both eyes, making fundus examination impossible. At 8 years old, the child presented with bilateral ptosis and had long curly eyelashes. The visual acuity was 20/250 in the right eye with high hyperopia and light perception in the left eye with an abnormal head position and with right eye fixating in adduction. The right cornea had cleared in the central part allowing the visualization of a normal iris (Fig. 1m) and a normal posterior pole (Fig. 1n) but not the left cornea (Fig. 1o). B-scan echography revealed a normal left posterior segment (Fig. 1p).
Subject 3
Subject 3 is currently a 13-year-old girl who was followed by medical genetics from an early age (Table 1, Fig. 1q). She was noted to have intrauterine growth restriction (IUGR) at 32 weeks on prenatal ultrasound. The pregnancy was further complicated by oligohydramnios. She was born at 37 weeks and weighed 2100 g and was 48 cm in length. Physical examination at birth was notable for micrognathia, arachnodactyly, joint hyperextensibility, and axial hypotonia. Her growth was slow and she had poor weight gain. At 9-months she underwent surgery for trigonocephaly. She also had unilateral renal pelvic ectasia and an atrial septal defect. Facial features included dolichocephaly, upslanting palpebral fissures, proptosis, and strabismus. MRI findings were generally nonspecific with only some generalized diminished white matter. She has mild to moderate global developmental delay with mild intellectual disability and speech delays.
Genetic analyses
In subject 1, Chromosome microarray revealed a maternally inherited deletion of uncertain significance at 7p21.3 of ∼127 kb (arr[hg19] 7p21.3(8592102_8719291)x1). The deleted interval only includes a single gene (NXPH1, a secreted neurexophilin) with no known role in human disease. In addition, GLI3 gene testing (Connective Tissue Gene Tests, Allentown, PA, USA) was performed due to suspicion for Greig cephalopolysyndactyly syndrome (MIM# 175700), but no variants were detected. WES was performed and revealed a heterozygous de novo likely pathogenic nonsense variant in exon 4 of BMP4 [Chr14(GRCh37): g.54416925G>C, NM_001202.5: c.1052C>G p.(Ser351*)] (Fig. 2b,c). This variant has not been reported in population frequency databases, such as ExAC or gnomAD or in any previously described cases. Given the presence of overlapping clinical features described in previously reported cases including high forehead, mid-face hyperplasia, ear anomalies, and polydactyly, this variant was considered likely causal of the patient’s phenotype. An additional maternally inherited heterozygous missense variant [Chr14(GRCh37): g.54417226G>A, NM_001202.5: c.751C>T, p.(His251Tyr) in exon 4] in the BMP4 gene was also detected. We were unable to determine whether this variant was in cis or trans with the c.1052C>G [p.(S351*)] variant. This variant was classified as a variant of uncertain significance and has a MAF of 0.001537% in East Asians in gnomAD (29/18870 East Asian alleles or 0.001537%; 32/277144 or 0.0001155% overall in gnomAD), and could be attributed to maternal Vietnamese ancestry [23]. The p.(His251Tyr) variant was previously reported in an individual of Chinese descent with bilateral microphthalmia and unilateral cataract [24]. In that study, the variant was also present in the proband’s healthy brother, and, by presumption, in at least one of the untested parents [24]. Our proband’s mother had no indication of dental or other craniofacial anomalies, and had no personal nor family history of ocular defects, thus reducing the likelihood that this second variant contributed to the proband’s phenotype. However, we are unable to completely rule out the possibility that the p.(His251Tyr) variant could exert a possible hypomorphic effect in the context of the de novo nonsense variant also identified in this patient.
Schematic diagram of BMP4 gene, transcript (NM_001202.5), and protein structure (NP_001193.2). a Previously reported deletions in 14q22.1q22.3 are shown as well as the de novo deletions identified in subjects 2 and 3. BMP4 is highlighted in yellow and falls within the minimally deleted region. Content sourced from—UCSC hg19 (NCBI Build 37). b, c Previously reported loss-of-function variants as well as variants found in subject 1 (red) are indicated. Exons are numbered as in NG_009215.1. Furin cleavage sites (S1 and S2 motifs) are also indicated within the overall protein structure. Cleavage of BMP4 occurs sequentially at the S1 furin recognition motif and then at the S2 motif to generate the free mature ligand, which can signal over larger distances
Subject 2 had a 2.4 Mb de novo deletion on chromosome 14 that includes 23 known genes including BMP4 (arr[hg19] 14q22.1q22.2(53001803_55443425)x1) (Fig. 2a). The deletion was confirmed in the patient by FISH using BAC probe RP11-299D05. FISH with the same probe was normal in both parents confirming the de novo status of the deletion.
