Abstract
Hepatocarcinogenesis is a multi-step process. However, the regulators of hepatocellular carcinoma (HCC) initiation are understudied. Adult liver-specific gene expression was globally downregulated in HCC. We hypothesize that adult liver-specific genes, especially adult liver-enriched transcription factors may exert tumor-suppressive functions in HCC. In this study, we identify ZBTB7B, an adult liver-enriched transcription factor as a permissive regulator of HCC initiation. ZBTB7B is highly expressed in hepatocytes in adult livers, compared to fetal livers. To evaluate the functions of ZBTB7B in hepatocarcinogenesis, we performed hepatocyte-specific ZBTB7B knockout in hydrodynamic oncogene transfer-induced mouse liver cancer models. Hepatocyte-specific knockout of ZBTB7B promotes activated Akt and N-Ras-induced HCC development. Moreover, ZBTB7B deficiency sensitizes hepatocytes to a single oncogene Akt-induced oncogenic transformation and HCC initiation, which is otherwise incompetent in inducing HCC. ZBTB7B deficiency accelerates HCC initiation by down-regulating adult liver-specific gene expression and priming livers to a fetal-like state. The molecular mechanism underlying ZBTB7B functions in hepatocytes was investigated by integrated transcriptomic, phosphoproteomic, and chromatin immunoprecipitation-sequencing analyses. Integrative multi-omics analyses identify c-Jun as the core signaling node in ZBTB7B-deficient liver cancer initiation. c-Jun is a direct target of ZBTB7B essential to accelerated liver cancer initiation in ZBTB7B-deficient livers. Knockdown of c-Jun expression or dominant negative c-Jun expression delays HCC development in ZBTB7B-deficient livers. In addition, ZBTB7B competes with c-Jun for chromatin binding. Ectopic ZBTB7B expression attenuates the tumor-promoting functions of c-Jun. Expression of ZBTB7B signature, composed of 140 genes co-regulated by ZBTB7B and c-Jun, is significantly downregulated in early-stage HCCs compared to adjacent normal tissues, correlates to liver-specific gene expression, and is associated with good prognosis in human HCC. Thus, ZBTB7B functions as a permissive regulator of HCC initiation by directly regulating c-Jun expression and function.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC), the major type of primary liver cancer, is unfortunately often diagnosed when the disease is already at advanced stages. It is therefore of special importance to fully understand how malignant transformation occurs in early-stage preneoplastic lesions. Hepatocarcinogenesis is a multi-step process progressing from regenerative nodules to early HCC (ref. [1, 2]). Genetic alterations driving HCC development, e.g., TERT promoter, TP53, are rarely found in preneoplastic lesions of HCC (ref. [3,4,5]), suggesting deregulated expression or functions of signaling molecules contribute to HCC initiation (ref. [2,3,4,5,6]). Intriguingly, transcriptomic and proteomic analyses of HCC reveal that genes downregulated in HCC are mostly those specifically expressed in differentiated hepatocytes (ref. [7, 8]). Global downregulation of the adult liver-specific proteins in HCC may be attributed to impaired functions of liver-specific transcription factors that dictate hepatocyte differentiation and regulate liver-specific functions. We hypothesize that deregulated functions of these transcription factors may have pivotal roles in the initiation of HCC.
Transcription factor zinc finger and BTB domain-containing 7b (ZBTB7B, also known as Th-POK or cKrox) is a critical regulator of cell fate determination in many biological processes. ZBTB7B is a key regulator in T cell lineage commitment (ref. [9,10,11]). Loss of ZBTB7B expression or impaired ZBTB7B function disrupts the development of CD4 T cells (ref. [12]). ZBTB7B expression is temporally induced during BMP4-induced reprogramming of mouse epiblast stem cells to naive pluripotency (ref. [13]). ZBTB7B facilitates the opening of naive pluripotent chromatin loci and the primed-to-naive transition (ref. [13]). ZBTB7B is a potent driver of brown fat development and cold-induced beige fat formation that activates thermogenic gene expression in adipocytes (ref. [14]). Besides cell fate specification, ZBTB7B is also involved in metabolic regulation. ZBTB7B regulates the expression of insulin receptor substrate-1 (IRS-1) (ref. [15]). ZBTB7B deficiency impairs insulin-induced Akt-mTOR-SREBP pathway and lipid biosynthesis in the mammary alveolar cells at lactation (ref. [15]).
In this study, we report that ZBTB7B is highly expressed in adult hepatocytes and is a negative regulator of HCC initiation. ZBTB7B deficiency primes hepatocytes to a fetal state, sensitizes hepatocytes to oncogenic transformation, and accelerates HCC initiation. ZBTB7B exerts tumor-suppressive functions by directly regulating c-Jun expression and function.
Results
ZBTB7B is an adult liver-enriched transcription factor
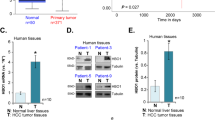
Adult liver-specific genes (ref. [16]) are globally downregulated in human HCC (Fig. S1A) (ref. [7, 8]). Combined overexpression of HA tagged-activated Akt (Myr-Akt-HA) and N-Ras (N-Ras V12) in the liver via hydrodynamic gene transfer induced rapid tumor development (ref. [17]). Similar to human HCC, adult liver-specific gene expression was substantially downregulated in Akt/N-Ras-induced tumors (Fig. S1B). We hypothesize that adult liver-specific genes, especially adult liver-enriched transcription factors may exert tumor-suppressive functions in HCC. Livers undergo postnatal maturation to establish functional organs. Clustering analysis of fetal and adult liver gene expression identified six hepatic gene clusters (Fig. S1C). Clusters 1, 2, 3, 5, and 6 represented genes that were highly expressed in stages of embryonic development, but were expressed at low levels in adult livers (Fig. S1C), including oncofetal genes Afp, Gpc3, Sall4 (cluster 2), H19 (Cluster 3) and Igf2 (Cluster 5), which are re-expressed in liver cancer (ref. [18,19,20,21,22]). Cluster 4 represented genes that were expressed at low levels in fetal livers, but high levels in adult livers (Fig. S1C). Top adult liver-specific transcription factors in cluster 4 included Nr1h3/Lxra and Rxra, transcription factors critical for adult liver functions (ref. [23]) (Fig. S1D). Cluster 4 also included Esr1 and Smad7, which were previously reported to be inhibitory to HCC development (ref. [24, 25]) (Fig. S1D). Interestingly, Zbtb7b, a transcription factor regulating cell fate determination, shared similar expression kinetics to Nr1h3/Lxra, Rxra, Esr1, and Smad7 (Fig. S1D, E). Hepatic ZBTB7B mRNA and protein levels were significantly increased during the postnatal maturation process (Fig. S1F–H). Expression of ZBTB7B was also increased in adult human liver, compared to human fetal liver (Fig. S1I).
ZBTB7B is a critical regulator of cell fate determination in many biological processes (ref. [9, 13, 14]). It is yet unknown whether ZBTB7B regulates hepatocyte differentiation, liver functions, and liver cancer development. Immunostaining on liver sections revealed that ZBTB7B was mainly expressed in hepatocytes in the liver, and ~90% of hepatocytes expressed ZBTB7B (Fig. S2). To investigate the functions of hepatic ZBTB7B, Zbtb7bf/f mice were crossed with Alb-Cre mice to specifically delete Zbtb7b in the hepatocytes (Zbtb7bΔli) (Fig. S2 and S3A). Despite no overt developmental abnormality in the hepatocyte Zbtb7b-ablated livers (Fig. S3A–E), phylogenetic analysis of liver transcriptomes revealed that Zbtb7bΔli livers had gene expression pattern resembling the fetal livers (Fig. 1A). Notably, expression of the adult liver-specific genes was significantly downregulated in the Zbtb7bΔli livers (Fig. 1B). Liver cancer gene signatures were enriched (Fig. 1C) and significantly more Ki67+ proliferating hepatocytes were present in the Zbtb7bΔli livers (Fig. S3F, G). Upon activated Akt/N-Ras expression, ZBTB7B expression was significantly downregulated (Fig. S3H). As deficiency of ZBTB7B, an adult liver-enriched cell fate regulator, in hepatocytes primed the livers to a fetal-like state and its expression was downregulated in the tumors, we next sought to investigate whether hepatocyte ZBTB7B would function as a tumor suppressor in regulating HCC development.
