Abstract
Antimicrobial resistance is a major threat for public health worldwide. Novel antimicrobial drugs are urgently needed to combat the growing prevalence of antimicrobial resistance. Nucleoside antibiotics represent a unique class of microbial natural products with distinctive structural features and diverse biological activities. We herein summarize recent findings on the biosynthesis of representative nucleoside antibiotics, and highlight recent advances in the discovery and rational generation of nucleoside antibiotics for the development of novel antimicrobial agents.
Similar content being viewed by others
Introduction
Antimicrobial resistance poses a serious threat to public health worldwide. To prioritize the research and development of new and effective antibiotic treatments, the World Health Organization has proposed a global list of antibiotic-resistant pathogens in three priority tiers of Priority I (critical), Priority II (high), and Priority III (medium) [1]. Of particular concern is the growing emergence of multidrug resistances in a number of key pathogens, such as Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacter species, and Mycobacterium tuberculosis [1, 2]. To combat antibiotic-resistant pathogens, it is of great importance to discover and create novel antimicrobial agents. Nucleoside antibiotics represent a unique class of microbial natural products with distinctive structural features and diverse biological activities. Over the past thirty years, several comprehensive reviews have focused on the structure, biosynthesis, regulation and biotechnology of nucleoside antibiotics [3,4,5,6,7,8]. This review is not intended to be comprehensive, but to summarize and highlight recent findings on the biosynthesis, combinatorial biosynthesis and chemical synthesis of representative nucleoside antibiotics toward the generation of novel antimicrobial drug candidates.
Representative nucleoside antibiotics
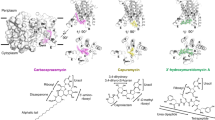
Nucleoside antibiotics are a large group of microbial natural products derived from nucleosides and nucleotides (Fig. 1). Considering that nucleosides and nucleotides play essential roles in most of the fundamental cellular metabolism, it is not surprising that nucleoside antibiotics are capable of targeting a variety of enzymes, such as enzymes involved in bacterial peptidoglycan biosynthesis, fungal chitin biosynthesis, and those involved in protein synthesis.
Structures of representative nucleoside antibiotics. a Structures of uridine-based nucleoside antibiotics targeting MraY, a phosphoglycosyl transferase involved in bacterial peptidoglycan biosynthesis. The uridine moieties are highlighted with pink colors. b Structures of nucleoside antibiotics targeting chitin synthase essential for fungal chitin biosynthesis. c Structures of nucleoside antibiotics targeting aminoacyl-tRNA synthetase and ribosomal peptidyl transferase required for protein synthesis
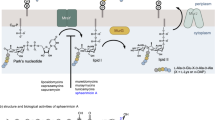
Peptidoglycan is a main component of cell wall in both Gram-positive and Gram-negative bacteria. Many enzymes are required for peptidoglycan biosynthesis that occurs in three stages from cytoplasm, cell membrane to cell wall. Among them, MraY is a phosphoglycosyl transferase that initiates the lipid cycle of peptidoglycan biosynthesis by transferring phospho-MurNAc-pentapeptide from the UDP-MurNAc pentapeptide onto a cell membrane-anchored lipid carrier, undecaprenyl phosphate [9]. Many nucleoside antibiotics function as inhibitors of MraY by mimicking the natural UDP-MurNAc-pentapeptide with uridine moiety as a key recognition element [10]. Thus, this group of nucleoside antibiotics is also referred to as uridine-based nucleosides. They have attracted increasing attention due to their strong efficacy against various pathogenic bacteria including M. tuberculosis, P. aeruginosa, methicillin-resistant S. aureus (MRSA), and vancomycin-resistant Enterococcus [8]. Representative members include pacidamycin, sansanmycin, and mureidomycin of uridyl peptide antibiotics, liposidomycin, caprazamycin and A-94964 of uridyl lipopeptide antibiotics, tunicamycin of uridyl liposaccharide antibiotics, and capuramycin of uridyl glycosylpeptide antibiotics (Fig. 1) [3]. Biosynthetic pathways of representative MraY inhibitors have been summarized in several comprehensive reviews [3, 7, 8]. A recent study has shown that a nonribosomal peptide synthetase (NRPS) CapU catalyzes the ATP-dependent amide bond formation in L-α-amino-ε-caprolactam, a unique unit in capuramycin-type nucleoside antibiotics [11]. In another study, a biosynthetic pathway for A-94964 biosynthesis has been proposed based on results from genetic studies and bioinformatics analysis [12]. Two recent structural studies have also been accomplished with MraY from Gram-negative bacterium Aquifex aeolicus in complex with muraymycin D2 [13], and MraY from Gram-positive bacterium Clostridium bolteae in complex with tunicamycin [14]. These studies promote our understanding of the biosynthesis of MraY inhibitors and help us gain insight into inhibition of MraY in bacterial cell wall synthesis.
