Abstract
Histone deacetylase enzymes (HDACs) regulate protein acetylation. HDAC1 is known to enhance ischemia/reperfusion (I/R) injury, but its underlying mechanism(s) of action have not been defined. Here, in vivo mouse models of myocardial I/R were used to investigate the role of HDAC1 during I/R myocardial injury. We show that HDAC1 enhances the inflammatory responses of I/R mice. Using a constructed macrophage H/R (hypoxia/ regeneration) injury model (Raw264.7 cells), we identified Nur77 as a HDAC1 target in macrophages. Nur77 deficient macrophages failed to downregulate IDH1 (isocitrate dehydrogenase 1) and accumulated succinic acid and other tricarboxylic acid (TCA) cycle-derived metabolites in a glutamine-independent manner. These data show that the inhibition of HDAC1 ameliorates H/R-inflammation in macrophages through the regulation of Nur77 and the TCA cycle.
Similar content being viewed by others
Introduction
Histone deacetylases (HDACs) play a key role in a range of pathophysiological conditions [1]. HDACs comprise classes I, II, III, and IV [2] and remove acetyl groups from lysine residues in histones and other cellular proteins to regulate gene expression [3]. The functional relevance of HDAC1 in cardiovascular disease remains uncharacterized.
Ischemic heart disease (IHD), specifically acute myocardial infarction, is a leading cause of mortality [4, 5]. Despite the acute nature of ischemia and re-perfusion injury (I/R), harmful effects of IHD on the myocardium are observed [6]. Low energy and oxygen levels lead to the depletion of ATP, the disruption of ionic homeostasis, increased glycolysis, disturbed oxidative phosphorylation in mitochondria, increased mitochondrial membrane permeability (MMP), and decreased myocardial contractility [7,8,9]. Previous studies have shown that the inhibition of HDAC1 preceding I/R injury is protective to left ventricular activity and the survival of myocardial cells [10,11,12].
Nur77 (NR4A1) is a transcription factor first observed in pheochromocytoma (PC12) tumor cells [13]. Nur77 participates in cell maturity, proliferation, and differentiation through its cooperation with epidermal growth factor (EGF) and nerve growth factor (NGF) [14]. Tumor cell apoptosis is facilitated by the relocalisation of Nur77 to the mitochondrial membrane [15, 16]. At the post-transcriptional level, Nur77 is regulated by acetylation. Histone deacetylase 1 (HDAC1) deactivates Nur77 leading to its ubiquitination [17]. HDAC1 mediated Nor1 and Nur77 inhibition promotes the survival of myeloid leukemia blast cells and stem cells [18]. The role of Nur77 and HDAC1 during I/R injury is poorly defined.
The activation of macrophages through inflammatory stimulation leads to mitochondrial metabolic re-programming and the production of proinflammatory cytokines. Metabolic reprogramming is marked by the down-regulation of IDH1, which regulates a range of inflammatory processes including macrophage polarization [19]. In this study, in vivo mouse models of myocardial I/R and a hypoxia/ regeneration (H/R) injury model in Raw264.7 cells were designed to investigate the role of HDAC1 during myocardial injury.
Results
HDAC1 promotes inflammation post-I/R
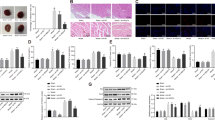
To investigate the role of HDAC1 in I/R, we constructed a myocardial ischemia-reperfusion mouse model. QRT-PCR and western blot analysis showed increased expression of HDAC1 compared to the control group (P < 0.01, P < 0.001 (Fig. 1A, B). Immunohistochemical analysis revealed higher levels of HDAC1 expression in infarct areas (Fig. 1C).
A qRT-PCR analysis of HDAC1 mRNA in control and I/R mice. B HDAC1 expression determined via western blotting. C HDAC1 expression in the infarct area analyzed by IHC. D qRT-PCR and E western blot analysis of HDAC1 expression in I/R mice injected with HDAC1-silencing adenovirus HDAC1 (Ad.HDAC1). F–I IL-6, TNF-a, IL-1β, and IL-10 levels in the supernatants of I/R mice assessed via ELISA. Data are the mean ± SD, n = 3; **P < 0.01. ***P < 0.001.
