Abstract
Mitochondrial dysfunction is a frequent participant in common diseases and a principal suspect in aging. To combat mitochondrial dysfunction, eukaryotes have evolved a large repertoire of quality control mechanisms. One such mechanism involves the selective degradation of damaged or misfolded mitochondrial proteins by mitochondrial resident proteases, including proteases of the ATPase Associated with diverse cellular Activities (AAA+) family. The importance of the AAA+ family of mitochondrial proteases is exemplified by the fact that mutations that impair their functions cause a variety of human diseases, yet our knowledge of the cellular responses to their inactivation is limited. To address this matter, we created and characterized flies with complete or partial inactivation of the Drosophila matrix-localized AAA+ protease Lon. We found that a Lon null allele confers early larval lethality and that severely reducing Lon expression using RNAi results in shortened lifespan, locomotor impairment, and respiratory defects specific to respiratory chain complexes that contain mitochondrially encoded subunits. The respiratory chain defects of Lon knockdown (LonKD) flies appeared to result from severely reduced translation of mitochondrially encoded genes. This translational defect was not a consequence of reduced mitochondrial transcription, as evidenced by the fact that mitochondrial transcripts were elevated in abundance in LonKD flies. Rather, the translational defect of LonKD flies appeared to be derived from sequestration of mitochondrially encoded transcripts in highly dense ribonucleoparticles. The translational defect of LonKD flies was also accompanied by a substantial increase in unfolded mitochondrial proteins. Together, our findings suggest that the accumulation of unfolded mitochondrial proteins triggers a stress response that culminates in the inhibition of mitochondrial translation. Our work provides a foundation to explore the underlying molecular mechanisms.
Similar content being viewed by others
Introduction
Mitochondria are responsible for most of the energy produced by a cell, but the generation of reactive oxygen species (ROS) as a byproduct of this activity can damage mitochondrial proteins, lipids, and DNA1,2,3. Also, while mitochondria contain their own genome, most mitochondrial proteins are encoded in the nucleus, and a stoichiometric imbalance between mitochondrial and nuclear encoded respiratory chain subunits can cause misfolding and aggregation of the unassembled proteins4,5. Fortunately, there are many surveillance pathways that oppose or reverse this damage, including the AAA+ family of mitochondrial proteases6,7,8. All of the AAA+ proteases form multimeric protein complexes and use ATP to unfold and transport substrates to an internal proteolytic cavity for degradation. In higher eukaryotes, there are five major mitochondrial AAA+ proteases that are distinguished by their subunit composition and mitochondrial localization. The most well-studied member of the AAA+ protease family is Lon.
Previous work indicates that Lon possesses three different activities, serving as a chaperone and DNA-binding protein in addition to its role in proteolysis9,10. Lon expression is regulated by multiple cellular stresses, including ROS and unfolded protein stress, and there is substantial support for the role of Lon in the degradation of oxidatively damaged and misfolded proteins11,12. Although few Lon substrates are known with certainty, a number of candidate Lon substrates have been identified from biochemical studies aimed at the identification of Lon-binding proteins, including the mitochondrial DNA (mtDNA) replication factors Twinkle, polymerase gamma, Tfam, and the chaperones Hsp60 and mtHsp7013,14,15,16,17,18,19. Subsequent studies aimed at validating the significance of these binding interactions indicated that Lon inactivation results in increased mtDNA copy number and destabilization of Hsp60 and mtHsp70 under environmental stress20,21,22. However, it is unclear whether these findings reflect direct effects of Lon inactivation, or downstream cellular responses to loss of Lon activity. Moreover, mutations in Lon have been shown to result in the recessive developmental disorder CODAS (cerebral, ocular, dental, auricular, and skeletal) syndrome, yet the mechanisms by which mutations in Lon cause this disease are currently unknown23.
To create a simple, genetically tractable model system to explore the biological role of Lon and the pathological consequences of Lon inactivation, we used CRISPR/Cas9-mediated gene targeting and RNAi to create Drosophila strains with complete and partial loss of Lon function. We found that Lon is an essential gene in Drosophila and that flies expressing RNAi against Lon (Lon knockdown flies, or LonKD flies) had shortened lifespan, defective locomotion, and altered respiratory chain activity. The respiratory chain deficits in LonKD flies were specific to respiratory chain complexes that contain subunits encoded by the mitochondrial genome, suggesting that altered expression of mitochondrially encoded components underlies this defect. Consistent with this conclusion, we found that reduced mitochondrial complex activity is accompanied by reduced complex abundance and diminished mitochondrial translation. The translational defect of LonKD flies was not a consequence of reduced mitochondrial transcription, but rather appeared to be a consequence of reduced association of mitochondrial transcripts on mitochondrial ribosomes and packaging of mitochondrial transcripts into highly dense ribonucleoparticles. The translational defect of LonKD flies was also accompanied by elevated abundance of unfolded mitochondrial proteins, and overexpression of another matrix protease, ClpP, partially rescued the defects associated with Lon inactivation, possibly by reducing the burden of unfolded mitochondrial proteins. Together, our findings strongly suggest that Lon inactivation results in the activation of an unfolded protein stress response that inhibits mitochondrial translation.
Results
Creation of Lon-deficient Drosophila strains
To explore the biological roles of Lon protease we used CRISPR/Cas9 technology to create a null allele of the Drosophila Lon gene (CG8798). Briefly, we constructed guide RNAs designed to create double-strand breaks flanking the Lon coding sequence (Suppl. Figure 1). We also created a donor vector construct consisting of the DsRed marker flanked by 5′ and 3′ untranslated sequences from the Lon coding region for use in homology-directed recombinational repair of the double-strand breaks. Flies expressing the DsRed marker were then subjected to whole-genome sequencing to verify correct targeting of DsRed to the Lon locus and complete deletion of the Lon gene. Flies heterozygous for this deletion were fully viable with no detectable phenotypes. However, homozygotes died at the second instar larval stage of development, demonstrating that Lon is essential for viability (Suppl. Figure 2a).
RNAi often reduces but does not completely eliminate target gene expression. Thus, we tested whether RNAi lines targeting Lon would circumvent the lethality conferred by a null mutation in Lon. We tested two different RNAi constructs targeting Lon (designated Lon-RNAi-1 and Lon-RNAi-2) that we used in previously published work to explore the influence of Lon on the abundance and activity of the mitophagy factor PINK124. Our previous work demonstrated that these RNAi lines differed in the efficiency by which they reduced Lon expression, with the Lon-RNAi-2 line resulting in greater reduction in Lon expression. We expressed each RNAi line using two GAL4 drivers simultaneously: the pan-neuronal elav-GAL4 driver and the ubiquitous da-GAL4 driver. Driving the stronger Lon-RNAi-2 line with this combination of GAL4 drivers failed to yield viable adult flies, but flies expressing the weaker Lon-RNAi-1 transgene were fully viable and fertile as young adults, and had no obvious morphological alterations. Western blot analyses performed on heads and whole flies indicated that Lon expression was significantly reduced in flies bearing the UAS-Lon-RNAi-1, elav-GAL4, and da-GAL4 transgenes compared to controls expressing RNAi against the exogenous mCherry sequence (UAS-mCherry-RNAi) (Fig. 1a and Suppl. Figure 2b). Flies bearing the Lon-RNAi-1, elav-GAL4, and da-GAL4 transgenes were used in most of the remaining studies and will hereafter be called LonKD.