Chitin, an essential component of cell walls of fungi, is required for the pathogenesis or survival of most pathogenic fungi. It consists of a β-1,4-linked homopolymer of N-acetylglucosamine (GlcNAc) synthesized from uridine diphosphate N-acetylglucosamine (UDP-GlcNAc) by chitin synthase [15]. Nucleoside antibiotics targeting chitin biosynthesis are represented by two promising antifungal agents, nikkomycin and polyoxin. Extensive genetic and biochemical studies have revealed biosynthetic pathways of nikkomycin and polyoxin [3, 4]. Three recent studies shed new light on the biosynthesis of nikkomycin and polyoxin. One study focuses on the complex interplay between the maturation process of NRPS NikP1, the recruitment of an auxiliary tailoring enzyme NikQ, and the regulation of early catalytic steps by L-His-NikP1, which are involved in the formation of 4-formyl-4-imidazolin-2-one base in nikkomycin biosynthesis [16]. Another study reveals the biosynthetic process of 3′-enolpyruvyl-UMP to octosyl acid, an unusual 8-carbon furanosyl nucleoside core intermediate during aminohexuronic acid formation in both nikkomycin and polyoxin biosynthesis [17]. The third study reveals that PolG functions as an ATP-dependent ligase catalyzing the amide bond formation via an activated acyl-phosphate intermediate in polyoxin biosynthesis [18]. Considering that SanS is the homolog of PolG [19], it is reasonable to propose that SanS catalyzes a parallel reaction in nikkomycin biosynthesis. These studies offer new opportunities for the combinatorial biosynthesis and pathway engineering of this group of nucleoside antibiotics.
Nucleoside antibiotics targeting protein synthesis are represented by inhibitors of aminoacyl-tRNA synthetase and ribosomal peptidyl transferase. Examples of the aminoacyl-tRNA synthetase inhibitors include albomycin, microcin C, and agrocin 84, while the peptidyl transferase inhibitors encompass blasticidin S, gougerotin, and amicetin. Here we only highlight recent findings on albomycin, microcin C, and blasticidin S. Albomycin consists of an iron-chelating ferrichrome siderophore and a serine-containing nucleosyl dipeptide termed SB-217452. The siderophore component allows albomycin to be actively transported via the iron uptake system into bacterial cells, where they are hydrolyzed by an unknown host peptidase to release SB-217452, an inhibitor of seryl-tRNA synthetase [20, 21]. Albomycin exhibits potent antimicrobial activities against many Gram-negative and Gram-positive bacteria with very low minimum inhibitory concentrations. It is a promising antibacterial drug candidate to treat S. pneumoniae and S. aureus infections [22]. The gene cluster for albomycin biosynthesis has been identified in Streptomyces griseus ATCC 700974 and a biosynthetic pathway has been proposed [23]. A recent study provides insight into the assembly of the thioheptose core during biosynthesis of albomycin involving a pyridoxal 5′-phosphate (PLP)-dependent transaldolase (AbmH), a PLP-dependent epimerase (AbmD) and a radical S-adenosyl-L-methionine (SAM) enzyme (AbmJ) [24]. Microcin C consists of a ribosomally synthesized heptapeptide attached to a modified adenosine. Of note is that the N-terminal methionine is formylated. The peptide component facilitates import of microcin C into susceptible bacterial strains via the inner membrane transporter YejABEF [25]. Once inside the bacterial cell, microcin C is then subjected to progressive digestion by peptide deformylase and any one of three cellular aminopeptidases, PepA, PepB, or PepN [26]. Upon hydrolysis of the peptide bond connecting the sixth and seventh residues of the peptide, a nonhydrolyzable analogue of aspartyl-adenylate is released to function as an inhibitor of aspartyl-tRNA synthetase [27]. Biosynthetic pathway of microcin C has been summarized by Severinov and Nair [28]. A recent study has suggested that MccB only efficiently processes the precursor heptapeptide that retains the N-formylated initiator Met, highlighting a rare example of the use of substrate inhibition in a natural product biosynthetic pathway [29]. For blasticidin S biosynthesis, two recent studies have suggested that a S-adenosyl methionine-dependent methyltransferase (BlsL) is responsible for the guanidine N-methylation of leucyldemethylblasticidin S [30], and a radical (SAM enzyme (BlsE) catalyzes the conversion of CGA to cytosylarabinopyranose [31]. These studies provide insights into the biosynthetic pathways of these protein inhibitors.
