Abstract
Medulloblastoma is the most common malignant brain tumour in children. Genomic studies have identified distinct disease subgroups: wnt/wingless (WNT), sonic hedgehog (SHH), and non-WNT/non-SHH, comprising group 3 and group 4. Alterations in WNT and SHH signalling form the pathogenetic basis for their subgroups, whereas those for non-WNT/non-SHH tumours remain largely elusive. Recent analyses have revealed recurrent in-frame insertions in the E3 ubiquitin ligase adaptor Kelch Repeat and BTB Domain Containing 4 (KBTBD4) in cases of group 3/4 medulloblastoma. Critically, group 3/4 tumours with KBTBD4 mutations typically lack other gene-specific alterations, such as MYC amplification, indicating KBTBD4 insertion mutations as the primary genetic driver. Delineating the role of KBTBD4 mutations thus offers significant opportunities to understand tumour pathogenesis and to exploit the underpinning mechanisms therapeutically. Here, we show a novel mechanism in cancer pathogenesis whereby indel mutations in KBTBD4 drive its recognition of neo-substrates for degradation. We observe that KBTBD4 mutants promote the recruitment and ubiquitylation of the REST Corepressor (CoREST), which forms a complex to modulate chromatin accessibility and transcriptional programmes. The degradation of CoREST promoted by KBTBD4 mutation diverts epigenetic programmes inducing significant alterations in transcription to promote increased stemness of cancer cells. Transcriptional analysis of >200 human group 3 and 4 medulloblastomas by RNA-seq, highlights the presence of CoREST and stem-like signatures in tumours with KBTBD4 mutations, which extend to a further sub-set of non-mutant tumours, suggesting CoREST alterations as a novel pathogenetic mechanism of wide relevance in groups 3 and 4. Our findings uncover KBTBD4 mutation as a novel driver of epigenetic reprogramming in non-WNT/non-SHH medulloblastoma, establish a novel mode of tumorigenesis through gain-of-function mutations in ubiquitin ligases (neo-substrate recruitment) and identify both mutant KBTBD4 and CoREST complexes as new druggable targets for improved tumour-specific therapies.
Similar content being viewed by others
Introduction
Cullin-RING Ligases (CRLs) represent a class of multi-subunit E3s, in which the cullin subunit forms a central scaffold to recruit interchangeable adaptors for substrate recruitment and ubiquitin-charged E2s for substrate ubiquitylation. The prototypical CRL1s assemble with Cul1, Skp1 and a variable substrate recruitment subunit called an F-box protein to form a Skp1-Cul1-F-box (SCF) ubiquitin ligase [1,2,3]. In SHH medulloblastoma, the substrate adaptor F-box/LRR-repeat protein 17 (Fbxl17) assembles into a functional SCFFbxl17 to promote the ubiquitylation-dependent degradation of the tumour suppressor Suppressor of fused homologue (SUFU), acting as a bona-fide oncoprotein [4]. However, the function of other CRL adaptors in medulloblastoma is largely unexplored.
Recurrent heterozygous in-frame insertion mutations in the Cullin RING ubiquitin Ligase 3 (CRL3) adaptor KBTBD4 are reported in children with non-WNT/non-SHH medulloblastoma [5], as well as in children and adults with Pineal Parenchymal Tumours of Intermediate Differentiation (PPTID) [6]. Recent analyses also reveal recurrent in-frame insertions in KBTBD4 in 40% of cases of adult group 3 medulloblastomas [7].
KBTBD4 forms a multi-subunit E3 using a BTB domain to engage Cul3 and a Kelch domain to recruit substrates. KBTBD4 mutations recur around the same conserved residues (G308 to R313) in the Kelch domain (Fig. 1A and Supplementary Fig. S1). CRL3 adaptors have critical roles in cancer pathogenesis and hotspot inactivating mutations have been identified in the substrate recruitment domain of Kelch-like protein 6 (KLHL6), favouring the growth of diffuse large B-cell lymphoma [8].
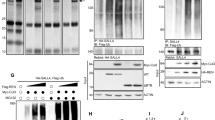
A Domain organisation of KBTBD4. The Kelch domain contains 6 Kelch repeat motifs (I-VI) that form the six blades of a β-propeller fold. Recurrent indel mutations KBTBD4R313PRR and KBTBD4P311PP map to the second Kelch repeat. B HEK293T cells transfected with empty vector (EV), Flag-KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP were immunoprecipitated and eluted with Flag-peptide. Eluates were digested and analysed by LC-MS/MS. Cells were treated with 100 nM MLN4924 for 4 h before harvesting. The table highlights interactors from the CRL3KBTBD4 and CoREST complexes (red and light blue shading, respectively). C Structural model of the KBTBD4 Kelch domain (left). Red star indicates the substrate-binding pocket. Indel mutations are predicted to lengthen the substrate-binding loop in blade II to support neo-substrate interaction (right). The inserted mutant amino acids (denoted by an arrow) lie on the outer edge of the loop that faces away from the central substrate pocket. N-terminal wild-type residues preceding the mutation site are therefore repositioned into the substrate pocket (Ile310 Cα atoms in each model are shown as spheres for reference). D HEK293T cells were co-transfected with HA-tagged CoREST and an Empty Vector, Flag-KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4R311PP and were treated with MLN4924. Whole-cell extracts were immunoprecipitated (IP) with anti-FLAG resin, and immunoprecipitates were immunoblotted as indicated. E HEK293T cells were co-transfected with HA-CoREST along with Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4R312RG as indicated (+). After treatment with MLN4924 cell lysates were immunoprecipitated by anti‐FLAG resin and immunoblotted as indicated. F HEK293T cells were co-transfected with MYC‐tagged ubiquitin and HA-CoREST along with Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP as indicated (+). HA‐CoREST was immunoprecipitated with anti‐HA resin under denaturing conditions and immunoblotted. Cells were treated with 5 μM MG132 for 4 h before harvesting. The poly-ubiquitylated CoREST (indicated by brackets) was detected by immunoblot with anti-Myc antibody.
Here, we present the first prototypical example of neomorphic mutation in an E3 ubiquitin ligase. We show that medulloblastoma-associated hotspot mutations in the Kelch domain of KBTBD4 promote the recruitment of CoREST as a neo-substrate for ubiquitylation and degradation. CoREST1/2/3 form critical scaffolds for chromatin modifying complexes, which incorporate lysine-specific histone demethylase 1A (LSD1) and histone deacetylase 1/2 (HDAC1/2) for the demethylation and deacetylation of histones at H3K4 in vitro and in vivo [9, 10]. We observe that degradation of CoREST by mutant KBTBD4 leads to increases in histone H3 methylation at lysine 4 (H3K4 mono- and di-methylation) and altered transcription. We further show that mutant KBTBD4 activates the transcription of CoREST targets favouring stemness, known to promote tumorigenesis in medulloblastoma [11].
Results
Medulloblastoma-associated KBTBD4 mutations drive recognition of the CoREST complex
As an initial strategy to identify substrates of KBTBD4, we immunoprecipitated KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP (the most recurrent mutations in group 3 and 4 medulloblastomas, respectively) (Fig. 1A) [5] from HEK293T cells and performed tryptic digestion and Liquid Chromatography/Tandem Mass Spectrometry (LC-MS/MS) to analyse their interacting partners. We identified specific components of the CRL3 machinery in KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP immunoprecipitates, including Cul3 and Rbx1. Strikingly, we could detect CoREST complex subunits only in KBTBD4R313PRR and KBTBD4P311PP immunoprecipitates, but not in that of KBTBD4WT, suggesting a putative mutant-specific neofunction (Fig. 1B). A Venn diagram comparing differential interacting partners of KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP identified in LC-MS/MS is presented in Supplementary Fig. S2A.
While the mutations in KBTBD4 are different in terms of amino acid composition and insertion length, the same CoREST complex was identified in the LC-MS/MS of different mutants, including CoREST, LSD1, HDAC1/2, Genetic Suppressor Element 1 (GSE1) and High Mobility Group 20A (HMG20A/B) (Fig. 1B). Using known Kelch domain structures [12,13,14], we mapped the 6 Kelch repeats in KBTBD4 and constructed 3D structural models for KBTBD4WT and oncogenic mutants. The recurrent indel mutations KBTBD4R313PRR and KBTBD4P311PP map to the same Kelch domain loop region for substrate recruitment (Fig. 1A). Here, the in-frame insertions are predicted to lengthen the substrate-binding loop to support neo-substrate interaction (Fig. 1C). Importantly, the inserted mutant amino acids lie on the outer edge of the loop that faces away from the central substrate pocket. It is therefore the N-terminal wild-type residues preceding the mutation site that are predicted to shift position to engage the CoREST substrates. This structural model therefore presents a possible explanation for how different in-frame insertions can modify the substrate-binding surface to promote similar neo-substrate recruitment (Fig. 1C).
To confirm the LC-MS/MS data and our model of neo-substrate recruitment, we first tested the interaction between KBTBD4WT, KBTBD4R313PRR, KBTBD4P311PP and CoREST by co-expressing these proteins in HEK293T cells. We could co-immunoprecipitate the CoREST complex with both KBTBD4R313PRR and KBTBD4P311PP, but not KBTBD4WT (Fig. 1D). This interaction was increased by treating the cells with MLN4924, which inhibits the activity of Nedd8-Activating Enzyme (NAE1). Neddylation of Cullins is required for the activity of CRL complexes; MLN4924 therefore blocks all CRL activity in cells and stabilises CRL-bound substrates [15]. Thus, the increased interaction between mutant KBTBD4 and CoREST in the presence of MLN4924 reflects the fact that KBTBD4R313PRR and KBTBD4P311PP can interact with CoREST, but not ubiquitylate or degrade it due to the inactivation of CRL3KBTBD4 by MLN4924 (Fig. 1D). Of note, KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP all bound equivalently to Cul3, showing that changes in CRL3 assembly were not related to the substrate specificity of KBTBD4 variants (Fig. 1D). Interaction of exogenously expressed CoREST was also observed with KBTBD4R313PRR and KBTBD4P311PP but not KBTBD4WT after immunoprecipitation of CoREST (Supplementary Fig. S2B).