a Immunoblot analysis from heads of 1-day-old control and LonKD flies using Lon and actin antibodies. The Lon band intensity is normalized against actin as a loading control. Significance was determined using Student’s t-test (***p < 0.0005 by Student’s t-test). The experiment was repeated at least three times. b Kaplan–Meier survival curves of LonKD flies (n = 167, 50% survival 31 days) and controls (n = 174, 50% survival 73 days) (****p < 0.0001 by Mantel–Cox log-rank test). c One-day-old LonKD flies exhibit a climbing defect. Error bars represent SEM (n = 135 for LonKD, 134 for control, ***p < 0.0005 by Student’s t-test). d One-day-old LonKD flies exhibit a significant decrease in flight index (n = 6 independent groups of 10–15 animals, **p < 0.005 from Student’s t-test). Control = UAS-mCherry-RNAi driven by elav-GAL4 and da-GAL4. LonKD = UAS-Lon-RNAi-1 driven by elav-GAL4 and da-GAL4
Partial inactivation of Lon results in shortened lifespan, locomotor defects, and altered respiratory chain activity
To further characterize the LonKD phenotypes, we subjected the flies to several standard Drosophila behavioral assays. Although LonKD flies appeared normal upon eclosion, they were significantly shorter-lived than the control flies (Fig. 1b). The maximal and median lifespans of LonKD flies were similar to those of flies bearing null mutations in genes encoding the other AAA+ mitochondrial protease family members dYME1L and SPG725,26. Young LonKD flies also exhibited defects in climbing and flight starting early in life (Fig. 1c, d).
Inactivation of Lon in other model systems results in reduced respiratory chain activity27. Thus, we tested whether LonKD flies exhibit similar respiratory chain defects, using established biochemical assays to quantify respiratory chain activity in young (1 day) and old (3 weeks) LonKD flies. We found that respiratory chain complexes I and IV had reduced activity in old LonKD flies compared to the controls, while young flies showed reduction only in complex I activity (Fig. 2a). However, complex II activity was increased significantly in both young and old LonKD flies. These alterations could result from a functional change in respiratory chain activity or from altered respiratory chain abundance. To distinguish between these possibilities while simultaneously confirming and extending our biochemical studies, we assessed the abundance of assembled complexes and their corresponding activities using blue native PAGE (BN-PAGE) and in-gel enzyme activity assays28. This work revealed that complexes I, III, and IV all exhibited both reduced activity and reduced abundance in LonKD flies, whereas complex II exhibited an increase in activity (Fig. 2b–d). Additionally, this work revealed the presence of a partially assembled but catalytically active F1 subunit–containing subcomplex of ATP synthase (complex V) (Fig. 2b, e). These alterations were also accompanied by a reduction in ATP content in LonKD flies (Suppl. Figure 3). Together, our findings indicate that partial inactivation of Lon results in a progressive age-dependent decline in oxidative phosphorylation capacity, and that this decline may underlie the shortened lifespan and behavioral deficits of LonKD flies.
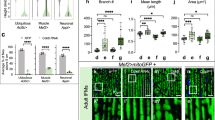
a The activity of respiratory chain complexes in control and LonKD flies at 1 day (left panel) and 21 days (right panel) of age (n = 2 independent groups of 500 adult flies, *p < 0.05, **p < 0.005 from Student’s t-test). b BN-PAGE analysis of mitochondrial protein extracts from 21-day-old control and LonKD flies. The red asterisk marks the location of the subcomplex containing F1 subunit of ATP synthase that is only detected in LonKD flies. Immunoblot of citrate synthase (bottom panel) was used as a loading control. c In-gel activity of mitochondrial complexes I and IV isolated from 21-day-old adults. SCs here refer to the supercomplexes. d In-gel activity of mitochondrial complex II isolated from 21-day-old flies. e In-gel activity of mitochondrial complex V isolated from 21-day-old adults. The red asterisk denotes the location of the subcomplex containing F1 subunit of ATP synthase in LonKD flies. Note that the protein molecular weight markers shown in (b) were used as reference to mark the gels in (c–e). For BN-PAGE and in-gel activity assays, the images shown are representative of two independent biological replicates. Control = UAS-mCherry-RNAi driven by elav-GAL4 and da-GAL4. LonKD = UAS-Lon-RNAi-1 driven by elav-GAL4 and da-GAL4
Lon knockdown results in increased mtDNA-encoded transcript abundance and reduced translation
All of the respiratory chain complexes exhibiting reduced abundance and assembly in LonKD flies (complexes I, III, IV, and V) contain one or more subunits encoded by mtDNA. Additionally, our BN-PAGE and in-gel activity assays in LonKD flies indicated that the ATP synthase F1 subunit containing subcomplex (consisting entirely of subunits encoded by nuclear DNA) accumulated at the expense of the FO subcomplex (which includes two subunits encoded by mtDNA). By contrast, complex II, which exhibited increased activity in LonKD flies, consists entirely of nuclear DNA–encoded components. These findings led us to hypothesize that reduced mtDNA abundance, reduced transcription of mtDNA-encoded subunits, and/or reduced translation of mitochondrial transcripts might account for the decreased expression of complexes I, III, IV, and V.
To begin to explore these hypotheses, we first compared mtDNA abundance in LonKD flies and the controls. While increased mtDNA abundance in response to Lon inactivation has been reported20, mtDNA copy number was unchanged in LonKD flies (Fig. 3a). This is consistent with our previous finding of unchanged mtDNA abundance when Lon was knocked down with elav-GAL4 alone24. We next analyzed the steady-state levels of several mitochondrial transcripts by qRT-PCR. We found that levels of all transcripts analyzed, including 12S and 16S rRNA and several mRNAs, were increased in old LonKD flies (Fig. 3b). Additionally, the transcript abundance of the mitochondrial transcription-promoting factors TFAM and mtTFB2 was unchanged in LonKD flies (Suppl. Figure 4). Together, these findings indicate that the respiratory chain defects in LonKD flies do not derive from reduced mitochondrial gene dosage or reduced mitochondrial transcription.