To further analyze the function(s) of HDAC1 in I/R models, mice were intramuscularly injected with empty-vector (Ad.Null) or an adenovirus silencing HDAC1 (Ad.HDAC1). As shown in Fig. 1D--E, HDAC1 expression was comparable between the Ad. Null+I/R and I/R groups, but decreased in the Ad group (P < 0.01, P < 0.001). When collected plasma samples were assayed for cytokine release by EILSA, higher levels of proinflammatory TNF-a, IL-6, and IL-1β were observed in the I/R group compared to the control (P < 0.001) or Ad groups. The Null+I/R group showed no difference to the I/R group. IL-6, TNF-a, and IL-1β concentrations were lower in the Ad-HDAC1 + I/R group compared to the I/R group (P < 0.01, P < 0.001) (Fig. 1F–H). In contrast, the levels of IL-10 in I/R mice were significantly lower than the control group, but increased in the Ad-HDAC1 + I/R group (P < 0.01, P < 0.001; Fig. 1I). These results confirmed that HDAC1 promotes inflammation in I/R mouse models.
Inhibition of HDAC1 improves H/R-induced macrophage inflammatory responses
We constructed an in vitro H/R injury model in RAW264.3 cells (hypoxia, 8 h; reoxygenation, 16 h). As shown in Fig. 2A, B, RAW264.3 cells in the H/R group showed higher HDAC1 expression compared to the control group (P < 0.001). HDAC1 levels were significantly lower in H/R mice treated with Ad.HDAC1 (P < 0.01, P < 0.001). ELISA analysis revealed lower levels of IL-6, IL-1β and TNF-α and increased levels of IL-10 in both H/R and Ad-HDAC1 groups compared to Null+H/R and control groups (P < 0.001; Fig. 2C–F). The levels of IL-6, TNF-α, and IL-1β were significantly lower in the H/R group treated with Ad. HDAC1, whilst the levels of IL-10 increased (P < 0.01, P < 0.001). All data were confirmed at the mRNA level via qRT-PCR analysis (P < 0.05, P < 0.01, P < 0.001, Fig. 2G–J). Immunofluorescent analysis showed increased expression of CD11b (a marker of M1 macrophages) and lower levels of CD206 (a marker of M2 macrophages) in H/R and Ad.Null+H/R groups compared to the control group. The Ad.HDAC1 + H/R group showed lower levels of CD11b and increased CD206 expression (Fig. 2K). Collectively, these data suggest that the inhibition of HDAC1 ameliorates macrophage inflammation in response to H/R.
A qRT-PCR and B western blot analysis of HDAC1 expression in H/R Raw264.7 cells transfected with Ad.HDAC1. C–F ELISA analysis of cytokine levels. G–J IL-6, IL-10, TNF-α, and IL-1β gene expression determined by qRT-PCR. K Immunofluorescence analysis of CD206 and CD11b expression. n = 3. Data are the mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
HDAC1 regulates metabolic reprogramming in macrophages following H/R
Metabolic reprogramming in macrophages has been linked to polarization and inflammation. Seahorse metabolic assays were performed to establish if the altered energy production was due to the changes in glycolysis or oxidative phosphorylation. RAW264.3 cells in the H/R group showed a significantly lower oxidative consumption rate (OCR) compared to control RAW264.3 cells. Ad.HDAC1 transfection led to a higher OCR compared to Ad. Null+H/R, indicative of increased levels of oxidative phosphorylation (Fig. 3A). In H/R cells, the extracellular acidification rate (ECAR) was higher compared to the control group. Ad.HDAC1 caused a decrease in the ECAR compared to the Ad.HDAC1 + H/R group, indicating reduced glycolytic capacity (Fig. 3B). ATP assays in RAW264.3 cells in the H/R group showed significantly lower levels of ATP production compared to the control group (P < 0.001). Ad.HDAC1 led to increased ATP production compared to H/R and Ad. Null+H/R groups (P < 0.01, Fig. 3C). In the H/R group, the activity of the respiratory chain complex I-V decreased in RAW264.3 cells, but increased in Ad.HDAC1 cells (P < 0.05, P < 0.01, P < 0.001; Fig. 3D–H). Assessment of the MMP using JC-1 (Fig. 3I) revealed a lower MMP in H/R RAW264.3 cells that was reversed by Ad.HDAC1. Together, these data suggest that the inhibition of HDAC1 leads to lower levels of mitochondrial damage.