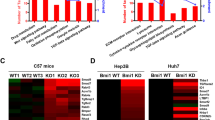
A Phylogenetic analysis of gene expression in the Zbtb7bf/f and Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) adult livers, and published gene expression profiles of adult and fetal livers. B GSEA comparison of liver-specific gene expression between the Zbtb7bf/f and Zbtb7bΔli livers. C GSEA comparison of liver cancer gene signatures between the Zbtb7bf/f and Zbtb7bΔli livers. D Study design. Hydrodynamic tail vein injection (HTVi) of Akt and N-Ras to induce tumor development in control (Alb-Cre or Zbtb7bf/f, Ctrl) or Zbtb7bΔli mice. E Kaplan–Meier survival analysis. Control (n = 6), Zbtb7bΔli (n = 7). F Gross liver images of control and Zbtb7bΔli mice 4 weeks after injection with vector (Vec) or Akt/N-Ras. G Liver/body weight ratio of control and Zbtb7bΔli mice 4 weeks after injection with vector or Akt/N-Ras. n = 5. H H&E staining of liver sections of control and Zbtb7bΔli mice 4 weeks after injection with vector or Akt/N-Ras. Magnification: ×4. Scale bar: 500 μm. I, J Numbers of tumors (I) and percentage of tumor area (J) in the livers of control and Zbtb7bΔli mice 4 weeks after Akt/N-Ras injection. n = 5. K Immunohistochemistry of HA-tag and Ki67 on liver sections of control and Zbtb7bΔli mice 4 weeks after Akt/N-Ras injection. Magnification: ×10. Scale bar: 200 μm. L Percentage of Ki67+ hepatocytes in the livers of control and Zbtb7bΔli mice 4 weeks after Akt/N-Ras injection. n = 5. Data are presented as mean ± SEM. Statistical analyses were performed with the Log-Rank test (E), two-way ANOVA followed by Šídák’s multiple comparison tests (G) or two-tailed unpaired student’s t test (I, J, L). *P < 0.05, ***P < 0.001. ns: not significant.
ZBTB7B deficiency accelerates activated Akt/N-Ras-induced hepatocarcinogenesis
Combined overexpression of activated Akt and N-Ras induced tumor development in the livers of Alb-Cre or Zbtb7bf/f (control, Ctrl) mice with median survival of 45.0 days (Fig. 1D, E) (ref. [17]). Ablation of Zbtb7b in hepatocytes significantly accelerated Akt/N-Ras-induced liver cancer development with median survival of 27.5 days (Fig. 1D, E). At 4 weeks after Akt/N-Ras oncogene transfer, Zbtb7bΔli mice had significantly more tumors with a much higher liver/body weight ratio than the control mice (Fig. 1F, G). Akt/N-Ras induced multifocal tumors in the liver (Fig. 1H). Significantly more tumors developed in the Zbtb7bΔli mice with larger tumor size, compared to the control mice (Fig. 1H–J). Immunostaining of the HA-tag showed more exogenous Akt-expressing cells in the Zbtb7bΔli livers (Fig. 1K). Zbtb7bΔli mice had significantly more Ki67+ proliferative hepatocytes (Fig. 1K, L). To further investigate whether ZBTB7B suppresses liver cancer development, ZBTB7B was expressed along with Akt/N-Ras oncogenes in the wild-type mice (Fig. S4A). Ectopic ZBTB7B expression alleviated tumor burden with reduced tumor numbers, tumor area, and proliferative hepatocyte numbers (Fig. S4). These data collectively suggested that ZBTB7B exerts tumor-suppressive functions in Akt/N-Ras oncogene-induced liver cancer development.
ZBTB7B deficiency accelerates liver cancer initiation and is permissive to single oncogene-induced hepatocarcinogenesis
Combined Akt/N-Ras oncogene expression drives stepwise hepatocarcinogenesis (ref. [17]). Consistent with histological changes, principal component analysis of gene expression in the livers transduced with Akt/N-Ras oncogenes showed stepwise global gene expression changes (Fig. 2A). The changes on gene expression in the Zbtb7bΔli livers preceded that in the Zbtb7bf/f livers (Fig. 2A), suggesting liver cancer initiation occurred at earlier time point in the Zbtb7bΔli livers than the Zbtb7bf/f livers. At 2 weeks after Akt/N-Ras oncogene transfer, liver cancer gene signatures were enriched in the Zbtb7bΔli livers compared to the Zbtb7bf/f livers (Fig. 2B). Zbtb7bΔli livers appeared paler than the Zbtb7bf/f livers with higher liver/body weight ratio (Fig. 2C, D). Despite no tumor nodule being evident in the Zbtb7bf/f livers at this time point (Fig. 2E, F), multiple HA-tag-positive, highly proliferative tumor nodules developed in the Zbtb7bΔli livers (Fig. 2E–G).
A Principal component analysis of gene expression in the livers of Zbtb7bf/f and Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) mice injected with vector or Akt/N-Ras. B GSEA comparison of liver cancer gene signatures between Zbtb7bf/f and Zbtb7bΔli livers 2 weeks after Akt/N-Ras injection. C Gross liver images of Zbtb7bf/f and Zbtb7bΔli mice 2 weeks after Akt/N-Ras injection. D Liver/body weight ratio of Zbtb7bf/f and Zbtb7bΔli mice 2 weeks after Akt/N-Ras injection. n = 5. E H&E staining and immunohistochemistry of HA-tag and Ki67 on the liver sections of Zbtb7bf/f and Zbtb7bΔli mice 2 weeks after Akt/N-Ras injection. Magnifications: ×10 and ×40. Scale bars: 200 μm. F, G Numbers of tumor nodules (F) and percentage of Ki67+ hepatocytes (G) in the livers of Zbtb7bf/f and Zbtb7bΔli mice 2 weeks after Akt/N-Ras injection. n = 5. Data are presented as mean ± SEM. Statistical analyses were performed with two-tailed unpaired student’s t test. *P < 0.05, ***P < 0.001.
Multiple genetic or epigenetic alternations are necessary for hepatocyte transformation. Single oncogene overexpression or tumor suppressor deletion is often insufficient to initiate liver cancer, e.g., activated Akt1 alone is incompetent in inducing liver cancer development (ref. [26]). As ZBTB7B deficiency primed the livers to the fetal-like state and accelerated Akt/N-Ras-induced liver cancer initiation, we next sought to investigate whether ZBTB7B-deficient hepatocytes would be susceptible to single oncogene-induced liver cancer. Consistent with previous reports, no overt tumor formation was observed in the Zbtb7bf/f mice up to 20 weeks after transducing Akt (Fig. 3A–C). At 20 weeks after Akt oncogene transfer, Zbtb7bf/f livers were largely normal in size, shape, and color (Fig. 3C–E). No tumor was noticeable on the liver surface (Fig. 3C). Histological inspection revealed a few small preneoplastic lesions in the Zbtb7bf/f livers (Fig. 3E–G). Immunostaining with HA-tag further confirmed only sporadic Akt-expressing hepatocytes with enlarged clear cytoplasm were present in the Zbtb7bf/f livers (Fig. 3E). Akt expression in Zbtb7bf/f livers had minimal effect on hepatocyte proliferation (Fig. 3E, H). In sharp contrast to the Zbtb7bf/f livers, Akt robustly induced tumor development in the livers of the Zbtb7bΔli mice with a median survival of 16 weeks (Fig. 3A–C). Zbtb7bΔli livers were pale in color with large tumors on the liver surface (Fig. 3C). Liver size was significantly augmented in Zbtb7bΔli mice (Fig. 3C, D). Histological inspection revealed large and multifocal tumors and large numbers of exogenous Akt-expressing and proliferating cells in the Zbtb7bΔli livers (Fig. 3E–H).
