Abstract
The role of mitophagy, a process that allows the removal of damaged mitochondria from cells, remains unknown in multiple sclerosis (MS), a disease that is found associated with dysfunctional mitochondria. Here we have qualitatively and quantitatively studied the main players in PINK1-mediated mitophagy in peripheral blood mononuclear cells (PBMCs) of patients with relapsing–remitting MS. We found the variant c.491G>A (rs550510, p.G140E) of NDP52, one of the major mitophagy receptor genes, associated with a MS cohort. Through the characterization of this variant, we discovered that the residue 140 of human NDP52 is a crucial modulator of NDP52/LC3C binding, promoting the formation of autophagosomes in order to drive efficient mitophagy. In addition, we found that in the PBMC population, NDP52 is mainly expressed in B cells and by ensuring efficient mitophagy, it is able to limit the production of the proinflammatory cytokine TNF-α following cell stimulation. In sum, our results contribute to a better understanding of the role of NDP52 in mitophagy and underline, for the first time, a possible role of NDP52 in MS.
Similar content being viewed by others
Introduction
Multiple sclerosis (MS) is an autoimmune disease with a chronic inflammatory response in the central nervous system that leads to demyelination and axonal damage resulting in motor, sensory, and cognitive lost. The causes of the inflammatory process are not yet known but a well-accepted theory indicates that several factors are involved, including genetic predisposition to develop the disease and environmental agents able to stimulate the immune system and trigger the autoimmune reaction. Among these hypotheses, there is the involvement of bacterial or viral infections [1, 2]. Immune responses are triggered and driven, upon microbial infection, by the recognition of pathogen-associated molecular patterns (PAMPs), by means of a broad spectrum of pattern-recognition receptors (PRRs) expressed on several innate and adaptive immune cells [3]. Toll-like receptors (TLRs) are generally perceived as the main PRRs and serve as immune sensors for the detection of specific PAMPs related to different pathogens [4].
Another characteristic of MS is the presence of mitochondrial dysfunctions associated with the pathology [5,6,7]. A mechanism called mitophagy selectively eliminates altered mitochondria, in order to regulate the functional quality of mitochondria and to reduce oxidative stress [8]. This removal mainly happens by a selective form of autophagy after activation of the kinase PINK1 and the E3 ubiquitin ligase PARKIN. Following the loss of mitochondrial membrane potential, PINK1 is stabilized on the outer mitochondrial membrane where it phosphorylates ubiquitin at Ser65 and recruits two crucial redundant mitophagy receptors OPTINEURIN (OPTN) and CALCOCO2/Nuclear dot protein 52 (NDP52) [9, 10]. These receptors bind ubiquitinated mitochondria through their ubiquitin-binding domain (UBD) and thanks to a LC3-interacting region (LIR) motif, they bind Atg8 (autophagy-related protein 8)/LC3 (microtubule-associated proteins 1A/1B light chain 3B) proteins onto autophagosome membranes in order to deliver mitochondria to the autophagosomes [11]. More recently, the LIR motif has been shown to serve as amplifiers of mitophagy by recruiting additional OPTN or NDP52 to growing phagophore membranes [12].
It has been demonstrated that ATG5, a protein involved in autophagy, and PARKIN increase in cerebrospinal fluid and serum of MS patients, highlighting these autophagy/mitophagy elements as new molecular markers of the disease’s active phases [13, 14]. However, the role of PINK1-mediated mitophagy pathway in MS remains obscure.
In the present work, we, quantitatively and qualitatively analyzed three indispensable proteins for the PINK1-mediated mitophagy pathway, i.e., PINK1, OPTN, and NDP52, in peripheral blood mononuclear cells (PBMCs) of relapsing–remitting (RR)-MS patients. We identified a possible protective variant of NDP52 associated with MS pathology, by promoting a more efficient clearance of damaged mitochondria with respect to the wild-type (WT) form. Finally, we discovered that, among PBMCs, NDP52 is mainly expressed in the B cell subpopulation ensuring effective mitophagy following B cell stimulation and reducing the release of the proinflammatory cytokine TNF-α.
Results
PINK1-mediated mitophagy genes are upregulated in PBMCs of RR-MS patients and damaged mitochondria are accumulated
We evaluated mRNA expression of PINK1, OPTN, and NDP52, the three indispensable PINK1-mediated mitophagy genes in PBMCs isolated from RR-MS patients (n = 35) and healthy donors (HD, n = 24). Our data indicate that PINK1 and NDP52 genes are upregulated in RR-MS patients with respect to controls (Fig. 1a, b). Of note, no particular increase in mRNA OPTN was observed (Fig. 1c). We found that both PINK1 and NDP52 protein levels were also increased in RR-MS subjects compared to HD (Fig. 1d, e). Again, OPTN protein levels were unchanged between HD and MS subjects (Fig. 1f).
a–c PINK1, NDP52, and OPTN mRNA levels, respectively, were analyzed in PBMCs of healthy donors (HD = 24) or multiple sclerosis patients (RR-MS = 35) by qPCR analysis. B2M was used as housekeeping gene. d PBMCs from HD or RR-MS people were lysated and immunoblotted for PINK1 and Vinculin antibodies (HD = 10; RR-MS = 28). The graph reports the PINK1/Vinculin ratio. e PBMCs from HD or RR-MS people were lysated and immunoblotted for NDP52 and Vinculin antibodies (HD = 10; RR-MS = 28). The graph reports the NDP52/Vinculin ratio. f Protein lysates from PBMCs were immunoblotted for OPTN (HD = 10; RR-MS = 26). Vinculin was used as loading control. The graphs show and OPTN/Vinculin ratio. g PBMCs from HD (n = 5) or RR-MS (n = 28) subjects were lysated and immunoblotted for the indicated antibodies. h The graph analyzed the mtDNA/nucDNA ratio in PBMCs of HD (n = 4) or RR-MS patients (n = 28) by qPCR analysis. i Four HD or 4 RR-MS PBMCs were stained with 10 nM of TMRM and analyzed by flow cytometry to measure the fluorescence intensity of the dye. The median FL-2A of HD patients were reported to 1 and compared to RR-MS patients. j Representative image of protein carbonylation analysis from HD or RR-MS PBMCs. The graph showing protein oxidation status (a.u.). All the quantifications resulting as the mean of at least three independent experiments (±SD). Statistical analysis was performed using Student t-test with Welch’s correction (*P < 0.05; **P < 0.01; ***P < 0.001, n.s.: not significant).
These findings provide evidence that the production of two main mitophagy elements of the PINK1-mediated mitophagy is induced in PBMCs of MS patients, suggesting that mitophagy is most likely stimulated in PBMCs of RR-MS patients.
Since mitophagy induction is typically characterized by a decreased mitochondrial mass, we thought to examine mitochondrial content in PBMCs by checking the levels of two inner mitochondrial membrane markers, the cytochrome c oxidase subunit II (COXII) and cytochrome c oxidase subunit IV (COXIV). We observed an increase in both COXII and COXIV protein levels in MS patients compared to HD (Fig. 1g), suggesting a putative block in mitochondrial clearance. Consistent with this increase, we observed an increase in the mitochondrial DNA (mtDNA)/nuclear DNA ratio (mtDNA:nucDNA ratio) in RR-MS patients compared to HD (Fig. 1h).