Discovery of novel nucleoside antibiotics
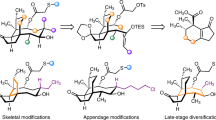
In recent years, there is a growing interest in the discovery of nucleoside antibiotics from underexplored sources, such as new isolates of terrestrial microorganisms and those from marine ecosystems. Several strategies, such as bioactivity-guided screening, genome mining, heterologous expression, and phosphopantetheinyl transferase (PPtase)-based approach, have been used to identify novel nucleoside antibiotics (Fig. 2). Examples include simamycin, a rare prenylated nucleoside from a soil-derived Streptomyces sp. TP-A0872 [32], and seven novel nucleoside antibiotics from marine-derived actinomycetes and marine sponge [33,34,35]. It is worth noting that five new nucleoside antibiotics, streptcytosines A-E, have been identified in a marine-derived Streptomyces sp. TPU1236A, of which streptcytosine A displays potent activity against Mycobacterium smegmatis [34]. Furthermore, a new blasticidin S analogue from the marine sponge Theonella swinhoei exhibits increased activity against a wide range of Gram-positive and Gram-negative bacteria, including laboratory and clinical strains with broad drug resistance [35]. In another study, the nucleoside analogue pseudouridimycin has been identified by screening a library of 3000 actinobacterial and fungal culture extracts. Of special note is that pseudouridimycin functions as an inhibitor of bacterial RNA polymerase and acts against both Gram-positive and Gram-negative bacteria, including multidrug-resistant (MDR) strains [36]. Another important source of new nucleoside antibiotics is cryptic biosynthetic pathways embedded in microbial genomes. Genome sequencing reveals four putative biosynthetic gene clusters (BGCs) dedicated to the biosynthesis of nucleoside antibiotics in Streptomyces griseochromogenes ATCC 14511 [37], and 1 BGC responsible for the biosynthesis of tunicamycin in Streptomyces sp. strain fd1-xmd [38]. Of special note is that bioinformatic analysis reveals the presence of microcin C-like gene clusters in the genomes and/or plasmids of diverse bacteria, indicating that microcin-like adenylated peptides are widespread in both Gram-negative and Gram-positive bacteria [39]. For illustration purpose, we herein only highlight several recent examples. Genome mining reveals a microcin C-like gene cluster in Bacillus amyloliquefaciens DSM7. Heterologous expression of the gene cluster in the surrogate host Bacillus subtilis 168 results in the production of a novel microcin C-like compound [40]. Genome mining reveals a cryptic mureidomycin biosynthetic gene cluster in the chromosome of Streptomyces roseosporus NRRL 15998. The gene cluster has been activated by constitutive expression of a foreign activator gene ssaA from sansanmycin biosynthetic gene cluster of Streptomyces sp. strain SS, leading to the discovery of eight acetylated mureidomycin analogues [41]. In another study, two new derivatives (puromycin B and C) have been identified through application of the phosphopantetheinyl transferase (PPtase)-based approach in Streptomyces alboniger NRRL B-1832 [42]. These studies suggest that microorganisms from underexplored habitats and uncharacterized biosynthetic pathways hidden in microbial genomes are two appealing sources for the discovery of new nucleoside antibiotics.