A Study design. Hydrodynamic tail vein injection (HTVi) of Akt to induce tumor development in Zbtb7bf/f or Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) mice. Mice were sacrificed 20 weeks (Zbtb7bf/f) after Akt injection or moribund (Zbtb7bΔli). n = 5. B Kaplan–Meier survival analysis. C Gross liver images of Zbtb7bf/f and Zbtb7bΔli mice 20 weeks after Akt injection. D Liver/body weight ratio of Zbtb7bf/f and Zbtb7bΔli mice. E H&E staining and immunohistochemistry of HA-tag and Ki67 on liver sections of Zbtb7bf/f and Zbtb7bΔli mice. Magnification: ×10. Scale bar: 200 μm. F, G Numbers of tumors (F) and percentage of tumor area (G) in the livers of Zbtb7bf/f and Zbtb7bΔli mice. H Percentage of Ki67+ hepatocytes in the livers of Zbtb7bf/f and Zbtb7bΔli mice. Data are presented as mean ± SEM. Statistical analyses were performed with the Log-Rank test (B) or two-tailed unpaired student’s t test (D, F, G, H). ***P < 0.001.
c-Jun is a core signaling node in accelerated liver cancer initiation in ZBTB7B-deficient hepatocytes
8804 genes were differentially expressed between the Zbtb7bf/f and Zbtb7bΔli livers during Akt/N-Ras oncogene-induced hepatocyte transformation (Fig. 4A). Clustering analysis of differentially expressed genes (DEGs) expression tendency identified 8 gene clusters (Fig. 4B and Table S1). Clusters 2, 4, 5, and 6 included genes synergistically regulated by ZBTB7B and Akt/N-Ras oncogenes that were upregulated (clusters 2 and 5) or downregulated (clusters 4 and 6) in the Zbtb7bΔli livers and further upregulated or downregulated upon Akt/N-Ras oncogene expression (Fig. 4B and Table S1). Whereas genes in clusters 1, 7, and 8 were mainly regulated by Akt/N-Ras oncogenes (Fig. 4B and Table S1). Transcription factor analysis showed that genes in each cluster were regulated by different sets of transcription factors, especially those involved in liver cancer development, including c-Jun, NF-κB, STAT1, and P53 (Fig. 4C and Table S2). KEGG pathway enrichment analysis suggested regulation of cell cycle, Ras-MAPK signaling, PI3K-Akt signaling, chemokine signaling, and NF-κB pathway were differentially activated (Fig. S5A). Quantitative gene set variation analysis (GSVA) further revealed Ras-MAPK, PI3K-Akt, and Hippo signaling were partially activated in the Zbtb7bΔli livers that were further activated upon Akt/N-Ras oncogene expression (Fig. S5B). Global phosphoproteomic profiling revealed 8459 differentially expressed phosphosites (DEPs) classed into 8 clusters (Fig. S5C, D, and Table S3). Cluster 2 DEPs represented phosphosites enriched in the Zbtb7bΔli livers, including phosphosites on P21, c-Jun, and Ki67, which were significantly upregulated upon Akt/N-Ras oncogene expression (Fig. S5D and Table S3). NetworKIN analysis (ref. [27]) enriched activities of 68 kinases from DEPs (Fig. S5E and Table S4). Transcription factor analysis enriched 13 TFs at the phosphoproteomic level (Table S2). To investigate the key regulators responsible for accelerated liver cancer initiation in the ZBTB7B-deficient livers, protein-protein interaction (PPI) network was constructed by integrated analysis of the functional intersection of DEGs and DEPs. 122 nodes and 368 edges were identified (Fig. S5F and Table S5). Functional genes, e.g., Akt1, Shc1, Prkce, Jun, and Hdac1, were highly related to each other and might play important regulatory roles in the accelerated liver cancer initiation in Zbtb7bΔli livers (Fig. S5F and Table S5).
A Hierarchical clustering analysis for differentially expressed genes (DEGs). B Temporal expression profiling of the DEGs using Mfuzz. C TRRUST analysis of transcriptional regulatory relationships. Top 20 transcription factors are shown. D Protein-protein interaction network synergistically regulated by ZBTB7B and Akt/N-Ras oncogenes. The shape and color of nodes represent the functional annotation of the gene. Mint blue diamonds: transcription factors, pink ellipses: kinases, dark green round rectangles: targets, purple round rectangles: substrates, yellow round rectangles: targets and substrates. Closed nodes represent direct targets of ZBTB7B. The edges represent the Pearson correlation coefficient calculated based on gene expression. The line type of edges indicated the data type. Solid line: RNA level, dash-dot line: phosphosite level. The color and width of the edges indicate the strength of the correlation. Blue to red: correlation coefficient from −1 to 1. Node size: vitality of the gene. E Genome view of the occupancy of ZBTB7B in Jun locus. F Quantitative RT-PCR analysis of c-Jun expression in Zbtb7bf/f and Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) livers. n = 5. G Immunohistochemistry of c-Jun on liver sections of Zbtb7bf/f and Zbtb7bΔli mice. Magnification: ×20. Scale bar: 100 μm. H Quantitation of c-Jun immunohistochemistry. n = 5. I Western blot analysis of c-Jun in liver lysates of Zbtb7bf/f and Zbtb7bΔli mice 2 weeks after injection with vector or Akt/N-Ras. J Western blot analysis of c-Jun in liver lysates of mice 4 weeks after Akt/N-Ras injection with GFP or ZBTB7B. Data are presented as mean ± SEM. Statistical analyses were performed with two-tailed unpaired student’s t test. **P < 0.01, ***P < 0.001.
To investigate which nodes were directly regulated by ZBTB7B, chromatin immunoprecipitation (ChIP)-sequencing was performed on the livers of the Zbtb7b-HA mice with an HA-tag in frame knocked-in before the stop codon of Zbtb7b gene (Fig. S6A, B). ZBTB7B is bound mainly to the promoter region vicinity of the transcription start site (Fig. S6C and S6D). ZBTB7B bound to the promoters of known ZBTB7B targets genes, e.g., Irs1 (ref. [15]), as well as genes important in liver cancer, including Jun, Akt1, Ccnd1, and Mki67 (Table S6). Functional enrichment analysis of ZBTB7B target genes suggested that ZBTB7B played a more prominent role in pathways in metabolism and cancer (Fig. S6E). Integration of ZBTB7B direct target genes into the PPI network revealed Jun, Akt1, Shc1, and Prkce as the top nodes with the highest degree correlation to ZBTB7B (Fig. S6F). PPI sub-network of DEGs and DEPs synergistically regulated by ZBTB7B and Akt/N-Ras oncogens (clusters 2, 4, 5, and 6) further revealed c-Jun as the central signaling node in accelerated liver cancer initiation in the Zbtb7bΔli livers (Fig. 4D). AP1 family transcription factor c-Jun, first identified as viral oncoprotein (ref. [28]), is required in diethylnitrosamine (DEN)-induced liver cancer initiation (ref. [29, 30]). c-Jun, identified in the cluster 2 DEGs (Fig. 4B and Table S1), was a direct target gene of ZBTB7B that ZBTB7B is directly bound to the Jun promoter (Fig. 4E). c-Jun expression was significantly increased in the Zbtb7bΔli livers (Fig. 4F–H). Expression of Akt/N-Ras oncogenes downregulated ZBTB7B expression and increased c-Jun expression (Fig. 4I). Increased c-Jun expression upon Akt/N-Ras expression was attenuated by ectopic ZBTB7B expression (Fig. 4J).
c-Jun is essential to accelerated hepatocarcinogenesis in ZBTB7B-deficient hepatocytes
As c-Jun was identified as a core signaling node in Akt/N-Ras-induced ZBTB7B-deficient liver cancer initiation (Fig. 4), we next sought to investigate whether ZBTB7B deficiency accelerates liver cancer initiation through c-Jun. Truncated c-Jun, which lacked the N-terminal transactivation domain, exerted dominant negative effects on c-Jun-mediated transcription (ref. [31]). Ectopic dominant negative (DN)-c-Jun expression inhibited Akt/N-Ras-induced tumor development in the Zbtb7bΔli livers (Fig. 5A, B) with significantly reduced liver/body weight ratio to the extent comparable to the Zbtb7bf/f livers (Fig. 5C). Histological inspection of DN-c-Jun-expressing Zbtb7bΔli livers revealed much-reduced tumor numbers, tumor area and Ki67+ proliferative cell numbers to the levels of the Zbtb7bf/f livers (Fig. 5D–G). The essential roles of c-Jun in accelerated hepatocarcinogenesis were further demonstrated by knocking down c-Jun expression in ZBTB7B-deficient hepatocytes (Fig. S7). ZBTB7B deficiency was permissive to single oncogene Akt-induced hepatocarcinogenesis (Fig. 3). DN-c-Jun expression completely abolished Akt-induced liver cancer development in the Zbtb7bΔli livers (Fig. S8), suggesting c-Jun is downstream of ZBTB7B mediating hepatocyte transformation and liver cancer development.