Our results show that mitochondria of RR-MS patients display a reduced membrane potential (ΔΨm) compared to HD (Fig. 1i), indicating that the accumulated mitochondria in RR-MS patients were corresponding to damaged mitochondria. We next thought to assess whether such mitochondrial dysfunction was associated with oxidative stress by evaluating the content of putative carbonyl groups introduced into proteins. As shown in Fig. 1j, protein extracts from PBMCs of RR-MS patients show an increase in protein carbonylation compared to HD.
Altogether, these data indicate that, even if PINK1 and NDP52 genes are induced in RR-MS patients, damaged mitochondria are found accumulated, participating most likely in oxidative stress production and in inflammation.
NDP52 G140E variant was associated and genetically protective in MS
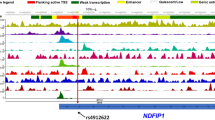
We next qualitatively analyzed the main mitophagy receptors involved in PINK1-mediated mitophagy in RR-MS patients. To this end, we extracted cDNA from PBMCs of RR-MS patients and amplified the two main functional fragments of both NDP52 and OPTN: the UBD and the LIR domains. We sequenced the extracted cDNA of 35 RR-MS patients in order to search for putative point mutations or specific polymorphisms in these functional fragments. We did not find any changes in the OPTN gene (data not shown). Instead, we observed the presence of c.491G>A (rs550510, p.G140E) variant in the LIR region of NDP52. We thus performed a genotyping analysis on a cohort of 203 RR-MS patients and 1000 control subjects. The frequency distribution of the alleles showed a higher frequency of the variant allele (A) in the control group with respect to cases. Similarly, the frequency distribution of the genotypes in our cohort showed a higher frequency of heterozygous (GA) and homozygous (AA) genotypes in the control group. The biostatistical analysis revealed a statistically significant association of c.491G>A variant with the disease (P = 0.007), with the A allele being protective (odds ratio, OR, A = 0.65, 95% CI: 0.48–0.89) for MS (Table 1). The allele association has been further confirmed by the genotype association, as shown by the P and OR of GA and AA genotypes, respectively (Table 1). Taken together, these results suggest that the NDP52 variant may confer protection to MS.
NDP52 G140E substitution increases the binding with LC3C
NDP52 is known to negatively regulate TLR-triggered NF-κB activation and to be degraded in the course of TLR activation [15]. Because enhanced immune response may benefit MS patients by eliminating microbes, we first anticipated that NDP52 G140E variant might be protective in MS by being more stable than the WT form of the protein after TLR3 stimulation and therefore by favoring activation of NF-kB. In order to verify this aspect in cells, we generated the NDP52 G140E mutant (hereafter called NDP52GE). We evaluated the stability of NDP52GE under Polyinosinic:polycytidylic acid treatment (Poly (I:C))-triggered TLR-activating conditions and whether NDP52GE was capable of dampening proinflammatory signaling. To this end, we transfected HeLa cells (that are known to express TLR3) with vectors encoding NDP52WT or NDP52GE and we treated cells with Poly (I:C) in order to mimic viral infection. TLR activation resulted in similar degradation of both NDP52WT and NDP52GE variants (Supplementary Fig. 1a). Moreover, by quantifying the ratio of nuclear/cytoplasmic presence of p65/NF-kB in NDP52-expressing cells, we did not observe differences of NF-kB activation between NDP52WT and NDP52GE expressing cells (Supplementary Fig. 1b). These data indicate that the substitution G140E of NDP52 did not affect its function in regulating TLR signaling.
At this point we hypothesized that NDP52GE may mitigate MS disease by clearing more efficiently damaged mitochondria compared to the WT form of the protein. Since the NDP52GE variant is subjected to an amino acid modification near its cLIR motif, we postulated that NDP52GE may favor the binding between LC3C, its preferential LC3 binding partner [16]. By performing co-immunoprecipitation experiments, we analyzed NDP52–LC3C interaction in basal conditions. We co-transfected HEK293 cells with a vector encoding HA-LC3C and a vector encoding FLAG-NDP52WT or FLAG-NDP52GE. We observed that G substitution in E at residue 140 enhanced the ability of NDP52 to interact with LC3C (Fig. 2a, b). We reinforced these data by performing a confocal microscopy analysis in which we analyzed the colocalization between NDP52 variants and LC3C. As shown on Fig. 2c, d, the colocalization between NDP52GE and LC3C is higher with respect to the one between NDP52WT and LC3C.
a Representative image of co-immunoprecipitation experiment in which HEK293 cells were co-transfected with Flag-NDP52WT or Flag-NDP52GE vectors in combination with HA-LC3C. b The graph represents the ratio between LC3C and NDP52 (n = 4). c HEK293 cells co-transfected as indicated in (a) were subjected to confocal immunofluorescence in order to evaluate LC3C-NDP52 colocalization. Scale bar 8 μm. d The graph indicates the colocalization value between LC3C and wild-type or mutant NDP52. e HEK293 cells co-transfected with pcDNA3 (empty vector) or Flag-NDP52WT or Flag-NDP52GE vectors in combination with HA-LC3C were treated with O/A (10 μM/4 μM) and CQ (20 μM) for the indicated times and subjected to co-immunoprecipitation analysis. f The graph represents the ratio between LC3C and NDP52 in O/A + CQ condition (n = 3). g HEK293 cells co-transfected as indicated in (a) were treated with O/A in order to induce mitophagy. Confocal immunofluorescence analysis evaluates LC3C-NDP52 colocalization. Scale bar 8 μm. h The graph indicates the colocalization (Pearson coefficient) between LC3C and wild-type or mutant NDP52, upon mitophagy induction. All the quantifications resulting as the mean of at least three independent experiments (±SD). Statistical analysis was performed using Student t-test (*P < 0.05; **P < 0.01).
To verify whether the highest ability of NDP52GE to bind LC3C occurs also following mitophagy induction, we checked for this binding following Oligomycin/Antimycin-A treatment (O/A), two agents commonly used to induce mitochondrial damage. We did not observe an increase of binding between NDP52 and LC3C following O/A (Supplementary Fig. 2a). We thus hypothesized that the stimulus of mitophagy induction plus the variant NDP52GE was too strong, accelerating the autophagy/mitophagy flux. We thus blocked the autophagy flux by adding Chloroquine (CQ) to O/A treatment and we found a stronger binding between NDP52GE and LC3C under mitochondrial stress compared with the NDP52WT condition (Fig. 2e, f). We confirmed these data by performing confocal microcopy analysis (Fig. 2g, h and Supplementary Fig. 2b).
Altogether, these results indicate that the amino acid on position 140 of NDP52 influences its interaction with LC3C both in basal condition and following mitophagy induction.