To further validate our model, we tested the interaction of another tumorigenic mutant of KBTBD4, namely KBTBD4R312RG. Again we detected interaction of KBTBD4R312RG with CoREST at comparable levels to the interaction detected with KBTBD4R313PRR (Fig. 1E). To confirm the formation of functional CRL3KBTBD4 directly, and to test whether CoREST is a genuine substrate, we measured the ubiquitylation activity of CRL3KBTBD4 on CoREST. Compared to empty vector background, both KBTBD4R313PRR and KBTBD4P311PP promoted ubiquitylation of CoREST, whereas KBTBD4WT did not, in accordance with the interaction data (Fig. 1F). The CoREST interaction with KBTBD4P311PP appeared weaker than with KBTBD4R313PRR, and the ubiquitylation levels of CoREST were consistent with the extent of interaction (Fig. 1D, F). We also observed that CoREST ubiquitylation was prevented by MLN4924 (Supplementary Fig. S2C) as predicted from the results of immunoprecipitation analysis (Fig. 1D).
Expression of KBTBD4 mutants in medulloblastoma cancer cells induces CoREST degradation and epigenetic rewiring
The experiments presented above used HEK293T cells. We subsequently derived several different medulloblastoma cell lines stably expressing KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP to confirm our findings in more physiologically relevant models. In DAOY medulloblastoma cell lines, we could detect interaction of endogenous CoREST with KBTBD4P311PP, but not with KBTBD4WT as in HEK293T (Supplementary Fig. S3A). Expression of mutant KBTBD4R313PRR or KBTBD4P311PP, but not KBTBD4WT, promoted a drastic reduction in CoREST levels, which could be restored by treating cells with the proteasome inhibitor MG132 (Supplementary Fig. S3B), showing that CoREST is targeted for proteasomal degradation by KBTBD4 mutants, but not KBTBD4WT, in accordance with the interaction data. MLN4924 similarly prevented the degradation of CoREST in DAOY cell lines expressing KBTBD4R313PRR or KBTBD4P311PP, demonstrating a role for CRL activity in controlling the levels of CoREST (Supplementary Fig. S3C).
While DAOY cells are the most cited medulloblastoma cell-based model, they do not represent the non-WNT/non-SHH medulloblastoma classes in which the KBTBD4 mutations are detected. We therefore sought to confirm our observations in other more representative cells. Medulloblastoma cell lines derived in the past 4 decades (n > 40), collectively show a surprising lack of genetic diversity and medulloblastoma cell lines containing mutant KBTBD4 have not been reported to date [16]. To establish suitable models to investigate KBTBD4, we assessed endogenous CoREST and LSD1 protein levels in five representative medulloblastoma cell lines of different subgroups: D283Med (group 3/4, without reported MYC amplification), HD-MB03 (group 3, MYC amplification), D425Med (group 3, MYC amplification), D458Med (metastatic counterpart of D425Med primary tumour) as well as DAOY (SHH subgroup). All lines showed detectable CoREST and LSD1 proteins (Supplementary Fig. S3D). We used lentiviral expression to stably introduce KBTBD4WT, KBTBD4R313PRR or KBTBD4P311PP in the group 3 medulloblastoma cell line D425Med. Cells expressing mutant KBTBD4 showed a reduction in CoREST protein level, whereas these levels were unchanged by expression of KBTBD4WT (Supplementary Fig. S3E), consistent with data from other medulloblastoma cell lines.
We reasoned that the D283Med cell line represented a particularly suitable model to investigate the role of KBTBD4 mutations in medulloblastoma as, in terms of transcriptional identity, D283Med are close to the identified subtype II of a more recent medulloblastoma classification of group 3 and 4 subtypes [17]. We created D283Med and D425Med lines in which the expression of KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP was controlled by addition of doxycycline. After induction of KBTBD4WT, KBTBD4R313PRR or KBTBD4P311PP expression in D283Med, we measured their respective interaction with endogenous CoREST complex. We detected interaction of CoREST and LSD1 with KBTBD4R313PRR and KBTBD4P311PP exclusively in cells treated with MLN4924 where the ubiquitylation activity of CRL3 was prevented (Fig. 2A).
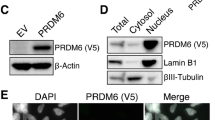
A D283Med cells expressing Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP after doxycycline induction (1 µg/mL for 40 h) were treated with 100 nM MLN4924 for 4 h as indicated (+). After lysis, extracts were immunoprecipitated with Flag-resin and immunoblotted as indicated. B D283Med cells expressing Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP were treated with doxycycline (1 µg/mL) for 16 h and lysed. Endogenous ubiquitylated proteins were isolated using GST fused ubiquitin binding domain UBAUbq and separated by SDS-PAGE. Isolated proteins were detected by immunoblot as indicated. All samples were treated with MG132 for 6 h. C D283Med cells containing Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP were treated with doxycycline (1 µg/mL) for the indicated time (h = hours). Samples were separated by SDS-page and immunoblotted as indicated. D, E Flow cytometry analysis. D (Top three panels) Representative images showing the gating procedure where Negative and Positive gates were determined using 5 controls: Compensation beads (green), unstained cells (orange), cells stained only with secondary antibody (not shown), cells treated with LSD1 inhibitor followed by incubation with either isotype (Blue) or H3K4me2 (red) primary antibodies and secondary antibody. The remaining panels are representative images showing the results related to flow cytometry analysis of H3K4me1 and H3K4me2 in D283Med cells expressing Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP after doxycycline induction (1 µg/mL for 4 days). E Bar charts showing the quantification of the analysis in D, plotted as percentage increase of the Mean Fluorescence Intensity (MFI) of positive cells where controls (- doxy) were set as 100% (dashed line). Data are presented as mean ±standard deviation (SD), with at least three independent experiments. P values (*P < 0.05 and **P < 0.005) were calculated by paired and two‐tailed t‐test.
To assess the capacity of KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP to ubiquitylate endogenous CoREST in the absence of overexpressed ubiquitin, we isolated endogenous ubiquitylated proteins using a GST fused ubiquitin binding domain UBAUbq previously reported [18] and detected the ubiquitylation formed on endogenous CoREST. KBTBD4R313PRR and KBTBD4P311PP promoted ubiquitylation of endogenous CoREST, while ubiquitylation of CoREST in parental D283Med cells or in the presence of KBTBD4WT was undetectable (Fig. 2B).
To determine the fate of ubiquitylated CoREST, we measured CoREST levels after addition of doxycycline. At comparable expression levels of KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP we observed a reduction in the levels of CoREST only upon expression of KBTBD4 mutants, consistent with CoREST degradation (Fig. 2C). Despite similar or higher levels of expression, KBTBD4WT did not affect CoREST levels. To exclude that the effect observed on CoREST was due to overexpression artefacts of KBTBD4, we tested the levels of CoREST after inducing KBTBD4WT and KBTBD4P311PP with increasing concentrations of doxycycline. Minimal induction of KBTBD4P311PP with the lowest doxycycline dose still promoted CoREST degradation (Supplementary Fig. S3F). By contrast, the highest expression of KBTBD4WT did not alter CoREST levels (Supplementary Fig. S3F). In addition to CoREST, LSD1 was also degraded, although at a seemingly reduced level compared to CoREST (Supplementary Fig. S3F).
CoREST exerts its activity through LSD1, which is known to demethylate mono- and di-methylated H3K4 in vitro and in vivo [9, 10]. We assessed the impact of KBTBD4R313PRR and KBTBD4P311PP expression on the chromatin modifications mediated by CoREST and observed an increase of H3K4 mono- and di-methylation in accordance with loss of CoREST and LSD1 activity (Fig. 2D, E). The increase in the epigenetic mark H3K4me2 was also confirmed by Western blotting in D283Med and DAOY cell lines (Supplementary Fig. S3G, H). Overall, the data presented highlight that KBTBD4R313PRR and KBTBD4P311PP can promote the degradation of CoREST complex and increase H3K4 mono- and di-methyl epigenetic marks. The data also exclude that KBTBD4WT targets CoREST under physiological conditions.
Expression of KBTBD4 mutants in medulloblastoma cells promotes the de-repression of CoREST target genes activating pro-tumorigenic programmes
The impact of mutant KBTBD4 on epigenetic markers suggested the potential for altered transcription through de-repression of CoREST targets. To identify the transcriptional programmes promoted by mutant KBTBD4, we compared total RNA-seq samples in quadruplicates from D283Med cells where expression of KBTBD4WT, KBTBD4R313PRR or KBTBD4P311PP was promoted by doxycycline versus untreated controls (Supplementary Fig. S4A, B). Although we detected a significant induction of KBTBD4WT, a volcano plot and heatmap of differentially expressed genes revealed that KBTBD4WT did not promote significant transcriptional changes (Fig. 3A and Supplementary Fig. S4B). By contrast, the expression of KBTBD4R313PRR and KBTBD4P311PP promoted increased expression of a specific genes, of which many were known CoREST target genes. Among 249 significantly upregulated genes, 130 genes in the KBTBD4R313PRR sample and 77 in KBTBD4P311PP were known CoREST targets (Fig. 3B, C and Supplementary Table S1). An increase in gene expression is expected following the loss of a transcriptional repressor like CoREST and establishes that mutant KBTBD4 alters transcription by controlling CoREST complex activity.