A Seahorse assays showing decreased oxidative consumption rates (OCR) in H/R (blue) compared to controls (black). Increased OCR in Ad.HDAC1 + H/R (purple) compared to H/R (blue) in response to oligomycin. B Seahorse assays demonstrating increased extracellular acidification rates (ECAR) in H/R (blue) vs. control groups (black), and decreased ECAR in Ad.HDAC1 + I/R (purple) vs. H/R groups (blue) in response to oligomycin. C ATP production in H/R following Ad.HDAC1 transfection. Complexes I (D), II (E), III (F), IV (G), and V (H) activity in RAW264.3 cells. I MMP in RAW264.3 cells. High-intensity red fluorescence and green fluorescence denote increased and decreased membrane potentials, respectively (×400, bar = 100 μm). n = 3. All data are mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
HDAC1 participates in mitochondrial metabolic reprogramming through the regulation of Nur77 deacetylation
As shown in Fig. 4AB, H/R and Ad. Null+H/R groups showed weaker Nur77 mRNA and protein expression compared to the control group. The levels of Nur77 were significantly higher in H/R cells transfected with Ad.HDAC1 (P < 0.01, P < 0.001). To confirm whether the acetylation of Nur77 was regulated by HDAC1 following H/R intervention, HDAC1 Co-immunoprecipitation (CO-IP) assays were performed. We found that the levels of Nur77 acetylation were significantly higher in the AD.HDAC1 + H/R group (Fig. 4C). The activity of Nur77 was enhanced by CsnB which increased ATP production in the I/R group (P < 0.05; Fig. 4D). In addition, CsnB could restore the levels of complexes I, II, III, IV, and V in H/R RAW264.3 cells (P < 0.01, P < 0.001) (Fig. 4E).
A qRT-PCR and B western blot analysis of Nur77 expression in H/R Raw264.7 cells transfected with Ad.HDAC1. C Acetylation (AC-K) of Nur77 regulated by HDAC1 evaluated through Co-IP assays. D ATP production in H/R following CsnB transfection. E Complex I, II, III, IV, and V activity in RAW264.3 cells (n = 3). Data represent the mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
Nur77 participates in inflammatory responses through enhanced transcriptional activity of IDH1
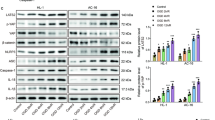
We next sought to determine whether Nur77 regulates the transcriptional activity of IDH1 in H/R RAW264.3 cells. Western blot and qRT-PCR analysis showed lower expression of IDH in the H/R group. However, IDH expression was significantly higher in the H/R group + CsnB (P < 0.05, P < 0.001; Fig. 5A, B). Dual-luciferase reporter assays showed increased luciferase activity in the CsnB+H/R group compared to the empty vector group (P < 0.001, Fig. 5C). Furthermore, IHD1 siRNA (siRNAIDH1) when co-transfected with CsnB into H/R RAW264.3 cells showed lower levels of ATP production and reduced complex I–V activity compared to the control group (P < 0.01, P < 0.001). CsnB alone enhanced ATP production and complex I-V activity in the H/R group (P < 0.001). siRNAIDH1 further restored the CsnB induced changes in H/R RAW264.3 cells (P < 0.05, Fig. 5D, E). JC-1 staining following IHC analysis showed similar effects of siRNAIDH1 on the MMP (Fig. 5F). Together, these findings demonstrate that Nur77 participates in inflammatory responses through its effects on the transcriptional activity of IDH1 in H/R cells.
A qRT-PCR and B western blot analysis of IDH1 expression in H/R Raw264.7 cells transfected with CsnB. C Luciferase reporter assays in CsnB transfected H/R Raw264.7 cells. D ATP production in H/R RAW264.3 cells co-transfected with CsnB and siRNAIDH1. E Complex I, II, III, IV, and V in H/R RAW264.3 cells following co-transfection of CsnB and siRNAIDH1. F Mitochondrial membrane potential assayed via JC-1 staining in H/R RAW264.3 cells co-transfected with CsnB and siRNAIDH1 (×400, bar = 100 μm). n = 3. Data represent the mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, #P < 0.05.
