Abstract
Cytochrome c oxidase subunit VIc (COX6c) is one of the most important subunits of the terminal enzyme of the respiratory chain in mitochondria. Numerous studies have demonstrated that COX6c plays a critical role in the regulation of oxidative phosphorylation (OXPHOS) and energy production. The release of COX6c from the mitochondria may be a hallmark of the intrinsic apoptosis pathway. Moreover, The changes in COX6c expression are widespread in a variety of diseases and can be chosen as a potential biomarker for diagnosis and treatment. In light of its exclusive effects, we present the elaborate roles that COX6c plays in various diseases. In this review, we first introduced basic knowledge regarding COX6c and its functions in the OXPHOS and apoptosis pathways. Subsequently, we described the regulation of COX6c expression and activity in both positive and negative ways. Furthermore, we summarized the elaborate roles that COX6c plays in various diseases, including cardiovascular disease, kidney disease, brain injury, skeletal muscle injury, and tumors. This review highlights recent advances and provides a comprehensive summary of COX6c in the regulation of OXPHOS in multiple diseases and may be helpful for drug design and the prediction, diagnosis, treatment, and prognosis of diseases.
Similar content being viewed by others
Facts
-
COX6c is one of the most important subunits of the terminal enzyme of the respiratory chain in mitochondria.
-
COX6c is closely related to a variety of diseases and can be used as a potential biomarker for diagnosis and treatment.
-
COX6c plays a pivotal role in the regulation of the OXPHOS pathway. Mitochondrial biogenesis can be stimulated in response to low temperature and is reflected in increases in the expression and activity of COX6c.
Open questions
-
What extent are the activity and expression of COX6c altered in various diseases?
-
Is the change in COX6c a secondary event associated with other pathways that might be the primary drivers of the functional outcomes or is it specific to the disease?
-
What are the specific mechanisms and processes that COX6c participates in during the development of disease?
Introduction
The mitochondrial respiratory chain, also known as the electron transport chain (ETC), is crucial to life. Cytochrome c oxidase (COX), the terminal enzyme of the respiratory chain in the mitochondrial inner membrane, has been acknowledged as a rate-limiting step of energy production in mitochondria. It not only catalyzes the electron transfer from reduced cytochrome c to molecular oxygen but also translocates protons across the mitochondrial membrane. The proton motive force generated by COX (Complex IV), along with the other two proton-pumping enzymes, namely, Complexes I (NADH: ubiquinone oxidoreductase) and III (cytochrome bc1), subsequently drives ATP synthase and facilitates energy production through oxidative phosphorylation (OXPHOS) [1]. In humans, COX biogenesis involves the coordinated assembly of 13 subunits, the largest 3 of which, I, II, and III, are catalytic subunits encoded by mitochondrial DNA (mtDNA) and synthesized within the mitochondria. The remaining 10 structural subunits are encoded by nuclear DNA and synthesized on cytosolic ribosomes, subsequently being translocated into the mitochondrial inner membrane mainly through mitochondrial targeting and import [2,3,4]. The mitochondrial-encoded subunits act in electron transfer, whereas the nuclear-encoded subunits appear to be involved in the structural integrity, regulation, and dimerization of the enzyme. The presence of isoforms may reflect the different energy requirements that depend on tissues, developmental stages, or growth conditions [5, 6]. For instance, COX4-2 is the lung-specific isoform, COX6B and COX8-3 are the two testis-specific isoforms, and COX6A, COX7A, and COX8A are the three liver/heart-type subunits. In addition, heart isoforms are expressed in tissues that possess a high aerobic capacity and an abundance of mitochondria, whereas liver isoforms are found in tissues that involve fewer mitochondria, such as the brain, liver, and kidney [7]. In particular, the nuclear gene that encodes subunit VIc, COX6c, has been shown to be the most altered in a variety of diseases.
COX6c is a 933-bp linear DNA located at human chromosome 8q22.2 that comprises 4 exons and 3 introns [8, 9]. Among all these COX subunits, COX6c has no paralogs in any vertebrate lineage, including mammals. COX6c polymorphisms in exons 2 and 3 form a haplotype [10]. The mature COX6c protein is 75 amino acids long and shows 69% homology between rats and humans as well as 73% between bovines and humans [11]. However, the structure in mice requires further elucidation. In addition, the mature protein-coding region of the COX6c gene has been shown to undergo accelerated evolution in amino acid replacement in the human lineage, suggesting an adaptive positive selection of functional significance [12]. Although mature COX6c protein possesses a property of the presequence, it does not have a cleavable presequence that targets mitochondria and has only two amino acids cleaved from the mature protein [3]. COX6c subunits can be assembled into subcomplexes in vitro, which may represent rate-limiting intermediates, according to in vitro introduction and assembly experiments [13]. Furthermore, ubiquitous COX6c has been reported to be constitutively expressed in almost all tissues, especially the heart and colon [14].
COX6c plays a pivotal role in the regulation of the OXPHOS pathway. Mitochondrial biogenesis can be stimulated in response to low temperature and is reflected in increases in the expression and activity of COX6c [15]. It has been well established that human sperm requires mitochondrial ATP through OXPHOS [16, 17]. Amaral and colleagues [18] noted that good-quality sperm expresses a significant amount of ETC proteins, including COXI and COX6c, compared with low-quality samples. The positive correlation between the expression of COX6c and sperm quality is more important for sperm concentration and motility than morphology. Notably, there is no correlation between the percentage of sperm expressing COX6c and mtDNA copy number. This result further confirms that other factors affect COX6c expression, such as nuclear respiratory factors (NRFs), which will be discussed below.