Strategies for the discovery and rational generation of novel nucleoside antibiotics. To find novel nucleoside antibiotics from underexplored sources, strategies used include bioactivity-guided screening, genome mining, heterologous expression, and phosphopantetheinyl transferase (PPtase)-based approach. To create novel nucleoside antibiotics, strategies used include precursor-directed biosynthesis, mutasynthesis, combinatorial biosynthesis, heterologous expression, and natural product-inspired chemical synthesis. Application of these strategies greatly expanded the repertoire of nucleoside antibiotics that may serve as drug candidates for the development of novel antimicrobial agents
Rational generation of novel nucleoside antibiotics
To expand structural diversity of nucleoside antibiotics, considerable efforts have also been directed toward rational design of new derivatives. Several strategies, such as precursor-directed biosynthesis, mutasynthesis, combinatorial biosynthesis, heterologous expression and natural product-inspired chemical synthesis, have been employed to generate new variants (Fig. 2). These novel derivatives of nucleoside antibiotics hold great promise for the development of potent antimicrobial agents.
Precursor-directed biosynthesis is a simple and effective method to generate new analogues in the native producing strain. This approach enables the generation of new analogues by incorporating biosynthetic precursor analogues into biosynthetic pathway of a nucleoside antibiotic. It is noteworthy that variations in the peptide backbone contribute to multiple components in uridyl peptide antibiotics, such as 10 pacidamycins, 16 sansanmycins, 4 mureidomycin, and 4 napamycins, suggesting that the NRPSs responsible for the assembly of peptide backbone have highly relaxed preference for different amino acid substrates [3, 43, 44]. The substrate promiscuity of NRPSs has been explored for the production of new analogues of pacidamycin and sansanmycin [43, 45]. Distinct from precursor-directed biosynthesis, mutasynthesis requires the generation of a mutant deficient in the formation of key biosynthetic intermediates, so that alternative intermediates fed to the mutant can be incorporated more efficiently to produce novel antibiotic analogues. With this strategy, 2 novel nikkomycin derivatives, 20 novel sansanmycin derivatives, and 7 new A201A derivatives have been generated [46,47,48]. It should be note that two derivatives (sansanmycin MX-2 and MX-4) exhibit promising activity against MDR and extensive-drug-resistant M. tuberculosis strains [47].
Combinatorial biosynthesis applies genetic engineering to modify biosynthetic pathways to natural products. This approach has been initially used to generate hybrid nucleoside antibiotics. For example, two hybrid antibiotics (polynik A and polyoxin N) have been generated after the introduction of genes required for CPOAA biosynthesis into a nikkomycin nonproducing mutant of Streptomyces ansochromogenes [19]. Similarly, two additional polyoxin analogs (polyoxin O and polyoxin P) have been obtained after the introduction of the complete polyoxin BGC into the same nikkomycin nonproducing mutant of S. ansochromogenes [49]. Inspired by these studies, polyoxin N and 6 hybrid antibiotics (nikkoxin B–G) has been produced by using similar strategies [50, 51]. Recently, genes for HPHT biosynthesis in the nikkomycin BGC have been replaced with those for CPOAA biosynthesis from the polyoxin BGC to generate a hybrid gene cluster of ~40 kb in Escherichia coli. The reconstructed cluster is then introduced into Streptomyces lividans TK23 for heterologous expression of hybrid antibiotic polynik A [52]. These studies provide an efficient strategy for gene cluster reconstruction that may facilitate further gene cluster modifications for the design and generation of more hybrid nucleoside antibiotics.