Volcano plots showing the differential gene expression analysis from D283Med cells expressing (A) Flag‐KBTBD4WT, (B) Flag-KBTBD4R313PRR or (C) Flag-KBTBD4P311PP treated with doxycycline (1 µg/mL) for four days compared to their untreated controls. Coloured dots represent differentially expressed genes between two groups with P-adjust <0.05. KBTBD4 (indicated in Purple) and known CoREST target genes (yellow) are highlighted. ChIP analysis of LSD1 occupancy on gene promoters in (D) D283Med and (E) D425Med cells expressing Flag‐KBTBD4WT or Flag-KBTBD4R313PRR upon doxycycline (1 µg/mL) treatment for 3 days. Data are expressed as percentage of input. F, G Dotplot of KEGG pathway enrichment analysis of genes differentially upregulated comparing either KBTBD4R313PRR or KBTBD4P311PP treated with doxycycline versus untreated controls. The X axis shows the gene ratio which is the percentage of genes enriched in a KEGG term. The Y axis represents the enriched pathways; the size of the node represents the number of enriched genes within a particular term.
To confirm that the transcriptional alterations detected are consistent in different models we also used the D425Med cell line, where expression of KBTBD4R313PRR or KBTBD4P311PP faithfully leads to CoREST degradation (Supplementary Fig. S2E). Using quantitative PCR (qPCR) in cells expressing KBTBD4R313PRR or KBTBD4P311PP we could detect the upregulation of multiple CoREST targets (Supplementary Fig. S4C), confirming the role of mutant KBTBD4 in different model cell lines.
Furthermore, to establish a direct relationship between alterations in gene transcription and CoREST complex ubiquitylation by KBTBD4R313PRR or KBTBD4P311PP, we performed Chromatin Immuno-Precipitation (ChIP) of LSD1 in D283Med and D425Med cell lines (Fig. 3D, E). Since LSD1 is also present in other chromatin remodelling complexes besides the ones formed by CoREST, we reasoned that, by using LSD1 ChIP, we could detect the recruitment and activity of the CoREST complex at the CoREST target genes in an unbiased way (i.e. independently of CoREST degradation). While in KBTBD4WT expressing cells we did not detect a difference, both D425Med and D283Med lines expressing KBTBD4R313PRR had significantly lower chromatin recruitment of LSD1 at four out of four CoREST target genes (Fig. 3D, E). Interestingly, the changes observed in gene expression after induction of KBTBD4 mutants could be in part recapitulated by siRNA mediated depletion of CoREST, reinforcing the idea that KBTBD4 mutants are mainly acting through CoREST depletion (Supplementary Fig. S3D).
To identify the transcriptional programmes contributing to tumorigenesis induced by KBTBD4R313PRR and KBTBD4P311PP expressing cells we also performed Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway analysis. We identified several cancer-related pathways (PI3K, RAP1 and proteoglycans in cancer) significantly upregulated in cells expressing KBTBD4 mutants (Fig. 3F, G) in accordance with pro-tumorigenic features. In addition, we detected a significant enrichment of pro-neurogenic programmes including axon guidance and glutamatergic synapses. While these programmes do not favour proliferation, they may be indicative of progenitor programmes specifying cellular identities favoured to grow in the cerebellum. Overall, the experiments presented above show that KBTBD4 mutants activate transcription by modulating CoREST complex occupancy and activity on chromatin, enabling the activation of specific cellular programmes for tumorigenesis and neurogenesis.
KBTBD4 mutants enhance stemness of medulloblastoma cancer cells
To establish a causal link between tumorigenesis and KBTBD4R313PRR or KBTBD4P311PP expression, we measured cell proliferation of D283Med and D425Med after induction of KBTBD4WT, KBTBD4R313PRR or KBTBD4P311PP. Surprisingly, we did not detect a significant difference in the proliferation of cells (Supplementary Fig. S5A), indicating that other features enable KBTBD4 mutants to promote tumorigenesis. In mice, genetic depletion of CoREST leads to the expansion of a progenitor stem cell population in the ventricular zone of the mouse brain with embryonic lethality [19]. Therefore, we hypothesised that CoREST degradation induced by KBTBD4 mutations could result in increased stemness of medulloblastoma cells, as described in other tumour types [20]. This hypothesis was supported by our RNA-seq data, wherein the expression of KBTBD4R313PRR or KBTBD4P311PP resulted in the enrichment of ES-like signature genes (Fig. 4A, B). Upon induction of KBTBD4R313PRR or KBTBD4P311PP, cells expressed genes controlled by pluripotency factors such as SRY-Box Transcription Factor 2 (Sox2), Octamer-Binding Protein 4 (OCT4) and Homeobox Transcription Factor Nanog (Nanog) (Fig. 4A, B and Supplementary Table S2). In addition, Gene Set Enrichment Analysis (GSEA) showed that epithelial to mesenchymal transition (EMT), a fundamental pathway for vertebrate embryonic development and often linked to cell migration and metastatic potential of cancer cells [21], was significantly enriched in cell lines expressing either KBTBD4 mutant (Supplementary Fig. S5B). In adherent medulloblastoma cells, we also could detect an increased migration of cells expressing KBTBD4P311PP (Supplementary Fig. S5C), supporting our GSEA analysis.
Volcano plot showing upregulated ES-like signature genes (yellow dots). RNA-seq upregulated genes (P-adjust <0.05) between (A) KBTBD4R313PRR and (B) KBTBD4P311PP treated with doxycycline compared to untreated samples. C Tumorsphere-forming capacity of D283Med cells in which KBTBD4WT, KBTBD4R313PRR or KBTBD4P311PP expression was induced with doxycycline (doxy+) compared to untreated controls (doxy−). Quantification of 3 independent experiments (bar chart) and representative pictures of doxycycline-induced cells (scale bar denotes 100 µm). D Aldehyde dehydrogenase (ALDH) activity, measured as percentage of ALDH positive cells, of D425Med parental cells or, D425Med cells expressing KBTBD4WT, KBTBD4R313PRR or KBTBD4P311PP+/− LSD1i treatment (0.5 µM for 24 h). [N,N-diethylaminobenzaldehyde (DEAB), ALDH inhibitor]. P values (**P < 0.005 and ****P < 0.00005) were calculated by paired and two‐tailed t‐test.
Most importantly, we assessed the effect of KBTBD4WT, KBTBD4R313PRR and KBTBD4P311PP expression on stemness using two well established assays for cancer stem cells, neurosphere formation ability and ALDH activity [22, 23]. Previous work showed that medulloblastoma initiating cells isolated based on ALDH activity had higher capacity for neurosphere formation, as well as higher expression of neural stem cell markers [24]. Additionally, these medulloblastoma initiating cells showed higher expression of pluripotency related genes and were able to be differentiated into multiple brain cell lineages [24]. Expression of KBTBD4R313PRR and KBTBD4P311PP induced increased neurosphere formation versus control D283Med cells, whereas expression of KBTBD4WT did not (Fig. 4C). Note that the increased tumorsphere formation can’t be ascribed to increased cell proliferation as this remains unchanged (Supplementary Fig. S5A).
In a second assay to measure stemness, we observed that KBTBD4R313PRR and KBTBD4P311PP expression more than doubled the percentage of D283Med cells presenting with high ALDH activity compared to cells expressing KBTBD4WT (Supplementary Fig. S5D). KBTBD4R313PRR and KBTBD4P311PP expression in D425Med cells also led to increased ALDH activity (Fig. 4D), showing that the stemness phenotype induced by KBTBD4R313PRR and KBTBD4P311PP overcomes MYC amplification present in the D425Med lines. More importantly, chemical inhibition of LSD1 (LSD1i) was found to mimic this phenotype and the combination of KBTBD4 mutant plus LSD1i did not show an additive effect (Fig. 4D and Supplementary Fig. S5C), showing that the increased stemness caused by KBTBD4R313PRR and KBTBD4P311PP is exerted via modulation of CoREST complex activity. Overall, our findings establish a potential mechanism of tumorigenesis through increased stemness in KBTBD4 mutant tumours.
KBTBD4 transcriptional signatures define a specific cluster of medulloblastomas
We next investigated the relevance of transcriptional programmes driven by KBTBD4R313PRR and KBTBD4P311PP in a large series of primary medulloblastomas. Using single sample Gene Set Enirchment Analysis (ssGSEA) of 223 primary MBGroup3/4 tumours (Fig. 5A), interrogation of KBTBD4 mutation-associated gene signatures identified in our models (including CoREST and ESC-related pathways) distinguished two clusters within these primary tumours. The first cluster displayed gene-set expression signatures similar to the KBTBD4WT models, while signatures of the second tumour cluster were similar to those of the KBTBD4 mutant models. The second cluster further resolved into two sub-clusters (a,b), with sub-cluster ‘a’ displaying the strongest mutation-associated signatures. As hypothesised, all KBTBD4 mutated primary tumours in the cohort (n = 3) resided in cluster 2 and demonstrated upregulation of KBTBD4 mutation-associated genesets. Cluster 2 represents 60% (134/223) of all MBGroup3/4 tumours examined, indicating a wider involvement of KBTBD4 mutation-associated transcriptional programmes and CoREST pathways in this tumour group, with KBTBD4-mutated cases representing a subset (3/134) of these cases. Moreover, whilst cluster 2 tumours are seen in each molecular subgroup and subtype, they are particularly frequent in MBGroup4 tumours (71% (110/156) of MBGroup4), and their associated subtypes (V-VIII) (Fig. 5B).
A Heatmap showing single sample Gene Set Enrichment Analysis (ssGSEA) scores for D283Med cell-based models and human primary medulloblastomas (Group3; n = 67, Group4; n = 156). Custom genesets represented are dysregulated by doxocycline-induced expression of KBTBD4 mutants (R313PRR and P311PP) in D283Med cell lines. + indicates doxycycline treatment, - indicates untreated controls. B Stacked bar plots representing percentage frequency of gene signature clusters across different subgroups (Group3 and Group4) and subtypes (I-VIII) of primary medulloblastomas.