NDP52 G140E substitution induces a conformational changes around LIR motif, enhancing the binding between NDP52 and LC3C
In order to characterize the interaction between NDP52 and LC3C protein, we performed isothermal titration calorimetry (ITC) experiments in which synthetic peptides spanning the short NDP52 cLIR motif were titrated into LC3C protein. We observed a very small increase in affinity for the mutated NDP52 peptide (KD values are 5.4 µM for NDP52128–155, and 4.5 µM for NDP52GE128–155) and decrease of unfavorable (positive) binding enthalpy (Fig. 3a). The same tendency was observed in LC3B-based ITC experiments (Supplementary Fig. 3a–d and Supplementary Table 2). The decrease in unfavorable enthalpy indicates that the “enthalpically” favorable interactions could play a role in the G140E-mediated NDP52 affinity increase to LC3C and LC3B proteins. ITC results are in agreement with the affinities reported previously and indicated preference of LC3C for interaction with NDP52 cLIR [16].
NDP52 G140E mutation enhances interaction between NDP52 and LC3C. a ITC analysis of the interaction between LC3C and peptides contained NDP52 LIR motif. Titrations of the NDP52128–155 peptide (NDP52 WT, left panel) and NDP52GE128–155 peptide (NDP52 G140E, right panel) to LC3C protein showed slight preference of LC3C to the NDP52GE128–155 (KD values and enthalpies for each interaction are indicated at the plots). The upper graph shows the raw measurement; the lower diagram represents the intergraded heat per injection (squares) and best fit function (solid line). All thermodynamic parameters are summarized in Supplementary Table 2. b NMR titration of 15N-labeled LC3C with NDP52128–224 and NDP52GE128–224 peptides. The representative area (the “fingerprint” area, containing LC3C K55 and L64 HN resonances) of the LC3C 15N-1H-TROSY-HSQC spectra upon titration with NDP52128–224 (left panel) and NDP52GE128–224 (right panel). Increasing concentrations of peptide are indicated with the rainbow color code from red (free LC3C) to violet (four times peptides excess). Identified CSP are highlighted by solid line arrows, the CSP exhibited difference between NDP52128–224 and NDP52GE128–224 peptides are showed by dashed arrows at the right plot. c CSP values (Δδ) for LC3C HN resonances at the last titration stages with NDP52128–224 are plotted against residues numbers. The bars for Δδ below standard deviation value (σ, calculated for all identified CSPs) are colored gray; for σ < Δδ < 2σ—yellow; and for Δδ above 2σ—red. Violet bars represent the residues that resonances disappear upon addition of NDP52128–224 and do not appear again (strong interaction with intermediate to slow exchange mode). Their Δδ values are set to the Δδ values of neighbor residues. The CSP values mapped on the LC3C protein structure (ribbon diagrams, PDB ID 1GNU) are shown in the upper right corner with the same color code. d Differences of CSP values (ΔΔδ) induced upon titration of NDP52128–224 and NDP52GE128–224 into 15N-labeled LC3C are mapped on the LC3C sequence and structure. The color code is the same as at c. e The cartoon representation shows one of the selected models of LC3C13–126-NDP52125–147 G140E as a reference with a gradient of colors from N- to C-terminal (light blue to dark blue for LC3C13–126 and yellow to green for NDP52125–147). The stick representation highlights the residue E140 of NDP52 and R76 of LC3C forming possible electrostatic interaction. The dotted line (hot pink) indicates the distance between the atoms Cδ of E140 and Cζ of R76. f The plot shows the one-dimensional free energy surface of the distance between E140 of NDP52 and R76 of LC3C as calculated by WTE-metaD simulation. The profile indicates two main minima, separated by less populated states. We highlighted the first minima (hot pink) in the range of distance (3–5 Å) compatible with the formation of a salt bridge for sake of clarity. The bottom panel highlights the side chain of E140 (green) and R76 (blue) and the distance between their Cδ and Cζ atoms (hot pink dotted line) that we used as a collective variable in the well-tempered metadynamics simulation.
In order to validate the ITC data and understand the mechanism of the affinity increase, we performed nuclear magnetic resonance (NMR) titration experiments, in which we titrated non-labeled NDP52128–224 and NDP52GE128–224 into the 15N-labeled LC3C sample (Fig. 3b–d). Both titration experiments showed that the chemical shift perturbations (CSPs) undergo intermediate to slow exchange modes, confirming low µM KD values for both NDP52 constructs. The resonances in the most relevant LC3C regions (β-strand 2 and in close proximity to it, HP1 and HP2) disappeared upon interaction with NDP52 and did not appeared again, indicating specific dynamics of the NDP52 constructs with equilibria between unfolded and α-helical conformations of the fragments observed by 1D NMR spectra (Supplementary Fig. 3e). It was possible to map CSPs on LC3C sequence and 3D structure (Fig. 3c). We observed few differences between the CSPs. The first difference is the CSP altitude—addition of NDP52GE128–224 induced either larger CSP (like R76 HN resonance, Fig. 3b) or different direction CSP (like V89 HN resonance, Supplementary Fig. 3f) for a set of LC3C residues. These differences are overviewed in Fig. 3d where the different CSP values (ΔΔδ) were mapped on the LC3C sequence and surface. The second difference is the dynamic behavior of the CSPs—a number of LC3C resonances showed more slow exchange mode upon titration with NDP52GE128–224 construct when with NDP52128–224 (I73 HN resonance in Fig. 3b). The larger CSP values and “slower” exchange mode indicate clear preference of LC3C protein to the NDP52GE variant, while different direction CSPs indicate changes in the resonance’s microenvironment upon addition of mutated NDP52 fragment.
Since the data suggest that the NDP52 G140E mutation affects intermolecular electrostatic interactions and conformational equilibria of close proximity to G/E140, we used molecular modeling and simulations to address this hypothesis. It is known that the C-terminal LIR extensions can modulate its binding affinity to human ATG8 proteins [17,18,19,20,21]. For example, FYCO1 LIR C-terminal extension was reported to form an α-helix and a key network of intermolecular electrostatic interactions was mediated by E1246, forming intermolecular hydrogen bonds with LC3B residue R70 [22]. G140 in NDP52 corresponds to E1246 in FYCO1 with respect to the position in sequence of the LIR motifs in these two proteins. We thus speculate that increased affinity of the NDP52GE variant to LC3C could be due to a similar scenario, including the formation of a salt bridge between E140 in NDP52GE and LC3C R76. G140 is located at the N-terminus of the predicted α-helical area (coiled-coil (cc) region) and the G140E substitution could also stabilize the helical structure.