A Study design. Hydrodynamic tail vein injection (HTVi) of Akt/N-Ras to induce tumor development in Zbtb7bf/f or Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) mice with pT3-N-FLAG vector control (Vec) or DN-c-Jun. Mice were sacrificed 4 weeks after oncogene injection. n = 6. B Gross liver images of Zbtb7bf/f and Zbtb7bΔli mice. C Liver/body weight ratio of Zbtb7bf/f and Zbtb7bΔli mice. D H&E staining and immunohistochemistry of HA-tag, FLAG-tag, and Ki67 on liver sections. Magnification: ×10. Scale bars: 200 μm. E, F Numbers of tumors (E) and percentage of tumor area (F). G Percentage of Ki67+ hepatocytes. Data are presented as mean ± SEM. Statistical analyses were performed with two-way ANOVA followed by Šídák’s multiple comparison test. ***P < 0.001. ns: not significant.
ZBTB7B negatively regulates c-Jun function in hepatocarcinogenesis
To determine the mechanistic role of c-Jun in mediating accelerated liver cancer initiation in Zbtb7b-deficient hepatocytes, ChIP-seq analysis of c-Jun genome-wide binding was performed. Zbtb7b deficiency led to significantly increased global c-Jun occupancy (Fig. 6A, B, Table S7 and S8). Upon ZBTB7B deletion, additional c-Jun binding peaks were detected in genes involved in regulating metabolism and cancer (Fig. 6C, D). We then investigated whether ZBTB7B can directly interfere with c-Jun chromatin occupancy by analyzing the relative position of the c-Jun and ZBTB7B-binding sites and revealed that the c-Jun peaks in both Zbtb7bf/f and Zbtb7bΔli livers largely overlapped with the corresponding ZBTB7B peaks in the wild-type livers (Fig. 6E). Interestingly, 32.1% (2595 of 8090) of ZBTB7B-binding sites overlapped with the c-Jun binding sites in the Zbtb7bf/f livers, while the vast majority of ZBTB7B-binding sites (70.9%, 5732 of 8090) overlapped with the c-Jun binding sites in the Zbtb7bΔli livers (Fig. 6F). In addition, c-Jun occupancy was significantly increased on 2526 c-Jun/ZBTB7B loci in the Zbtb7bΔli livers, compared to the Zbtb7bf/f livers (Fig. 6G), suggesting ZBTB7B directly antagonizes c-Jun chromatin occupancy. Furthermore, 50% of the nodes in the PPI network, including Zbtb7b, Jun, and Shc1, were co-regulated by ZBTB7B and c-Jun (Fig. 6H and Table S5).
A–G ChIP-seq analysis of genome-wide c-Jun occupancy in Zbtb7bf/f and Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) livers. A, B Heatmaps (A) and distribution of c-Jun binding intensity (B) centered at transcription start site (TSS). C Mean-average (MA) plot of differential c-Jun ChIP-seq peak density in the Zbtb7bf/f and Zbtb7bΔli livers. Log fold change (M) and log of the mean (A) of normalized read density are shown. D KEGG pathway analysis of genes corresponding to ChIP-seq peaks unique in the Zbtb7bΔli livers. E Enrichment profiles representing the c-Jun ChIP-seq reads density from the Zbtb7bf/f and Zbtb7bΔli livers centered at ZBTB7B peaks. F Venn diagram showing the overlaps among c-Jun peaks from Zbtb7bf/f and Zbtb7bΔli livers and ZBTB7B peaks. G c-Jun ChIP signals from the 2526 c-Jun/ZBTB7B sites in the Zbtb7bf/f and Zbtb7bΔli livers. H Protein-protein interaction network synergistically regulated by ZBTB7B and c-Jun. Node color denotes the upstream transcription factors. The shape of the nodes, the color, and the line type of the edges are the same as in Fig. 4D. Data are presented as mean ± SEM. Statistical analyses were performed with two-tailed paired student’s t test. ****P < 0.0001.
We next sought to investigate whether ZBTB7B suppresses liver cancer development by blocking c-Jun function, in addition to regulating c-Jun expression. Phosphorylation of the c-Jun transactivation domain on serines 63 and 73 increases its transcription activity (ref. [32, 33]). Expression of active c-Jun (c-Jun S63D/S73D) accelerated Akt/N-Ras oncogene-induced liver cancer initiation (Fig. 7A–C) and liver cancer development (Fig. 7D–G). In contrast to more preneoplastic nodules at 2 weeks (Fig. 7B, C) and more tumors and proliferative hepatocytes at 4 weeks after active c-Jun expression (Fig. 7D–G), simultaneous overexpression of ZBTB7B completely abolished the tumor-promoting effects of active c-Jun (Fig. 7). ZBTB7B overexpression reduced the numbers of preneoplastic nodules, tumor numbers, tumor area, and Ki67+ proliferative hepatocyte numbers to the levels of vector control (Fig. 7).
A Study design. Hydrodynamic tail vein injection (HTVi) of Akt and N-Ras oncogenes with active c-Jun (c-Jun S63D/S73D) and ZBTB7B. Mice were sacrificed 2 weeks or 4 weeks after hydrodynamic injection. n = 5. B H&E staining on liver sections of wild-type mice 2 weeks after Akt/N-Ras injection with active c-Jun and ZBTB7B. Magnification: ×10. Scale bar: 200 μm. C Numbers of tumor nodules 2 weeks after Akt/N-Ras injection with active c-Jun and ZBTB7B. D H&E staining and immunohistochemistry of HA-tag and Ki67 on liver sections of wild-type mice 4 weeks after Akt/N-Ras injection with active c-Jun and ZBTB7B. Magnification: ×10. Scale bars: 200 μm. E–G Numbers of tumors (E), percentage of tumor area (F), and Ki67+ hepatocytes (G) in the livers 4 weeks after Akt/N-Ras injection with active c-Jun and ZBTB7B. Data are presented as mean ± SEM. Statistical analyses were performed with one-way ANOVA followed by Tukey’s multiple comparison test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
ZBTB7B signature is associated with liver cancer prognosis
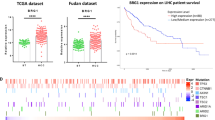
To determine to which extent ZBTB7B might be associated with human liver cancer phenotypes, we queried the Cancer Genome Atlas hepatocellular carcinoma (TCGA_LIHC) dataset. 1105 of 2166 genes with overlapping ZBTB7B and c-Jun binding were differentially expressed in which 176 genes were downregulated in the Zbtb7bΔli livers and further downregulated upon Akt/N-Ras oncogene expression (cluster 4 and 6, P < 0.001) (Fig. S9A). 140 human orthologous genes were used as ZBTB7B signature to assess the ZBTB7B activity in liver cancer samples (Fig. S9A and Table S9). Expression of the 140-gene ZBTB7B signature was significantly lower in HCC tumors (Fig. 8A). Comparison of paired tumor samples and adjacent normal liver tissues further supported that ZBTB7B activity was downregulated in the liver cancer (Fig. 8B). ZBTB7B activity was significantly lower in all stage/grade liver cancers, compared to the adjacent normal tissues (Fig. 8C, D). A comparison of liver cancers at different stages and grades revealed that advanced liver cancer only had a moderate further decrease of ZBTB7B activity (Fig. 8C, D). Association and linear regression analyses demonstrated a tight correlation between ZBTB7B signature and liver-specific gene expression levels in human HCC (Fig. 8E), consistent with the finding that ZBTB7B was an adult liver-enriched transcription factor (Fig. S1) and expression of adult liver-specific genes were significantly downregulated in the Zbtb7bΔli livers (Fig. S1B). In line with increased expression and activity of c-Jun in the Zbtb7bΔli livers (Fig. 4), ZBTB7B signature and c-Jun target gene expression levels negatively correlated to each other in human HCCs (Fig. 8F). Patients with high ZBTB7B signature (HR: 0.50 (0.35–0.71), P < 0.0001) and high liver-specific gene expression (HR: 0.52 (0.36–0.73), P = 0.0008) had better prognosis, while patients with high c-Jun target gene expression (HR: 1.94 (1.34–2.81), P < 0.0001) had significantly shorter overall survival (Fig. 8G–I). In a second cohort of 244 liver cancer patients, ZBTB7B activity was similarly lower in the liver cancers compared to the adjacent normal tissues (Fig. S9B, C), but comparable among different stages (Fig. S9D–F). ZBTB7B signature was correlated to the liver-specific gene expression levels but negatively correlated to the c-Jun target gene expression (Fig. S9G, H). ZBTB7B signature and liver-specific genes predicted a good prognosis, while c-Jun target genes predicted a poor prognosis (Fig. S9I–K).