Novel function of COX6c beyond OXPHOS
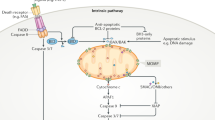
The release of COX6c from mitochondria may be a telling sign of the intrinsic apoptotic pathway, which is consistent with the significant function that COX6c plays in modulating OXPHOS (Fig. 1). Interleukin (IL)-24 controls the production of several genes involved in cell proliferation and death, according to research by Hadife and colleagues [19]. Within the first 6 h, the transcription of genes related to DNA replication and metabolism, such as cell division cycle 6 (CDC6), is inhibited in the B-cell differentiation model. In the later 36 h, IL-24 stimulates COX6c expression and triggers the mitochondrial apoptotic pathway. IL-24-mediated apoptosis in B-cells has been shown to be dependent on p53. Several lines of evidence suggest that p53 can cripple the Warburg effect, where cancer cells exhibit a preferential utilization of glycolysis over aerobic respiration to produce ATP. The related mechanisms involve the regulation of the expression of synthesis of COX (SCO2), a copper-binding protein that is essential for the assembly of COX [20, 21]. Moreover, COX6c expression is remarkably suppressed by glucose, which is also supported by evidence that cells grown in the presence of lactate exhibit approximately 5 times higher COX6c mRNA levels than those grown in glucose [22, 23]. It will be of interest to determine whether COX6c expression is further boosted by p53 under stress conditions, such as glucose deprivation.
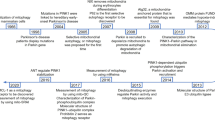
COX6c is regulated by multiple factors. (1) On the one hand, APE1 promotes the binding of NRFs to TFAM and regulates mitochondrial biogenesis. In addition, the expression of COX6c is regulated by the redox-dependent coactivator NRFs of APE1; (2) On the other hand, PGC-1α binds to NRFs and ERRα, respectively, to promote the expression of COX6c, which in turn promotes apoptosis and oxidative phosphorylation. During the whole process, COX6c is negatively regulated by miR-4276, Tat, DAZAP1.
Othumpangat and colleagues [24] observed that during the first 3 h after the influenza virus exposure, host cells boost COX6c mRNA expression by silencing miR-4276. Moreover, COX6c in turn inhibits viral replication via activation of caspase-9. This indicates an initial cellular defense that eliminates infected cells by driving them into an apoptotic state. Noticeably, when the virus takes control of the cell, miR-4276 induction represses the expression of COX6c and caspase-9, allowing the cell to transform into an antiapoptotic state and finally promoting viral replication and spread. Since the expression of miR-4276 and COX6c is not specific to the viral strain and there is an antigenic shift and drift of the virus, miRNA inhibitors may be more broadly applicable than vaccines that target a specific virus. Thus, these data strongly render a more effective approach to antiviral therapy.
Regulation of COX6c
Peroxisome proliferator-activated receptor γ (PPARγ) coactivator 1 (PGC-1α), the energetic regulator, is acknowledged as a critical signaling pathway regulating COX6c expression [25]. As a powerful regulator of cellular energy metabolism, PGC-1α is primarily expressed in tissues and organs rich in mitochondria, such as the heart, liver, kidney, brain, brown adipose tissue, and skeletal muscle. PGC-1α initiates the expression of a wide range of coactivated genes involved in virtually all aspects of mitochondrial energy metabolism [26]. NRF-1, one of the most important coactivated targets of PGC-1α, remarkably activates the expression of the gene that encodes COX6c through binding to its promoter [27, 28]. Moreover, NRF-1-binding sites are also identified in the promoter of mitochondrial transcription factor A (TFAM), which subsequently binds to mtDNA and exerts significant effects on mtDNA replication, transcription, and maintenance [26]. The transcriptional activity of both NRF-1 downstream target genes, COX6c and TFAM, is enhanced by an increase in NRF-1 activity upon hydrogen peroxide-induced oxidative stress [29, 30]. The other isoform, NRF-2, plays a parallel role in the transcriptional regulation of COX6c [31, 32]. Furthermore, the human apurinic/apyrimidinic (AP) endonuclease 1 (APE1) has been demonstrated to play an additional regulatory part in the DNA-binding and transcriptional activity of NRF-1 through its redox function, further modulating mitochondrial function after oxidative stress. Knockdown or redox mutation of APE1 impairs NRF-1 DNA-binding activity, consequently reducing the expression of COX6c [29]. Hence, these results suggest that APE1 could be either the redox-dependent coactivator of NRF-1 or the redox-dependent mediator between PGC-1α and NRF-1, which needs to be further clarified. (Fig. 1)
The expression of COX6c is also induced by the estrogen-associated receptor, another PGC-1 partner, and ERRα target genes have a conserved ERR responder (ERRE) in their promoter [33, 34]. In addition, the binding of ERRα to ERRE in its own promoter stimulates its transcription in a positive autoregulatory loop once activated by PGC-1α [26]. COX6c can be used as an endpoint of ERRα activity in contracting cardiomyocytes by measuring changes in the corresponding mRNA level in response to hypoxia. As shown by Cunningham and colleagues [33], contracting adult cardiomyocytes’ adaptive response to persistent hypoxia includes the stimulation of the PGC-1/ERR/COX6c-signaling pathway. Together, the PGC-1α-signaling pathway is essential for the transcriptional regulation of COX6c and further helps maintain mitochondrial function.
COX6c activity and expression can also be negatively modulated. The human immunodeficiency virus type 1 (HIV-1) transactivator of transcription (Tat) protein is an important contributor to the HIV-induced pathogenesis of acquired immunodeficiency syndrome (AIDS), including the apoptosis of various cell types [35]. Tat protein is able to suppress COX activity in homogenates from various human tissues and organs, including the liver, heart, brain, and skeletal muscle [36]. This increases the potential of the viral protein as a COX6c inhibitor. The transcriptional activity of COX6c can be regulated at the posttranslational level by miRNAs, such as miR-4276, which has been discussed above. Moreover, modifications of COX6c expression include a decrease in COX6c pre-mRNA splicing efficiency by DAZAP1, which is an evolutionarily conserved RNA-binding protein that can regulate gene transcription and translation [37, 38]. This is demonstrated by the overexpression of DAZAP1, which directly contributes to the accumulation of unspliced COX6c pre-mRNA, further inhibiting COX6c. It is notable that DAZAP1 loads onto COX6c mRNA with just the last intron of the COX6c gene present, indicating the occurrence of specific intron-dependent mRNA binding. In addition, DAZAP1 knockdown causes the overexpression of COX6c, which ultimately slows the proliferation of HEK293 cells [9]. Overall, DAZAP1 is identified as a negative regulator of COX6c expression and may control energy production in mitochondria.