We thus built a model for the LC3C8–127-NDP52GE125–147 complex and carried out unbiased molecular dynamics (MD) simulations (see GitHub repository) (Fig. 3e). We used MD simulations to see if conformational changes that favor the formation of a salt bridge or other electrostatic interactions in proximity of R76 are likely to occur. During unbiased hundreds ns MD simulations, we observed a low tendency for intermolecular salt bridge or hydrogen bond formation for E140 with LC3C residues. For example, we only found a salt bridge interaction between the charged groups of R76 and E140, in less than 1% of the MD structures (see GitHub repository). This is likely due to inherent limitations in the sampling of the conformational space that classical MD can provide [23], as our starting structures did not feature the salt bridge in question. We thus applied another approach based on the metadynamics algorithm in the well-tempered ensemble flavor to boost the sampling of the conformational space [24, 25]. We observed two main free energy minima separated by other less populated states. The first one is in the range of 3–5 Å, which is compatible with the salt bridge formation between E140 and R76 and the second one is in the range of 16–18 Å, which is similar to the distance between the two groups at the beginning of the simulation. The latter is likely to be overestimated due to the starting conformation of our model (Fig. 3f).
Taken together, these results give a proof of concept that the point mutation in G140 in “E” induces a specific conformation of the region near the NDP52-LIR domain, which favor its interaction with LC3C (and most likely with all mATG8 proteins).
NDP52GE promotes a more efficient initiation of mitophagy compared to the wild-type form of NDP52
Several studies have demonstrated the importance of the LIR motif for selective autophagy [26, 27]. More recently, the LIR motif has been shown to mediate the translocation of autophagy receptors to mitochondria independently of the ubiquitin binding [12]. Since NDP52 plays a key role during autophagosome initiation by promoting the recruitment of the ULK1 complex to mitochondria, thanks to its LC3 binding [28], we hypothesized that NDP52GE, by binding more efficiently LC3C, could be localized at the mitochondrial surface at higher levels when compared to its WT isoform. We thus checked, by performing confocal microscopy analysis, NDP52 localization on mitochondria in HEK293. Cells were transfected with vectors encoding NDP52WT or NDP52GE and mitochondria were stained with TOM20 antibody. We found a higher localization of NDP52GE to mitochondria respect to its WT form in basal condition (Fig. 4a–c). Moreover, we noticed that the mitochondrial network was reduced in NDP52GE positive cells compared to the NDP52WT positive cells, suggesting a basal mitophagy induction in NDP52GE positive cells.
a HEK293 cells transfected Flag-NDP52WT or Flag-NDP52GE vectors were fixed and immunostained with antibodies against TOM20 in order to label mitochondria and Flag to detect NDP52 transfected cells. Scale bar 5 μm. b The graph represents the colocalization (Pearson coefficient) between mitochondrial marker TOMM20 and NDP52 evaluated on in single transfected cells from three independent experiments. c The graph represents mitochondrial content in Flag-NDP52WT or Flag-NDP52GE transfected cells (27 cells in total from three independent experiments) following mitophagy induction. d, e Representative confocal microscopy images in which HEK293 cells transfected and treated as indicated in (a). Scale bar 8 μm. The graphs report the number of ATG13 (d) or ATG16L (e) foci per cell (n = 23 cells), respectively. f HEK293 cells downregulated for NDP52 using specific siRNA were next transfected with pcDNA3, Flag-NDP52WT or Flag-NDP52GE vectors and treated with O/A (10 μM/4 μM) for 8 h. Mitochondrial markers COXII and COXIV were detected by western blot. HSP90 was used as loading control. The graphs represent the COXII/HSP90 or COXIV/HSP90 ratio in mitophagy condition, resulting as the mean of three independent experiments (±SD). g Cell treated as in (f) were stained with the MitoTracker Green probe and analyzed by FACS analysis. All statistical analysis were performed using Student t-test (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001).
These data indicate that NDP52GE, which binds LC3C with a higher affinity, is more translocated to mitochondria compared to the WT form.
We next postulated that NDP52GE may favor the recruitment of the machinery necessary for autophagosome formation. We thus quantified the formation of autophagosome initiation dots containing the ULK1 complex subunit ATG13 [28, 29] in cells expressing NDP52WT or NDP52GE after O/A treatment. Our data indicate that the overall number of ATG13 dots in cells expressing NDP52GE was significantly higher than those expressing NDP52WT (Fig. 4d). Consistent with this, the recruitment of ATG16L1, a downstream autophagy effector, was also enhanced in cells expressing NDP52GE (Fig. 4e). We next assessed the mitophagy activity of LIR-mutant NDP52GE and NDP52WT in endogenous NDP52-downregulated HEK293 cells. We found that NDP52GE significantly degraded COXII and COXIV compared to NDP52WT (Fig. 4f). In agreement, we found that NDP52GE is degraded to a higher level than the WT form of the protein following O/A treatment (Supplementary Fig. 4). We confirmed these data by evaluating the mitochondrial mass using the MitoTracker-Green dye (Fig. 4g).
These results demonstrate that NDP52GE recruits, with more efficiency, the platform necessary for autophagosome formation during mitophagy and promotes a rapid clearance of damaged mitochondria compared to the WT form of NDP52.
NDP52, predominantly expressed in B cells, ensures effective mitophagy and reduces secretion of the proinflammatory cytokine TNF-α following B cell stimulation
To better understand the role of NDP52 in blood cells, we checked in which subpopulation of PBMCs NDP52 was expressed. We discovered that NDP52 was highly expressed in B cells compared to T cells or monocytes (Fig. 5a, b). In order to exclude that the increase of NDP52 expression found in MS patients (Fig. 1e) was due to an increase in B cells population, we compared the number of B cells in healthy patients versus MS patients. As illustrated in Supplementary Fig. 5, no difference between the two populations was observed, confirming that NDP52 is upregulated in B cells of MS patients compared to healthy controls. Since B cells play a key role in MS disease [30], we checked the levels of NDP52 after stimulation of B cells and found that the expression of NDP52 was increased after stimulation with PMA and Ionomycin (PMA/Iono) (Fig. 5c, d).
a Total PBMCs, monocytes, B or T cells from healthy donors (HD) were lysated and immunoblotted for NDP52 and Vinculin antibodies (HD = 3). b The graph reports the NDP52/Vinculin ratio of three independent experiments. c Immunoblotting image of B cells from HD were stimulated with PMA (20 ng/ml) and Ionomycin (200 ng/ml) treatment for 12 h. d The graph reports the NDP52/Vinculin ratio in basal or PMA/Iono condition of three independent experiments. e LCL cells were downregulated or not for NDP52 using specific siRNA and then stimulated with PMA (20 ng/ml) and Ionomycin (200 ng/ml) treatment for 12 h. The levels of NDP52, COXII and Vinculin (as a loading control) have been detected by western blot. f The graph reports the NDP52/Vinculin ratio of three independent experiments. g The graph reports the COXII/Vinculin ratio of three independent experiments. h The graphs report the results of ELISA test performed to detect TNF-α in the supernatant of LCL cells (n = 4). All quantifications resulting as the mean of at least three independent experiments (±SD). Statistical analysis was performed using Student t-test (*P < 0.05; **P < 0.01).