Gene set variation analysis (GSVA) scores of ZBTB7B signature genes, liver-specific genes (HSIAO_LIVER_SPECIFIC_GENES), and c-Jun target genes (MATTHEWS_AP1_TARGETS) were calculated for each HCC patient samples in the TCGA_LIHC cohort. A Comparison of ZBTB7B signature scores in tumors and adjacent normal tissues. B Comparison of ZBTB7B signature scores in paired tumors and adjacent normal tissues. C, D Comparison of ZBTB7B signature scores in HCC tumors of different grades (C) and stages (D). E, F Pearson correlation of ZBTB7B signature scores to expression levels of liver-specific genes (E) and c-Jun target genes (F). G–I Probabilities of overall survival of HCC patients according to the expression level of the ZBTB7B signature (G), liver-specific genes (H), and c-Jun target genes (I). Statistical analyses were performed with two-tailed unpaired (A) or paired (B) student’s t test, one-way ANOVA followed by Tukey’s multiple comparison tests (C, D), or Log-Rank test (G–I). ***P < 0.001, ****P < 0.0001.
Discussion
In this study, we identify transcription factor ZBTB7B as a suppressive regulator of HCC initiation. ZBTB7B-deficient hepatocytes are susceptible to oncogenic transformation and HCC initiation. ZBTB7B suppresses HCC initiation by inhibiting c-Jun expression and activity, a key transcription factor in regulating HCC initiation.
ZBTB7B is a potential permissive regulator in HCC initiation. Hepatocarcinogenesis is the consequence of the accumulation of genetic and epigenetic alterations (ref. [1, 5]). Co-occurrence of functional driver mutations provides liver cancer cells growth advantage (ref. [5]). These gene mutations alter key signaling pathways in liver cancer cells, e.g., telomere maintenance, Wnt/β-catenin, P53/cell cycle regulation, Akt/mTOR, MAP kinase and oxidative stress (ref. [7, 34, 35]). However, activation of a single signaling pathway is generally insufficient in inducing HCC development. Overexpression of activated Akt1 can only induce liver cancer development with very long latency in the mouse models (ref. [26]). In contrast, Akt1 robustly induces HCC development in the background of ZBTB7B deficiency with much-shortened latency. Although mutation or deletion of ZBTB7B is not a frequent event in HCC, ZBTB7B expression is downregulated at the early-stage of Akt/N-Ras-induced HCC. Epigenetic dysregulation plays a crucial role in hepatocarcinogenesis (ref. [1, 5]). Expression and/or transcription activity of ZBTB7B might be regulated at early stages in hepatocarcinogenesis which provides hepatocytes permissive signals to oncogenic transformation. Consistent with this notion, genes regulated by ZBTB7B are already downregulated in very early-stage HCC (Fig. 8 and S9).
Transcriptomic profiling of HCC tumors reveals a subset of HCCs that are less differentiated expressing progenitor markers IGF2, EpCAM, and AFP (ref. [36,37,38]). Hepatocytes might dedifferentiate into progenitor-like cells before transforming into HCC (ref. [39, 40]). ZBTB7B is an adult liver-enriched transcription factor and is associated with adult liver features. In line with the functions as a cell lineage commitment regulator in multiple developmental processes (ref. [12,13,14]), ablation of ZBTB7B attenuates adult liver-specific gene expression (Fig. 1). Interestingly, the ZBTB7B gene signature (Fig. S9 and Table S9) significantly overlaps with the signatures of hepatoblastoma (ref. [41]) (42 of 140 in ZBTB7B gene signature, FDR q value 4.69E-26), liver-specific genes (ref. [16]) (23 of 140 in ZBTB7B gene signature, FDR q-value 1.32E-22), HCC subclass characterized by increased proliferation, high AFP levels and chromosomal instability (ref. [42]) (18 of 140 in ZBTB7B gene signature, FDR q value 5.61E-18), and HCC subtype associated with hepatocyte differentiation (ref. [37]) (18 of 140 in ZBTB7B gene signature, FDR q value 3.30E-15). Attenuated ZBTB7B expression and/or transcriptional activity might therefore maintain hepatocytes at the dedifferentiated state that is more susceptible to oncogenic transformation and HCC development.
Integrated analyses reveal that c-Jun, a direct target gene of ZBTB7B, functions as the core signaling node in ZBTB7B-deficient HCC initiation (Fig. 4D). c-Jun is required in DEN-induced mouse liver cancer initiation (ref. [29, 30]). c-Jun activation is predicted to play key roles in the development of a hepatoblast subtype HCC arising from hepatic progenitor cells (ref. [43]). ZBTB7B not only regulates c-Jun expression, but also regulates c-Jun transcriptional activity in the early-stage of Akt/N-Ras-induced HCC. The majority of ZBTB7B-binding sites in the genome overlap with c-Jun binding sites and ZBTB7B deficiency increases c-Jun chromatin binding (Fig. 6). It is noteworthy that 50% of the signaling nodes in the regulatory network are co-regulated by ZBTB7B and c-Jun (e.g., Insr, Pik3r1, Shc, and Braf), and about one-third of the signaling nodes are genes directly regulated by c-Jun, but not ZBTB7B (e.g., Mapk3, Iqgap1, Sos, Tsc1, and Rps6ka3) (Fig. 6H). The signaling nodes also contain genes regulated by transcription factors other than ZBTB7B and c-Jun, e.g., Mecp2, which binds to methylated DNA and mediates transcriptional repression through interaction with HDACs and the SIN3A corepressor. Ras/MAPK and PI-3 kinase/Akt signaling were substantially more activated in the Zbtb7b-deficient livers upon Akt/N-Ras oncogene expression (Fig. S5B), suggesting that ZBTB7B may regulate HCC initiation at least partly by regulating MAPK and Akt function in a c-Jun-dependent manner. ZBTB7B deficiency also significantly increases the activity of NF-κB and STAT1, key transcription factors mediating inflammatory responses (Fig. S6F). Chronic liver inflammation and immune regulation play important roles in HCC development (ref. [44]). Inflammatory cytokines, e.g., IL-6 and TNF-α, synergistically regulate hepatocyte transformation and HCC development (ref. [45, 46]). Damage-associated molecular patterns released from damaged hepatocytes activate innate immunity signaling in hepatocytes and the microenvironment and accelerate HCC development (ref. [44]). ZBTB7B may therefore orchestrate c-Jun-inflammatory signal crosstalk and regulate HCC initiation through a broader network of signaling pathways and effector molecules.