Subject 3 had a 2.4 Mb de novo deletion on chromosome 14 that includes 28 genes including BMP4 (arr[hg19] 14q22.1q22.3(53854436_56266758)x1) (Fig. 2a). The patient also had a co-occurring de novo 6.4 Mb deletion on 17q21.3 (arr[hg19] 17q21.3(43706876_44345048)x1) that includes the KANSL1 gene the critical gene in the autosomal dominant Koolen de Vries syndrome (MIM# 610443). The de novo status of these deletion variants was confirmed by array in both parents. The phenotype associated with Koolen de Vries syndrome partially overlaps that associated with MCOPS6 (Supplementary Table 1).
Discussion
BMP4 dosage is tightly controlled and haploinsufficiency has been shown to result in developmental/patterning defects in both animal models and humans [4]. In this report, 27 individuals harboring 14 BMP4 loss-of-function variants were reviewed, including 24 previously reported cases and three new cases. Among the evaluated variants, a novel nonsense variant [c.1052C>G, p.(S351*)] falling within the mature ligand was discovered (Patient 1, Fig. 2c). While truncating loss-of-function variants have been described within the propeptide domain [6, 9, 12], no truncating variants have been described in the mature domain fragment before, making this variant a relevant addition to the catalogue of published BMP4 variants. Deletions of the 14q22q23 region have been reported in several individuals within the MAC spectrum disorders, but only 24 cases of deletions or loss-of-function variants restricted to BMP4 (and not including OTX2 or SIX6), have been described to date (Supplementary Table 2). Of these, 17 are familial cases [7,8,9]. In our study, seven unique truncating variants due to deletions were analyzed, with a wide-range of phenotypic consequences involving eye, brain, and limb abnormalities.
While microphthalmia is the most commonly observed eye-related finding in patients with deletions or loss-of-function variants in BMP4, the clinical presentation is extremely variable even within affected families. This is similar to the observations in mice, where Bmp4 is critical in dorsoventral patterning of the optic vesicle [25] and Bmp4 +/− heterozygous mice have a variety of eye defects [25,26,27], however, the penetrance and severity of the ocular phenotype is strongly influenced by the background strain [28]. Altogether, ophthalmological findings were present in a majority of the individuals reviewed in this study, with the most common presentations being exophthalmia (5), anophthalmia (4), sclerocornea (4), and microphthalmia (3) (Fig. 3, Supplementary Table 2). In addition, variable features including Peters anomaly, Rieger anomaly, microcornea, congenital glaucoma, strabismus, nystagmus, among others, were also identified (Supplementary Table 2). Only two of the newly described individuals in this study had eye-related findings: subject 2 presented with Peters anomaly and sclerocornea, while subject 3 presented with exophthalmia and strabismus. Interestingly, a nonsense variant (c.130G>T, p.[Gly44*]) upstream of previously reported truncating variants was recently described in a family diagnosed with autosomal dominant Stickler syndrome. In this family, five affected individuals from two generations presented with variable features including megalophthalmos, high myopia, bilateral retinal detachments, and congenital hypoplastic vitreous [29]. This variant falls nearby the previously described c.171dupC, p.(Glu58Argfs*17) associated with anophthalmia and postaxial polydactyly, which were not present in the family with Stickler-like syndrome [9].
Summary of clinical features reported for the BMP4 cohort. Phenotypic abnormalities are indicated by an arrow and text, and the ratios indicate the number of cases observed with the described anomaly. The most common ophthalmological findings were exophthalmia (5), anophthalmia (4), sclerocornea (4), and microphthalmia (3). Other clinical features not included in this figure are hyperextensible joints (4), diaphragmatic hernia (2), congenital heart anomalies (1), and neoplasms/cysts (3)
Bmp4 is also involved in mouse central nervous system neurogenesis and is required for development of the roof plate and commissural plate which gives rise to the corpus callosum [30]. In our study and review of the literature, various individuals presented with brain-related abnormalities, including abnormal brain MRI findings (7), dilatation of the ventricles (4), diminished white matter sometimes suggestive of periventricular leukomalacia (3), hypoplastic corpus callosum (2), among others (Fig. 3, Supplementary Table 2). Four individuals presented with mild to moderate intellectual disability, while others had normal intellect despite evidence of structural brain defects. Neither feature appeared to correlate with the type of variant (large deletion vs. loss-of-function variant in BMP4). In addition, several individuals were reported to have delayed motor milestones. Interestingly, subjects 2 and 3 both had deletions involving the GCH1 gene, which was recently implicated in disease in a family with autosomal dominant or recessive dopa-responsive dystonia in addition to eye and skeletal anomalies attributed to BMP4 haploinsufficiency [15]. However, dystonia was not reported in either of the newly reported cases described in this study.