Discussion
We have identified a novel pathogenetic mechanism in non-WNT/non-SHH medulloblastoma unveiling an actionable target. We find that oncogenic mutations in the ubiquitin ligase KBTBD4 in group 3/4 medulloblastomas lead to the recruitment and ubiquitylation of epigenetic regulatory complexes containing CoRESTs and LSD1. The lack of CoREST1/2 promotes the hyperproliferation of neuronal progenitors in the developing mouse brain [19]. Therefore, we surmise that the degradation of CoRESTs by neomorphic KBTBD4 mutants induces epigenetic remodelling that blocks neuronal precursor differentiation in favour of progenitor expansion (Fig. 6). This model uncovers a novel driver of epigenetic reprogramming in paediatric brain tumours, establishes a novel mode of tumorigenesis through gain-of-function mutations in ubiquitin ligases (neo-substrate recruitment) and identifies a new druggable target for tumour-specific therapies.
During normal development, CoREST complexes containing LSD1, HDAC1 or 2, and CoRESTs (also known as RCORs) silence chromatin by deacetylating and demethylating histone H3 at position Lys4 (K4). In medulloblastoma, mutations allow KBTBD4 to destabilise CoREST complexes thereby preventing chromatin repression leading to the increased transcription of CoREST targets and stemness genes. The activation of stemness genes is a potential driver of medulloblastoma tumorigenesis.
While mutations in ubiquitin ligases classically act in cancer by disrupting substrate recruitment, this study has defined the first prototypic example where E3 mutations promote the recruitment of neo-substrates. The change of substrate specificity in KBTBD4 raises the intriguing possibility that neomorphic mutations could be present in other E3 ligases, which would further expand the variety of degrons participating in cancer development that have been identified through deep learning algorithms [25]. In addition to the broad implications this observation has for the study of E3 ligases in cancer, it highlights a gain–of–function mutant E3 for the potential targeting of PROTAC drug modalities with tumour cell-specific activity.
Remarkably, the KBTBD4 mutations phenocopy the effects of the investigational drug UM171 [26], which is currently investigated in phase II clinical trials [27]. The authors proposed that UM171 may potentiate a weak interaction of the CoREST/LSD1 complex with the endogenous KBTBD4WT. However, our data obtained with 4 different models of medulloblastoma and HEK293T cells did not confirm binding of CoREST to KBTBD4WT, nor its ubiquitylation, suggesting a neomorphic effect in medulloblastoma cells resulting from KBTBD4 mutation.
CoREST acts as a core scaffold to recruit epigenetic remodellers and certain DNA-binding factors. We observed significant differences in the epigenetic marks H3K4 mono- and di-methylation consistent with altered LSD1 activity. We also observed a strong correlation between the KBTBD4-dependent degradation of CoREST and the transcriptional activation of CoREST target genes. While CoREST was first described as a partner of the RE1-silencing transcription factor REST, it has since been shown to assemble independently [28], leading to changes in its DNA site preferences and repressor activity. The precise function of CoREST therefore depends on the CoREST partner genes expressed and the subcomplexes formed. It has been shown previously that Growth Factor Independent Protein 1 (GFI1) and GFI1B are transactivated by enhancers in group 4 medulloblastoma and form a complex with LSD1 and CoREST that promotes tumour growth [29]. These observations are in stark contrast with the example of KBTBD4 where the loss of CoREST by ubiquitin-mediated degradation promotes stemness. However, the identified complex recruited by KBTBD4 did not include GFI1 or GFI1B likely explaining their different mechanisms. While GFI1 and GFI1B overexpressing tumours show high LSD1 activity that could be blocked using specific chemical inhibitors [30], our findings show that inhibition of LSD1 increases stemness (Fig. 4D and Supplementary Fig. S5D), raising caution over the use of LSD1 inhibitors in the treatment of specific subtypes of medulloblastoma. Our data also indicate that KBTBD4 mutation leads cells into an intermediate state of differentiation, keeping ES-like and EMT gene signatures (Fig. 4A, B and Supplementary Fig. S5A) while expressing differentiated features of several synaptic pathways (Fig. 3D, E). This is in line with evidence showing that embryonic stem cells lacking LSD1 fail to fully differentiate [31].
Data from our human primary tumour cohorts support our cell-based findings. Human medulloblastomas with CoREST signatures showed an increase in expression of well-established stemness-associated transcription factor gene-sets (e.g. Nanog, Oct4, Sox2) (Fig. 5A). Moreover, these expression signatures define a cluster of tumours which include, but extend beyond, KBTBD4 mutant cases to characterise a wider disease group. Together, these initial findings suggest a wider involvement, and open further lines of investigation, of the role of CoREST in non-WNT/non-SHH medulloblastoma.
Neomorphic mutations are increasingly implicated as a molecular driver of tumorigenesis, including brain tumours. Perhaps the most established examples are the mutations found in in Isocitrate Dehydrogenase 1 (IDH1) and IDH2 that cause a major subset of gliomas. These mutations alter the enzyme activity to convert alpha-ketoglutarate into 2-hydroxyglutarate, which acts as an epigenetic remodeller to inhibit glioma stem cell differentiation [32]. The discovery of neomorphic mutations in KBTBD4 extends the spectrum of neomorphic mutations in cancer, highlighting KBTBD4 and the CoREST complex for further investigations of the origins and therapeutic opportunities in group 3/4 medulloblastomas.
Materials and Methods
Cell Lines
HEK293T cells were cultured in high glucose Dulbecco’s Modified Eagle’s Medium (GIBCO #41965039) with 5% Penicillin Streptomycin (ThermoFisher Scientific # 15070063) and 10% Fetal Bovine Serum (Sigma-Aldrich #F9665) inside a 5% CO2 incubator at 37 °C. D283Med (ATCC® HTB-185™) and DAOY (ATCC® HTB-186™) cell lines were purchased from ATCC. D425Med, D458Med and HDMB-03 were gifts from Professor Beth Coyle, the University of Nottingham. All medulloblastoma cells were cultured in Minimum Essential Medium (MEM) (Thermo Fisher Scientific #11095080) supplemented with 10% Fetal Bovine Serum (Sigma-Aldrich #F9665) inside a 5% CO2 incubator at 37 °C. D283Med and D425Med cells infected with pCW57.1 were cultured in MEM (Thermo Fisher Scientific #11095080) supplemented with 10% Tetracycline-Free Fetal Bovine Serum (PAN Biotech # Fetal Bovine Serum).
Plasmids and gene transfer
For studies conducted in HEK293T cells, full length human KBTBD4 (Uniprot Q9NVX7 isoform 1) was cloned into pcDNA3.1(+) with the 3xFLAG epitope tagged to its N-terminus using standard restriction enzyme cloning. The mutant KBTBD4 constructs (R313PRR and P311PP) were created in the same vector by site directed mutagenesis. DNA sequences were verified by Source Bioscience Ltd. CoREST1 (pCMV5 HA RCOR1; NM_015156.4) construct was purchased from the MRC Protein Phosphorylation and Ubiquitylation Unit (https://mrcppureagents.dundee.ac.uk). Myc-His-Ubiquitin plasmid was used as previously described [1, 2]. For studies conducted in D283Med and D425Med cell lines, Flag-KBTBD4 wild type and mutants were first cloned into pENTR3C (ThermoFisher Scientific #A10464) and then transferred through GATEWAY (Thermo Fisher Scientific #11791100) into pCW57.1 (Addgene #41393). For studies conducted in the DAOY cell line, Flag-KBTBD4 wild type and mutants were cloned into MIGR1 (Addgene #27490). pCW57.1 was a gift from David Root (Addgene plasmid # 41393; http://n2t.net/addgene:41393; RRID:Addgene_41393).
pCW57.1 and MIGR1 virus were produced in HEK293 cells using packaging plasmids psPAX2 and pMD2G and Transfection Reagent TransIT®-Lenti (MIR 6610). Media was replaced after 16 h and virus production was allowed for 48 h. Media containing virus was collected, filtered, supplemented with 10 ng/mL polybrene and medulloblastoma cells lines were infected. After 16 h, media was replaced by complete media and cells were allowed to recover for 48 h before antibiotic selection.
Sample preparation for mass spectrometry proteomics
Flag-KBTBD4 wild type or mutants along with an empty vector control were transfected into HEK293T cells at 60% confluency with PEI-MAX 40kDA (11 µg DNA with 66 µg PEI per 15 cm dish). 40 h post transfection, cells were treated with 100 nM MLN4924 (Calbiochem, 951950–33–7) for 4 h before harvesting. Cells were then washed with ice-cold PBS and lysed in lysis buffer (50 mM Tris pH 7.5, 150 mM NaCl, 10% glycerol, 0.1% NP-40, 1 mM EDTA and 5 mM MgCl2) containing protease inhibitors (Sigma-Aldrich), beta-glycerolphosphate, DTT, PMSF and okadaic acid. 25 mg protein extract per sample was precleared with protein A agarose (Invitrogen), and then immunoprecipitated by ANTI-FLAG M2 Affinity Gel (Sigma-Aldrich). Flag-KBTBD4 and its interactors were eluted with 3xFLAG peptide (Sigma-Aldrich). The FLAG eluates were first incubated with 5 mM DTT for 30 min for reduction and then with 20 mM iodoacetamide for 30 min for alkylation. Then the proteins in the eluates were precipitated by methanol/chloroform to remove organic impurities. Precipitated proteins were reconstituted in 20 mM HEPES pH 8, 8 M urea, and then diluted to 20 mM HEPES pH 8, 1 M urea before adding immobilised trypsin resin (Pierce 20230) for 16 h at 37 °C. Trypsin digestion was terminated by adding 1% TFA and trypsin resin removed by centrifugation. Tryptic-digested peptides were desalted using the SOLA HRP SPE cartridges (Thermo Fisher Scientific) and dried using a vacuum spinner (Thermo Fisher Scientific).