ZBTB7B functions as both a transcriptional activator and repressor by interaction with different epigenetic regulators. ZBTB7B interacts with class I (HDACs 1, 2, 3) and class II (HDACs 4, 5, 10) HDACs (ref. [47, 48]). ZBTB7B may form multicomponent complexes with different HDAC-interacting transcriptional corepressors, e.g., HDAC1/2-interacting Sin3a and HDAC3-interacting NcoR2 (Fig. S6F) (ref. [48]). HDAC-dependent histone deacetylation is required for ZBTB7B-mediated Cd8 transcription silencing during CD4 T cell differentiation (ref. [47]). ZBTB7B also interacts with acetyltransferase p300 and KAT5 (ref. [49, 50]) and may promote histone acetylation and gene transcription at a subset gene loci. p300 catalyzes histone crotonylation and succinylation (ref. [51, 52]), while Class I HDACs are major histone decrotonylases and desuccinylases (ref. [53, 54]). It raises the interesting possibility that ZBTB7B may regulate gene transcription by modulating histone crotonylation and succinylation, which play significant roles in regulating chromatin structure and gene transcription. Besides histone acylation, ZBTB7B may directly or indirectly interact with other epigenetic regulators, e.g., CBX3, CTBP1/2, UHRF1, as well as WDR5 (ref. [48]), which suppress or activate gene transcription. ZBTB7B is a multi-domain protein. The N-terminal BTB/POZ domain is responsible for ZBTB7B dimerization and interaction with transcriptional co-regulators, while the C-terminal zinc finger domains are responsible for DNA binding. Both BTB/POZ and zinc finger domains are required for ZBTB7B-mediated transcriptional regulation and function (ref. [47, 55]). It warrants further investigation whether ZBTB7B interacts with different epigenetic regulators and transcriptional co-regulators, which may result in changes of histone modification and chromatin accessibility to other transcription factors, e.g., c-Jun, and ultimately result in gene transcriptional activation or repression in a gene loci-specific manner.
In summary, ZBTB7B is an adult hepatocyte-enriched transcription factor and is a permissive regulator of HCC initiation. ZBTB7B deficiency primes hepatocytes to a fetal state, sensitizes hepatocytes to oncogenic transformation, and accelerates HCC initiation. ZBTB7B exerts tumor-suppressive functions by directly regulating c-Jun expression and competing with c-Jun for chromatin binding.
Materials and methods
Animal experiments
All mice were maintained in a C57/BL6 background and housed in a specific pathogen-free environment and treated in strict accordance with protocols approved by the Institutional Animal Care and Use Committee (Approval number: SIBCB-NAF-15-003-S325-006). Zbtb7bf/f mice (ref. [56]) were crossed with Alb-cre mice to generate Alb-Cre, Zbtb7bf/f (Zbtb7bΔli) mice. Zbtb7b-HA knockin mice were generated as described (ref. [57]) by the Genome Tagging Project (GTP) Center at the Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences. pT3-myr-Akt1-HA, pT3-N-Ras V12, and sleeping beauty transposase (SB) expression plasmids were generously provided by Dr. Xin Chen (UCSF). The coding sequences of Zbtb7b and Jun were amplified using mouse liver cDNA as a template and cloned into a pT3 vector. 6–8-week-old male wild-type, Zbtb7bf/f, and Zbtb7bΔli mice were randomly allocated into groups and hydrodynamic injection was performed to induce liver cancer as described (ref. [17, 58]). No blinding was performed.
Histology and immunohistochemistry
Mice were anesthetized and livers were isolated and fixed in 4% paraformaldehyde followed by embedding in paraffin. Paraffin-embedded livers were sectioned and stained with hematoxylin and eosin (H&E) or subjected to immunohistochemical staining as previously described (ref. [59]). The primary antibodies used are listed in Supplementary Table S10. Images were captured at 3 random fields at ×4, ×10, ×20, or ×40 magnifications with a digital camera (DP71; OLYMPUS, Inc.) and Digital Acquire software (DPController; OLYMPUS, Inc.).
RNA-seq and ChIP-seq analyses
Total RNA was extracted and purified from livers using TRIZOL (Invitrogen). Three biological replicates were subjected to complementary DNA library preparation and sequencing according to the Illumina standard protocol. RNA-seq data was mapped to the mouse mm10 reference genome by STAR (v.2.9). R package DESeq2 (v.1.24.0) was used to perform normalization and differential expression analysis. Gene ontology analysis was performed using g:profiler (ref. [60]). GSEA was performed on the GSEA application (v.4.0.3) with gene signatures obtained from the MSigDB database (v7.5.1). Statistical significance was assessed by comparing the enrichment score to enrichment results generated from 1,000 random permutations of the gene set to obtain a P-value (nominal P-value) (ref. [61]).
Perfused and formaldehyde cross-linked liver tissues were homogenized and cells were collected by centrifugation. Chromatin at an average size of 200–500 bp was incubated with HA antibody, c-Jun antibody, or normal rabbit IgG. Immunoprecipitated chromatin DNA was subjected to library preparation using a library preparation kit (#ND607, Vazyme) according to the manufacturer’s instructions. The libraries were sequenced on Illumina Novaseq. ChIP-seq data was mapped to the mouse mm10 reference genome by Bowtie2 (v.2.3.1). Peak detection was performed using the MACS2 (v.2.1.1). The downstream visualization and annotation were performed by IGV (v.2.11.2) and R package ChIPseeker (v.1.30.3) respectively. Peak overlapping was computed by using the default bedtools intersect intervals functions with at least 1 base pair overlap. MA plot was generated by MAnorm (v.1.3.0) (ref. [62]).
Mass spectrometry analysis
Phosphopeptide enrichment in mouse liver was performed according to the previous report (ref. [7]). With a two-hour liquid chromatography gradient elution method, the phosphopeptides were identified in data independent analysis (DIA) mode by Thermo Scientific Q Exactive HF-X Hybrid Quadrupole-Orbitrap mass spectrometer and analyzed using SpectronautTM 15 (Biognosys Inc.). Peptide Collapse in Perseus (1.6.8.0) (ref. [63]) was used to combine Spectronaut precursor quantifications into consensus phosphosites. The differentially expressed phosphosites (DEPs) were obtained by analysis of variance (ANOVA P value < 0.05) and were divided into 8 clusters according to their expression patterns. The kinases estimated by NetworKIN 3.0 (ref. [27]) and transcription factors estimated by TRRUST (Built into Metascape, http://www.metascape.org/) of each cluster were obtained by kinase-substrate enrichment analysis and transcription factor enrichment analysis, respectively. All possible protein-protein interactions (PPI) were obtained using known databases (MIPPIE, TRRUST, UniProt, and STRING), and their Pearson correlation coefficients were calculated. A simplified phosphoproteome-level PPI network was obtained. Transcriptome-level PPI network was treated in the same way. Network relationships were presented by Cytoscape (Version 3.8.0, http://www.cytoscape.org/).
Patient survival analysis
Mouse liver (PRJNA66167) and human liver (PRJNA280600) transcriptome data were downloaded from GEO and mapped to mm10 and hg19 reference genomes, respectively. Raw counts were normalized to FPKM. R package Mfuzz was used to perform cluster analysis on default parameters. Gene set enrichment scores of ZBTB7B signature (Table S9), Hsiao liver-specific genes (ref. [16]) and Matthews AP1 targets (ref. [64]) were calculated for each sample in previously published hepatocellular carcinoma (TCGA (ref. [35]) and GSE14520 (ref. [65]) gene expression data sets using GSVA (ref. [66]). Survival analysis was performed using the Kaplan–Meier (Log-Rank test) method to evaluate the correlation of the signature GSVA scores with overall survival in these two data sets. ROC analyses were applied to detect the optimal cutoff point.