Nur77 activator (CsnB) improves inflammatory responses following I/R in vivo
To further investigate the role of Nur77 in the regulation of I/R by HDAC1, I/R mice were treated with CsnB. ELISA analysis in the I/R group showed increased concentrations of IL-6, TNF-α, and IL-1β and a decreased concentration of IL-10 in the plasma of mice (P < 0.001). As shown in Fig. 6A–D, CsnB treatment could reverse these changes (P < 0.01, P < 0.001), revealing its ability to alleviate inflammatory responses post-I/R.
Discussion
I/R injury occurs following coronary artery disease for which the treatment options remain limited [9, 20]. HDAC inhibitors can preserve myocardial function in a range of animal models [21]. Here, we developed an I/R mouse and H/R injury model in RAW264.3 cells to demonstrate that I/R-induced myocardial injury can be partially alleviated by HDAC1 inhibition.
Previous studies have reported the detrimental effects of HDACs on I/R injury [10, 12, 21]. We found that HDAC1 promotes inflammation in I/R mice and that HDAC1 inhibition suppresses inflammation. We further demonstrated that H/R leads to inflammation and mitochondrial damage.
Macrophages are key to innate immune responses due to their role in tissue development, host defenses and cellular homeostasis [22]. In activated macrophages, the inflammatory response leads to profound reprogramming during cellular metabolism [23]. M1 and M2 macrophages display distinct functions [24]. M1 macrophages express high levels of CD11b [25, 26] and produce proinflammatory cytokines. M2 macrophages are characterized by high CD206 and IL-10 expression [27, 28]. We found that HDAC1 inhibition increased the polarization of M2 macrophages but reduced M1 polarization in H/R RAW264.3 cells. Furthermore, the inhibition of HDAC1 increased both the glycolytic capacity and energy efficiency of RAW264.3 cells, increasing ATP production and mitochondrial complex I-V activity. The inhibition of HDAC1 also decreased the H/R-induced loss of MMP. Oxidative phosphorylation and metabolic reprogramming are characteristic of activated inflammatory M1 macrophages [29]. Glycolysis, oxidative phosphorylation, pentose phosphate pathway, fatty acid oxidation and the metabolism of amino acids represent key metabolic pathways [30, 31]. Ischemic preconditioning activates mitochondrial Src, regulating complex I activity and the levels of mitochondrial reactive oxygen species (ROS) to counter myocardial I/R [32]. In line with these studies, mitochondrial dysfunction and oxidative stress were observed in H/R RAW264.3 cells, which increased following HDAC1 silencing. HDAC1 therefore contributed to oxidative stress and mitochondrial dysfunction through the progression of I/R.
HDAC1 prevents the acetylation and subsequent expression of Nur77 [18]. In RAW264.3 cells, HDAC1 was found to interact with Nur77 and inhibit its acetylation. The inhibition of HDAC1 enhanced the levels of AcK, a promotor driven by Nur77, implying that the inhibition of HDAC1 facilitated the hyperacetylation of Nur77. CsnB has been identified as a ligand for Nur77 [33, 34] and promoted oxidative stress in H/R RAW264.3 cells. Moreover, CsnB could improve the inflammatory response in I/R mice. Collectively, these data suggest that HDAC1 participates in mitochondrial metabolic reprogramming through the regulation of Nur77 deacetylation.
IDH1 participates in the TCA cycle, catalyzing oxidative isocitrate decarboxylation for the production of alpha-ketoglutarate (AKG) and CO2 [35]. Activated macrophages are glycolytic and produce high levels of ROS and succinate [36]. We found that siRNAIDH1 enhanced the effects of CsnB in H/R RAW264.3 cells (P < 0.05, Fig. 5D, E). These data suggest that Nur77 participates in inflammatory responses through its regulation of the transcriptional activity of IDH1.
In summary, we identify the HDAC-mediated deacetylation of Nur77 as a novel mechanism that is critical for macrophage function following I/R injury. This identifies the HDAC-Nur77 axis as potential therapeutic target for much-needed I/R therapeutics.
Materials and methods
Animals studies and I/R grouping
Six-week-old BALB/c nude mice (22–25 g weight) were purchased from Charles River, Beijing, China. Mice were fed an identical diet and provided free access to water. All procedures were approved by the Animal Ethics Committee of the Tianjin Chest Hospital (TJCH-2022002). Mice were divided into 4 groups (n = 6 per-group): [1] Control; [2] I/R; [3] Ad.HDAC1 + I/R; and [4] Ad.Null+I/R groups. Mice were euthanatized using 40 mg/kg sodium pentobarbital (i.p.). Blood and tissue samples were collected for subsequent analysis.