COX6c and diseases
COX6c plays a key part in endocrine, urinary, reproductive, circulatory, respiratory, and other systems. This section mainly introduces its relationship with diseases of different systems, such as cardiovascular disease, and diabetic nephropathy (DN).
COX6c and cardiovascular disease
Cardiovascular diseases such as atherosclerosis are common diseases of the human circulatory system, which affect the life quality of human beings. Atherosclerosis is an inflammatory progressive disease attributed to the deposition of lipids, namely, cholesterol and cholesterol esters, to the intima of the arterial wall. This vascular disease is associated with various risk factors, such as hypercholesterolemia, hypertension, and diabetes [39]. The analysis of KEGG signaling revealed significant enrichment of differentially expressed genes, including the COX6c gene in the OXPHOS pathway in familial hypercholesterolemia patients [40]. Since mitochondria are also a prime cellular source of reactive oxygen species (ROS), the underlying mechanism may entail the ROS accumulation in COX6c, which contributes to the increase in oxidative stress, ultimately hastening the evolution of hypercholesterolemia and atherosclerosis [40, 41]. Moreover, the downregulation of COX6c expression in blood samples from familial hypercholesterolemia patients suggests that aberrant expression of COX6c may result in the development of atherosclerosis through the OXPHOS pathway [40]. These data strongly indicate that the COX6c gene may function as a potential marker for the prediction and treatment of atherosclerosis (Table 1).
COX6c and kidney disease
Diabetes is a complex polygenic disease with increasing global prevalence. COX6c has been proven to use as a significant gene in both fulminant type 1 diabetes [42] and type 2 diabetes mellitus [43] to provide new therapeutic targets. It is closely related that DN is a serious endocrine system disease, which has become the single most common cause of end-stage renal disease (ESRD) in adults. Approximately 30% of patients with diabetes mellitus develop DN. Zhang et al. [44] discovered that the expression of COX6c is noticeably elevated in the glomeruli of DN rats, implying that the OXPHOS pathway may be vital to the emergence of DN. By artificially deflating COX6c expression and blocking the OXPHOS pathway, they proved that telmisartan ultimately improves renal function in diabetic rats [45]. Additionally, through PPAR, a crucial coactivated partner of PGC-1, telmisartan moderates renal function [45, 46]. Telmisartan has also been shown to down-regulate TGF-1-induced expression of α-smooth muscle actin and collagen IV in mesangial cells by activating PPAR [45]. Furthermore, telmisartan also inhibits angiotensin II receptor (AT-1R) expression and drives the expression of ROS detoxifying enzymes, such as superoxide dismutase (SOD), through the PGC-1α/PPARγ pathway, further improving kidney function together with the OXPHOS pathway [26, 46, 47].
ESRD patients need renal replacement therapies (RRTs), including hemodialysis (HD) and peritoneal dialysis (PD), to ensure their survival. It has been demonstrated that the expression of COX6c and its upstream gene PGC-1α is significantly downregulated in ESRD patients on HD therapy (ESRD/HD), indicating a reduction in OXPHOS activity [25]. Notably, the downregulation of PGC-1α is independently associated with the development of cardiovascular disease in HD patients. The pharmacological modulation of PGC-1α regulates its downstream effectors, including COX6c, NRF-1, and TFAM, and might be a promising therapeutic tool to reduce oxidative stress-related clinical complications in ESRD/HD patients. Hashad and colleagues [28] confirmed that the expression of NRF-1 and COX6c is downregulated in ESRD/HD patients. Moreover, there is a negative correlation between oxidative stress and the expression of both the NRF-1 and COX6c genes, and COX6c appears more relevant. In addition, Zaza and colleagues [48] found that the expression levels of PGC-1α, NRF-1, and their downstream target gene COX6c are significantly downregulated in ESRD/PD patients in association with increased oxidative stress. On the other hand, nuclear factor erythroid 2-related factor 2 (Nrf2), a transcription factor that induces the expression of the antioxidant enzyme SOD2, is upregulated in ESRD/PD patients [48, 49]. SOD2 then binds to the superoxide byproducts of OXPHOS and converts them to hydrogen peroxide and diatomic oxygen [48]. Together, these data strongly suggest that the decrease in mitochondrial OXPHOS activity reduces ROS accumulation and creates antioxidant feedback, and COX6c and its upstream regulators might be indicators of mitochondrial oxidative metabolism.
COX6c and brain disorder
Decreased COX6c expression and activity have been observed in temporal and parietal cortices in patients with Alzheimer’s disease [10, 50]. However, it is worth noting that this decrease may not be specific to brain disorders but as a secondary event in brain areas associated with impaired metabolic activity and neuronal loss. As one of the components of Complex IV in the ETC, the suppression of COX6c due to a lack of oxygen is also a core initial process of brain ischemic injury. A new study using MALDI imaging mass spectrometry showed increased expression of COX6c in ischemic tissue compared to healthy opposite-brain areas [51]. The underlying mechanism of this compensatory effect involves an elevated requirement of mitochondrial function to restore OXPHOS after ischemia, which might be promoted by the increase of total mitochondrial mass. Therefore, COX6c might represent a potential diagnostic biomarker or target molecule for brain disorders.
COX6c and skeletal muscle injury
Skeletal muscle injury is closely related to the motor system. Du and colleagues [52] found that the COX6c gene in combination with sTnI is more accurate than other genes in estimating the age of wounds following skeletal muscle contusion in rats. Sun and colleagues [53] confirmed that the COX6c expression level in contused skeletal muscle increases within 6 h after contusion when the main pathological feature includes edema and hemorrhage in myocytes without inflammatory cells or fibrous proliferation, and it decreases after 6 h when myocyte degeneration and necrosis occur. These results suggest that the level of COX6c is regular after contusion and may act as a suitable indicator for estimating wound age in combination with sTnI expression or pathological features.