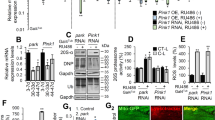
a Mitochondrial DNA abundance was compared in LonKD and control flies using qPCR to compare the ratio of mtDNA-encoded mt:Cyt-b to that of nuclear-encoded Act79b (n = 3 independent groups of 40–45 fly heads). Statistical significance was determined using Student’s t-test. b qRT-PCR was used to quantify steady-state abundance of the indicated mitochondrial RNAs in 21-day-old adult fly heads. Mitochondrial RNA abundance was normalized to the abundance of the nuclear-encoded Act79b transcript (n = 3 independent groups of 40–45 fly heads). Error bars indicate mean ± SEM. Student’s t-test was applied, *p < 0.05, **p < 0.005, ***p < 0.0005. c Immunoblot analysis of third instar larvae to confirm knockdown of Lon using actin as a loading control. n = 3 independent groups of five third instar larvae. Significance was determined using Student’s t-test, **p < 0.005. d In organello translation was performed using mitochondria isolated from third instar larvae. Mitochondria were labeled by incubating with 35S-methionine for 1 h. Positions of individual mitochondrially encoded proteins are indicated (left panel). Coomassie-stained gel (right panel) was used as a loading control. Control = UAS-mCherry-RNAi driven by elav-GAL4 and da-GAL4. LonKD = UAS-Lon-RNAi-1 driven by elav-GAL4 and da-GAL4
To test whether LonKD flies manifest a translational defect, we used 35S-labeled methionine to perform in organello labeling of mitochondria prepared from adult flies. However, we detected very little labeling of mitochondrial proteins in our control flies (Suppl. Figure 5). This finding may reflect the fact that mitochondrial proteins are extremely long-lived29 and that there is thus a decreased need for mitochondrial translation during the adult stage of development. However, previous work has detected robust labeling of mitochondrial proteins using mitochondria obtained from larvae, so we repeated our in organello labeling studies using mitochondria from third instar larvae30. Our western blot analysis confirmed that Lon expression was greatly reduced in LonKD larvae relative to the controls (Fig. 3c). This experiment revealed a substantial decrease in de novo labeling of mitochondrial translation products in LonKD larvae relative to the controls (Fig. 3d), suggesting that the decreased abundance of complexes I, III, IV, and V in LonKD flies is a direct consequence of a translational defect. The increased abundance of mtDNA-encoded transcripts and nuclear DNA–encoded complex II may represent compensatory responses to alleviate this translational deficiency31,32,33.
Mitochondrial transcripts are packaged into untranslated particles in LonKD flies
To investigate the mechanism by which inactivation of Lon impairs mitochondrial translation, we assessed the state of mitochondrial ribosome assembly by performing sucrose density gradient sedimentation analyses. Mitochondrial extracts from LonKD flies and controls were subjected to sucrose density gradient analysis as previously described30. Fractions from the sucrose gradient were then subjected to qRT-PCR to quantify the distribution of the 12S and 16S mitochondrial ribosomal RNAs, which mark the small (28S) and large (39S) mitochondrial ribosomal subunits, respectively. Co-localization of the 12S and 16S mitochondrial ribosomal RNAs within the same fraction is diagnostic for fully assembled and actively translating mitochondrial ribosomes (55S). This analysis revealed that the abundance of 12S and 16S ribosomal RNAs in fully assembled mitochondrial ribosome fractions (fraction 15–17) was decreased by 5.8% and 6.4%, respectively, in LonKD flies relative to the controls (Fig. 4).
Mitochondrial lysates from 21-day-old control and LonKD flies were subjected to sucrose density gradient fractionation to assess the state of assembly of mitochondrial ribosomes. The relative proportions of the small (28S) subunit, large (39S) subunit, and fully assembled (55S) mitochondrial ribosomes were assessed by subjecting the density gradient fractions to qRT-PCR using primer sets specific to 12S rRNA, which marks the small subunit, and to 16S rRNA, which marks the large subunit. Co-localization of the 12S and 16S rRNAs is diagnostic of fully assembled and actively translating ribosomes. Fractions containing the small (28S) subunit, large (39S) subunit, and fully assembled (55S) ribosome are shaded cyan, pink, and yellow, respectively. The relative abundance of a given rRNA in each fraction was calculated as the percentage relative to the total RNA abundance in all fractions after normalizing to a luciferase control RNA that was spiked into each of the fractions prior to RNA isolation. Control = UAS-mCherry-RNAi driven by elav-GAL4 and da-GAL4. LonKD = UAS-Lon-RNAi-1 driven by elav-GAL4 and da-GAL4
The mild decrease in mitochondrial ribosome assembly appeared insufficient to account for the translational defect of LonKD flies, especially in light of the fact that mitochondrial transcripts are elevated in abundance relative to the controls. Thus, we examined other possible explanations for the translational defect of Lon-deficient flies. One possible explanation of our findings was that mitochondrial transcripts are translated at lower efficiency in LonKD flies. To test this model, we used sucrose density gradient centrifugation to quantify the relative distribution of four different mitochondrial transcripts, including mt:ND5, mt:Cyt-b, mt:CoII, and ATPase8/6, in gradient fractions (Fig. 5). All of these mRNAs displayed a predominant sedimentation peak co-migrating with the 55S ribosome in control samples. By contrast, in LonKD flies a smaller proportion of these transcripts co-migrated with the 55S ribosome, and this decrease was accompanied by a striking increase in the proportion of these transcripts that sedimented to the bottom of the gradient (Fig. 5). This finding suggests that a substantial proportion of the mitochondrial transcripts in LonKD flies are packaged into large untranslated particles.
qRT-PCR analysis of individual sucrose gradient fractions was used to characterize the distribution of the indicated mtDNA-encoded mRNAs relative to the 28S and 39S subunits, and fully assembled 55S mitochondrial ribosomes. Mitochondrial homogenates for sucrose density fractionation were prepared from 21-day-old adult LonKD and control flies. Relative abundance represents the fraction of the mRNA in any given fraction relative to the total after normalizing to a luciferase control RNA that was added to each of the fractions prior to RNA isolation. Control = UAS-mCherry-RNAi driven by elav-GAL4 and da-GAL4. LonKD = UAS-Lon-RNAi-1 driven by elav-GAL4 and da-GAL4
Lon knockdown flies accumulate unfolded mitochondrial proteins and trigger the mitochondrial unfolded protein response
Lon is believed to play a critical role in the degradation of unfolded mitochondrial proteins34,35,36. The accumulation of unfolded proteins in multiple cellular compartments has been shown to activate a stress response known as the unfolded protein response (UPR) that is specific to the compartment in which the unfolded proteins reside37,38,39. The cytosolic and endoplasmic reticulum (ER) UPR restore protein homeostasis through a two-tiered system consisting of increased expression of chaperones to facilitate the refolding of misfolded proteins, and phosphorylation-mediated inactivation of the cytosolic translation-initiation factor eIF2α to attenuate translation40. While previous work has established that unfolded protein stress in the mitochondria triggers the induction of chaperones and proteases, whether mitochondrial translation is inhibited by unfolded protein stress is less clear. Our findings that Lon inactivation appears to result in the sequestration of mitochondrially encoded mRNAs into translationally inactive particles, coupled with the severe defect in mitochondrial translation, led us to hypothesize that the mitochondrial UPR triggers translational inhibition.
To explore our hypothesis we first analyzed whether LonKD flies accumulate unfolded mitochondrial proteins and whether they activate the mitochondrial UPR. To test whether unfolded mitochondrial proteins accumulate in LonKD flies, we prepared fly head protein extracts using Triton X-100. We then subjected Triton-soluble and -insoluble proteins to western blot analysis using antisera to selected mitochondrial proteins to quantify the proportion of each protein in the soluble and insoluble fractions. LonKD flies had higher levels of insoluble mitochondrial proteins than the control flies at both day 1 and day 21 of age (Fig. 6a and Suppl. Figure 6a), indicating that LonKD flies accumulate unfolded mitochondrial proteins. We next examined the abundance of the mitochondrial UPR markers Hsp60 and Hsc70-5 in LonKD flies. This analysis revealed a marked increase in both Hsp60 and Hsc70-5 in LonKD flies at day 1 and at day 21 of age (Fig. 6b and Suppl. Figure 6b). Together, these findings indicate that LonKD flies accumulate unfolded proteins and trigger the mitochondrial UPR.