We thus hypothesized that NDP52 may be a key element for correct mitophagy occurrence in B cells, reducing the release of proinflammatory cytokines that trigger inflammation in MS patients. To test this, in order to manipulate NDP52 expression, we moved to a lymphoblastoid cell line (LCL) derived from a healthy patient. First, we checked whether NDP52 could be also modulated following stimulation of these cells. As illustrated in Fig. 5c, the PMA/Iono treatment was sufficient to induce an upregulation of NDP52. In addition, we found that NDP52 increase was accompanied by a decrease of COXII protein, indicating induction of mitophagy. By contrast, downregulation of NDP52 favors COXII accumulation thus inhibiting mitochondrial clearance (Fig. 5e–g). Also, by performing enzyme-linked immunosorbent assay (ELISA), we found that human proinflammatory cytokine TNF-α was increased in LCL cells downregulated for NDP52 compared to control cells stimulated with PMA/Iono (Fig. 5h).
In summary, our data underline NDP52 as a novel regulator of the immune response in B cells. In particular, we found that NDP52, by ensuring correct mitochondrial clearance following B cell stimulation, limits proinflammatory cytokine production.
Altogether our data suggest that NDP52GE variant may be protective in MS thanks to its higher ability to induce mitophagy respect to the WT form and we can assume that, by acting on B cells, this pro-mitophagic activity may reduce inflammation in MS patients (Fig. 6).
The PBMC inflammatory state in RR-MS patients induces a robust production of PINK1-mediated mitophagy elements (PINK1 and NDP52). Despite this fact, we noticed a block in mitochondrial clearance and thus an accumulation of dysfunctional mitochondria. The defect in mitochondrial clearance may thus favor a feedback inflammation loop in PBMCs of RR-MS patients. The substitution G140E of NDP52 (NDP52GE, protective variant in MS) improves the efficiency of mitophagy compared to the WT (NDP52WT) form of the protein (by binding more LC3C and thus favoring phagophore formation) and this allows most likely B cells to decrease their proinflammatory cytokine production thus reducing inflammation.
Discussion
Our study reveals that PINK1 and NDP52 expressions are induced in PBMCs of RR-MS patients. Since RR-MS patients displayed an accumulation of NDP52 and a block in mitochondrial clearance, we suggest that mitophagy induction is not sufficient to remove damaged mitochondria. In fact, we found an accumulation of mitochondria accompanied by oxidative stress in PBMCs of RR-MS patients. Interestingly, an increase of PINK1 protein has been found in MS brain lesions [31]. These data suggest that most likely, PINK1-mediated mitophagy may also be stimulated in the brain of MS patients. Of note, we did not find positive correlation between high expression of PINK1 and NDP52 in PBMCs of single MS patients (Supplementary Fig. 6). Moreover, we cannot exclude that an increase in the OPTN mRNA levels is necessary for correct mitophagy induction in PBMCs of RR-MS patients. Therefore, further investigation of this aspect may explain why mitophagy is not sufficient to remove damaged mitochondria in PBMCs of RR-MS subjects.
Our results reveal a significant association of NDP52 c.491G>A (rs550510, p.G140E) with a decreased susceptibility to MS. Interestingly, a risk polymorphism in the NDP52 gene has been found to be associated with Crohn’s disease [32]. Functionally, the authors found that the NDP52 risk variant is more stable compared to the WT form following TLR signaling stimulation, favoring inflammation. We did not find differences between our variants in regulating TLR signaling but our data showed that the NDP52GE variant is more efficient in mitochondria removal. We have identified an amino acid in NDP52 sequence that is crucial in order to extend the cLIR motif and favor its binding with LC3C, inducing a positive feedback loop on phagophore formation and thus on mitophagy. Our data support the ubiquitin-independent mechanism of autophagy receptor recruitment discovered by Padman et al. [12]. The exact mechanism of the affinity enhancement still needs a careful investigation. It is necessary to understand the role of the G140E mutation in the stabilization of the predicted NDP52 cc-region C-terminally to the cLIR motif. Interestingly, another non-canonical LIR-like motif is located in the cc-regions (residues 203-DYWE-206, [33]). This LIR-like motif showed preference for LC3B protein and seems could prompt autophagosome maturation. Indeed, NDP52 constructs possessing both cLIR and LIR-like motifs showed more complex ITC profile (Supplementary Fig. 3c). We cannot exclude that other signals could affect the length and stability of the cc-region in order to induce a stronger pro-mitophagy activity of NDP52GE. A more comprehensive computational study should be carried to quantitatively estimate the propensity for hydrogen bonds or/and salt bridge formation.
Of note, available data on rs550510 variant (reported on 1000Genomes and GnomAD databases of genetic variants annotation) indicate that the protective allele is more common in some populations (i.e., East Asian) than others (i.e., Europe). Since genetic features, environmental factors and habits influence variant distribution, it is not a surprise that, like other genetic polymorphism, the NDP52GE variant shows a variable frequency distributions among the worldwide populations. It should be interesting to investigate, in the future, why such an efficient variant has not been selected too much in the European population.
Our results highlight that NDP52 protein is mainly expressed in B cell subset. We found that the absence of NDP52 is associated with a block in mitochondrial clearance and an increased production of the proinflammatory cytokine TNF-α following B cell stimulation, underlying a critical role of NDP52 in regulating mitophagy in B cells. Interestingly, defects in mitochondrial clearance predispose human monocytes to interleukin-1β hyper-secretion [34]. In the future, it would be interesting to check whether B cells isolated from MS patients bearing the NDP52GE variant exhibit a particular cytokines profile. Since we found that NDP52GE is more “mitophagic” than the WT form of the protein, we believe that this variant may prevent the release of proinflammatory cytokines (and maybe mitochondrial factors) that trigger inflammation in MS patients and most likely in other autoimmune diseases. However, we cannot exclude that, as a mitophagy and xenophagy receptor, NDP52 has other protective effects. Hence, additional studies will be needed to clarify this point.
Our work indicates that small molecule development to treat complex diseases, such as MS, may focus on compounds enhancing/stabilizing LIR-dependent interactions between NDP52 and human ATG8-family proteins, thereby opening an important path for the generation of therapeutic agents in addition to compounds that can enhance autophagy/mitophagy process.
Materials and methods
Human samples
Blood samples were collected from 238 untreated patients (females and males, age mean: 35 years) admitted to the neurological clinic of Santa Lucia Foundation of Rome or already stored in its biobank, and diagnosed as suffering from RR-MS, according to internationally established criteria [34]. The diagnosis of MS was established at the end of the diagnostic protocol by clinical, laboratory, and magnetic resonance imaging parameters, and matched published criteria. The Expanded Disability Status Scale scores were always <3 for RR-MS. All RR-MS patients were treatment-naive, and blood collection was performed at least 1 month after the last corticosteroid therapy [35]. All peripheral blood samples were collected after obtaining the written informed consent of the study participants, in accordance with the Helsinki Declaration. Most of the samples were stored in the Biobank and a small part of them comes from the project CE/AG4-PROG.246–121. The study protocol was approved by the local ethic committee of the recruitment center.
PBMCs were separated by density gradient over Ficoll-Hypaque, according to standard procedures. Briefly, heparinized blood was diluted with one volume of PBS and gently layered over the Ficoll. Following centrifugation at 660 g for 30 min, cells at the interface of the gradient were collected and washed three times.