In vivo regional I/R surgery
Anesthesia, intubation and ventilation were performed as previously described [37, 38]. Regional I/R was performed through the ligation of LAD for 30 min and its release for a single day with a left lateral thoracotomy.
Cell culture and H/R
Raw264.7 cells were purchased from ATCC and maintained in DMEM (Gibco, USA) supplemented with 10% FBS and 100 U/mL streptomycin-penicillin. Raw264.7 cells were assigned to the following groups: [1] Control; [2] H/R; [3] Ad.Null+H/R; [4] Ad.HDAC1 + H/R and [5] CsnB+H/R (Nur77 activator, Cytosporine B (CsnB) 10 μg/ml). For H/R, cells were cultured in hypoxic buffer (5% CO2; 1% O2; 94% N2) for 30 min at 37 °C in a hypoxia/anaerobic incubator [39]. Cells were then incubated in DMEM + 10% FBS for 1 h to reoxygenate the cells.
Adenovirus transduction
Mice were infected with 2.4 × 107 PFU/ml recombinant adenovirus for HDAC1 silencing (Genchem, Changzhou, China). For I/R, the heart was exposed between the 4th and 5th ribs of the left chest, and microinjections were performed into the left ventricle anterior wall using a 30 G (Gauge) needle (Ad.Null or Ad.HDAC1). For in vitro experiments, Raw264.7 cells were seeded into 6-well plates and transfected with lentiviral plasmids carrying Ad.HDAC1 using Lipofectamine 3000 (Invitrogen, USA).
Macrophage polarization assays
Immunofluorescence analyses were performed as previously described [40, 41]. Briefly, Raw264.7 cells were seeded into 24 well plates and fixed for 30 min in 4% paraformaldehyde. Cells were permeabilsed in 0.05% Triton X-100 for 10 min, blocked in 1% BSA in PBS for 10 min and probed with anti-CD11b (1:100) and anti-CD206 (1:200) primary antibodies overnight at 4 °C. Cells were labeled with fluorescently conjugated goat anti-mouse IgG (Abcam, UK). Nuclei were stained with DAPI. Cells were imaged on a confocal laser scanning microscopy (CLSM, LSM 510 META; Germany).
RT-PCR
Cells were lysed in TRIzol (Invitrogen, Waltham, MA, USA) and total RNA was isolated. Samples were DNase-I treated and subject to first-strand cDNA synthesis using oligo-dT primers (Invitrogen, USA) and M-MuLV reverse transcriptase (Fermentas, USA). Primers for RT-PCR were as follows: HDAC1 (F: 5′-CCGCATGACTCATAATTTGCTG-3′, R: 5′-ATTGGCTTTGTGAGGGCGATA-3′), Nur77 (F: 5′- GTTGATGTTCCCGCCTTTGCC-3′, R: 5′- TCAGAAAGACAATGTGTCCAT-3′), IDH1 (F: 5′-TGCCACCAACGACCAAGTCA-3′, R: 5′-TGTGTTGAGATGGACGCCTATTTG-3′), and β-actin (reference gene) (F: 5′-CATGTACGTTGCTATCCAGGC-3′, R: 5′-CTCCTTAATGTCACGCACGAT-3′).
Western blotting
Cells were washed in PBS and lysed in Radio-Immunoprecipitation Assay (RIPA) buffer on ice. Lysates were clarified by centrifugation and proteins were resolved by SDS-PAGE electrophoresis. Samples were transferred onto nitrocellulose membranes and probed with the following primary antibodies: anti-HDAC1, anti-Nur77, and anti-IDH1 (1:1000) for 24 h at 4 °C. Membranes were labeled with horseradish peroxidase (HRP)-conjugated anti-mouse antibodies for 1 h. Luminal agent was used for protein detection (SC-2048, Santa Cruz Biotechnology, CA, USA).
ELISA
Blood samples were collected and serum IL-6, TNF-a, IL-10, and IL-1β levels were evaluated by ELISA (USCN Life Science, Inc., China). Absorbances were read on a microplate reader (Bio-Tek Instruments, Inc., USA).