Heterologous expression of BGCs in an alternative host is a useful way to enhance the diversity of nucleoside antibiotics. For example, heterologous expression of the caprazamycin gene cluster in Streptomyces coelicolor M512 led to the production of nonglycosylated derivatives of caprazamycin [53], while heterologous expression of blasticidin S biosynthetic gene cluster in S. lividans resulted in the production of deaminohydroxyblasticidin S [54]. Heterologous expression of polyoxin biosynthetic gene cluster resulted in the production of polyoxin H, and introduction of a thymine-7-hydroxylase gene from Streptomyces avermitilis leads to the production of polyoxin A and H [55, 56]. Production of these variants is most likely attributed to the presence or absence of a specific-modifying enzyme in the heterologous host.
Other than strategies that utilize nature’s biosynthetic machinery, chemical synthesis is also of great interest for the generation of natural occurring nucleoside antibiotics. Total synthesis has been achieved with several nucleoside antibiotics, including muraymycins [57, 58], polyoxins [59], plicacetin, and streptcytosine A [60], albomycin [22], and A201A [61]. More importantly, chemical synthesis is an alternative to generate new derivatives of nucleoside antibiotics. A library of sansanmycin analogues have been generated and subjected to biological evaluation. The synthetic compounds are nanomolar inhibitors of MraY and exhibit potent and selective inhibition of M. tuberculosis [62]. Fluorinated analogues of polyoxin J and L have been generated via chemical synthesis. It is noteworthy that the fluorinated polyoxin J displays potent activities against pathogenic fungi, while the fluorinated polyoxin L is active toward MRSA and VRSA [59]. Inspired by tunicamycin and mureidomycin A, inhibitors targeting phosphoglycosyl transferase PglC have been designed to serve as lead compounds for future development of selective and effective inhibitors of diverse phosphoglycosyl transferase [63]. Of special note is pyrrolocytosines RX-04 A-D that are designed to bind to the bacterial 50S ribosomal subunit based on modeling of the ribosomal interactions of natural occurring blasticidin and amecitin, and synthetic TAB-1057A/B. Evaluation of pyrrolocytosines RX-04 A-D against a panel of 96 Gram-negative clinical isolates shows that all four compounds exhibit broad anti-Gram-negative activity with RX-04 A as the most active analogue [64]. These studies substantially expand the repertoire of nucleoside antibiotics that will ultimately speed up the search for new therapeutic agents (Table 1).
Conclusion
Novel antimicrobial drugs are urgently needed to combat the growing threat of antimicrobial resistance. Nucleoside antibiotics represent a unique class of microbial natural products with great promise for the development of novel antimicrobial drugs. Further efforts should be geared to gain access to previously underexplored microbial sources, such as the underrepresented genera of actinomycetes, microorganisms from marine ecosystems, and those in associations with mammals, invertebrates and plants. Over the past several decades, extensive studies have been carried out to reveal the complex biosynthetic machinery of nucleoside antibiotics. Based on knowledge from these studies, considerable efforts have been directed to create new derivatives. These derivatives of nucleoside antibiotics can be subjected to systematic structure–activity relationship studies that may aid the design and generation of novel analogues for the development of novel antimicrobial agents.
References
Tacconelli E, et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis. 2018;18:318–27.
Rice LB. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: no ESKAPE. J Infect Dis. 2008;197:1079–81.
Niu G, Tan H. Nucleoside antibiotics: biosynthesis, regulation, and biotechnology. Trends Microbiol. 2015;23:110–9.
Niu G, Zheng J, Tan H. Biosynthesis and combinatorial biosynthesis of antifungal nucleoside antibiotics. Sci China Life Sci. 2017;60:939–47.
Isono K. Current progress on nucleoside antibiotics. Pharmacol Ther. 1991;52:269–86.
Isono K. Nucleoside antibiotics: structure, biological activity, and biosynthesis. J Antibiot. 1988;41:1711–39.