Sorting of cells
To separate the different immune populations present in the peripheral blood, we used magnetic beads for cell isolation of Miltenyi Biotec. Specifically, we used anti human antibodies CD19 PE, CD3 PE, and CD14 PE followed by anti-PE Microbeads to separate B lymphocytes, T lymphocytes, and monocytes, respectively. After separation the samples were analyzed by CytoFLEX cytometer (Beckman Coulter) and only samples with a purity greater than 90% were considered for the study.
Count of B cells
Antibodies against CD45, CD3, CD19 and CD20 were added to 1 ml of whole blood from 31 HD and 24 patients with MS at the predetermined concentrations. After lysis of the red blood cells they were resuspended in 1 ml of PBS and analyzed by CytoFLEX (Beckman Coulter) flow cytometer that allows to have the event count per microliter.
Quantitative real-time PCR (qRT-PCR)
RNA from thawed PBMCs samples of MS and HD subjects was isolated and then retro-transcribed using the ReliaPrep RNA Cell Miniprep System (Promega) according to the manufacturer’s instruction.
The quantitative analysis of cDNA in PBMCs was performed using quantitative real-time polymerase chain reaction (PCR) for PINK1 (Primer FW: GGGAGTATGGAGCAGTCAC; Primer RW: AGCACATCAGGGTAGTCG), NDP52 (Primer FW: CCTTTCCAATTCCGTCCAG; Primer RW: TGAGTTCTGCTTCTGGAGG), OPTN (Primer FW: GGAGCACAGAGAAAGAGAATG; Primer RW: CGCTGAGTTTTGTATGAGCC), and the housekeeping gene B2M (Primer FW: CTCCGTGGCCTTAGCTGTG; Primer RW: TCTCTGCTGGATGACGTGAG). All genes content were calculated with the ΔΔCt method after normalization on B2M.
mtDNA levels assessment
We checked for the mtDNA/nuclear DNA ratio (mtDNA:nucDNA ratio) levels in PBMCs by performing RT-PCR for mitochondrial COXII (Primer FW: GTCCTGTATGCCCTTTTCCTAACACTC; Primer RV: GACCTCGTCTGTTATGTAAAGGATGCG). We used B2M as housekeeping gene (Primer FW: CTCCGTGGCCTTAGCTGTG; Primer RV: TCTCTGCTGGATGACGTGAG). DNA were isolated from PBMCs using the ReliaPrep RNA Cell Miniprep System (Promega) according to the manufacturer’s instruction. COXII content was calculated with the ΔΔCt method after normalization on B2M.
Measurement of the mitochondrial membrane potential and mitochondrial mass
Mitochondrial membrane potential (ΔΨm) was estimated by probing the fluorescent intensity. PBMCs were washed and incubated at 37 °C with 10 nM of TMRM+ for 30 min in DPBS. After the incubation, cells were washed with PBS and trypsinized off and resuspended in 0.5 ml of DPBS. TMRM+ were excited at 488 nm, and fluorescence emitted at 588 nm were measured with (FL2) channel by FACS using the CytoFLEX flow cytometer (Beckman Coulter). For each sample, approximately 300,000 lymphocytes were selected based on physical size (FSC) and grain size (SSC) parameters, and analysis was conduced using the FlowJo v10.5 software (TreeStar, Ashland, OR). The data were expressed in mean fluorescence intensity. Mitochondrial mass was analyzed by FACS analysis by using the MitoTracker Green FM dye (Thermo Fisher). HEK293 cells were exposed to 150 nM MitoTracker Green FM for 30 min at 37 °C, washed in PBS and analyzed by flow cytometry. FACS analysis were conducted using the FlowJo v10.5 software (TreeStar, Ashland, OR).
Protein oxidation assay
PBMCs from healthy or MS subjects were lysed in RIPA buffer as described above. The DPN-derivatized proteins containing carbonyl groups were evaluated by using the OxyBlot Protein Oxidation Detection Kit (Millipore S7150), according to the manufacture’s protocol.
Cell lines and primary cells cultures
HeLa and HEK293 cells were cultured in Dulbecco modified Eagle medium (GIBCO 41966-029) supplemented with 10% fetal bovine serum (FBS) (Thermo Fisher Scientific, 10270-106) at 37 °C under 5% CO2. These cells were coming from an in-house stock. All cell lines were routinely tested for mycoplasma contamination. B cells (isolated by FACS from healthy patients) and LCL cell line (kind gift form Dr Battistini Lab, IRCCS Fondazione Santa Lucia) were cultured in RPMI 1640 medium (Thermo Fisher 21875) supplemented with 10% FBS.
Cells transfection, cell interference, and treatment
Transient transfections of WT or mutant plasmids or siRNA oligo were performed using Lipofectamine 2000 according to the supplier’s instructions (Invitrogen, 11668019). Cells were treated with the combined treatment O/A [10 μM Oligomycin (Calbiochem)/4 μM Antimycin A (Sigma)] in order to induce mitophagy. Cells were treated with PMA/Iono (Sigma, 20 and 200 ng/ml, respectively) in order to stimulate B cells or with Poly (I:C) (Sigma, 50 μg/ml) in order to mimic viral infection in HeLa cells.
Plasmid, cloning, and siRNA
3x-Flag-NDP52 plasmid into p3xFLAG CMV10 vector, kindly donated by Prof. Honglin Luo (Vancouver, Canada) was subjected to mutagenesis in order to generate the 3x-FLAG-NDP52GE mutant according to the manufacture’s protocol of QuickChange site-directed mutagenesis kit (Stratagene).
(Primer FW: CAATCTCTTCCACCTCTTCCTGAGTGGTAACAACC; Primer RV: GGTTGTTACCACTCAGGAAGAGGTGGAAGAGATTG). The sequence was verified using the primer: GCATGTCTGAGTTCTGCTTCTGGAG by DNA sequencing (BioFab Research, Rome, Italy). The oligonucleotides used for mutagenesis, PCR, and DNA sequencing were purchased from Sigma. HA-LC3C vector was a gift from Dr Christian Berhendts (Boston, USA). PcDNA3 empty vector was used as control plasmid. The oligo siRNA-NDP52 was purchased from Santa Cruz Biotechnology (sc-93738).