Skeletal muscle function plays a central role in the survival and the prediction of mortality risk in patients with chronic obstructive pulmonary disease (COPD) [54]. COX6c is a hallmark for muscle function loss when acute COPD worsens [55]. The relevant mechanisms could involve the downregulation of the OXPHOS pathway and the upregulation of the apoptosis pathway, both of which are modulated by COX6c under conditions of excessive ROS production within the muscle, ultimately contributing to the alteration of myofilament contractile properties. Hence, the therapeutic intervention of COX6c might counteract the detrimental effects of exacerbation on these pathways and limit the loss of muscle function.
COX6c and tumors
COX6c and prostate cancer
Prostate cancer is the most common form of cancer in males. In situ hybridization has shown that COX6c mRNA expression is upregulated in prostate tumor tissue and is the highest in scattered epithelial cells of prostate carcinoma. Besides that, the mRNA level of COX6c is relatively less in adjacent issues [56]. It has been found that the citrate level is extremely low in cancerous tissue compared with normal prostate tissue [57]. This could be due to the Krebs cycle, which is the mechanism used to produce energy from the metabolism of glucose and lipids, oxidizing citric acid [56]. This meets the increased energy requirement of the cancerous state for the process of malignancy, which may be supported by elevated COX6c. Together, these data indicate that the expression pattern of COX6c may serve as a useful marker to study the alteration of energy metabolism in cancer cells and help the diagnosis of prostate cancer (Table 1).
COX6c and breast cancer
Breast cancer, the most common cancer in females, is responsive to a wide range of chemotherapies. Mitoxantrone (MX), for instance, has attained clinical approval and is routinely used alone or in combination [58]. However, there is still a significant obstacle to the successful therapy of breast cancer due to the development of chemoresistance to multidrug resistance [59]. ATP-binding cassette subfamily G member 2 (ABCG2) is considered the major contributor to drug resistance in MCF-7/MX cells. ABCG2, as well as COX6c and ATP synthase expression, exhibite significantly higher in the MCF-7/MX cell line compared to normal MCF7 cell lines, indicating that all three of these genes increase MCF7 cell tolerance to MX [60]. The increase in COX6c expression appears to be a compensating mechanism to maintain mitochondrial function to produce a large amount of ATP to help ABCG2 pump the MX out of cells against a concentration gradient and to serve as a survival mechanism to overcome MX treatment. Taken together, these data prove that COX6c is a crucial factor in the development of drug resistance in cancer cells and may be critical for the reasonable design and use of new treatment strategies to effectively confront cancers.
Mid-region parathyroid hormone-related protein has been shown to remarkably restrain proliferation and contribute to striking toxicity and accelerate cell death in breast cancer cells [61]. The underlying molecular mechanisms may involve transcriptional reprogramming, especially downregulation of the expression of COX6c and apoptotic genes [62, 63]. Since anticancer therapeutic agents are diverse and highly targeted, there is a necessity for identifying molecularly defined subtypes of breast cancer as well as predictive and prognostic biomarkers. It has been demonstrated that COX6c plays a crucial role in the identification of hormone-responsive breast cancer or estrogen receptor (ER)+ subtypes [64, 65]. Additionally, in predicting patient survival, models incorporating COX6c protein expression together with three other proteins (GATA3, NAT1, and ESR1) significantly outperform baseline models (age, tumor size, and nuclear grade) according to a quantitative assessment of protein expression levels by Emerson and colleagues [65]. This model also offers equivalent prognostic value as lymph node status alone, but the best model is to combine the protein expression data with the nodal status. These results will help further research studies of drug responsiveness and clinical studies of patient prognosis.
COX6c and other tumors
COX6c helps to understand not only the etiology of prostate and breast cancers but also other cancers, including follicular thyroid cancer, uterine myomas, and retroperitoneal lipoma, which aids in tumor diagnosis. Previously, a novel COX6cC/DERL2 translocation in follicular thyroid carcinoma was detected [66]. This somatic alteration is associated with the endoplasmic reticulum and cellular metabolism. Furthermore, the new gene fusion of COX6c to HMGIC is one of the tumorigenic mechanisms in the development of uterine myomas. The first three exons of the HMGIC gene at 12q15, which encode the three DNA-binding domains, are fused to exon 2 of the COX6c gene at 8q22.2, according to nucleotide sequences of the fusion transcript [67]. The fact that Mine and colleagues [68] validated the COX6c/HMGIC translocation in uterine myomas suggested that the fusion partner COX6c gives rise to the detachment of the DNA-binding domains of HMGIC from the spacer and the acidic carboxyl-terminal regulatory domain, thereby promoting tumor growth. Moreover, COX6c has been proposed as a potential participant in the chromosomal organization in a retroperitoneal lipoma instance [69]. Yun et al. also found that COX6c is a key gene in bladder cancer tumors by detecting bladder cancer chromosomal copy number aberrations in urinary sediment [70].
Extracellular vesicles (EVs) obtained from metastatic tissue from melanoma patients have higher levels of mitochondrial membrane proteins, including MT-CO2 (encoded by the mitochondrial genome) and COX6c (encoded by the nuclear genome), than EVs isolated from non-melanoma-derived cells [71]. In gastric cancer, mRNA for LncMI-AS1 and NDUFA4 is upregulated. The expression levels of crucial indicators in the oxidative phosphorylation pathway, such as COX6C, COX5B, and NDUFA8, are observably elevated when lncMIF-AS1 or NDUFA4 are overexpressed in AGS cells [72]. Altogether, these data suggest the involvement of COX6c in gene rearrangement during tumorigenesis, providing additional tools for tumor diagnostics.
Conclusions and perspectives
This review described the pivotal role of COX6c in various diseases. COX6c is one of the most important subunits of the terminal enzyme of the ETC in mitochondria that regulates the OXPHOS pathway and energy production. Moreover, the release of COX6c from the mitochondria may be a hallmark of the intrinsic apoptosis pathway. Notably, we emphasize the promising future of COX6c in tumor therapy and highlight the potential of COX6c in organ protection.