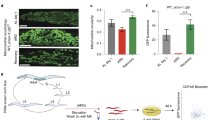
a Triton-insoluble mitochondrial proteins detected by western blot in heads from 1-day-old control and LonKD flies, using antibodies to complex Vβ, PDHα, aconitase, and NDUFS3. The results were quantified by ratio to actin and normalized to control levels. The experiment was repeated at least three times. *p < 0.05, **p < 0.005, ***p < 0.0005 by Student’s t-test. b Immunoblot analysis of head proteins using antibodies to Hsp60A and Hsc70-5. Protein was extracted from heads of day 1 control and LonKD flies using RIPA buffer. Quantification was performed as in (a). Experiments were repeated at least three times. *p < 0.05, **p < 0.005 by Student’s t-test. Control = UAS-mCherry-RNAi driven by elav-GAL4 and da-GAL4. LonKD = UAS-Lon-RNAi-1 driven by elav-GAL4 and da-GAL4
Overexpression of ClpP protease ameliorates the defect of Lon knockdown flies
Protein quality control in the mitochondrial matrix is regulated by an elaborate network of proteases and chaperones, including the AAA+ protease family member Clp protease. To test whether the accumulation of unfolded proteins in LonKD flies is responsible for the phenotypes associated with Lon inactivation, we co-expressed the proteolytic subunit of the Drosophila Clp protease (CG5045, hereafter ClpP) along with the Lon-RNAi-1 RNAi construct and examined whether ClpP overexpression is capable of rescuing the locomotor defect of LonKD flies. To account for possible titration of GAL4 protein in the presence of two UAS transgenes, we compared Lon knockdown flies overexpressing ClpP to Lon knockdown flies expressing mCherry RNAi. However, strong overexpression of ClpP in LonKD flies worsened the locomotor defect (Suppl. Figure 7). Thus, we tested whether ClpP expression could rescue a climbing defect caused by expressing the Lon-RNAi-1 transgene using just the elav-GAL4 driver (designated as LonKD-elav). We first confirmed that driving the UAS-ClpP transgene with elav-GAL4 resulted in the production of detectable ClpP protein (Fig. 7a) and use of the elav-GAL4 driver in conjunction with the Lon-RNAi-1 transgene reduced Lon expression (Suppl. Figure 8) and caused a climbing defect (Fig. 7b). Results of this analysis revealed that ClpP overexpression rescued the locomotor defect of Lon knockdown flies, thus supporting the hypothesis that the accumulation of unfolded proteins is responsible for the phenotypes of Lon-deficient flies (Fig. 7b).
a Immunoblot analysis of fly heads to confirm expression of FLAG-tagged ClpP in flies bearing a UAS-ClpP-FLAG-HA transgene and the elav-GAL4 pan-neuronal driver. b Climbing was measured in 1-day-old control flies (n = 94), LonKD-elav flies (n = 91), and LonKD-elav flies co-expressing ClpP protease (n = 83). Control = UAS-mCherry-RNAi driven by elav-GAL4. LonKD-elav; mCherry RNAi = UAS-Lon-RNAi-1 and UAS-mCherry-RNAi driven by elav-GAL4. LonKD-elav; UAS-ClpP = UAS-Lon-RNAi-1 and UAS-ClpP-FLAG-HA driven by elav-GAL4. Error bars represent SEM. ***p < 0.0005, ****p < 0.0001 by Student’s t-test
Discussion
Mitochondria are particularly prone to damage. They are the primary source and recipient of damaging ROS and continual replication of the mitochondrial genome throughout life culminates in mutation frequencies often orders of magnitude greater than that of the nuclear genome4. Moreover, expression of the mitochondrial and nuclear genomes must be coordinated to ensure proper stoichiometry of mitochondrial and nuclear-encoded respiratory chain components. An imbalance in this coordination can result in the accumulation of misfolded proteins and mitochondrial dysfunction39. The AAA+ mitochondrial protease family is believed to play an important role in maintaining mitochondrial competence by degrading and thereby facilitating the replacement of oxidatively damaged and misfolded proteins. However, we know little of the cellular responses and the pathogenic mechanisms of diseases caused by mutations in the genes that encode the AAA+ proteases. Our current work advances our understanding of these matters by showing that inactivation of the AAA+ protease Lon results in respiratory chain defects that appear to result from translational inhibition. Our findings further indicate that the translational defect caused by Lon inactivation results at least in part from the sequestration of mtDNA-encoded mRNAs into translationally inactive particles. Finally, we find that insoluble mitochondrial matrix proteins accumulate in LonKD animals, suggesting that mitochondrial protein aggregation triggers translational inactivation as a stress response. Our work provides a foundation to further explore the mechanisms by which mitochondrial unfolded protein stress inhibits mitochondrial translation.
Protein unfolding in the ER and cytoplasm induce well-characterized stress responses37,38,40,41,42,43. Each of these stress pathways has two arms: increased expression of chaperones to refold proteins, and translational inhibition to limit further protein synthesis. Translational inhibition is mediated by phosphorylation of eIF2α and culminates in the formation of translationally stalled ribonucleoprotein/mRNA complexes known as stress granules44. More recently, work in a variety of experimental systems has established the existence of a mitochondria-specific version of the unfolded protein response (UPRmt)45,46,47,48,49. Although the degree to which this pathway is evolutionarily conserved remains an active area of investigation, it has been shown in both vertebrate and invertebrate models that mitochondrial unfolded protein stress triggers the transcriptional activation of genes involved in protein folding and degradation45,47. However, previous work has not clearly established whether the UPRmt also results in inhibition of mitochondrial translation. Recently published findings in vertebrate cell culture have revealed that inactivation of the matrix chaperone Trap1 results in inhibition of mitochondrial translation as a consequence of a pre-RNA processing defect50. Our findings that Lon knockdown flies have both an excess of mitochondrial unfolded proteins and an impairment of mitochondrial translation raise the possibility that the UPRmt, like other unfolded protein stress pathways, has a second arm involving translational inhibition. However, the precise mechanism by which mitochondrial unfolded protein stress in Lon-deficient animals inhibits mitochondrial translation will require further investigation.
Although most of the respiratory chain complexes exhibited reduced activity and abundance upon Lon inactivation, complex II activity was increased. This increase may simply represent a compensatory response, as complex II has no mitochondrially encoded subunits, and increased complex II activity has previously been observed in mutants with defects in mitochondrial gene expression30,51,52. However, it is also possible that increased complex II activity is a consequence of reduced degradation of the complex II assembly factor Sdh5, given that previous work has shown that Lon mediates the turnover of Sdh553. Lon knockdown might thus reduce the degradation of Sdh5 and promote complex II assembly. The increased abundance of mitochondrially encoded transcripts upon Lon inactivation may likewise reflect either a general compensatory response or a specific change due to altered turnover of a possible Lon substrate. Increased mitochondrial transcript abundance has been reported previously in mutants with defective mitochondrial RNA processing and translation, suggesting that increasing the abundance of mitochondrial mRNA is a common response to inadequate mitochondrial translation30,31,52. However, the accumulation of mitochondrially encoded RNAs in LonKD flies could also be explained by the finding that these flies accumulate the RNA-binding protein Lrpprc1 (leucine-rich pentatricopeptide repeat motif-containing 1 protein; Suppl. Figure 9), which is known to stabilize mitochondrial transcripts51,54,55. Finally, Lon is a known component of mitochondrial nucleoids, and could therefore influence the turnover and abundance of other proteins involved in the synthesis or stabilization of mitochondrial transcripts56. Future work will be required to address these matters.