LC‐MS/MS and data analysis
Dried tryptic-digested peptides were re-constituted in 15 µL of LC-MS grade water containing 1% acetonitrile and 0.1% TFA. Samples were analysed by liquid chromatography-tandem mass spectrometry (LC-MS/MS) using a Dionex Ultimate 3000 UPLC coupled to a Q-Exactive mass spectrometer (Thermo Fisher Scientific). The peptides were loaded onto a trap column (PepMapC18; 300 µm × 5 mm, 5 µm particle size, Thermo Fischer Scientific) for 1 min at 20 μL/min flowrate. The loaded peptides were separated on a 50 cm-long chromatographic EasySpray column (ES803, Thermo Fischer Scientific) with a gradient of 2 to 35% acetonitrile in 0.1% formic acid and 5% DMSO at 250 nL/min flow rate for 60 min. The Q-Exactive was operated in a data-dependent acquisition (DDA) mode to automatically switch between full MS-scan and MS/MS acquisition. Survey-full MS scans were acquired in the Orbitrap mass analyser over an m/z window of 380–1500 and at a resolution of 70k (AGC target at 3e6 ions). Prior to MSMS acquisition, the top fifteen most intense precursor ions (charge state ≥ 2) were sequentially isolated in the Quad (m/z 1.6 window) and fragmented on the HCD cell (normalised collision energy of 28). MS/MS data were obtained in the orbitrap at a resolution of 17.5 K with a maximum acquisition time of 128 ms, an AGC target of 1e5 and a dynamic exclusion of 27 s.
The raw data were searched against the Human UniProt-SwissProt database (Jan 2018; containing 20,259 human sequences) using the Mascot data search engine. The search was carried out by enabling the Decoy function, whilst selecting trypsin as enzyme (allowing 1 missed cleavage), peptide charge of +2, +3, +4 ions, peptide tolerance of 10 ppm and MS/MS of 0.05 Da; #13 C at 1; Carboamidomethyl (C) as fixed modification; and Deamidated (NQ), Oxidation (M), as a variable modification. MASCOT outputs were filtered using an ion score cut off of 20 and a false discovery rate (FDR) of 1%. The mass spectrometry raw data included in this paper had been deposited to the Proteome eXchange Consortium via the PRIDE partner repository with the dataset identifier PXD031683 [3].
Immunoprecipitation and immunoblotting
HEK293T cells were transfected as indicated at 60% confluency (5 µg DNA per plate using PEI-MAX 40kDA at ratio 6:1). After 20 h, cells were treated with 0.1 µM MLN4924 (Calbiochem, 951950–33–7) for 4–5 h. Cells were washed twice with ice-cold PBS, harvested into lysis buffer (50 mM Tris pH 7.5, 150 mM NaCl, 10% glycerol, 0.1% NP-40, 1 mM EDTA and 5 mM MgCl2) containing protease inhibitors (Sigma-Aldrich), beta-glycerolphosphate, DTT, PMSF and okadaic acid, incubated on ice for 30 min and centrifuged for 30 minutes at 20 000 g at 4 °C. The supernatant was incubated with ANTI-FLAG M2 Affinity Gel (Sigma-Aldrich) or HA affinity gel (E6779–1ML, Sigma-Aldrich) at 4 °C for 1 h before being washed 4 times with lysis buffer. Co-immunoprecipitated samples were eluted using 2× LDS buffer (Life Technologies) supplemented with β-mercaptoethanol and boiled for 5 min at 95 °C.
D283med cells harbouring pCW57.1 KBTBD4 variants were induced with 1 µg/mL doxycycline for 40 hours before 6 h 0.1 µM MLN4924 (Calbiochem, 951950–33–7) treatment as indicated in the figure. For each dish, adherent cells were scraped off and combined with cells in suspension collected by centrifugation. Cells were washed and lysed the same way as described above. The supernatant was incubated with ANTI-FLAG M2 Affinity Gel (Sigma-Aldrich) at 4 °C for 3 h. After 4 times of washes with lysis buffer, co-immunoprecipitated samples were eluted in LDS buffer (Life Technologies) supplemented with β-mercaptoethanol and boiled for 5 min at 95 °C. Whole-cell lysates were obtained by lysing cells in 2xSB (625 mM Bis-Tris pH 6.8, 20% (w/v) glycerol, 4% (w/v) SDS), boiled, sonicated and protein concentration was evaluated using BCA protein kit (Thermo Fisher Scientific) in most cases. Cell lysate or immunoprecipitate was resolved in 10% Bis-Tris gels, transferred to nitrocellulose membranes (Millipore) and immunoblotted with corresponding antibodies. The same membranes were probed with Anti-GAPDH (Thermo Fisher Scientific MA5–15738) or Anti-alpha Tubulin (TU-02) antibody (Santa Cruz Biotechnology, # sc-8035) for loading control.
These techniques have been described previously and are detailed above [33,34,35]. Uncropped Western blots are provided in Supplementary Material.
Recombinant protein production of GST-UBAUbq
An expression plasmid encoding the UBA domain of Ubiquilin-1 (Q9UMX0, isoform 1, 536–589 a.a.) fused with N-terminal GST and C-terminal hexahistidine tags in pGEX6P1 was obtained as a gift from Prof Mads Gyrd-Hansen’s lab in the University of Oxford [4]. The plasmid was transformed into E.coli strain BL21(DE3)-R3-pRARE2. Cells were cultured in TB medium at 37 °C until OD600 reached 2. Recombinant UBAUbq expression was then induced by addition of 0.4 mM isopropyl β-D-1thiogalactopyranoside, followed by 18 h continuous shaking at 18 °C. Cells were harvested by centrifugation and lysed by sonication in binding buffer (50 mM HEPES pH 7.5, 500 mM NaCl, 5% glycerol, 5 mM imidazole) supplemented with 0.5 mM TCEP. UBAUbq was captured on nickel sepharose resin, washed with binding buffer and eluted by a stepwise gradient of 30–250 mM imidazole. The protein was further cleaned up by size exclusion chromatography using a HiLoad 16/60 S200 Superdex column buffered in 50 mM HEPES pH 7.5, 300 mM NaCl, 0.5 mM TCEP. Protein masses were confirmed by intact LC-MS mass spectrometry.
Ubiquitylation assay
HEK293T cells at 60% confluency were transfected with 1 µg HA-CoREST1, 1 µg Myc-Ubiquitin and 1 µg Flag KBTBD4 wild type or mutants along with empty vector control as indicated per 10 cm dish. 30 hours after transfection, cells were washed with ice-cold PBS once. Cells were then thoroughly lysed and boiled in 500 μL denaturing lysis buffer (2% SDS, 150 mM NaCl, 10 mM Tris-HCl pH 7.4, supplemented with protease inhibitors). After cooling down to room temperature, cell lysates were then subjected to sonication until losing viscosity. Lysates were then diluted 10 times with dilution buffer (10 mM Tris-HCl pH 7.4, 150 mM NaCl, 2 mM EDTA, 1% Triton x-100) and incubated at 4 °C for 30 min. The diluted lysates were centrifuged at 14,000 rpm for 10 min to remove insoluble debris. Supernatant was used for immunoprecipitation with HA affinity gel (E6779–1ML, Sigma-Aldrich, 10 μL beads per sample) overnight at 4 °C. HA affinity gel was collected via centrifugation at 200 × g for 90 s and washed with 1 mL wash buffer (10 mM Tris-HCl pH 7.4, 1 M NaCl, 1 mM EDTA, 1% NP-40) 4 times before being mixed with 50 μL LDS buffer (Life Technologies) and boiled. Results were analysed by immunoblotting (Anti-Flag antibody, Sigma-Aldrich #F1804; Anti-HA antibody, Roche #12013819001; Anti-Myc antibody, Cell Signaling Technology #9B11).
D283Med cells harbouring Flag-KBTBD4 variants were induced with 1 μg/mL doxycycline for 10 hours where indicated followed by 6 h of 5 μM MG132 treatment along with the parental D283Med cells. Cells were lysed in buffer containing 20 mM sodium phosphate buffer pH 7.4, 1% NP40, 2 mM EDTA supplemented with 5 mM N-ethylmaleimide, 50 μM PR619 and protease inhibitors. 150 µg/mL UBAUbq was incubated with glutathione sepharose 4B resin (GE Healthcare) for at least 1 hour and then the unbound protein was washed away.
Endogenous ubiquitylated proteins were pulled down by glutathione sepharose resin pre-bound with UBAUbq and eluted with LDS buffer. Results were analysed by immunoblotting (Anti-Flag antibody, Sigma-Aldrich #F1804; Anti-GAPDH antibody, Thermo Fisher Scientific MA5–15738; Anti-CoREST1, Cell Signaling Technology #14567).
RNA-sequencing
1 × 106 D283Med cells harbouring Flag‐KBTBD4WT, Flag-KBTBD4R313PRR or Flag-KBTBD4P311PP+/− doxycycline (1 µg/mL) were cultured for 4 days before harvesting. After washing once in phosphate-buffered saline, cell pellets were resuspended in TRIzol Reagent (Thermo Fisher Scientific cat # 15596026), snap frozen and preserved at −80 °C. RNA extraction, library preparation and sequencing were outsourced to NOVOGENE.
Small interfering RNAs transfection
Silencer® Select Small interfering RNAs (siRNA) (Thermo Fisher Scientific) targeting CoREST1 (siRNA 1 ID s23229 and siRNA 2 ID s23230) were transfected in D283Med and D425Med cells using HiPerFect Transfection Reagent (Qiagen). Cells were harvested 72 h after transfection and used for WB analysis of knockdown efficiency and Total RNA extraction for gene expression analysis.
Total RNA extraction and cDNA synthesis
Total RNA from medulloblastoma cell lines was isolated using RNeasy Plus Micro Kit (Qiagen, #74034) following the manufacturer’s protocol. RNA with an absorbance read 260/280 ratio between 1.8 and 2.0, assessed on a Nanodrop2000 spectrophotometer (Thermo Fisher Scientific), was used for reverse transcription. For every sample, 0.5 µg of total RNA was used for cDNA synthesis using SuperScript IV VILO Master Mix (Thermo Fisher Scientific, #11756050) following the manufacturer’s protocol.
qPCR
For real time gene expression quantification, PowerTrack™ SYBR Green Master Mix (Thermo Fisher Scientific, # A46113) was used. Specific pair of primers were designed for each gene transcript using Primer designing tool - NCBI (http://bioinfo.ut.ee/primer3–0.4.0/https://www.ncbi.nlm.nih.gov/tools/primer-blast/), and pair of primers analysis to predict secondary structures formation was done by using Beacon Designer tool (http://www.premierbiosoft.com/qOligo/Oligo.jsp?PID=1). The list of primers used in this study is reported below. All reactions were run in QuantStudio™ 5 Real-Time PCR System, 384-well (Thermo Fisher Scientific, #A28140). Each sample was tested in triplicate. Data analysis was performed with Design & Analysis software version 2.5.1 (Thermo Fisher Scientific). Relative changes in expression (i.e doxycycline treated relative to untreated controls) were evaluated by using the 2ΔCt formula.