PGC-1α, as well as its coactivated partners, NRFs and ERRs, is essential for the transcriptional regulation of COX6c and further helps maintain mitochondrial function. COX6c activity and expression can also be negatively modulated by multiple molecules. The upregulation of COX6c expression has been determined to meet the increased energy requirement for the process of malignancy or restoration through the OXPHOS pathway. The downregulation of COX6c expression is mainly associated with decreased energy production along with oxidative stress. According to a recent study, COX6c is also a candidate biomarker for liver metabolism, which entails the addition or exposure of functional groups to the substrate through oxidation, reduction, and hydroxylation to elevate the substrate’s solubility in water [73]. There are also important involvements of COX6c in gene rearrangement during tumorigenesis and in the development of drug resistance in cancer cells. Therefore, the expression pattern of COX6c can be chosen as a useful marker of the OXPHOS pathway to study the alteration of energy metabolism and to help the prediction, diagnosis, treatment, and prognosis of various diseases.
Irradiation appears to induce a regularizing balance in cancer cell function and decrease lymph metastases. Notably, COX6c is also associated with ionizing irradiation, as shown by the considerably increased levels of COX6c found in human esophageal cancer cells following radiation exposure [74]. In another study, low-intensity laser irradiation enhanced cellular activity and boosted ATP generation in a human skin fibroblast cell type. COX6c is increased in irradiation injured and diabetic wounded cells, promoting wound repair [75]. This effect is achieved by increases in DNA, RNA, and thus protein synthesis, which is supported by an increase in ATP. Thus, COX6c modulated in the early response to irradiation may be helpful for understanding the molecular basis of radiotherapy. More work is required to elucidate the role of COX6c targeted by irradiation before developing strategies to augment its beneficial effects or establishing novel alternative adjuvant therapies.
In addition, other targeting strategies, for example, COX6c knockout or inhibitors may provide a key role in disease treatment, but there are currently no studies to support this strategy. Cyclooxygenase-2(COX-2) is also an important target in cardiovascular disease and plays a similar role to COX6c in inflammation-related diseases. Nonsteroidal anti-inflammatory drugs (NSAIDs) that are non-selective inhibitors of COX-2 inhibit inflammation while reducing angiogenesis and form a certain anti-cancer role [76]. However, when NSAIDs inhibit COX-2, they will destroy COX-1 which maintains the physiological function of cells, tissues, and organs [77, 78]. NSAIDs provide an important direction for the development of suitable inhibitors targeting COX6c for the treatment of cardiovascular diseases.
As was already established, aberrant COX6c expression is seen in various disorders, including familial hypercholesterolemia, chronic renal disease, diabetes, breast cancer, prostate cancer, uterine fibroids, follicular thyroid cancer, melanoma tissue [43, 67, 71], suggesting its potential as a treatment target. However, the lack of elucidation of COX6c and its molecular regulation mode has limited clinical treatment. In addition, conventional treatment drugs have pharmacokinetic limitations, and the efficacy is not obvious. It is thought-provoking that some studies have used nanomaterials to encapsulate drugs to target intracellular proteins to achieve the purpose of treating diseases, which has the advantages of high targeting and high circulation efficiency. For example, solid lipid nanoparticles are solid lipid-based nano-drug delivery vehicles with good physical stability and tolerance and can protect drugs from degradation and control drug release at non-targeted sites [79].
In addition, a new bioactive component—vesicle-like nanoparticles—in honey (H-VLNs) are membrane-bound nano-scale particles that contain lipids, proteins, and small-sized RNAs. H-VLNs impede the formation and activation of the nucleotide-binding domain and leucine-rich repeat-related (NLR) family, pyrin domain containing 3 (NLRP3) inflammasome, which is a crucial inflammatory signal platform in the innate immune system [80]. This study provides an important idea for the treatment of inflammation-related diseases by targeting COX6c with nano-drug delivery vehicles.
However, much of this evidence is in preclinical testing, and many problems remain unresolved. For instance, to what extent are the activity and expression of COX6c altered in various diseases? Is the change in COX6c a secondary event associated with other pathways that might be the primary drivers of the functional outcomes or is it specific to the disease? What are the specific mechanisms and processes that COX6c participates in during the development of disease? Is the expression pattern of COX6c the same in diverse tissues and cells? How do one set standards for evaluating the COX6c level to predict and diagnose disease? In conclusion, further work regarding the roles of COX6c in various diseases will promote it as a potential biomarker for the prediction, diagnosis, treatment, and prognosis of various diseases.
References
Sazanov LA. A giant molecular proton pump: structure and mechanism of respiratory complex I. Nat Rev Mol Cell Biol. 2015;16:375–88.
Hofmann S, Lichtner P, Schuffenhauer S, Gerbitz KD, Meitinger T. Assignment of the human genes coding for cytochrome c oxidase subunits Va (COX5A), VIc (COX6C) and VIIc (COX7C) to chromosome bands 15q25, 8q22->q23 and 5q14 and of three pseudogenes (COX5AP1, COX6CP1, COX7CP1) to 14q22, 16p12 and 13q14->q21 by FISH and radiation hybrid mapping. Cytogenet Cell Genet. 1998;83:226–7.
Lenka N, Vijayasarathy C, Mullick J, Avadhani NG. Structural organization and transcription regulation of nuclear genes encoding the mammalian cytochrome c oxidase complex. Prog Nucleic Acid Res Mol Biol. 1998;61:309–44.
Rehling P, Brandner K, Pfanner N. Mitochondrial import and the twin-pore translocase. Nat Rev Mol Cell Biol. 2004;5:519–30.
Szuplewski S, Terracol R. The cyclope gene of Drosophila encodes a cytochrome c oxidase subunit VIc homolog. Genetics 2001;158:1629–43.
Kadenbach B, Huttemann M. The subunit composition and function of mammalian cytochrome c oxidase. Mitochondrion 2015;24:64–76.
Timon-Gomez A, Nyvltova E, Abriata LA, Vila AJ, Hosler J, Barrientos A. Mitochondrial cytochrome c oxidase biogenesis: recent developments. Semin Cell Dev Biol. 2018;76:163–78.