There are many unanswered questions regarding the biological roles of the AAA+ protease family and the mitochondrial response to unfolded protein stress. For example, mutations in the human gene encoding Lon cause a developmental disorder known as CODAS syndrome23. However, the underlying mechanisms causing disease are completely unknown. Although Lon is well known to promote the degradation of oxidatively damaged and unfolded proteins, the specific protein substrates of Lon are largely unknown. Whether unfolded protein stress in the mitochondria generally triggers translation inhibition, or this feature is specific to Lon inactivation is also unclear. Our study provides a foundation to address these questions and the mechanisms underlying CODAS syndrome in future work.
Materials and methods
Fly stocks and maintenance
Drosophila stocks were maintained on cornmeal-molasses food at 25 °C on a 12 h:12 h light–dark cycle. The UAS-Lon-RNAi-1 construct P{GD14030}-v36036 was obtained from the Vienna Drosophila Resource Center. The w1118, UAS-mCherry RNAi (P{VALIUM20-mCherry}attP2), UAS-Lon-RNAi-2 (P{TRiP.HMS01060}attP2), elav-GAL4, and da-GAL4 driver lines were obtained from the Bloomington Stock Center (Bloomington, IN, USA). The UAS-ClpP-FLAG-HA transgenic line was obtained from the Fly Facility, National Centre for Biological Sciences, Bangalore, India.
The Lon knockout allele was created using CRISPR/Cas9-mediated gene editing according to a published procedure25,57. Briefly, we replaced the Lon (CG8798) coding sequence with DsRed through homology-mediated repair. The following primer sequences were used for guide RNAs targeting the 5′ and 3′ UTR regions of Lon:
5′-Guide RNA
Sense oligo: 5′-CTTCGATAATCACTCACCACACATT-3′
Antisense oligo: 5′-AAACAATGTGTGGTGAGTGATTATC-3′
3′-Guide RNA
Sense oligo: 5′-CTTCGGGGTGTTGCGGGTGTTGAT-3′
Antisense oligo: 5′-AAACATCAACACCCGCAACACCCC-3′
Sequences flanking the Lon coding region were amplified from genomic DNA to facilitate homology-directed repair using the following primer sequences:
5′-Homology arm
Forward: 5′-CCGGCACCTGCGGCCTCGCAGTGCTCCGATCACGTTGGGAATGGG-3′
Reverse: 5′-CCGGCACCTGCGGCCCTACGTGTGGTGAGTGATTATGTGACGGCTGGTG-3′
3′-Homology arm
Forward: 5′-GGCCGCTCTTCATATGATAGGTTTTATAAATATCTATCGTTATCAGG-3′
Reverse: 5′-CCGGGCTCTTCTGACTTTCCCCGCCTCACCGGTGGACGGCC-3′
These homology arms were then cloned into the pHD-DsRed-attP vector containing the eye-specific 3xP3 promoter fused with DsRed and the resulting construct was microinjected into Cas9-expressing embryos by a commercial service (Rainbow Transgenic Flies Inc.). Flies bearing the Lon deletion were identified by screening the offspring of injected adults for expression of red fluorescence in the compound eye. Flies expressing the DsRed marker were then further subjected to whole-genome sequencing to verify deletion of the Lon gene.
Lifespan and behavioral analyses
Lifespan and behavioral analyses were performed using male flies. Age in all these experiments refers to the number of days following eclosion. Longevity assays were performed at 25 °C and involved 20 flies per vial. Flies were transferred to fresh food every 2–3 days and the number of dead flies was recorded during each transfer. Kaplan–Meier lifespan curves were generated using GraphPad Prism v5, and we used the Mantel–Cox log-rank test to determine the statistical significance of differences in survival between tested genotypes.
For climbing and flight assays, flies were anesthetized with CO2 and allowed to recover for at least 24 h before the experiment. Climbing behavior was assessed using the Rapid Iterative Negative Geotaxis (RING) assay at day 1 according to a previously published protocol with minor modifications58. In particular, height climbed was measured on still images from video recordings rather than on photographs. Briefly, 15 flies were transferred into plastic vials and these vials were then loaded onto the RING apparatus. The apparatus was tapped down to initiate the climbing response and the height climbed by each fly after 3 s was recorded. The climbing assay was repeated three times for each group.
Flight assays were performed according to a previously published protocol59,60. Briefly, an acetate sheet was divided into five equal parts, coated with grease and inserted into a 2-liter graduated cylinder. One-day-old flies were tapped into a funnel at the top of the cylinder and became stuck to the vacuum grease where they alighted. The number of flies that alighted in each of the five sections was counted and multiplied by the number corresponding to each section (0-4, labeled from bottom to top). The flight index was calculated by summing these values and dividing this sum by the maximum possible score (four times the number of flies used in the assay). At least 100 flies of each genotype were used for a given experiment.
Mitochondrial respiratory chain activity assay
Mitochondrial respiratory chain activity assays were performed according to a published procedure with several minor modifications25. Briefly, 1000 adult flies were homogenized in isolation buffer (5 mM Tris (pH 7.4), 250 mM sucrose, and 2 mM EGTA) with 1% (w/v) fatty acid free bovine serum albumin. The lysate was subjected to centrifugation at 600 g for 10 min to remove cellular debris. The supernatant was then subjected to further centrifugation at 7000 g for 10 min to pellet mitochondria. Mitochondria were washed twice in isolation buffer, resuspended in the same buffer, flash frozen in liquid nitrogen, and stored at −80 °C. Roughly 100 µg of mitochondria was used for each assay. Complex I activity was determined spectrophotometrically by monitoring the oxidation of NADH at 340 nm using ubiquinone-1 as an electron acceptor. Nonspecific activity was determined using 10 μM rotenone and subtracted to calculate complex I-specific activity. Complex II activity was determined by monitoring the reduction of 2,6-dichlorophenolindophenol at 600 nm in the presence of succinate and decylubiquinone. Background activity was determined using 10 mM malonate and subtracted to calculate complex II-specific activity. Complex III activity was determined by monitoring the reduction of cytochrome c at 550 nm in a reaction mixture containing decylubiquinol and cytochrome c. Nonspecific activity was determined using antimycin A and subtracted to calculate complex III-specific activity. Complex IV activity assay was performed by monitoring the oxidation of reduced cytochrome c at 550 nm. Background activity was determined using potassium cyanide and used to calculate complex IV-specific activity. All activities were normalized to citrate synthase activity, which was determined by following the reduction of 5,5′-dithiobis (2-nitrobenzoic acid) at 412 nm in the presence of acetyl-coenzyme A and oxaloacetate.