Metabolic parameters
Cells were treated with Seahorse XF assay medium (pH = 7.4) supplemented with 2 mM glutamine, 1 mM pyruvate, and 10 mM glucose. Cells were incubated in CO2 free conditions for 1 h hour at 37 °C. Oxygen consumption (OCR) and extracellular acidification (ECAR) rates were measured following the addition of glucose, 2-deoxy-d-glucose (2DG) and oligomycin. Cells were further assessed following treatment with carbonyl cyanide 4-trifluoromethoxyphenylhydrazone (FCCP), oligomycin, glucose, 2-DG, rotenone, and antimycin A (Rot/AA). Adenosine triphosphate (ATP) levels and metabolic parameters were calculated using a Seahorse XF96 Analyzer (Agilent, Santa Clara, CA, USA).
Mitochondrial respiratory chain complex activity
Mitochondrial respiratory activity was measured using commercial kits (ab110419, Abcam, Cambridge, UK) as per the manufacturer’s instructions. Absorbances were measured on a BioRad microplate reader (BioRad, Hercules, CA).
Mitochondrial membrane potential
JC-1 was used as a marker of MMP. When Dwm is high, JC-1 produces red fluorescence. Green fluorescence is produced by JC-1 monomers. For actin staining, cells were permeabilized in 0.1% Triton X-100 for 5 min, blocked in 10% goat serum in PBS for 1 h and labeled with mouse anti-a-actin (Sigma, St. Louis, USA, 1:1000) and Cy3 labeled secondary antibodies for 24 h at 4 °C. Nuclei were counterstained with DAPI.
Co-immunoprecipitation analysis
Cells were washed in PBS and resuspended in BC300 buffer (0.2 mM EDTA, 0.2 mM PMSF, 20 mM Tris-HCl, 300 mM NaCl, 0.2% Tween-20, and 10% glycerol), followed by sonication for 10 min on ice. Lysates were incubated with primary antibodies (anti-Nur77, ab283264, Abcam; acetylated-lysine antibody (ACK) #9441, CST) for 24 h at 4 °C. Anti-Rabbit IgG antibodies were added as a negative control. Lysates were incubated with A/G agarose beads (Cell Signaling Technology, USA) for 3 h and centrifuged. Beads were washed in BC300 buffer, boiled and analyzed on 15% SDS-PAGE gels.
Statistical analysis
SPSS 20.2 and GraphPad Prism 7 were used for data analysis. Data represent the mean ± SD. A Student’s t-test (two-tailed) or a one-way ANOVA were used for single or multiple group comparisons, respectively. P < 0.05 was deemed statistically significant.
Data availability
All the data used to support the findings of this study are included within the article.
References
Gao Y, Gao J, Mu G, Zhang Y, Huang F, Zhang W, et al. Selectively enhancing radiosensitivity of cancer cells via in situ enzyme-instructed peptide self-assembly. Acta Pharm Sin B. 2020;10:2374–83.
Wang Y, Wang L, Su Z, Sun W, Zhang M, Yang C, et al. POU2AF1 promotes MSCs adipogenesis by inhibiting HDAC1 expression. Adipocyte. 2021;10:251–63.
Bäckdahl L, Bushell A, Beck S. Inflammatory signalling as mediator of epigenetic modulation in tissue-specific chronic inflammation. Int J Biochem Cell Biol. 2009;41:176–84.
Roth GA, Johnson C, Abajobir A, Abd-Allah F, Abera SF, Abyu G, et al. Global, regional, and national burden of cardiovascular diseases for 10 causes, 1990 to 2015. J Am Coll Cardiol. 2017;70:1–25.
Hausenloy DJ, Yellon DM. Ischaemic conditioning and reperfusion injury. Nat Rev Cardiol. 2016;13:193–209.
Hausenloy DJ, Yellon DM. Myocardial ischemia-reperfusion injury: a neglected therapeutic target. J Clin Invest. 2013;123:92–100.
de Groot H, Rauen U. Ischemia-reperfusion injury: processes in pathogenetic networks: a review. Transpl Proc. 2007;39:481–4.
Wu XJ, Sun XH, Wang SW, Chen JL, Bi YH, Jiang DX. Mifepristone alleviates cerebral ischemia-reperfusion injury in rats by stimulating PPAR γ. Eur Rev Med Pharm Sci. 2018;22:5688–96.
Lesnefsky EJ, Hoppel CL. Ischemia-reperfusion injury in the aged heart: role of mitochondria. Arch Biochem Biophys. 2003;420:287–97.