Otsuka M, Mizuno Y, Yoshida M, Kagawa Y, Ohta S. Nucleotide sequence of cDNA encoding human cytochrome c oxidase subunit VIc. Nucleic Acids Res. 1988;16:10916.
Sasaki K, Ono M, Takabe K, Suzuki A, Kurihara Y. Specific intron-dependent loading of DAZAP1 onto the cox6c transcript suppresses pre-mRNA splicing efficacy and induces cell growth retardation. Gene 2018;657:1–8.
Lu J, Wang K, Rodova M, Esteves R, Berry D, E L, et al. Polymorphic variation in cytochrome oxidase subunit genes. J Alzheimers Dis. 2010;21:141–54.
Little AG, Kocha KM, Lougheed SC, Moyes CD. Evolution of the nuclear-encoded cytochrome oxidase subunits in vertebrates. Physiol Genom. 2010;42:76–84.
Schmidt TR, Goodman M, Grossman LI. Amino acid replacement is rapid in primates for the mature polypeptides of COX subunits, but not for their targeting presequences. Gene. 2002;286:13–9.
Lazarou M, Smith SM, Thorburn DR, Ryan MT, McKenzie M. Assembly of nuclear DNA-encoded subunits into mitochondrial complex IV, and their preferential integration into supercomplex forms in patient mitochondria. FEBS J. 2009;276:6701–13.
Sheehan TE, Kumar PA, Hood DA. Tissue-specific regulation of cytochrome c oxidase subunit expression by thyroid hormone. Am J Physiol Endocrinol Metab. 2004;286:E968–74.
Duggan AT, Kocha KM, Monk CT, Bremer K, Moyes CD. Coordination of cytochrome c oxidase gene expression in the remodelling of skeletal muscle. J Exp Biol. 2011;214:1880–7.
Ruiz-Pesini E, Diez C, Lapena AC, Perez-Martos A, Montoya J, Alvarez E, et al. Correlation of sperm motility with mitochondrial enzymatic activities. Clin Chem. 1998;44:1616–20.
Ruiz-Pesini E, Lapena AC, Diez-Sanchez C, Perez-Martos A, Montoya J, Alvarez E, et al. Human mtDNA haplogroups associated with high or reduced spermatozoa motility. Am J Hum Genet. 2000;67:682–96.
Amaral A, Ramalho-Santos J, St John JC. The expression of polymerase gamma and mitochondrial transcription factor A and the regulation of mitochondrial DNA content in mature human sperm. Hum Reprod. 2007;22:1585–96.
Hadife N, Nemos C, Frippiat JP, Hamade T, Perrot A, Dalloul A. Interleukin-24 mediates apoptosis in human B-cells through early activation of cell cycle arrest followed by late induction of the mitochondrial apoptosis pathway. Leuk Lymphoma. 2013;54:587–97.
Assaily W, Benchimol S. Differential utilization of two ATP-generating pathways is regulated by p53. Cancer Cell. 2006;10:4–6.
Matoba S, Kang JG, Patino WD, Wragg A, Boehm M, Gavrilova O, et al. p53 regulates mitochondrial respiration. Science 2006;312:1650–3.
Fontanesi F, Soto IC, Horn D, Barrientos A. Assembly of mitochondrial cytochrome c-oxidase, a complicated and highly regulated cellular process. Am J Physiol Cell Physiol. 2006;291:C1129–47.
Wright RM, Rosenzweig B, Poyton RO. Organization and expression of the COX6 genetic locus in Saccharomyces cerevisiae: multiple mRNAs with different 3’ termini are transcribed from COX6 and regulated differentially. Nucleic Acids Res. 1989;17:1103–20.
Othumpangat S, Noti JD, Beezhold DH. Lung epithelial cells resist influenza A infection by inducing the expression of cytochrome c oxidase VIc which is modulated by miRNA 4276. Virology. 2014;468-470:256–64.
Elsayed ET, Nassra RA, Naga YS. Peroxisome proliferator-activated receptor-gamma-coactivator 1alpha (PGC-1alpha) gene expression in chronic kidney disease patients on hemodialysis: relation to hemodialysis-related cardiovascular morbidity and mortality. Int Urol Nephrol. 2017;49:1835–44.
Di W, Lv J, Jiang S, Lu C, Yang Z, Ma Z, et al. PGC-1: the energetic regulator in cardiac metabolism. Curr Issues Mol Biol. 2018;28:29–46.
Evans MJ, Scarpulla RC. NRF-1: a trans-activator of nuclear-encoded respiratory genes in animal cells. Genes Dev. 1990;4:1023–34.
Hashad D, Elgohry I, Dwedar F. Nuclear respiratory factor-1 (NRF-1) gene expression in chronic kidney disease patients undergoing hemodialysis and mitochondrial oxidative dysregulation. Clin Lab. 2016;62:2149–54.
Li M, Vascotto C, Xu S, Dai N, Qing Y, Zhong Z, et al. Human AP endonuclease/redox factor APE1/ref-1 modulates mitochondrial function after oxidative stress by regulating the transcriptional activity of NRF1. Free Radic Biol Med. 2012;53:237–48.
Dhar SS, Ongwijitwat S, Wong-Riley MT. Nuclear respiratory factor 1 regulates all ten nuclear-encoded subunits of cytochrome c oxidase in neurons. J Biol Chem. 2008;283:3120–9.
Ongwijitwat S, Liang HL, Graboyes EM, Wong-Riley MT. Nuclear respiratory factor 2 senses changing cellular energy demands and its silencing down-regulates cytochrome oxidase and other target gene mRNAs. Gene 2006;374:39–49.
Ongwijitwat S, Wong-Riley MT. Is nuclear respiratory factor 2 a master transcriptional coordinator for all ten nuclear-encoded cytochrome c oxidase subunits in neurons? Gene 2005;360:65–77.
Cunningham KF, Beeson GC, Beeson CC, Baicu CF, Zile MR, McDermott PJ. Estrogen-Related Receptor alpha (ERRalpha) is required for adaptive increases in PGC-1 isoform expression during electrically stimulated contraction of adult cardiomyocytes in sustained hypoxic conditions. Int J Cardiol. 2015;187:393–400.