Total ATP determination
Total ATP was determined from whole flies according to a previously published procedure61. Briefly, five 21-day-old flies were homogenized in 100 μL of homogenization buffer (6 M guanidine HCL, 100 mM Tris (pH 7.8), and 4 mM EDTA). The samples were boiled for 5 min and subjected to centrifugation for 3 min at 21,000 g to remove debris. The supernatant was then diluted to 1:750 and total ATP content was measured using an ATP determination kit (A22066, Molecular Probes). Total ATP content was determined by comparing the luminescence measurements for each sample to the ATP standard curve and normalized to the total number of flies used in the assay.
Blue native PAGE (BN-PAGE) analysis and in-gel activity assay
BN-PAGE and in-gel activity assays were performed according to a previously published protocol62. Briefly, 100 μg of mitochondria prepared from 3-week-old adult flies was solubilized in a buffer containing a digitonin/protein (w/w) ratio of 8 and subjected to centrifugation at 20,000 g for 10 min at 4 °C. Coomassie G-250 was added to the supernatant and the sample was analyzed by native PAGE. Following electrophoresis, the resulting gel was subjected to in-gel activity assays as described below.
The combined complex I and IV in-gel activity assay was performed by first incubating the gel in a solution containing 1 mg/ml of the complex IV substrate cytochrome c along with 0.5 mg/ml 3,3′-diaminobenzidine and 45 mM phosphate buffer (pH 7.4) for 40 min. After the appearance of brown reaction products, the gel was washed with water and incubated in a solution containing 0.1 mg/ml of the complex I substrate NADH along with 2 mM Tris (pH 7.4) and 2.5 mg/ml nitrotetrazolium blue chloride for 20 min. The reaction was quenched with 10% acetic acid upon the appearance of the violet color indicative of complex I activity.
The complex II in-gel activity assay was performed by incubating the gel in a solution consisting of 5 mM Tris (pH 7.4), 20 mM sodium succinate, 2.5 mg/ml nitrotetrazolium blue chloride, and 0.2 mM phenazine methosulfate for 40 min. The reaction was quenched with 10% acetic acid upon the appearance of the violet color indicative of complex II activity.
The complex V in-gel activity assay was carried out by incubating the gel in a solution containing 35 mM Tris, 270 mM glycine, 14 mM magnesium sulfate, 10 mM adenosine triphosphate, and 0.2% lead (II) nitrate for 16 h. The reaction was stopped using 50% methanol upon the appearance of silver bands indicative of complex V activity.
Immunoblotting
Fly heads were homogenized in RIPA buffer with protease inhibitor cocktail (Roche) for 20 min on ice, and after centrifugation at 21,000 × g for 20 min the supernatant was collected and subjected to western blot analysis. The antibodies used were as follows: rabbit anti-LONP1 1:500 (NBP1-81734, Novus Biologicals); mouse anti-Actin 1:50,000 (MAB1501, Chemicon/Bioscience Research Reagents); mouse anti-FLAG 1:1000 (F3165, Sigma); rabbit anti-citrate synthase 1:1000 (CISY11-A, Alpha Diagnostics), rabbit anti-Hsp60 1:500 (D307, Cell Signaling Technology); and rabbit anti-GRP 75 (H155) 1:1000 (sc-13967, Santa Cruz) in Phosphate buffered saline with 0.1% Tween-2024. The secondary antibodies (anti-mouse HRP and anti-rabbit HRP (Sigma)) were used at a dilution of 1:10,000. Western blot images were quantified using ImageJ software (NIH) and normalized to actin. Each experiment was performed in triplicate.
To analyze unfolded mitochondrial proteins, protein was extracted from fly heads as previously described, except that 0.5% rather than 1% Triton X-100 was used in accordance with other work on mitochondrial proteins63,64. Antibodies for assaying mitochondrial protein folding status were used as follows: mouse anti-NDUFS3 1:500 (ab14711, Abcam), mouse anti-Complex V Beta 1:2000 (A-21351, Invitrogen), mouse anti-PDH E1α (clone 8D10E6) 1:1000 (45-660-0, Fisher Scientific), and rabbit anti-aconitase (ACO2) 1:2000 (AP1936C, Abgent).
Genomic DNA isolation and mitochondrial DNA copy number estimation
Mitochondrial and nuclear DNA was isolated from 40–50 heads obtained from 21-day-old adult flies using the DNeasy Blood & Tissue kit (Qiagen). A total 5 ng of genomic DNA was used as a template to perform qPCR using iTaq Universal SYBR Green Supermix (Bio-Rad). Mitochondrial DNA levels were estimated by using primers (Supplemental Table 1) to amplify the mt:Cyt-b gene, and normalized to levels of the nuclear gene Act79b. The relative fold change was determined through the 2−ΔΔCt method30.
RNA isolation and quantitative reverse transcription-PCR (qRT-PCR)
RNA isolation and qRT-PCR was performed according to a previously published procedure24,30,51. Total RNA was isolated from 40–50 heads obtained from 21-day-old adult flies using the Direct-zol RNA MiniPrep kit (Zymo Research). The RNA was reverse transcribed to cDNA using the iScript cDNA Synthesis Kit (Bio-Rad). For RNA quantification, qRT-PCR experiments were performed using iTaq Universal SYBR Green Supermix and a LightCycler 480 (Roche). Each sample was analyzed in triplicate and normalized to Act79b transcript abundance. The relative fold change was determined by the 2−ΔΔCt method. All primers used for qRT-PCR are listed in Supplemental Table 1.
In organello translation
De novo labeling of mitochondrial translation products was performed as previously described30,51. Approximately 750 µg of mitochondria isolated from third instar larvae or adult flies was resuspended in 500 µl of translation buffer (100 mM mannitol, 10 mM sodium succinate, 80 mM potassium chloride, 5 mM magnesium chloride, 1 mM potassium phosphate, 25 mM HEPES (pH 7.4), 60 μg/ml all amino acids except methionine, 5 mM ATP, 0.2 mM GTP, 6 mM creatine phosphate, and 60 μg/mL creatine kinase) supplemented with 500 μCi/ml of 35S-methionine (Perkin–Elmer). Following incubation at 30 °C for 1 h, mitochondria were washed four times using isolation buffer and resuspended in SDS sample buffer. Roughly 300 µg of mitochondria was subjected to SDS-PAGE. Following electrophoresis, the gel was dried and exposed to a phosphor screen. The phosphor screen was scanned using a gel imaging scanner (GE Typhoon FLA 9000). Mitochondrial translation profile was compared to a previously published study done in Drosophila larvae30. Roughly 75 µg of mitochondria was loaded on gel and stained with Coomassie to use as a loading control.