Statistical analysis
Variance similarity between the groups is being statistically compared. Data meet the assumptions of the tests. No data were excluded from the analysis. The sample size was based on previous research experience. Sample numbers were indicated in the figure legends. Statistical significance between conditions was assessed by unpaired or paired two-tailed Student’s t-tests, one-way ANOVA followed by Tukey’s post-hoc analysis, or two-way ANOVA followed by Šídák’s multiple comparison test using GraphPad Prism. Error bars represent SEM.
Data availability
RNA-seq and ChIP-seq data are deposited to the National Omics Data Encyclopedia (NODE, https://www.biosino.org/node, accession number OEP003425).
References
Craig AJ, von Felden J, Garcia-Lezana T, Sarcognato S, Villanueva A. Tumour evolution in hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol. 2020;17:139–52.
Desjonqueres E, Campani C, Marra F, Zucman-Rossi J, Nault JC. Preneoplastic lesions in the liver: molecular insights and relevance for clinical practice. Liver Int. 2022;42:492–506.
Zucman-Rossi J, Villanueva A, Nault JC, Llovet JM. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology. 2015;149:1226–1239.e1224.
Schulze K, Imbeaud S, Letouze E, Alexandrov LB, Calderaro J, Rebouissou S, et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat Genet. 2015;47:505–11.
Rebouissou S, Nault JC. Advances in molecular classification and precision oncology in hepatocellular carcinoma. J Hepatol. 2020;72:215–29.
Nault JC, Martin Y, Caruso S, Hirsch TZ, Bayard Q, Calderaro J, et al. Clinical impact of genomic diversity from early to advanced hepatocellular carcinoma. Hepatology. 2020;71:164–82.
Gao Q, Zhu H, Dong L, Shi W, Chen R, Song Z, et al. Integrated proteogenomic characterization of HBV-related hepatocellular carcinoma. Cell. 2019;179:561–577.e522.
Chen X, Cheung ST, So S, Fan ST, Barry C, Higgins J, et al. Gene expression patterns in human liver cancers. Mol Biol Cell. 2002;13:1929–39.
He X, Park K, Kappes DJ. The role of ThPOK in control of CD4/CD8 lineage commitment. Annu Rev Immunol. 2010;28:295–320.
Keefe R, Dave V, Allman D, Wiest D, Kappes DJ. Regulation of lineage commitment distinct from positive selection. Science. 1999;286:1149–53.
Naito T, Taniuchi I. The network of transcription factors that underlie the CD4 versus CD8 lineage decision. Int Immunol. 2010;22:791–6.
He X, He X, Dave VP, Zhang Y, Hua X, Nicolas E, et al. The zinc finger transcription factor Th-POK regulates CD4 versus CD8 T-cell lineage commitment. Nature. 2005;433:826–33.
Yu S, Zhou C, Cao S, He J, Cai B, Wu K, et al. BMP4 resets mouse epiblast stem cells to naive pluripotency through ZBTB7A/B-mediated chromatin remodelling. Nat Cell Biol. 2020;22:651–62.
Li S, Mi L, Yu L, Yu Q, Liu T, Wang GX, et al. Zbtb7b engages the long noncoding RNA Blnc1 to drive brown and beige fat development and thermogenesis. Proc Natl Acad Sci USA. 2017;114:E7111–E7120.
Zhang R, Ma H, Gao Y, Wu Y, Qiao Y, Geng A, et al. Th-POK regulates mammary gland lactation through mTOR-SREBP pathway. PLoS Genet. 2018;14:e1007211.
Hsiao LL, Dangond F, Yoshida T, Hong R, Jensen RV, Misra J, et al. A compendium of gene expression in normal human tissues. Physiol Genomics. 2001;7:97–104.
Ho C, Wang C, Mattu S, Destefanis G, Ladu S, Delogu S, et al. AKT (v-akt murine thymoma viral oncogene homolog 1) and N-Ras (neuroblastoma ras viral oncogene homolog) coactivation in the mouse liver promotes rapid carcinogenesis by way of mTOR (mammalian target of rapamycin complex 1), FOXM1 (forkhead box M1)/SKP2, and c-Myc pathways. Hepatology. 2012;55:833–45.
Smith JA, Francis TI, Edington GM, Williams AO. Immunofluorescent localisation of human alpha feto-protein in fetal and neonatal livers and cultured cells from hepatocellular carcinoma. Br J Cancer. 1971;25:343–9.
Tietze L, Kessler SM. The good, the bad, the question-H19 in hepatocellular carcinoma. Cancers (Basel). 2020;12:1261.
Martinez-Quetglas I, Pinyol R, Dauch D, Torrecilla S, Tovar V, Moeini A, et al. IGF2 is up-regulated by epigenetic mechanisms in hepatocellular carcinomas and is an actionable oncogene product in experimental models. Gastroenterology. 2016;151:1192–205.
Morford LA, Davis C, Jin L, Dobierzewska A, Peterson ML, Spear BT. The oncofetal gene glypican 3 is regulated in the postnatal liver by zinc fingers and homeoboxes 2 and in the regenerating liver by alpha-fetoprotein regulator 2. Hepatology. 2007;46:1541–7.
Yong KJ, Gao C, Lim JS, Yan B, Yang H, Dimitrov T, et al. Oncofetal gene SALL4 in aggressive hepatocellular carcinoma. N Engl J Med. 2013;368:2266–76.
Russo-Savage L, Schulman IG. Liver X receptors and liver physiology. Biochim Biophys Acta Mol Basis Dis. 2021;1867:166121.
Bigsby RM, Caperell-Grant A. The role for estrogen receptor-alpha and prolactin receptor in sex-dependent DEN-induced liver tumorigenesis. Carcinogenesis. 2011;32:1162–6.
Moon H, Ju HL, Chung SI, Cho KJ, Eun JW, Nam SW, et al. Transforming growth factor-beta promotes liver tumorigenesis in mice via up-regulation of snail. Gastroenterology. 2017;153:1378–1391.e1376.
Calvisi DF, Wang C, Ho C, Ladu S, Lee SA, Mattu S, et al. Increased lipogenesis, induced by AKT-mTORC1-RPS6 signaling, promotes development of human hepatocellular carcinoma. Gastroenterology. 2011;140:1071–83.
Horn H, Schoof EM, Kim J, Robin X, Miller ML, Diella F, et al. KinomeXplorer: an integrated platform for kinome biology studies. Nat Methods. 2014;11:603–4.
Eferl R, Wagner EF. AP-1: a double-edged sword in tumorigenesis. Nat Rev Cancer. 2003;3:859–68.
Min L, Ji Y, Bakiri L, Qiu Z, Cen J, Chen X, et al. Liver cancer initiation is controlled by AP-1 through SIRT6-dependent inhibition of survivin. Nat Cell Biol. 2012;14:1203–11.
Eferl R, Ricci R, Kenner L, Zenz R, David JP, Rath M, et al. Liver tumor development. c-Jun antagonizes the proapoptotic activity of p53. Cell. 2003;112:181–92.
Hennigan RF, Stambrook PJ. Dominant negative c-jun inhibits activation of the cyclin D1 and cyclin E kinase complexes. Mol Biol Cell. 2001;12:2352–63.
Smeal T, Binetruy B, Mercola DA, Birrer M, Karin M. Oncogenic and transcriptional cooperation with Ha-Ras requires phosphorylation of c-Jun on serines 63 and 73. Nature. 1991;354:494–6.
Pulverer BJ, Kyriakis JM, Avruch J, Nikolakaki E, Woodgett JR. Phosphorylation of c-jun mediated by MAP kinases. Nature. 1991;353:670–4.
Qiu Z, Li H, Zhang Z, Zhu Z, He S, Wang X, et al. A pharmacogenomic landscape in human liver cancers. Cancer Cell. 2019;36:179–193.e111.
Cancer Genome Atlas Research Network. Electronic address: wheeler@bcm.edu; Cancer Genome Atlas Research Network Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell. 2017;169:1327–1341.e1323.