Finally, growth and morphogenesis of the limb and digits in mouse is controlled by secreted signals including Bmp2, Bmp4, Bmp7, and Shh via morphogen gradients. Increased Shh activity in the context of Bmp4 loss has been shown to result in polydactyly [31]. Paradoxically, conditional inactivation of Bmp4 can lead to an overall increase in the amount of BMP signaling [32]; as BMP expression increases, these gradients merge and can result in syndactyly [33]. Five individuals in our study had postaxial polydactyly involving either both hands (4) or one hand (1), four individuals had short hands and/or feet, four individuals had postaxial polydactyly of one or both feet (one individual with partial duplication of the fifth toe), three individuals had either unilateral or bilateral toe syndactyly (two with 4th and 5th toes affected, one with 4th, 5th, and 6th toes affected), two individuals had proximal syndactyly between the 2nd and 3rd fingers, and one individual had small nails on the fifth finger and toe bilaterally (Fig. 3, Supplementary Table 2). Similar to the variability in eye abnormalities, it is also likely that BMP4’s function in morphogen-mediated developmental patterning leads to increased genotype–phenotype complexity as this developmental process is partially stochastic [34, 35]. SHH and other BMP protein expression in these individuals will be a subject for future studies to functionally tie BMP4 variants to the type of limb malformations.
As has been shown throughout this study, BMP4 is an important developmental gene with truncating and loss-of-function variants generating a wide clinical spectrum. As only a few cases have been described, the actual frequency of associated clinical phenotypes in MCOPS6 is unknown and genotype-first unbiased genomic approaches will likely help further clarify the phenotypic spectrum associated with loss-of-function variants in BMP4 [36].
References
Williamson KA, FitzPatrick DR. The genetic architecture of microphthalmia, anophthalmia and coloboma. Eur J Med Genet. 2014;57:369–80.
Richardson R, Sowden J, Gerth-Kahlert C, Moore AT, Moosajee M. Clinical utility gene card for: non-syndromic microphthalmia including next-generation sequencing-based approaches. Eur J Hum Genet. 2017;25:e1–6.
Bragdon B, Moseychuk O, Saldanha S, King D, Julian J, Nohe A. Bone morphogenetic proteins: a critical review. Cell Signal. 2011;23:609–20.
Wang RN, Green J, Wang Z, Deng Y, Qiao M, Peabody M, et al. Bone morphogenetic protein (BMP) signaling in development and human diseases. Genes Dis. 2014;1:87–105.
Derynck R, Zhang YE. Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature. 2003;425:577–84.
Bakrania P, Efthymiou M, Klein JC, Salt A, Bunyan DJ, Wyatt A, et al. Mutations in BMP4 cause eye, brain, and digit developmental anomalies: overlap between the BMP4 and hedgehog signaling pathways. Am J Hum Genet. 2008;82:304–19.
Lumaka A, Van Hole C, Casteels I, Ortibus E, De Wolf V, Vermeesch JR, et al. Variability in expression of a familial 2.79 Mb microdeletion in chromosome 14q22.1-22.2. Am J Med Genet A. 2012;158A:1381–7.
Martínez-Fernández ML, Bermejo-Sánchez E, Fernández B, MacDonald A, Fernández-Toral J, Martínez-Frías ML. Haploinsufficiency of BMP4 gene may be the underlying cause of Frías syndrome. Am J Med Genet A. 2014;164A:338–45.
Reis LM, Tyler RC, Schilter KF, Abdul-Rahman O, Innis JW, Kozel BA, et al. BMP4 loss-of-function mutations in developmental eye disorders including SHORT syndrome. Hum Genet. 2011;130:495–504.
Martínez-Frías ML, Toral JF, López-Grondona F, Mendioroz J, Bermejo E. Growth deficiency, facial anomalies, and brachydactyly (Frías syndrome): a second family. Am J Med Genet A. 2005;137A:288–91.
Nolen LD, Amor D, Haywood A, St Heaps L, Willcock C, Mihelec M, et al. Deletion at 14q22-23 indicates a contiguous gene syndrome comprising anophthalmia, pituitary hypoplasia, and ear anomalies. Am J Med Genet A. 2006;140:1711–8.
Suzuki S, Marazita ML, Cooper ME, Miwa N, Hing A, Jugessur A, et al. Mutations in BMP4 are associated with subepithelial, microform, and overt cleft lip. Am J Hum Genet. 2009;84:406–11.
Weber S, Taylor JC, Winyard P, Baker KF, Sullivan-Brown J, Schild R, et al. SIX2 and BMP4 mutations associate with anomalous kidney development. J Am Soc Nephrol. 2008;19:891–903.
Lubbe SJ, Pittman AM, Matijssen C, Twiss P, Olver B, Lloyd A, et al. Evaluation of germline BMP4 mutation as a cause of colorectal cancer. Hum Mutat. 2011;32:E1928–38.