Mitochondrial ribosomal profiling using sucrose density gradient assay
Mitochondrial ribosomal profiling was performed according to a previously published procedure with minor modifications30. Freshly isolated mitochondria (2 mg) were incubated on ice in lysis buffer (260 mM sucrose, 100 mM NH4Cl, 10 mM MgCl2, 30 mM Tris-HCl pH 7.5, 40 U/ml Protector RNase Inhibitor, and 1% Triton X-100) supplemented with EDTA-free complete protease inhibitor cocktail (Roche) and PhosSTOP phosphatase inhibitor cocktail (Roche). Mitochondrial lysates were cleared by pelleting the debris through centrifugation at 10,000 g for 45 min at 4 °C. The supernatant was then loaded onto a 7–47% linear sucrose gradient made in a buffer containing 100 mM NH4Cl, 10 mM MgCl2, 30 mM Tris-HCl pH 7.5, and EDTA-free complete protease inhibitor cocktail (Roche). Samples were then subjected to centrifugation at 39,000 rpm for 6 h at 4 °C. Fractions of 500 μl from the sucrose gradient sedimentation were collected and RNA was extracted from each fraction using TRIzol LS Reagent (Invitrogen) and the Direct-zol RNA MiniPrep kit. RNA was reverse transcribed and qRT-PCR analyses were performed to quantify RNAs as described above.
Statistics
All findings are presented as mean ± SEM. Unless otherwise stated, statistical significance was calculated using two-tailed Student’s t-test in GraphPad Prism v5.
Change history
10 July 2019
Due to a technical error, content intended for publication in Volume 4 (2018) published in Volume 5 (2019). The content has been moved into the correct volume, and the citation information was updated accordingly.
References
Kauppila, T. E. S., Kauppila, J. H. K. & Larsson, N. G. Mammalian mitochondria and aging: an update. Cell Metab. 25, 57–71 (2017).
Sun, N., Youle, R. J. & Finkel, T. The mitochondrial basis of aging. Mol. Cell 61, 654–666 (2016).
Pickrell, A. M. & Youle, R. J. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85, 257–273 (2015).
Gustafsson, C. M., Falkenberg, M. & Larsson, N. G. Maintenance and expression of mammalian mitochondrial DNA. Annu. Rev. Biochem. 85, 133–160 (2016).
Hallberg, B. M. & Larsson, N. G. Making proteins in the powerhouse. Cell Metab. 20, 226–240 (2014).
Patron, M., Sprenger, H. G. & Langer, T. m-AAA proteases, mitochondrial calcium homeostasis and neurodegeneration. Cell Res. 28, 296–306 (2018).
Rugarli, E. I. & Langer, T. Mitochondrial quality control: a matter of life and death for neurons. EMBO J. 31, 1336–1349 (2012).
Quiros, P. M., Langer, T. & Lopez-Otin, C. New roles for mitochondrial proteases in health, ageing and disease. Nat. Rev. Mol. Cell Biol. 16, 345–359 (2015).
Pinti, M. et al. Emerging role of Lon protease as a master regulator of mitochondrial functions. Biochim. Biophys. Acta 1857, 1300–1306 (2016).
Pinti, M. et al. Mitochondrial Lon protease at the crossroads of oxidative stress, ageing and cancer. Cell. Mol. Life Sci. 72, 4807–4824 (2015).
Nystrom, T. Role of oxidative carbonylation in protein quality control and senescence. EMBO J. 24, 1311–1317 (2005).
Bota, D. A. & Davies, K. J. Lon protease preferentially degrades oxidized mitochondrial aconitase by an ATP-stimulated mechanism. Nat. Cell Biol. 4, 674–680 (2002).
Wagner, I., Arlt, H., van Dyck, L., Langer, T. & Neupert, W. Molecular chaperones cooperate with PIM1 protease in the degradation of misfolded proteins in mitochondria. EMBO J. 13, 5135–5145 (1994).
Savel’ev, A. S. et al. ATP-dependent proteolysis in mitochondria. m-AAA protease and PIM1 protease exert overlapping substrate specificities and cooperate with the mtHsp70 system. J. Biol. Chem. 273, 20596–20602 (1998).
Major, T., von Janowsky, B., Ruppert, T., Mogk, A. & Voos, W. Proteomic analysis of mitochondrial protein turnover: identification of novel substrate proteins of the matrix protease pim1. Mol. Cell Biol. 26, 762–776 (2006).
Bayot, A. et al. Identification of novel oxidized protein substrates and physiological partners of the mitochondrial ATP-dependent Lon-like protease Pim1. J. Biol. Chem. 285, 11445–11457 (2010).
Li, L. et al. Changes in specific protein degradation rates in Arabidopsis thaliana reveal multiple roles of Lon1 in mitochondrial protein homeostasis. Plant J.: Cell Mol. Biol. 89, 458–471 (2017).
Song, J. Y., Marszalek, J. & Craig, E. A. Cysteine desulfurase Nfs1 and Pim1 protease control levels of Isu, the Fe-S cluster biogenesis scaffold. Proc. Natl Acad. Sci. USA 109, 10370–10375 (2012).
Ciesielski, S. J., Schilke, B., Marszalek, J. & Craig, E. A. Protection of scaffold protein Isu from degradation by the Lon protease Pim1 as a component of Fe-S cluster biogenesis regulation. Mol. Biol. Cell. 27, 1060–1068 (2016).
Matsushima, Y., Goto, Y. & Kaguni, L. S. Mitochondrial Lon protease regulates mitochondrial DNA copy number and transcription by selective degradation of mitochondrial transcription factor A (TFAM). Proc. Natl Acad. Sci. USA 107, 18410–18415 (2010).
Lu, B. et al. Phosphorylation of human TFAM in mitochondria impairs DNA binding and promotes degradation by the AAA+Lon protease. Mol. Cell 49, 121–132 (2013).
Kao, T. Y. et al. Mitochondrial Lon regulates apoptosis through the association with Hsp60-mtHsp70 complex. Cell Death Dis. 6, e1642 (2015).
Strauss, K. A. et al. CODAS syndrome is associated with mutations of LONP1, encoding mitochondrial AAA+Lon protease. Am. J. Hum. Genet. 96, 121–135 (2015).
Thomas, R. E., Andrews, L. A., Burman, J. L., Lin, W. Y. & Pallanck, L. J. PINK1-Parkin pathway activity is regulated by degradation of PINK1 in the mitochondrial matrix. PLoS Genet. 10, e1004279 (2014).
Pareek, G., Thomas, R. E. & Pallanck, L. J. Loss of the Drosophila m-AAA mitochondrial protease paraplegin results in mitochondrial dysfunction, shortened lifespan, and neuronal and muscular degeneration. Cell death & Dis. 9, 304 (2018).
Qi, Y., Liu, H., Daniels, M. P., Zhang, G. & Xu, H. Loss of Drosophila i-AAA protease, dYME1L, causes abnormal mitochondria and apoptotic degeneration. Cell Death Differ. 23, 291–302 (2016).
Quiros, P. M. et al. ATP-dependent Lon protease controls tumor bioenergetics by reprogramming mitochondrial activity. Cell Rep. 8, 542–556 (2014).
Garcia, C. J., Khajeh, J., Coulanges, E., Chen, E. I. & Owusu-Ansah, E. Regulation of mitochondrial complex I biogenesis in Drosophila flight muscles. Cell Rep. 20, 264–278 (2017).
Vincow, E. S. et al. The PINK1-Parkin pathway promotes both mitophagy and selective respiratory chain turnover in vivo. Proc. Natl Acad. Sci. USA 110, 6400–6405 (2013).
Baggio, F. et al. Drosophila melanogaster LRPPRC2 is involved in coordination of mitochondrial translation. Nucleic Acids Res. 42, 13920–13938 (2014).