Western blot analysis
PBMCs of RR-MS or healthy subjects were lysed with 100 μl of RIPA buffer (50 mM Tris HCl pH 7.4, 1% Triton X-100, 0.5% NP40, 150 mM NaCl, 10% Glycerol) plus protease and phosphatase inhibitor cocktail (Sigma-Aldrich). After 15 min on ice, cells were centrifuged at 11,000 g for 10 min at 4 °C. HeLa and HEK293 cells lysed in RIPA buffer plus protease inhibitor cocktail (Sigma-Aldrich, P8340), Na4VO3 0.1 mM (Sigma-Aldrich, S6508), NaF 1 mM (Sigma-Aldrich, S7920), β-Glycerophosphate 5 mM (Sigma-Aldrich, G6376) were centrifuged at 13,000 rpm for 10 min at 4 °C. PBMCs or cell lines protein concentrations were determined with the Bio-Rad Protein Assay Kit (Bio-Rad, 5000001). Proteins were separated by SDS-PAGE and transferred into PVDF membranes (Millipore). We next incubated membranes with the following antibodies: anti-PINK1 (Novus #BC100-49455), anti-NDP52 (Cell Signaling, #60732), anti-OPTN (Abcam, Ab23666), anti-FLAG (Sigma-Aldrich, F7425), anti-HA (Sigma-Aldrich, H3663), anti-TOMM20 (Santa Cruz Biotechnology, sc-FL145), anti-HSP90 (Santa Cruz Biotechnology, sc-7947), anti-COXII (Abcam, ab110258) anti-COXIV (Abcam, Ab33985), anti-Vinculin (Santa Cruz Biotechnology, sc-73614).
Densitometry analyses of the western blot were performed using the ImageJ software.
Co-Immunoprecipitation (Co-IP)
After HEK293 cell lysis, proteins were incubated with primary antibody (anti-Flag rabbit) for 24 h. Then 20 µl of protein A agarose beads (Roche, 11719408001) was added for 1 h. The beads were collected by centrifugation and washed three times with the RIPA buffer. Finally, immuno-complexes were eluted with 20 µl of SDS sample buffer and heated to 95 °C for 10 min.
Immunofluorescence analysis
HEK293 cells were washed in PBS and fixed with 4% paraformaldehyde in PBS for 10 min. After permeabilization with 0.4% Triton X-100 (Sigma-Aldrich, X-100) in PBS for 5 min, cells were incubated at 4 °C with primary antibodies (24 h) with 2% normal goat serum (Sigma-Aldrich, G9023). Cells were then washed with PBS (GIBCO, BE17-512F) and incubated for 1 h with labeled anti-mouse (Thermo Fisher Scientific, A11017-A21425) or anti-rabbit (Thermo Fisher Scientific, A11070-A21430-A31573) secondary antibodies. Then cells were stained with 1 μg/ml DAPI in order to detect the nuclei and analyzed with Zeiss LSM 700 63x oil-immersion objective (CLSM700; Jena, Germany). Colocalization measurement were performed for each transfected individual cells using ImageJ colocalization 2 plugin (Pearson value). In order to measure cellular mitochondrial area, RGB images captured were processed using ImageJ software that calculates total area occupied by mitochondria particles in the cell of interest using the outline algorithm of the “Analyse particles.” The count of ATG13 and ATG16L foci was performed using the same “analysis particles” with ImageJ software. All acquisitions were performed in non-saturated single z-confocal planes.
Genotyping and association analysis
Samples from 203 RR-MS patients and 1000 healthy controls have been subjected to automated DNA extraction, in accordance with the Declaration of Helsinki. All subjects were asked to provide written and signed informed consent. The extracted DNA was employed for the genotyping analysis through a predesigned TaqMan assay on QuantStudio® 5 Real-Time PCR System (Applied Biosystems). Each real-time PCR run was performed using a negative control and three positive samples that were previously tested by direct sequencing (BigDye Terminator v3.1) and run on ABI3130xl (Applied Biosystems) [36]. The genotyping results have been subsequently tested for biostatistical analysis, in order to evaluate the association and the effect of the identified variants. The Hardy–Weinberg equilibrium was confirmed in both cases’ and controls’ cohorts. In particular, the obtained data were evaluated by calculating the P value through a 2 × 2 (allele association) and 2 × 3 (genotype association) contingency tables. The statistical associations were considered significant for P < 0.05 with a 95% confidence interval. The strength of association was established by calculating the OR [37].
ITC and NMR samples preparation
For ITC and NMR experiments, the non-labeled or 15N-labeled LC3C and LC3B proteins were obtained based on the protocols described elsewhere [38]. The shortest NDP52 LIR-contained peptides (with Trp residue added to the N-terminus of each peptide in order to calculate peptides concentration):
NDP52128–152WENEEDILVVTTQGEVEEIEQHNKEL
NDP52128–155WENEEDILVVTTQGEVEEIEQHNKELCKE
NDP52GE128–152WENEEDILVVTTQEEVEEIEQHNKEL
NDP52GE128–155WENEEDILVVTTQEEVEEIEQHNKELCKE
were purchased from GenScript Inc (NJ, USA). The longer NDP52 fragments (WT or with mutation) were subcloned into pNIC28_Ub19_Bsa4 vector by ligation-independent cloning technology. The expression, isolation, and purification of the NDP52 fragments were performed based on the protocols described elsewhere [37]. The NDP52 constructs used in this study are overviewed in Supplementary Table 1 and presented schematically at Supplementary Fig. 3a. Before experiments, all proteins and peptides were equilibrated with a buffer containing 25 mM HEPES, 100 mM NaCl, pH 7.0; and supplied with 5 mM protease inhibitors cocktail.
Isothermal titration microcalorimetry (ITC)
The ITC experiments were performed at 25 °C using a MicroCal VP-ITC microcalorimeter (Malvern Instruments Ltd., UK). The peptides at concentrations of 0.3 mM were titrated into 0.015 mM LC3 proteins in 21 steps. The ITC data were analyzed with the ITC-Origin 7.0 software with a “one-site” binding model. The proteins and peptides concentrations were calculated from the UV-absorption at 280 nm by Nanodrop spectrophotometer (Thermo Fisher Scientific, DE, USA).
Nuclear magnetic resonance spectroscopy
All NMR experiments were performed at a sample temperature of 298 K on Bruker Avance III spectrometer operating at a proton frequencies of 950 MHz. Titration experiments were performed with a 0.190 mM 15N-labeled LC3C protein samples to which the non-labeled NDP52128–224 and NDP52GE128–224 constructs were added stepwise until four times excess to LC3C protein. CSP analysis was done according to the most recent review [39]. CSP values were calculated for each individual backbone HN resonance using the formula: Δδ = [ΔδH2 + (ΔδN/5)2]1/2. The obtained CSP values were mapped on LC3C (PDB ID 3WAM [40]) structure.
Molecular modeling
We modeled the three-dimensional structure of the complex between LC3C8–127 and the cLIR of NDP52 (NDP52125–147) using a multi-step procedure based on MODELLER version 9.15 [41]. In the first step, we collected a model of the complex with WT NDP52 using the X-ray structure of LC3C13–126-NDP52125–141 as partial template (PDB entry 3VVW, [16]) to reconstruct the missing coordinates for the N- and C-terminal residues of LC3C. We generated a set of structures and selected one, as detailed in the GitHub repository associated with the publication. We performed another round of modeling using the model collected at the first step for LC3C8–127-NDP52125–141 and the X-ray structure of the complex between FYCO11235–1253 and LC3B4–120 (PDB entry 5WRD, [21]) as templates. The alignment between LC3B and LC3C to guide the modeling was provided by Clustal Omega [42]. The crystal asymmetric unit of 5WRD contains two conformations of the FYCO1-LC3C complex; therefore, we carried out this procedure as two independent MODELLER runs using each of them as templates and generating 1000 models for each. We then selected the model having the shortest distance between the Cα atom of G140 in NDP52 and the Cζ atom of R76 in LC3C from each run. We introduced the mutation of interest (G140E) in both structures using PyMOL, selecting the rotamer of E140 characterized by the lowest number of interatomic clashes with the surrounding atoms.