Boyault S, Rickman DS, de Reynies A, Balabaud C, Rebouissou S, Jeannot E, et al. Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology. 2007;45:42–52.
Hoshida Y, Nijman SM, Kobayashi M, Chan JA, Brunet JP, Chiang DY, et al. Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res. 2009;69:7385–92.
Calderaro J, Couchy G, Imbeaud S, Amaddeo G, Letouze E, Blanc JF, et al. Histological subtypes of hepatocellular carcinoma are related to gene mutations and molecular tumour classification. J Hepatol. 2017;67:727–38.
Tarlow BD, Pelz C, Naugler WE, Wakefield L, Wilson EM, Finegold MJ, et al. Bipotential adult liver progenitors are derived from chronically injured mature hepatocytes. Cell Stem Cell. 2014;15:605–18.
Seehawer M, Heinzmann F, D’Artista L, Harbig J, Roux PF, Hoenicke L, et al. Necroptosis microenvironment directs lineage commitment in liver cancer. Nature. 2018;562:69–75.
Carrillo-Reixach J, Torrens L, Simon-Coma M, Royo L, Domingo-Sabat M, Abril-Fornaguera J, et al. Epigenetic footprint enables molecular risk stratification of hepatoblastoma with clinical implications. J Hepatol. 2020;73:328–41.
Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B, et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res. 2008;68:6779–88.
Lee JS, Heo J, Libbrecht L, Chu IS, Kaposi-Novak P, Calvisi DF, et al. A novel prognostic subtype of human hepatocellular carcinoma derived from hepatic progenitor cells. Nat Med. 2006;12:410–6.
Leone V, Ali A, Weber A, Tschaharganeh DF, Heikenwalder M. Liver inflammation and hepatobiliary cancers. Trends Cancer. 2021;7:606–23.
Wang X, Sun W, Shen W, Xia M, Chen C, Xiang D, et al. Long non-coding RNA DILC regulates liver cancer stem cells via IL-6/STAT3 axis. J Hepatol. 2016;64:1283–94.
Naugler WE, Sakurai T, Kim S, Maeda S, Kim K, Elsharkawy AM, et al. Gender disparity in liver cancer due to sex differences in MyD88-dependent IL-6 production. Science. 2007;317:121–4.
Rui J, Liu H, Zhu X, Cui Y, Liu X. Epigenetic silencing of CD8 genes by ThPOK-mediated deacetylation during CD4 T cell differentiation. J Immunol. 2012;189:1380–90.
Srivastava A, Mishra RK. Interactome of vertebrate GAF/ThPOK reveals its diverse functions in gene regulation and DNA repair. J Biosci. 2020;45:38.
Li Y, Tsun A, Gao Z, Han Z, Gao Y, Li Z, et al. 60-kDa Tat-interactive protein (TIP60) positively regulates Th-inducing POK (ThPOK)-mediated repression of eomesodermin in human CD4+ T cells. J Biol Chem. 2013;288:15537–46.
Zhang M, Zhang J, Rui J, Liu X. p300-mediated acetylation stabilizes the Th-inducing POK factor. J Immunol. 2010;185:3960–9.
Sabari BR, Tang Z, Huang H, Yong-Gonzalez V, Molina H, Kong HE, et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol Cell. 2015;58:203–15.
Zorro Shahidian L, Haas M, Le Gras S, Nitsch S, Mourao A, Geerlof A, et al. Succinylation of H3K122 destabilizes nucleosomes and enhances transcription. EMBO Rep. 2021;22:e51009.
Li J, Lu L, Liu L, Ren X, Chen J, Yin X, et al. HDAC1/2/3 are major histone desuccinylases critical for promoter desuccinylation. Cell Discov. 2023;9:85.
Wei W, Liu X, Chen J, Gao S, Lu L, Zhang H, et al. Class I histone deacetylases are major histone decrotonylases: evidence for critical and broad function of histone crotonylation in transcription. Cell Res. 2017;27:898–915.
Lee WS, Lavery L, Rousseaux MWC, Rutledge EB, Jang Y, Wan YW, et al. Dual targeting of brain region-specific kinases potentiates neurological rescue in Spinocerebellar ataxia type 1. EMBO J. 2021;40:e106106.
Vacchio MS, Wang L, Bouladoux N, Carpenter AC, Xiong Y, Williams LC, et al. A ThPOK-LRF transcriptional node maintains the integrity and effector potential of post-thymic CD4+ T cells. Nat Immunol. 2014;15:947–56.
Zhong C, Yin Q, Xie Z, Bai M, Dong R, Tang W, et al. CRISPR-Cas9-mediated genetic screening in mice with haploid embryonic stem cells carrying a guide RNA library. Cell Stem Cell. 2015;17:221–32.
Liu F, Song Y, Liu D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther. 1999;6:1258–66.
Wu Y, Cao Y, Xu K, Zhu Y, Qiao Y, Wu Y, et al. Dynamically remodeled hepatic extracellular matrix predicts prognosis of early-stage cirrhosis. Cell Death Dis. 2021;12:163.
Raudvere U, Kolberg L, Kuzmin I, Arak T, Adler P, Peterson H, et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019;47:W191–W198.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Shao Z, Zhang Y, Yuan GC, Orkin SH, Waxman DJ. MAnorm: a robust model for quantitative comparison of ChIP-Seq data sets. Genome Biol. 2012;13:R16.
Bekker-Jensen DB, Bernhardt OM, Hogrebe A, Martinez-Val A, Verbeke L, Gandhi T, et al. Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral libraries. Nat Commun. 2020;11:787.
Matthews CP, Birkholz AM, Baker AR, Perella CM, Beck GR Jr., Young MR, et al. Dominant-negative activator protein 1 (TAM67) targets cyclooxygenase-2 and osteopontin under conditions in which it specifically inhibits tumorigenesis. Cancer Res. 2007;67:2430–8.
Roessler S, Long EL, Budhu A, Chen Y, Zhao X, Ji J, et al. Integrative genomic identification of genes on 8p associated with hepatocellular carcinoma progression and patient survival. Gastroenterology. 2012;142:957–966.e912.
Hanzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7.
Acknowledgements
The authors acknowledge Dr. Lijian Hui (CEMCS, CAS, China) for kindly providing Alb-Cre mice, Dr. Lie Wang (Zhejiang University, China) for kindly providing Zbtb7bf/f mice, Dr. Jinsong Li (CEMCS, CAS, China) for helping with Zbtb7b-HA knockin mice, Dr. Xin Chen (UCSF, USA) for kindly providing myr-Akt/N-Ras G12V and SB plasmids, the Center for Single-Cell Omics (CSCOmics) Bioinformatics Core (Shanghai Jiao Tong University School of Medicine) for providing the bioinformatics and high-performance computing services, and Dr. Baojin Wu (CEMCS, CAS, China) for technical support. This work was supported by the Ministry of Science and Technology of China (2020YFA0803203 to GG), the National Natural Science Foundation of China (82372665, 32070789 to GG, 82373446 to L.C., 31900514 to YG), the Shanghai Municipal Science and Technology Major Project (to GG) and Research Grants (21DZ2201700, 23ZR1435800 to LC), and the innovative research team of high-level local universities in Shanghai (to LC). Zbtb7b-HA mice were supplied by the Genome Tagging Project (GTP) Center supported by the Shanghai Municipal Commission for Science and Technology Grant (19411951800).
Author information
Authors and Affiliations
Contributions
YZ, QW, CL, and GG designed the research; YZ, QW, XX, CM, YQ, YZ, YW, and YG performed the research; JJ, XL, and JC contributed new reagents/analytic tools; YZ, QW, XX, CM, and YQ analyzed the data; YZ, QW, CL, and GG wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study did not require ethical approval.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Yufang Shi
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, Y., Wang, Q., Xie, X. et al. ZBTB7B is a permissive regulator of hepatocellular carcinoma initiation by repressing c-Jun expression and function. Cell Death Dis 15, 55 (2024). https://doi.org/10.1038/s41419-024-06441-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06441-y