Szczepanowska, K. et al. CLPP coordinates mitoribosomal assembly through the regulation of ERAL1 levels. EMBO J. 35, 2566–2583 (2016).
Camara, Y. et al. MTERF4 regulates translation by targeting the methyltransferase NSUN4 to the mammalian mitochondrial ribosome. Cell Metab. 13, 527–539 (2011).
Dogan, S. A. et al. Tissue-specific loss of DARS2 activates stress responses independently of respiratory chain deficiency in the heart. Cell Metab. 19, 458–469 (2014).
Bernstein, S. H. et al. The mitochondrial ATP-dependent Lon protease: a novel target in lymphoma death mediated by the synthetic triterpenoid CDDO and its derivatives. Blood 119, 3321–3329 (2012).
Suzuki, C. K., Suda, K., Wang, N. & Schatz, G. Requirement for the yeast gene LON in intramitochondrial proteolysis and maintenance of respiration. Science 264, 891 (1994).
Bezawork-Geleta, A., Brodie, E. J., Dougan, D. A. & Truscott, K. N. LON is the master protease that protects against protein aggregation in human mitochondria through direct degradation of misfolded proteins. Sci. Rep. 5, 17397 (2015).
Ron, D. & Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 8, 519–529 (2007).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Shpilka, T. & Haynes, C. M. The mitochondrial UPR: mechanisms, physiological functions and implications in ageing. Nat. Rev. Mol. Cell Biol. 19, 109–120 (2018).
Pakos-Zebrucka, K. et al. The integrated stress response. EMBO Rep. 17, 1374–1395 (2016).
Balch, W. E., Morimoto, R. I., Dillin, A. & Kelly, J. W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Lamech, L. T. & Haynes, C. M. The unpredictability of prolonged activation of stress response pathways. J. Cell Biol. 209, 781–787 (2015).
Akerfelt, M., Morimoto, R. I. & Sistonen, L. Heat shock factors: integrators of cell stress, development and lifespan. Nat. Rev. Mol. Cell Biol. 11, 545–555 (2010).
Protter, D. S. & Parker, R. Principles and properties of stress granules. Trends Cell Biol. 26, 668–679 (2016).
Nargund, A. M., Fiorese, C. J., Pellegrino, M. W., Deng, P. & Haynes, C. M. Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPR(mt). Mol. Cell 58, 123–133 (2015).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012).
Zhao, Q. et al. A mitochondrial specific stress response in mammalian cells. EMBO J. 21, 4411–4419 (2002).
Owusu-Ansah, E., Song, W. & Perrimon, N. Muscle mitohormesis promotes longevity via systemic repression of insulin signaling. Cell 155, 699–712 (2013).
Siegelin, M. D. et al. Exploiting the mitochondrial unfolded protein response for cancer therapy in mice and human cells. J. Clin. Invest. 121, 1349–1360 (2011).
Munch, C. & Harper, J. W. Mitochondrial unfolded protein response controls matrix pre-RNA processing and translation. Nature 534, 710–713 (2016).
Bratic, A. et al. The bicoid stability factor controls polyadenylation and expression of specific mitochondrial mRNAs in Drosophila melanogaster. PLoS Genet. 7, e1002324 (2011).
Rackham, O. et al. Hierarchical RNA Processing Is Required for Mitochondrial Ribosome Assembly. Cell Rep. 16, 1874–1890 (2016).
Bezawork-Geleta, A., Saiyed, T., Dougan, D. A. & Truscott, K. N. Mitochondrial matrix proteostasis is linked to hereditary paraganglioma: LON-mediated turnover of the human flavinylation factor SDH5 is regulated by its interaction with SDHA. FASEB J. 28, 1794–1804 (2014).
Matsushima, Y. et al. Drosophila protease ClpXP specifically degrades DmLRPPRC1 controlling mitochondrial mRNA and translation. Sci. Rep. 7, 8315 (2017).
Siira, S. J. et al. LRPPRC-mediated folding of the mitochondrial transcriptome. Nat. Commun. 8, 1532 (2017).
Gilkerson, R. et al. The mitochondrial nucleoid: integrating mitochondrial DNA into cellular homeostasis. Cold Spring Harb. Perspect. Biol. 5, a011080 (2013).
Gratz, S. J. et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics 196, 961–971 (2014).
Gargano, J. W., Martin, I., Bhandari, P. & Grotewiel, M. S. Rapid iterative negative geotaxis (RING): a new method for assessing age-related locomotor decline in Drosophila. Exp. Gerontol. 40, 386–395 (2005).
Greene, J. C. et al. Mitochondrial pathology and apoptotic muscle degeneration in Drosophila parkin mutants. Proc. Natl Acad. Sci. USA 100, 4078–4083 (2003).
Burman, J. L. et al. A Drosophila model of mitochondrial disease caused by a complex I mutation that uncouples proton pumping from electron transfer. Dis. Models Mech. 7, 1165–1174 (2014).
Tennessen, J. M., Barry, W. E., Cox, J. & Thummel, C. S. Methods for studying metabolism in Drosophila. Methods 68, 105–115 (2014).
Jha, P., Wang, X. & Auwerx, J. Analysis of mitochondrial respiratory chain supercomplexes using blue native polyacrylamide gel electrophoresis (BN-PAGE). Curr. Protoc. Mouse Biol. 6, 1–14 (2016).
Pimenta de Castro, I. et al. Genetic analysis of mitochondrial protein misfolding in Drosophila melanogaster. Cell Death Differ. 19, 1308–1316 (2012).
Davis, M. Y. et al. Glucocerebrosidase deficiency in Drosophila results in alpha-synuclein-independent protein aggregation and neurodegeneration. PLoS Genet. 12, e1005944 (2016).
Acknowledgements
We thank Dr. Scott Kennedy (Department of Pathology, University of Washington) for assistance with whole genome sequencing of the Londel mutant; Dr. Dana Miller for allowing us to use her lab space for experiments involving radioactive isotopes; and Dr. Evan Eichler, Dr. Christine Queitsch, and Dr. Stanley Fields for providing access to their lab facilities. We thank Dr. Paul M. Macdonald for providing an antibody against Lrpprc protein. This work was supported by a grant from the National Institutes of Health to LP (R21NS094901).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pareek, G., Thomas, R.E., Vincow, E.S. et al. Lon protease inactivation in Drosophila causes unfolded protein stress and inhibition of mitochondrial translation. Cell Death Discov. 4, 51 (2018). https://doi.org/10.1038/s41420-018-0110-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-018-0110-1
This article is cited by
-
Hypoxia-induced mitochondrial stress granules
Cell Death & Disease (2023)
-
Evidence for mitochondrial Lonp1 expression in the nucleus
Scientific Reports (2022)
-
Exercise alters the mitochondrial proteostasis and induces the mitonuclear imbalance and UPRmt in the hypothalamus of mice
Scientific Reports (2021)
-
Insights from Drosophila on mitochondrial complex I
Cellular and Molecular Life Sciences (2020)
-
Inactivation of Lon protease reveals a link between mitochondrial unfolded protein stress and mitochondrial translation inhibition
Cell Death & Disease (2018)