Chen W, et al. Natural and engineered biosynthesis of nucleoside antibiotics in actinomycetes. J Ind Microbiol Biotechnol. 2016;43:401–17.
Chen S, Kinney WA, Van Lanen S. Nature’s combinatorial biosynthesis and recently engineered production of nucleoside antibiotics in Streptomyces. World J Microbiol Biotechnol. 2017;33:66.
Typas A, Banzhaf M, Gross CA, Vollmer W. From the regulation of peptidoglycan synthesis to bacterial growth and morphology. Nat Rev Microbiol. 2011;10:123–36.
Walsh CT, Zhang W. Chemical logic and enzymatic machinery for biological assembly of peptidyl nucleoside antibiotics. ACS Chem Biol. 2011;6:1000–7.
Liu X, et al. The role of a nonribosomal peptide synthetase in L-lysine lactamization during capuramycin biosynthesis. Chembiochem. 2016;17:804–10.
Shiraishi T, Nishiyama M, Kuzuyama T. Biosynthesis of the uridine-derived nucleoside antibiotic A-94964: identification and characterization of the biosynthetic gene cluster provide insight into the biosynthetic pathway. Org Biomol Chem. 2019;17:461–6.
Chung BC, et al. Structural insights into inhibition of lipid I production in bacterial cell wall synthesis. Nature. 2016;533:557–60.
Hakulinen JK, et al. MraY-antibiotic complex reveals details of tunicamycin mode of action. Nat Chem Biol. 2017;13:265–7.
Lenardon MD, Munro CA, Gow NA. Chitin synthesis and fungal pathogenesis. Curr Opin Microbiol. 2010;13:416–23.
Wise CE, Makris TM. Recruitment and regulation of the non-ribosomal peptide synthetase modifying cytochrome P450 involved in nikkomycin biosynthesis. ACS Chem Biol. 2017;12:1316–26.
He N, et al. Construction of an octosyl acid backbone catalyzed by a radical S-adenosylmethionine enzyme and a phosphatase in the biosynthesis of high-carbon sugar nucleoside antibiotics. Chem Sci. 2017;8:444–51.
Gong R, et al. An ATP-dependent ligase with substrate flexibility involved in assembly of the peptidyl nucleoside antibiotic polyoxin. Appl Environ Microbiol. 2018;84:e00501–18.
Li J, Li L, Tian Y, Niu G, Tan H. Hybrid antibiotics with the nikkomycin nucleoside and polyoxin peptidyl moieties. Metab Eng. 2011;13:336–44.
Stefanska AL, Fulston M, Houge-Frydrych CS, Jones JJ, Warr SR. A potent seryl tRNA synthetase inhibitor SB-217452 isolated from a Streptomyces species. J Antibiot. 2000;53:1346–53.
Pramanik A, Braun V. Albomycin uptake via a ferric hydroxamate transport system of Streptococcus pneumoniae R6. J Bacteriol. 2006;188:3878–86.
Lin Z, et al. Total synthesis and antimicrobial evaluation of natural albomycins against clinical pathogens. Nat Commun. 2018;9:3445.
Zeng Y, et al. Biosynthesis of albomycin δ2 provides a template for assembling siderophore and aminoacyl-tRNA synthetase inhibitor conjugates. ACS Chem Biol. 2012;7:1565–75.
Ushimaru R, Liu HW. Biosynthetic origin of the atypical stereochemistry in the thioheptose core of albomycin nucleoside antibiotics. J Am Chem Soc. 2019;141:2211–4.
Novikova M, et al. The Escherichia coli Yej transporter is required for the uptake of translation inhibitor microcin C. J Bacteriol. 2007;189:8361–5.
Kazakov T, et al. Escherichia coli peptidase A, B, or N can process translation inhibitor microcin C. J Bacteriol. 2008;190:2607–10.
Metlitskaya A, et al. Aspartyl-tRNA synthetase is the target of peptide nucleotide antibiotic microcin C. J Biol Chem. 2006;281:18033–42.