Lohmann K, Redin C, Tönnies H, Bressman SB, Subero JIM, Wiegers K, et al. Complex and dynamic chromosomal rearrangements in a family with seemingly non-Mendelian inheritance of Dopa-responsive dystonia. JAMA Neurol. 2017;74:806–12.
Posey JE, Harel T, Liu P, Rosenfeld JA, James RA, Coban Akdemir ZH, et al. Resolution of disease phenotypes resulting from multilocus genomic variation. N Engl J Med. 2017;376:21–31.
Yang Y, Muzny DM, Reid JG, Bainbridge MN, Willis A, Ward PA, et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med. 2013;369:1502–11.
Yang Y, Muzny DM, Xia F, Niu Z, Person R, Ding Y, et al. Molecular findings among patients referred for clinical whole-exome sequencing. J Am Med Assoc. 2014;312:1870–9.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Challis D, Yu J, Evani US, Jackson AR, Paithankar S, Coarfa C, et al. An integrative variant analysis suite for whole exome next-generation sequencing data. BMC Bioinf. 2012;13:8. 2009 10:1
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Vallespín E, Palomares Bralo M, Mori MÁ, Martín R, García-Miñaúr S, Fernández L, et al. Customized high resolution CGH-array for clinical diagnosis reveals additional genomic imbalances in previous well-defined pathological samples. Am J Med Genet A. 2013;161:1950–60.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Zhang X, Li S, Xiao X, Jia X, Wang P, Shen H, et al. Mutational screening of 10 genes in Chinese patients with microphthalmia and/or coloboma. Mol Vis. 2009;15:2911–8.
Furuta Y, Hogan BLM. BMP4 is essential for lens induction in the mouse embryo. Genes Dev. 1998;12:3764–75.
Winnier G, Blessing M, Labosky PA, Hogan BLM. Bone morphogenetic protein-4 is required for mesoderm formation and patterning in the mouse. Genes Dev. 1995;9:2105–16.
Dunn NR, Winnier GE, Hargett LK, Schrick JJ, Fogo AB, Hogan BL. Haploinsufficient phenotypes in Bmp4 heterozygous null mice and modification by mutations in Gli3 and Alx4. Dev Biol. 1997;188:235–47.
Goldman DC, Donley N, Christian JL. Genetic interaction between Bmp2 and Bmp4 reveals shared functions during multiple aspects of mouse organogenesis. Mech Dev. 2009;126:117–27.
Nixon TRW, Richards A, Towns LK, Fuller G, Abbs S, Alexander P, et al. Bone morphogenetic protein 4 (BMP4) loss-of-function variant associated with autosomal dominant Stickler syndrome and renal dysplasia. Eur J Hum Genet. 2018;25:1389.
Chizhikov VV, Millen KJ. Control of roof plate development and signaling by Lmx1b in the caudal vertebrate CNS. J Neurosci. 2004;24:5694–703.
Selever J, Liu W, Lu M-F, Behringer RR, Martin JF. Bmp4 in limb bud mesoderm regulates digit pattern by controlling AER development. Dev Biol. 2004;276:268–79.
Bénazet J-D, Bischofberger M, Tiecke E, Gonçalves A, Martin JF, Zuniga A, et al. A self-regulatory system of interlinked signaling feedback loops controls mouse limb patterning. Science. 2009;323:1050–3.
Badugu A, Kraemer C, Germann P, Menshykau D, Iber D. Digit patterning during limb development as a result of the BMP-receptor interaction. Sci Rep. 2012;2:991.
Salazar-Ciudad I, Jernvall J. How different types of pattern formation mechanisms affect the evolution of form and development. Evol Dev. 2004;6:6–16.
Fish JL. Developmental mechanisms underlying variation in craniofacial disease and evolution. Dev Biol. 2016;415:188–97.
Stessman HA, Bernier R, Eichler EE. A genotype-first approach to defining the subtypes of a complex disease. Cell. 2014;156:872–7.
Acknowledgements
We would like to thank the subjects and their families for participating in this study. We would also like to thank the Mayo Clinic Center for Individualized Medicine (CIM) for supporting this research through the CIM Investigative and Functional Genomics Program. This study makes use of data generated by the DECIPHER Consortium. A full list of centers who contributed to the generation of the data is available from https://decipher.sanger.ac.uk/ and via email from decipher@sanger.ac.uk. Funding for DECIPHER was provided by the Wellcome Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Blackburn, P.R., Zepeda-Mendoza, C.J., Kruisselbrink, T.M. et al. Variable expressivity of syndromic BMP4-related eye, brain, and digital anomalies: A review of the literature and description of three new cases. Eur J Hum Genet 27, 1379–1388 (2019). https://doi.org/10.1038/s41431-019-0423-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-019-0423-4
This article is cited by
-
DNA-based eyelid trait prediction in Chinese Han population
International Journal of Legal Medicine (2021)