Dufour CR, Wilson BJ, Huss JM, Kelly DP, Alaynick WA, Downes M, et al. Genome-wide orchestration of cardiac functions by the orphan nuclear receptors ERRalpha and gamma. Cell Metab. 2007;5:345–56.
Romani B, Engelbrecht S, Glashoff RH. Functions of Tat: the versatile protein of human immunodeficiency virus type 1. J Gen Virol. 2010;91:1–12.
Lecoeur H, Borgne-Sanchez A, Chaloin O, El-Khoury R, Brabant M, Langonne A, et al. HIV-1 Tat protein directly induces mitochondrial membrane permeabilization and inactivates cytochrome c oxidase. Cell Death Dis. 2012;3:e282.
Hsu LC, Chen HY, Lin YW, Chu WC, Lin MJ, Yan YT, et al. DAZAP1, an hnRNP protein, is required for normal growth and spermatogenesis in mice. RNA. 2008;14:1814–22.
Smith RW, Anderson RC, Smith JW, Brook M, Richardson WA, Gray NK. DAZAP1, an RNA-binding protein required for development and spermatogenesis, can regulate mRNA translation. RNA 2011;17:1282–95.
Tall AR, Yvan-Charvet L. Cholesterol, inflammation and innate immunity. Nat Rev Immunol. 2015;15:104–16.
Wang HX, Zhao YX. Prediction of genetic risk factors of atherosclerosis using various bioinformatic tools. Genet Mol Res. 2016;15:gmr7347.
Scherz-Shouval R, Elazar Z. ROS, mitochondria and the regulation of autophagy. Trends Cell Biol. 2007;17:422–7.
Ye X, Zeng T, Kong W, Chen LL. Integrative analyses of genes associated with fulminant type 1 diabetes. J Immunol Res. 2020;2020:1025857.
Zhu X, Qiu Z, Ouyang W, Miao J, Xiong P, Mao D, et al. Hepatic transcriptome and proteome analyses provide new insights into the regulator mechanism of dietary avicularin in diabetic mice. Food Res Int. 2019;125:108570.
Zhang Q, Xiao X, Li M, Li W, Yu M, Zhang H, et al. Gene expression profiling in glomeruli of diabetic nephropathy rat. Exp Biol Med. 2012;237:903–11.
Zhang Q, Xiao X, Li M, Li W, Yu M, Zhang H, et al. Telmisartan improves kidney function through inhibition of the oxidative phosphorylation pathway in diabetic rats. J Mol Endocrinol. 2012;49:35–46.
Lv J, Jiang S, Yang Z, Hu W, Wang Z, Li T, et al. PGC-1alpha sparks the fire of neuroprotection against neurodegenerative disorders. Ageing Res Rev. 2018;44:8–21.
Imayama I, Ichiki T, Inanaga K, Ohtsubo H, Fukuyama K, Ono H, et al. Telmisartan downregulates angiotensin II type 1 receptor through activation of peroxisome proliferator-activated receptor gamma. Cardiovasc Res. 2006;72:184–90.
Zaza G, Granata S, Masola V, Rugiu C, Fantin F, Gesualdo L, et al. Downregulation of nuclear-encoded genes of oxidative metabolism in dialyzed chronic kidney disease patients. PLoS ONE. 2013;8:e77847.
Jiang S, Deng C, Lv J, Fan C, Hu W, Di S, et al. Nrf2 weaves an elaborate network of neuroprotection against stroke. Mol Neurobiol. 2017;54:1440–55.
Kish SJ, Mastrogiacomo F, Guttman M, Furukawa Y, Taanman JW, Dozic S, et al. Decreased brain protein levels of cytochrome oxidase subunits in Alzheimer’s disease and in hereditary spinocerebellar ataxia disorders: a nonspecific change? J Neurochem. 1999;72:700–7.
Llombart V, Trejo SA, Bronsoms S, Morancho A, Feifei M, Faura J, et al. Profiling and identification of new proteins involved in brain ischemia using MALDI-imaging-mass-spectrometry. J Proteom. 2017;152:243–53.
Du QX, Wang XW, Zhang L, Li SQ, Gao CR, Wang YY, et al. Relative expression of indicators for wound age estimation in forensic pathology. Fa Yi Xue Za Zhi. 2015;31:81–4.
Sun JH, Zhang L, Wang XW, Du QX, Lu J, Wang YY. [Relation between injury time and the expression of COX6C mRNA in skeletal muscle of rats after contusion]. Fa Yi Xue Za Zhi. 2015;31:177–80.
Swallow EB, Reyes D, Hopkinson NS, Man WD, Porcher R, Cetti EJ, et al. Quadriceps strength predicts mortality in patients with moderate to severe chronic obstructive pulmonary disease. Thorax. 2007;62:115–20.
Crul T, Testelmans D, Spruit MA, Troosters T, Gosselink R, Geeraerts I, et al. Gene expression profiling in vastus lateralis muscle during an acute exacerbation of COPD. Cell Physiol Biochem. 2010;25:491–500.
Wang FL, Wang Y, Wong WK, Liu Y, Addivinola FJ, Liang P, et al. Two differentially expressed genes in normal human prostate tissue and in carcinoma. Cancer Res. 1996;56:3634–7.
Marberger H, Marberger E, Mann T, Lutwak-Mann C. Citric acid in human prostatic secretion and metastasizing cancer of prostate gland. Br Med J. 1962;1:835–6.
Kim J, Lee J, Chang E, Suh K, Lee C, Jee J, et al. Prognostic factors in patients with stage ii/iii breast cancer treated with adjuvant extension of neoadjuvant chemotherapy: A Retrospective Cohort Study with Ten-Years of Follow-Up Data. J Breast Cancer. 2011;14:39–45.
Lage H. An overview of cancer multidrug resistance: a still unsolved problem. Cell Mol Life Sci. 2008;65:3145–67.
Chang FW, Fan HC, Liu JM, Fan TP, Jing J, Yang CL, et al. Estrogen enhances the expression of the multidrug transporter gene ABCG2-increasing drug resistance of breast cancer cells through estrogen receptors. Int J Mol Sci. 2017;18:163.