Severinov K, Nair SK. Microcin C: biosynthesis and mechanisms of bacterial resistance. Future Microbiol. 2012;7:281–9.
Dong SH, et al. Biosynthesis of the RiPP trojan horse nucleotide antibiotic microcin C is directed by the N-formyl of the peptide precursor. Chem Sci. 2019;10:2391–5.
Wang X, Du A, Yu G, Deng Z, He X. Guanidine N-methylation by BlsL is dependent on acylation of Beta-amine arginine in the biosynthesis of blasticidin S. Front Microbiol. 2017;8:1565.
Liu L, et al. A mechanistic study of the non-oxidative decarboxylation catalyzed by the radical S-adenosyl-l-methionine enzyme BlsE involved in blasticidin S biosynthesis. Chem Commun. 2017;53:8952–5.
Igarashi Y, Kyoso T, Kim Y, Oikawa T. Simamycin (5’-O-geranyluridine): a new prenylated nucleoside from Streptomyces sp. J Antibiot. 2017;70:607–10.
Aksoy SÇ, Uzel A, Bedir E. Cytosine-type nucleosides from marine-derived Streptomyces rochei 06CM016. J Antibiot. 2016;69:51–6.
Bu YY, Yamazaki H, Ukai K, Namikoshi M. Anti-mycobacterial nucleoside antibiotics from a marine-derived Streptomyces sp. TPU1236A. Mar Drugs. 2014;12:6102–12.
Davison JR, et al. A new natural product analog of blasticidin S reveals cellular uptake facilitated by the NorA multidrug transporter. Antimicrob Agents Chemother. 2017;61:e02635–16.
Maffioli SI, et al. Antibacterial nucleoside-analog inhibitor of bacterial RNA Polymerase. Cell. 2017;169:1240–8.
Wu L, Chen G, Feng G. Complete genome sequence of Streptomyces griseochromogenes ATCC 14511T, a producer of nucleoside compounds and diverse secondary metabolites. J Biotechnol. 2017;249:16–19.
Yu Y, et al. Identification of the streptothricin and tunicamycin biosynthetic gene clusters by genome mining in Streptomyces sp. strain fd1-xmd. Appl Microbiol Biotechnol. 2018;102:2621–33.
Bantysh O, et al. Enzymatic synthesis of bioinformatically predicted microcin C-like compounds encoded by diverse bacteria. mBio. 2014;5:e01059–14.
Serebryakova M, et al. A trojan-horse peptide-carboxymethyl-cytidine antibiotic from Bacillus amyloliquefaciens. J Am Chem Soc. 2016;138:15690–8.
Jiang L, et al. Identification of novel mureidomycin analogues via rational activation of a cryptic gene cluster in Streptomyces roseosporus NRRL 15998. Sci Rep. 2015;5:14111.
Yan X, et al. Puromycin A, B and C, cryptic nucleosides identified from Streptomyces alboniger NRRL B-1832 by PPtase-based activation. Synth Syst Biotechnol. 2018;3:76–80.
Gruschow S, et al. New pacidamycin antibiotics through precursor-directed biosynthesis. Chembiochem. 2009;10:355–60.
Xie Y, et al. NRPS substrate promiscuity leads to more potent antitubercular sansanmycin analogues. J Nat Prod. 2014;77:1744–8.
Zhang N, et al. Precursor-directed biosynthesis of new sansanmycin analogs bearing para-substituted-phenylalanines with high yields. J Antibiot. 2016;69:765–8.
Feng C, et al. Novel nikkomycin analogues generated by mutasynthesis in Streptomyces ansochromogenes. Micro Cell Fact. 2014;13:59.
Shi Y, et al. Improving the N-terminal diversity of sansanmycin through mutasynthesis. Micro Cell Fact. 2016;15:77.
Zhu Q, Song Y, Huang H, Li Q, Ju J. Characterization of MtdV as a chorismate lyase essential to A201A biosynthesis and precursor-directed biosynthesis of new analogs. Org Biomol Chem. 2019;17:3760–4.