Aune SE, Herr DJ, Mani SK, Menick DR. Selective inhibition of class I but not class IIb histone deacetylases exerts cardiac protection from ischemia reperfusion. J Mol Cell Cardiol. 2014;72:138–45.
Zhao TC, Cheng G, Zhang LX, Tseng YT, Padbury JF. Inhibition of histone deacetylases triggers pharmacologic preconditioning effects against myocardial ischemic injury. Cardiovasc Res. 2007;76:473–81.
Zhang L, Qin X, Zhao Y, Fast L, Zhuang S, Liu P, et al. Inhibition of histone deacetylases preserves myocardial performance and prevents cardiac remodeling through stimulation of endogenous angiomyogenesis. J Pharm Exp Ther. 2012;341:285–93.
Maruoka H, Sasaya H, Shimamura Y, Nakatani Y, Shimoke K, Ikeuchi T. Dibutyryl-cAMP up-regulates nur77 expression via histone modification during neurite outgrowth in PC12 cells. J Biochem. 2010;148:93–101.
Liu Y, Zhang J, Yi B, Chen M, Qi J, Yin Y, et al. Nur77 suppresses pulmonary artery smooth muscle cell proliferation through inhibition of the STAT3/Pim-1/NFAT pathway. Am J Respir Cell Mol Biol. 2014;50:379–88.
Lee SO, Jin UH, Kang JH, Kim SB, Guthrie AS, Sreevalsan S, et al. The orphan nuclear receptor NR4A1 (Nur77) regulates oxidative and endoplasmic reticulum stress in pancreatic cancer cells. Mol Cancer Res. 2014;12:527–38.
To SK, Zeng WJ, Zeng JZ, Wong AS. Hypoxia triggers a Nur77-β-catenin feed-forward loop to promote the invasive growth of colon cancer cells. Br J Cancer. 2014;110:935–45.
Kang SA, Na H, Kang HJ, Kim SH, Lee MH, Lee MO. Regulation of Nur77 protein turnover through acetylation and deacetylation induced by p300 and HDAC1. Biochem Pharmacol. 2010;80:867–73.
Zhou L, Ruvolo VR, McQueen T, Chen W, Samudio IJ, Conneely O, et al. HDAC inhibition by SNDX-275 (Entinostat) restores expression of silenced leukemia-associated transcription factors Nur77 and Nor1 and of key pro-apoptotic proteins in AML. Leukemia. 2013;27:1358–68.
Haschemi A, Kosma P, Gille L, Evans CR, Burant CF, Starkl P, et al. The sedoheptulose kinase CARKL directs macrophage polarization through control of glucose metabolism. Cell Metab. 2012;15:813–26.
Wu MY, Yiang GT, Liao WT, Tsai AP, Cheng YL, Cheng PW, et al. Current mechanistic concepts in ischemia and reperfusion injury. Cell Physiol Biochem. 2018;46:1650–67.
Xie M, Kong Y, Tan W, May H, Battiprolu PK, Pedrozo Z, et al. Histone deacetylase inhibition blunts ischemia/reperfusion injury by inducing cardiomyocyte autophagy. Circulation. 2014;129:1139–51.
Wynn TA, Vannella KM. Macrophages in tissue repair, regeneration, and fibrosis. Immunity. 2016;44:450–62.
Viola A, Munari F, Sánchez-Rodríguez R, Scolaro T, Castegna A. The metabolic signature of macrophage responses. Front Immunol. 2019;10:1462.
Murray PJ, Wynn TA. Protective and pathogenic functions of macrophage subsets. Nat Rev Immunol. 2011;11:723–37.
Abumaree MH, Al Jumah MA, Kalionis B, Jawdat D, Al Khaldi A, Abomaray FM, et al. Human placental mesenchymal stem cells (pMSCs) play a role as immune suppressive cells by shifting macrophage differentiation from inflammatory M1 to anti-inflammatory M2 macrophages. Stem Cell Rev Rep. 2013;9:620–41.
Park HJ, Kim J, Saima FT, Rhee KJ, Hwang S, Kim MY, et al. Adipose-derived stem cells ameliorate colitis by suppression of inflammasome formation and regulation of M1-macrophage population through prostaglandin E2. Biochem Biophys Res Commun. 2018;498:988–95.