Luparello C, Romanotto R, Tipa A, Sirchia R, Olmo N, Lopez de Silanes I, et al. Midregion parathyroid hormone-related protein inhibits growth and invasion in vitro and tumorigenesis in vivo of human breast cancer cells. J Bone Min Res. 2001;16:2173–81.
Sirchia R, Priulla M, Sciandrello G, Caradonna F, Barbata G, Luparello C. Mid-region parathyroid hormone-related protein (PTHrP) binds chromatin of MDA-MB231 breast cancer cells and isolated oligonucleotides “in vitro”. Breast Cancer Res Treat. 2007;105:105–16.
Sirchia R, Luparello C. Mid-region parathyroid hormone-related protein (PTHrP) and gene expression of MDA-MB231 breast cancer cells. Biol Chem. 2007;388:457–65.
West M, Blanchette C, Dressman H, Huang E, Ishida S, Spang R, et al. Predicting the clinical status of human breast cancer by using gene expression profiles. Proc Natl Acad Sci USA. 2001;98:11462–7.
Emerson JW, Dolled-Filhart M, Harris L, Rimm DL, Tuck DP. Quantitative assessment of tissue biomarkers and construction of a model to predict outcome in breast cancer using multiple imputation. Cancer Inf. 2009;7:29–40.
Swierniak M, Pfeifer A, Stokowy T, Rusinek D, Chekan M, Lange D, et al. Somatic mutation profiling of follicular thyroid cancer by next generation sequencing. Mol Cell Endocrinol. 2016;433:130–7.
Kurose K, Mine N, Doi D, Ota Y, Yoneyama K, Konishi H, et al. Novel gene fusion of COX6C at 8q22-23 to HMGIC at 12q15 in a uterine leiomyoma. Genes Chromosomes Cancer. 2000;27:303–7.
Mine N, Kurose K, Nagai H, Doi D, Ota Y, Yoneyama K, et al. Gene fusion involving HMGIC is a frequent aberration in uterine leiomyomas. J Hum Genet. 2001;46:408–12.
Foa C, Mainguene C, Dupre F, Coindre JM, Huguet C, Kober C, et al. Rearrangement involving chromosomes 1 and 8 in a retroperitoneal lipoma. Cancer Genet Cytogenet. 2002;133:156–9.
Cai YX, Yang X, Lin S, Xu YW, Zhu SW, Fan DM, et al. Low-coverage sequencing of urine sediment DNA for detection of copy number aberrations in bladder cancer. Cancer Manag Res. 2021;13:1943–53.
Jang SC, Crescitelli R, Cvjetkovic A, Belgrano V, Olofsson Bagge R, Sundfeldt K, et al. Mitochondrial protein enriched extracellular vesicles discovered in human melanoma tissues can be detected in patient plasma. J Extracell Vesicles. 2019;8:1635420.
Li L, Li Y, Huang Y, Ouyang Y, Zhu Y, Wang Y, et al. Long non-coding RNA MIF-AS1 promotes gastric cancer cell proliferation and reduces apoptosis to upregulate NDUFA4. Cancer Sci. 2018;109:3714–25.
Drag M, Skinkyte-Juskiene R, Do DN, Kogelman LJA, Kadarmideen HN. Differential expression and co-expression gene networks reveal candidate biomarkers of boar taint in non-castrated pigs. Sci Rep. 2017;7:12205.
Bo H, Ghazizadeh M, Shimizu H, Kurihara Y, Egawa S, Moriyama Y, et al. Effect of ionizing irradiation on human esophageal cancer cell lines by cDNA microarray gene expression analysis. J Nippon Med Sch. 2004;71:172–80.
Masha RT, Houreld NN, Abrahamse H. Low-intensity laser irradiation at 660 nm stimulates transcription of genes involved in the electron transport chain. Photomed Laser Surg. 2013;31:47–53.
Davies G, Martin LA, Sacks N, Dowsett M. Cyclooxygenase-2 (COX-2), aromatase and breast cancer: a possible role for COX-2 inhibitors in breast cancer chemoprevention. Ann Oncol. 2002;13:669–78.
Zhu L, Xu C, Huo X, Hao H, Wan Q, Chen H, et al. The cyclooxygenase-1/mPGES-1/endothelial prostaglandin EP4 receptor pathway constrains myocardial ischemia-reperfusion injury. Nat Commun. 2019;10:1888.
Marnett LJ. The COXIB experience: a look in the rearview mirror. Annu Rev Pharm Toxicol. 2009;49:265–90.
Mulik R, Mahadik K, Paradkar A. Development of curcuminoids loaded poly(butyl) cyanoacrylate nanoparticles: physicochemical characterization and stability study. Eur J Pharm Sci. 2009;37:395–404.
Chen X, Liu B, Li X, An TT, Zhou Y, Li G, et al. Identification of anti-inflammatory vesicle-like nanoparticles in honey. J Extracell Vesicles. 2021;10:e12069.
Funding
This work was supported by the National Natural Science Foundation of China (81871607 and 82070422), Youth Science and Technology Rising Star Project of Shaanxi Province (2020KJXX-036), Key Research and Development Program of Shaanxi (2020ZDLSF04-03), Major Research Projects of Xi'an Science and Technology Plan (201805104YX12SF38(2)), and Innovation Capability Strong Foundation Plan of Xi’an City (Medical Research Project, 21YXYJ0037).
Author information
Authors and Affiliations
Contributions
CYW, JJL, XCX, JWL, YQL, DNX, and YTJ performed substantial contributions to conception, design, and assembled the figure. SJ, MHZ, YY, and SFZ drafted the article or revised it critically for important intellectual content and approved the final version to be published.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, C., Lv, J., Xue, C. et al. Novel role of COX6c in the regulation of oxidative phosphorylation and diseases. Cell Death Discov. 8, 336 (2022). https://doi.org/10.1038/s41420-022-01130-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-01130-1
This article is cited by
-
Superresolved spatial transcriptomics transferred from a histological context
Applied Intelligence (2023)