Assay | Gene symbol | Accession Number | Forward primer 5’-3’ | Reverse primer 5’-3’ |
|---|---|---|---|---|
qPCR | ALPL | NM_001127501 | AACATCAGGGACATTGACGTG | GTATCTCGGTTTGAAGCTCTTCC |
AMIGO2 | NM_181847 | CCTGGGAACCTTTTCAGACTG | GCAAACGATACTGGAATCCACT | |
CLU | NM_001831 | CTACTTCTGGATGAATGGTGACC | CGGGTGAAGAACCTGTCCT | |
CLVS2 | NM_001010852 | TACACACTGGTGGATATTTTGCG | TTTAGAGGCTTGCTTGAAAGTGA | |
COL6A2 | NM_058174 | GAACGGGACCGATGGACAG | CCCTTGGCCCCGATTTCTC | |
HEPH | NM_001130860 | TGCGATATGAAGCCTTTCAAGAT | GGAGGCACGGTTGTAGAAGA | |
RAB3B | NM_002867 | CAACAGCCTATTACCGTGGGG | TAGCCCAGTCTTGGACAGCA | |
SCN1A | NM_001202435 | ATGTGGAAATAGCTCTGATGCAG | AGCCCAACTGAAGGTATCAAAG | |
SCN9A | NM_002977 | ATTCGTGGCTCCTTGTTTTCTG | CTACTGGCTTGGCTGATGTTAC | |
SFRP1 | NM_003012.5 | CTCCATAGCCACGCTCCAAA | TCTCACTTTCCGCCCAATCC | |
SPARC | NM_003118 | CCCATTGGCGAGTTTGAGAAG | CAAGGCCCGATGTAGTCCA | |
STC1 | NM_003155 | CACGAGCTGACTTCAACAGGA | GGATGTGCGTTTGATGTGGG | |
TGM2 | NM_004613 | CAAGGCCCGTTTTCCACTAAG | GAGGCGATACAGGCCGATG | |
TJP3 | NM_014428 | GCTTTGGCATTGCGATCTCTG | GATGTGGTCGCCTGTCTGTAG | |
ChIP-qPCR | CLU | NM_001831 | GAGCCAGCACAGCTATTCGT | CCAAAGAATGCCGCGGAAAG |
CLVS2 | NM_001010852 | CTTGCACACAAAGGGCGAAG | GGTAGGACGGCAATTTGGGT | |
COL6A2 | NM_058174 | CTGAGCAAGCCGGACACA | GACTCGCCCCTTGGTAGC | |
HEPH | NM_001130860 | AAGGGGAAGAGGTGGTGAGA | GGGGGTGGGTTGTACTTCTG | |
SCN9A | NM_002977 | TCGCTCCTACCAGCTCTGAA | GGGGAAAGAAACGTGGGGAA |
Chromatin immunoprecipitation
D283Med and D425Med cells harbouring Flag‐KBTBD4WT or Flag-KBTBD4R313PRR −/+doxycycline (1 µg/mL) were cultured for 3 days before the assay. Approximately 4 × 106 cells for each immunoprecipitation was processed using SimpleChIP® Plus Enzymatic Chromatin IP Kit (Cell Signalling Technology, #9005) following the manufacturer’s recommended protocol. We performed the optimisation of chromatin digestion recommended and 0.5 µL of Micrococcal Nuclease per IP prep was used (20 min at 37oC). Normal Rabbit IgG (#2729), Histone H3 (#4620) and LSD1 (#2184) antibodies from Cell Signalling Technology were used at the manufacturer’s recommended concentration. qPCR was performed and analysed as described above using 2 µL of purified ChIP DNA. Data were plotted as percentage of input.
Tumorsphere assays
Tumorsphere formation assays were performed as described previously [22]. Tumoursphere formation assays were performed in Ultra-Low Attachment 6-wells plates (Costar® #CLS3471) or cell-repellent surface 6-wells plates (Cellstar® #657970). D283Med cells were plated at a density of 5000 cells per well in triplicate, with or without doxycycline (1 µg/mL) in 1% methylcellulose containing media (NeuroCultTM #05751, supplemented with MethoCultTM #04100, STEMCELL Technologies). Tumourspheres were imaged and quantified 7 days after plating using a Celigo Image Cytometer (Nexcelom Bioscience LLC). Spheres with a diameter equal or higher than 40 μm were deemed tumourspheres. Experiments were repeated at least three times.
ALDH activity
ALDH activity was measured in intact D283Med cells using the ALDEFLUOR™ (STEMCELL Technologies, #01700) following the manufacturer’s protocol. One million D283Med cells were resuspended in 1 mL of Aldefluor assay buffer and mixed with 5 µL of BODIPY-aminoacetaldehyde (BAAA) ALDH substrate. Immediately after the addition of BAAA, half of the cells were transferred into a tube containing 10 µL of the ALDH inhibitor N,N-diethylaminobenzaldehyde (DEAB). After a 30-min incubation period, cells were pelleted to discard supernatant and then resuspended in 250 µL of Aldefluor assay buffer. Cells were analysed using a CytoFLEX LX Flow Cytometer, and results were generated using FlowJo software. Experiments were repeated at least three times.
Flow cytometry
For the detection of H3K4 mono and di-methylation in D283Med, cells were dissociated to single cells, pelleted and washed once in phosphate-buffered saline. Fixation and permeabilization were performed using the True Nuclear fixation kit (BioLegend, Cat #424401) following the manufacturer recommendations. Cells were then stained with rabbit anti-H3K4me1 (Cell Signaling Technology, Catalog #5326), rabbit anti-H3K4me2 (Cell Signaling Technology, Catalog #9725), normal rabbit IgG (Isotype control, sc-2027) and Alexa Fluor 594 (Thermo Fisher Scientific # A-21207). Flow cytometry analysis was performed using BD LSR Fortessa X20 (Becton Dickinson, San Jose, CA, USA) using the 488 nm laser (60 mW) for FSC (380 V) and SSC (212 V), and 561 nm laser (50 mW) using a 610/20 nm bandpass filter and a PMT voltage of 390 for AF594 excitation/detection. Results were generated using FlowJo software. Negative and Positive gates were determined using 5 controls: 1. Compensation beads (AbC™ Total Antibody Compensation Bead Kit, Thermo Fisher Scientific #A10513) composed by: a) polystyrene microspheres that have primary/secondary fluorescent antibody conjugate capture capacity (positive compensation beads) to define a clear positive signal and; b) inert beads (negative compensation beads) to provide a clear negative signal; 2. Unstained cells used to detect “auto-fluorescence” of the cells of interest and define negative gating; 3. Cells stained only with secondary antibody used to detect background staining and define negative gating; 4. Cells treated with LSD1i following incubation either with: a) isotype primary antibody and secondary antibody (used to define negative gating) or; b) H3K4me2 primary antibody and secondary antibody (used to define positive gating). Additionally, species and concentration matching isotype control was used for all primary antibodies and sample type.
Structural modelling
Homology models of the Kelch domain of KBTBD4 and its mutants were prepared using the ICM software package (Molsoft) [5] based on crystal structures of previously solved Kelch domains. The initial model was refined by energy minimisation and side chain optimisation in ICM-Pro (Molsoft).
Migration assay
A scratch wound assay was used to measure the migration capacity of DAOY cells overexpressing KBTBD4 wild type or mutants. Cells were seeded at 3 × 105 per well into a 12-wells plate in starvation medium with 1% FBS to form a confluent cell monolayer. Then, using a sterile 20 μL plastic pipette tip, two linear scratches (cross shape) were generated in each well. Media were carefully aspirated to remove floating cells generated by the scratch and replaced by a complete growth media (MEM supplemented with 10% FBS). Images were acquired every 2 h for a period of 24 h using a Zeiss LSM 780 confocal microscope equipped with live cell imaging incubator system.
Quantification and statistical analysis
RNA sequencing reads were aligned to the reference GRCh38 using HISAT2 v2.0.5. StringTie v1.3.1 was used to assemble transcripts and estimate transcript abundance in each sample. Gene FPKMs (fragments per kilo-base of exon per million fragments mapped) were computed by summing the FPKMs of transcripts in each gene group. Differential expression analysis was calculated using Ballgown at transcript level [36]. Transcripts with P-adjust < 0.05 were described as differentially expressed between any two groups. Gene level counts were computed using StringTie and DESeq2 v1.32.0R package was used for differential expression analysis [37]. KEGG pathway analysis were performed using the clusterProfiler v 3.13.0 [8] KEGG with Benjamini Hochberg (BH) corrected P value >0.05 were considered as significantly enriched. EnrichR was used to perform transcription factor analysis and identify CoREST and SOX2/NANOG/OCT4 lists of genes differentially expressed (upregulated) in KBTBD4 mutants. P values <0.05 were considered significant [38,39,40].
Analysis of human medulloblastoma tumours was carried out using RNA sequencing datasets [41]. For clustering and visualisation, read counts were normalised and a variance stabilising transformation applied using ‘vst’ function (DESeq2, R/Bioconductor). Single sample Gene Set Enrichment Analysis (ssGSEA) was performed using the implementation within GSVA (R/Bioconductor) and heatmaps produced using ‘pheatmap’ (R/Bioconductor). Mutations within KBTBD4 hotspot Kelch motif region (codons 308–313) were identified by direct PCR, Sanger sequencing and visual inspection of electropherograms.