Anzai A, Anzai T, Nagai S, Maekawa Y, Naito K, Kaneko H, et al. Regulatory role of dendritic cells in postinfarction healing and left ventricular remodeling. Circulation. 2012;125:1234–45.
Perego C, Fumagalli S, Zanier ER, Carlino E, Panini N, Erba E, et al. Macrophages are essential for maintaining a M2 protective response early after ischemic brain injury. Neurobiol Dis. 2016;96:284–93.
Vijayan V, Pradhan P, Braud L, Fuchs HR, Gueler F, Motterlini R, et al. Human and murine macrophages exhibit differential metabolic responses to lipopolysaccharide - A divergent role for glycolysis. Redox Biol. 2019;22:101147.
O’Neill LA, Kishton RJ, Rathmell J. A guide to immunometabolism for immunologists. Nat Rev Immunol. 2016;16:553–65.
Zarrinpar A, Bensinger SJ. The therapeutic potential of T cell metabolism. Am J Transpl. 2017;17:1705–12.
Ge H, Zhao M, Lee S, Xu Z. Mitochondrial Src tyrosine kinase plays a role in the cardioprotective effect of ischemic preconditioning by modulating complex I activity and mitochondrial ROS generation. Free Radic Res. 2015;49:1210–7.
Liu TY, Yang XY, Zheng LT, Wang GH, Zhen XC. Activation of Nur77 in microglia attenuates proinflammatory mediators production and protects dopaminergic neurons from inflammation-induced cell death. J Neurochem. 2017;140:589–604.
Ismail H. Vascular endothelial growth factor and angiopoietin-1 regulate leukocyte adhesion to endothelial cells through the nuclear receptor Nur77. Canada: McGill University; 2012.
Chang YC, Tsai HF, Huang SP, Chen CL, Hsiao M & Tsai WC. Enrichment of aldolase C correlates with low non-mutated IDH1 expression and predicts a favorable prognosis in glioblastomas. Cancers. 2019;11:1238.
Balic JJ, Albargy H, Luu K, Kirby FJ, Jayasekara WSN, Mansell F, et al. STAT3 serine phosphorylation is required for TLR4 metabolic reprogramming and IL-1β expression. Nat Commun. 2020;11:3816.
Quan N, Sun W, Wang L, Chen X, Bogan JS, Zhou X, et al. Sestrin2 prevents age-related intolerance to ischemia and reperfusion injury by modulating substrate metabolism. FASEB J. 2017;31:4153–67.
Wang J, Yang L, Rezaie AR, Li J. Activated protein C protects against myocardial ischemic/reperfusion injury through AMP-activated protein kinase signaling. J Thromb Haemost. 2011;9:1308–17.
Chu Q, Zhang Y, Zhong S, Gao F, Chen Y, Wang B, et al. N-n-butyl haloperidol iodide ameliorates oxidative stress in mitochondria induced by hypoxia/reoxygenation through the mitochondrial c-Jun N-terminal kinase/Sab/Src/reactive oxygen species pathway in H9c2 Cells. Oxid Med Cell Longev. 2019;2019:7417561.
Xu R, Zhang F, Chai R, Zhou W, Hu M, Liu B, et al. Exosomes derived from pro-inflammatory bone marrow-derived mesenchymal stem cells reduce inflammation and myocardial injury via mediating macrophage polarization. J Cell Mol Med. 2019;23:7617–31.
Liu N, Kong T, Chen X, Hu H, Gu H, Liu S, et al. Ubiquitin-specific protease 14 regulates LPS-induced inflammation by increasing ERK1/2 phosphorylation and NF-κB activation. Mol Cell Biochem. 2017;431:87–96.
Acknowledgements
The study was supported by Tianjin Education Commission scientific research project (2022YGYB10) and Tianjin Health Science and Technology Project (No. ZC20147) to ZW.
Author information
Authors and Affiliations
Contributions
ZW and YPB wrote the manuscript. YQ and CC collected the data. YJ, YBB, and ZG analyzed the data. All authors revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, Z., Bai, Y., Qi, Y. et al. HDAC1 disrupts the tricarboxylic acid (TCA) cycle through the deacetylation of Nur77 and promotes inflammation in ischemia-reperfusion mice. Cell Death Discov. 9, 10 (2023). https://doi.org/10.1038/s41420-023-01308-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01308-1