Li J, Li L, Feng C, Chen Y, Tan H. Novel polyoxins generated by heterologously expressing polyoxin biosynthetic gene cluster in the sanN inactivated mutant of Streptomyces ansochromogenes. Micro Cell Fact. 2012;11:135.
Zhai L, et al. Engineering of an industrial polyoxin producer for the rational production of hybrid peptidyl nucleoside antibiotics. Metab Eng. 2012;14:388–93.
Qi J, et al. Metabolic engineering of an industrial polyoxin producer for the targeted overproduction of designer nucleoside antibiotics. Biotechnol Bioeng. 2015;112:1865–71.
Zhuo J, et al. Reconstruction of a hybrid nucleoside antibiotic gene cluster based on scarless modification of large DNA fragments. Sci China Life Sci. 2017;60:968–79.
Kaysser L, et al. Identification and manipulation of the caprazamycin gene cluster lead to new simplified liponucleoside antibiotics and give insights into the biosynthetic pathway. J Biol Chem. 2009;284:14987–96.
Li L, Wu J, Deng Z, Zabriskie TM, He X. Streptomyces lividans blasticidin S deaminase and its application in engineering a blasticidin S-producing strain for ease of genetic manipulation. Appl Environ Microbiol. 2013;79:2349–57.
Zhao C, Huang T, Chen W, Deng Z. Enhancement of the diversity of polyoxins by a thymine-7-hydroxylase homolog outside the polyoxin biosynthesis gene cluster. Appl Environ Microbiol. 2010;76:7343–7.
Chen W, et al. Characterization of the polyoxin biosynthetic gene cluster from Streptomyces cacaoi and engineered production of polyoxin H. J Biol Chem. 2009;284:10627–38.
Mitachi K, Aleiwi BA, Schneider CM, Siricilla S, Kurosu M. Stereocontrolled total synthesis of muraymycin D1 having a dual mode of action against Mycobacterium tuberculosis. J Am Chem Soc. 2016;138:12975–80.
Katsuyama A, Ichikawa S. Synthesis and medicinal chemistry of muraymycins, nucleoside antibiotics. Chem Pharm Bull. 2018;66:123–31.
Fujino H, et al. Unified total synthesis of polyoxins J, L, and fluorinated analogues on the basis of decarbonylative radical coupling reactions. Angew Chem Int Ed Engl. 2017;56:11865–9.
Fu J, Laval S, Yu B. Total synthesis of nucleoside antibiotics plicacetin and streptcytosine A. J Org Chem. 2018;83:7076–84.
Nie S, Li W, Yu B. Total synthesis of nucleoside antibiotic A201A. J Am Chem Soc. 2014;136:4157–60.
Tran AT, et al. Sansanmycin natural product analogues as potent and selective anti-mycobacterials that inhibit lipid I biosynthesis. Nat Commun. 2017;8:14414.
Walvoort MT, Lukose V, Imperiali B. A modular approach to phosphoglycosyltransferase inhibitors inspired by nucleoside antibiotics. Chemistry. 2016;22:3856–64.
Vickers A, Mushtaq S, Woodford N, Doumith M, Livermore DM. Activity of RX-04 pyrrolocytosine protein synthesis inhibitors against multidrug-resistant Gram-negative bacteria. Antimicrob Agents Chemother. 2018;62:e00689–18.
Acknowledgements
This paper is dedicated to Prof. Kiyoshi Isono for his tremendous contribution to the study of nucleoside antibiotics. This work was supported in part by the National Natural Science Foundation of China (31870061, 31571281, and 31771378), grants from Chongqing Science and Technology Commission (cstccxljrc201904, cstc2017jcyjAX0467 and cstc2018jcyjAX0066).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Niu, G., Li, Z., Huang, P. et al. Engineering nucleoside antibiotics toward the development of novel antimicrobial agents. J Antibiot 72, 906–912 (2019). https://doi.org/10.1038/s41429-019-0230-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-019-0230-8