Materials availability
Reagents generated in this study will be made available on request. A completed Materials Transfer Agreement may be requested if there is potential for commercial application.
Table of reagents
Reagent or resource | Source | Identifier |
|---|---|---|
Antibodies | ||
Anti-FLAG antibody | Sigma-Aldrich | Cat#F1804; RRID:AB_262044 |
Anti-FLAG M2 Affinity Gel | Sigma-Aldrich | Cat#A2220; RRID:AB_10063035 |
Anti-HA-Peroxidase antibody | Roche | Cat#12013819001, RRID:AB_390917 |
EZviewTM Red Anti-HA Affinity Gel | Sigma-Aldrich | Cat#E6779, RRID:AB_10109562 |
Anti-Myc antibody | Cell Signaling Technology | Cat#2276, RRID:AB_331783 |
Anti-LSD1 antibody | Cell Signaling Technology | Cat#2184, RRID:AB_2070132 |
CoREST1 (D6I2U) Rabbit mAb | Cell Signaling Technology | Cat#14567, RRID:AB_2798514 |
alpha Tubulin (TU-02) antibody | Santa Cruz Biotechnology | Cat# sc-8035, RRID:AB_628408 |
Mono-Methyl-Histone H3 (Lys4) (D1A9) Rabbit mAb | Cell Signaling Technology | Cat# 5326, RRID:AB_10695148 |
Di-Methyl-Histone H3 (Lys4) (C64G9) Rabbit mAb | Cell Signaling Technology | Cat# 9725, RRID:AB_10205451 |
Histone H3 (D1H2) XP® Rabbit mAb | Cell Signaling Technology | Cat# 4499, RRID:AB_10544537 |
Normal Rabbit IgG | Cell Signaling Technology | Cat#2729, RRID:AB_1031062 |
Normal rabbit IgG (Isotype control) | Santa Cruz Biotechnology | Cat# sc-2027, RRID:AB_737197 |
Donkey anti-Rabbit IgG (H + L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 594 | Thermo Fisher Scientific | Cat# A-21207, RRID:AB_141637 |
Anti-GAPDH antibody | Thermo Fisher Scientific | Cat#MA5–15738, RRID:AB_10977387 |
Anti-Cul3 antibody | Bethyl | Cat#A301–109A, RRID:AB_873023 |
Bacterial and virus strains | ||
Subcloning Efficiency™ DH5α Competent Cells | Thermo Fisher Scientific | Cat# 18265017 |
One Shot™ ccdB Survival™ 2 T1R Competent Cells | Thermo Fisher Scientific | Cat# A10460 |
One Shot™ Stbl3™ Chemically Competent E. coli | Thermo Fisher Scientific | Cat# C737303 |
Chemicals, peptides, and recombinant proteins | ||
3xFLAG peptide | Sigma-Aldrich | Cat#F4799 |
Immobilised trypsin | Pierce | Cat#20230 |
Neddylation inhibitor MLN4924 | Calbiochem | Cat#951950–33–7 |
Proteasome inhibitor MG132 | ApexBio Technology | Cat#A2585 |
Doxycycline | Sigma-Aldrich | Cat#D3447 |
Puromycin | Life technologies | Cat#A1113803 |
PR-619 | Selleck Chem | Cat#S7130 |
NEM | Sigma-Aldrich | Cat#E3876 |
TUBE Agarose resin | Lifesensors | Cat# UM-0402 |
GSK-LSD1 2HCl (LSD1 inhibitor) | Selleckchem | Cat#S7574 |
PEI Max 40KDA | Polysciences | Cat# 24765 |
HiPerFect Transfection Reagent | Qiagen | Cat# 301704 |
OptiMEM | GIBCO | Cat#31985070 |
NeuroCultTM Neural Cell Culture Media | STEMCELL Technology | Cat#05751 |
MethoCultTM | STEMCELL Technology | Cat#04100 |
MEM (Minimum Essential Medium) | Thermo Fisher Scientific | Cat#11095080 |
Dulbecco’s Modified Eagle’s Medium | GIBCO | Cat#41965039 |
Fetal Bovine Serum | Sigma-Aldrich | Cat#F9665 |
Fetal bovine serum, EU approved, Tetracycline Free | PAN Biotech | Cat# P30–3602 |
Critical commercial assays | ||
ALDEFLUOR™ | STEMCELL Technologies | Cat#01700 |
True Nuclear fixation kit | BioLegend | Cat # 424401 |
AbC™ Total Antibody Compensation Bead Kit | Thermo Fisher Scientific | Cat#A10513 |
Q5 Site-Directed Mutagenesis Kit | New England Biolabs | Cat# E0554S |
TransIT®-Lenti | MirusBio | Cat# MIR 6610 |
Gateway™ LR Clonase™ II Enzyme mix | Thermo Fisher Scientific | Cat#11791100 |
SimpleChIP® Plus Enzymatic Chromatin IP Kit | Cell Signaling Technology | Cat#9005 |
PowerTrack™ SYBR Green Master Mix | Thermo Fisher Scientific | Cat# A46113 |
RNeasy Plus Micro Kit | Qiagen | Cat#74034 |
SuperScript IV VILO Master Mix | Thermo Fisher Scientific | Cat#11756050 |
Deposited data | ||
Structures used for KBTBD4 Kelch domain homology modelling | PDB: 1U6D, 4ASC, 4CH9, 6GY5 | |
Experimental models: cell lines | ||
HEK293T | ATCC | Cat#CRL-3216; RRID:CVCL_0063 |
D283Med | ATCC | Cat# HTB-185, RRID:CVCL_1155 |
D425Med | Millipore | Cat# SCC290, RRID:CVCL_1275 |
D458Med | ||
HD-MB03 | DSMZ | Cat# ACC-740, RRID:CVCL_S506 |
DAOY | ATCC | Cat# HTB-186, RRID:CVCL_1167 |
Oligonucleotides | ||
KBTBD4 pcDNA3.1(+) fwd: CCGGCCGGATCCATGGAATCACCAGAGGAGCCTGG | Eurofins Genomics | N/A |
KBTBD4 pcDNA3.1(+) rev: GCATACGTCGACTTAGGCCAACACAAACTGCAAATTG | Eurofins Genomics | N/A |
KBTBD4 R313PRR site direct mutagenesis fwd: TTGTATGTGGTGGGAGGGTCCATCCCACGGCCACGGCGCATGTGGAAGT | Eurofins Genomics | N/A |
KBTBD4 R313PRR site direct mutagenesis rev: ACTTCCACATGCGCCGTGGCCGTGGGATGGACCCTCCCACCACATACAA | Eurofins Genomics | N/A |
KBTBD4 P311PP site direct mutagenesis fwd: TTGTATGTGGTGGGAGGGTCCATCCCACCACGGCGCATGTGGAAGT | Eurofins Genomics | N/A |
KBTBD4 P311PP site direct mutagenesis rev: ACTTCCACATGCGCCGTGGTGGGATGGACCCTCCCACCACATACAA | Eurofins Genomics | N/A |
KBTBD4 MIGR1 fwd: CCGGCCGGATCCATGGACTACAAAGACGATGACGACAAGGAATCACCAGAGGAGCCTGG | Thermo Fisher Scientific | N/A |
KBTBD4 MIGR1 rev: GCATACGTCGACTTAGGCCAACACAAACTGCAAATTG | Thermo Fisher Scientific | N/A |
KBTBD4 pCW57.1 fwd: GGGGGGTCGACCCACCATGGACTACAAAGACGATGACGACAAGGAAT | This paper | N/A |
KBTBD4 pCW57.1 Rev: GGGGGCTCGAGTTAGGCCAACACAAACTGCAAATTGGTAAGC | This paper | N/A |
Silencer® Select Small interfering RNAs (siRNA) targeting CoREST1 | Thermo Fisher Scientific | siRNA 1 ID s23229 siRNA 2 ID s23230 |
Recombinant DNA | ||
pcDNA3.1 (+) FLAG-KBTBD4 full-length wild type | This paper | N/A |
pcDNA3.1 (+) FLAG-KBTBD4 full-length R313PRR | This paper | N/A |
pcDNA3.1 (+) FLAG-KBTBD4 full-length P311PP | This paper | N/A |
pCMV5 HA RCOR1 (custom) | MRC Protein Phosphorylation and Ubiquitylation Unit | N/A |
MIGR1 FLAG-KBTBD4 full-length wild type | This paper | N/A |
MIGR1 FLAG-KBTBD4 full-length R313PRR | This paper | N/A |
MIGR1 FLAG-KBTBD4 full-length P311PP | This paper | N/A |
pCW57.1 FLAG-KBTBD4 full-length wild type | This paper | N/A |
pCW57.1 FLAG-KBTBD4 full-length R313PRR | This paper | N/A |
pCW57.1 FLAG-KBTBD4 full-length P311PP | This paper | N/A |
pENTR3C FLAG-KBTBD4 full-length wild type | This paper | N/A |
pENTR3C FLAG-KBTBD4 full-length R313PRR | This paper | N/A |
pENTR3C FLAG-KBTBD4 full-length P311PP | This paper | N/A |
Software and algorithms | ||
Molsoft ICM-Pro | Molsoft LLC. | |
Mascot search engine | Matrix Science | |
Data availability
LC/MS-MS raw data are available through https://www.ebi.ac.uk/pride with the following accession number: PXD031683. RNA-seq data have been deposited in GEO with the following accession number: GSE197240.
References
Harper JW, Schulman BA. Cullin-RING ubiquitin ligase regulatory circuits: a quarter century beyond the f-box hypothesis. Annu Rev Biochem. 2021;90:403–29.
Horn-Ghetko D, Krist DT, Prabu JR, Baek K, Mulder MPC, Klugel M, et al. Ubiquitin ligation to F-box protein targets by SCF-RBR E3-E3 super-assembly. Nature. 2021;590:671–6.
Skaar JR, D’Angiolella V, Pagan JK, Pagano M. SnapShot: F Box proteins II. Cell. 2009;137:1358, e1.
Raducu M, Fung E, Serres S, Infante P, Barberis A, Fischer R, et al. SCF (Fbxl17) ubiquitylation of Sufu regulates Hedgehog signaling and medulloblastoma development. EMBO J. 2016;35:1400–16.
Northcott PA, Buchhalter I, Morrissy AS, Hovestadt V, Weischenfeldt J, Ehrenberger T, et al. The whole-genome landscape of medulloblastoma subtypes. Nature. 2017;547:311–7.
Lee JC, Mazor T, Lao R, Wan E, Diallo AB, Hill NS, et al. Recurrent KBTBD4 small in-frame insertions and absence of DROSHA deletion or DICER1 mutation differentiate pineal parenchymal tumor of intermediate differentiation (PPTID) from pineoblastoma. Acta Neuropathol. 2019;137:851–4.
Wong GC, Li KK, Wang WW, Liu AP, Huang QJ, Chan AK, et al. Clinical and mutational profiles of adult medulloblastoma groups. Acta Neuropathol Commun. 2020;8:191.
Choi J, Lee K, Ingvarsdottir K, Bonasio R, Saraf A, Florens L, et al. Loss of KLHL6 promotes diffuse large B-cell lymphoma growth and survival by stabilizing the mRNA decay factor roquin2. Nat Cell Biol. 2018;20:586–96.
Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 2004;119:941–53.
Lee MG, Wynder C, Cooch N, Shiekhattar R. An essential role for CoREST in nucleosomal histone 3 lysine 4 demethylation. Nature. 2005;437:432–5.
Selvadurai HJ, Luis E, Desai K, Lan X, Vladoiu MC, Whitley O, et al. Medulloblastoma arises from the persistence of a rare and transient Sox2(+) granule neuron precursor. Cell Rep. 2020;31:107511.
Canning P, Cooper CD, Krojer T, Murray JW, Pike AC, Chaikuad A, et al. Structural basis for Cul3 protein assembly with the BTB-Kelch family of E3 ubiquitin ligases. J Biol Chem. 2013;288:7803–14.
Schumacher FR, Sorrell FJ, Alessi DR, Bullock AN, Kurz T. Structural and biochemical characterization of the KLHL3-WNK kinase interaction important in blood pressure regulation. Biochem J. 2014;460:237–46.
Chen Z, Picaud S, Filippakopoulos P, D’Angiolella V, Bullock AN. Structural basis for recruitment of DAPK1 to the KLHL20 E3 ligase. Structure. 2019;27:1395–404 e4.
Soucy TA, Smith PG, Milhollen MA, Berger AJ, Gavin JM, Adhikari S, et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature. 2009;458:732–6.
Ivanov DP, Coyle B, Walker DA, Grabowska AM. In vitro models of medulloblastoma: choosing the right tool for the job. J Biotechnol. 2016;236:10–25.
Schwalbe EC, Lindsey JC, Nakjang S, Crosier S, Smith AJ, Hicks D, et al. Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohort study. Lancet Oncol. 2017;18:958–71.
Hrdinka M, Fiil BK, Zucca M, Leske D, Bagola K, Yabal M, et al. CYLD Limits Lys63- and Met1-linked ubiquitin at receptor complexes to regulate innate immune signaling. Cell Rep. 2016;14:2846–58.
Monaghan CE, Nechiporuk T, Jeng S, McWeeney SK, Wang J, Rosenfeld MG, et al. REST corepressors RCOR1 and RCOR2 and the repressor INSM1 regulate the proliferation-differentiation balance in the developing brain. Proc Natl Acad Sci USA. 2017;114:E406–E15.
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A, et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet. 2008;40:499–507.
Lambert AW, Weinberg RA. Linking EMT programmes to normal and neoplastic epithelial stem cells. Nat Rev Cancer. 2021;21:325–38.
Ioris RM, Galie M, Ramadori G, Anderson JG, Charollais A, Konstantinidou G, et al. SIRT6 suppresses cancer stem-like capacity in tumors with PI3K activation independently of its deacetylase activity. Cell Rep. 2017;18:1858–68.
Vassalli G. Aldehyde dehydrogenases: not just markers, but functional regulators of stem cells. Stem Cells Int. 2019;2019:3904645.
Choi SA, Lee JY, Phi JH, Wang KC, Park CK, Park SH, et al. Identification of brain tumour initiating cells using the stem cell marker aldehyde dehydrogenase. Eur J Cancer. 2014;50:137–49.
Tokheim C, Wang X, Timms RT, Zhang B, Mena EL, Wang B, et al. Systematic characterization of mutations altering protein degradation in human cancers. Mol Cell. 2021;81:1292–308 e11.
Fares I, Chagraoui J, Gareau Y, Gingras S, Ruel R, Mayotte N, et al. Cord blood expansion. Pyrimidoindole derivatives are agonists of human hematopoietic stem cell self-renewal. Science. 2014;345:1509–12.
Cohen S, Roy J, Lachance S, Delisle JS, Marinier A, Busque L, et al. Hematopoietic stem cell transplantation using single UM171-expanded cord blood: a single-arm, phase 1-2 safety and feasibility study. Lancet Haematol. 2020;7:e134–e45.
Maksour S, Ooi L, Dottori M. More than a corepressor: the role of CoREST proteins in neurodevelopment. eNeuro. 2020;7:1–12.
Northcott PA, Lee C, Zichner T, Stutz AM, Erkek S, Kawauchi D, et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 2014;511:428–34.
Lee C, Rudneva VA, Erkek S, Zapatka M, Chau LQ, Tacheva-Grigorova SK, et al. Lsd1 as a therapeutic target in Gfi1-activated medulloblastoma. Nat Commun. 2019;10:332.
Whyte WA, Bilodeau S, Orlando DA, Hoke HA, Frampton GM, Foster CT, et al. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature. 2012;482:221–5.
Ward PS, Patel J, Wise DR, Abdel-Wahab O, Bennett BD, Coller HA, et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. Cancer Cell. 2010;17:225–34.
D’Angiolella V, Donato V, Forrester FM, Jeong YT, Pellacani C, Kudo Y, et al. Cyclin F-mediated degradation of ribonucleotide reductase M2 controls genome integrity and DNA repair. Cell. 2012;149:1023–34.
Fung E, Richter C, Yang HB, Schaffer I, Fischer R, Kessler BM, et al. FBXL13 directs the proteolysis of CEP192 to regulate centrosome homeostasis and cell migration. EMBO Rep. 2018;19:e44799.
Mavrommati I, Faedda R, Galasso G, Li J, Burdova K, Fischer R, et al. beta-TrCP- and casein kinase II-mediated degradation of cyclin F controls timely mitotic progression. Cell Rep. 2018;24:3404–12.
Pertea M, Kim D, Pertea GM, Leek JT, Salzberg SL. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc. 2016;11:1650–67.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinforma. 2013;14:128.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44:W90–7.
Williamson D, Schwalbe EC, Hicks D, Aldinger KA, Lindsey JC, Crosier S, Richardson S, et al. Medulloblastoma group 3 and 4 tumors comprise a clinically and biologically significant expression continuum reflecting early human cerebellar development. Cell Rep. In revision.
Acknowledgements
Mass spectrometry analysis was performed in the Discovery Proteomics Facility (headed by Roman Fischer) which is part of the TDI MS Laboratory (led by Benedikt Kessler).
Funding
This study was possible thanks to the support of a Medical Research Council (MRC) grant MC_UU_00001/7 to V. D’A and a Cancer Research UK (CRUK) grant DRCNPG-May21\100002 to V.D’A., A.N.B. and S.C.C. This work was also supported by The Brain Tumour Charity through an Expanding Theories award GN-000559 to V. D’A. We acknowledge further support by a John Fell (133/075) and Wellcome Trust grant (097813/Z/11/Z) to B.M.K. A.N.B. acknowledges funding from the Innovative Medicines Initiative 2 Joint Undertaking (JU) under grant agreement No 875510 (EUbOPEN). The JU receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA and Ontario Institute for Cancer Research, Royal Institution for the Advancement of Learning McGill University, Kungliga Tekniska Hoegskolan, Diamond Light Source Limited. A.N.B. also acknowledges support from a Wellcome strategic award [106169/ZZ14/Z]. F.M.B. acknowledges support from European Research Council Award microC 772970. Z.C. acknowledges support from the China Scholarship Council - Nuffield Department of Medicine Scholarship. R.M.I. acknowledges a PhD scholarship – CAPES, Science without borders program (DPE 99999.013236/2013–00), as well as a fellowship from the Swiss National Science Foundation - Early Postdoc. Mobility program (P2GEP3_188295). S.C.C. acknowledges Programme Grant funding from Cancer Research UK.
Author information
Authors and Affiliations
Contributions
RMI and ZC performed all the biochemical experiments. IV and BMK conducted the LC-MS/MS and analysed the data. LB, AV and FMB analysed and processed the RNA-seq data from cell lines. SCC and SR analysed and processed the transcriptomic data from human medulloblastoma. ANB, RMI and VDA analysed all experiments, managed the overall study and supervised manuscript preparation and submission. All authors have contributed to the final version of the manuscript and rebuttal.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Melino
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Z., Ioris, R.M., Richardson, S. et al. Disease-associated KBTBD4 mutations in medulloblastoma elicit neomorphic ubiquitylation activity to promote CoREST degradation. Cell Death Differ 29, 1955–1969 (2022). https://doi.org/10.1038/s41418-022-00983-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-022-00983-4
This article is cited by
-
AMPK attenuates SHH subgroup medulloblastoma growth and metastasis by inhibiting NF-κB activation
Cell & Bioscience (2023)
-
Histopathology and molecular pathology of pediatric pineal parenchymal tumors
Child's Nervous System (2023)
-
Molecular characterisation defines clinically-actionable heterogeneity within Group 4 medulloblastoma and improves disease risk-stratification
Acta Neuropathologica (2023)