Notably, as the C-terminal region of FYCO1 corresponding to our region of interest in NDP52 with respect to the core LIR motif (residues 1235–1239) is in a helical conformation which is stabilized by intermolecular electrostatic interactions [21], we constrained the helical conformation for NDP52 C-terminal region (residues 138–147) during the modeling procedure.
Molecular dynamics and metadynamics simulations
Using the two models selected above for the complex of LC3C13–126-NDP52125–141, we carried out all-atom 250-ns MD simulations in explicit solvent using GROMACS version 5.1.5 [43]. We employed the CHARMM22* force field [44] together with the TIPS3P water model [45]. To avoid artificial interactions formed by charged N- and C- terminal groups, we used a N- and C-terminal capping for both LC3C and NDP52 (see GitHub repository for details). We performed the MD simulations employing a dodecahedral box of water to solvate the protein, with a minimum distance between protein atoms and the box edges of 18 Å using periodic boundary conditions. We applied different steps of energy minimization, solvent equilibration, thermalization, and pressurization to prepare the systems for simulation (see GitHub repository). We performed the productive MD simulations using an integration step of 2 fs in the canonical ensemble with the Bussi thermostat [46] at a temperature of 298 K. We used a 9-Å cutoff for both Van Der Waals and short-range Coulomb electrostatic interactions. We also applied the particle mesh Ewald scheme for electrostatics [47]. Additional details are reported in the input files in the GitHub repository.
We used the well-tempered metadynamics (WTE-metaD) [23] to enhance the sampling of the free energy surface of salt bridge formation by adding the history-dependent bias potential on the distance between E140 Cδ of NDP52 and R76 Cζ of LC3C. Gaussians with width of 0.1 nm and height of 2.5 KJ/mol were added every 500 simulation steps (1 ps). The bias factor (γ = 10) and temperature (298 K) were kept constant throughout the simulations. A restraining potential was used to limit the sampling of the CV below 1.7 nm, with a force constant of 10,000 kJ/(mol nm2). We collected two WTE-metaD simulations starting from the two selected models of the complex LC3C8–127-NDP52125–147, to ensure convergence—in fact, two independent simulations starting from different starting structures should converge to the same free energy surface. Simulations were performed using GROMACS 2019.4 and PLUMED 2.5.3 [48].
Simulations analyses
We estimated the occurrence, in the unbiased MD ensembles, of intermolecular salt bridge and hydrogen bonds using PyInteraph [49]. We applied a distance cutoff of 4.5 and 3 Å, respectively. We also tested higher cutoffs for salt bridges (i.e., 5–6 Å) to rule out that the charged groups of E140 and positively charged residues of LC3C could be overlooked.
The free energy landscape was reconstructed from the deposited bias during the metadynamics simulation using the sum_hills tool available in PLUMED using a stride of 1000. We have used three different strategies to assess the convergence of our metadynamics simulations. The details are reported in the GitHub repository.
Detection of human TNF-α by ELISA
Human TNF-α Quantikine ELISA kits (Invitrogen) was used according to the manufacturer’s protocol. Absorbance was measured by Thermo Multiskan EX Microplate Photometer (Thermo Fisher Scientific) at 450 nm with wavelength correction set at 540 nm.
Statistical analysis
All statistical calculations were performed and graphed using GraphPad Prism 6. Comparisons between two groups were analyzed using unpaired Student t-test. Significance were defined as *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Data availability
All the software used is freely available. The scripts, input, and outputs files generated during the study will be freely available in a GitHub repository associated with our publication (https://github.com/ELELAB/LC3C_NDP52_GE_mutant). The MD trajectories will be available in Open Science Framework (OSF): https://osf.io/48wzq/.
References
Compston A, Coles A. Multiple sclerosis. Lancet. 2002;359:1221–31.
Noseworthy JH, Lucchinetti C, Rodriguez M, Weinshenker BG. Multiple sclerosis. N Engl J Med. 2000;343:938–52.
Dendrou CA, Fugger L, Friese MA. Immunopathology of multiple sclerosis. Nat Rev Immunol. 2015;15:545–58.
Brown J, Wang H, Hajishengallis GN, Martin M. TLR-signaling networks: an integration of adaptor molecules, kinases, and cross-talk. J Dent Res. 2011;90:417–27.
Su K, Bourdette D, Forte M. Mitochondrial dysfunction and neurodegeneration in multiple sclerosis. Front Physiol. 2013;4:169.
Chiurchiu V. Novel targets in multiple sclerosis: to oxidative stress and beyond. Curr Top Med Chem. 2014;14:2590–9.
Gonzalo H, Nogueras L, Gil-Sánchez A, Hervás JV, Valcheva P, González-Mingot C, et al. Impairment of mitochondrial redox status in peripheral lymphocytes of multiple sclerosis patients. Front Neurosci. 2019;13:938.
Lou G, Palikaras K, Lautrup S, Scheibye-Knudsen M, Tavernarakis N, Fang EF. Mitophagy and neuroprotection. Trends Mol Med. 2020;26:8–20.
Koyano F, Okatsu K, Kosako H, Tamura Y, Go E, Kimura M, et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature. 2014;510:162–6.
Kazlauskaite A, Kondapalli C, Gourlay R, Campbell DG, Ritorto MS, Hofmann K, et al. Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem J. 2014;460:127–39.
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL, et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature. 2015;524:309–14.
Padman BS, Nguyen TN, Uoselis L, Skulsuppaisarn M, Nguyen LK, Lazarou M. LC3/GABARAPs drive ubiquitin-independent recruitment of Optineurin and NDP52 to amplify mitophagy. Nat Commun. 2019;10:408.
Patergnani S, Castellazzi M, Bonora M, Marchi S, Casetta I, Pugliatti M, et al. Autophagy and mitophagy elements are increased in body fluids of multiple sclerosis-affected individuals. J Neurol Neurosurg Psychiatry. 2018;89:439–41.
Castellazzi M, Patergnani S, Donadio M, Giorgi C, Bonora M, Fainardi E, et al. Correlation between auto/mitophagic processes and magnetic resonance imaging activity in multiple sclerosis patients. J Neuroinflammation. 2019;16:131.
Inomata M, Niida S, Shibata KI, Into T. Regulation of toll-like receptor signaling by NDP52-mediated selective autophagy is normally inactivated by A20. Cell Mol Life Sci. 2012;69:963–79.
von Muhlinen N, Akutsu M, Ravenhill BJ, Foeglein Á, Bloor S, Rutherford TJ, et al. LC3C, bound selectively by a noncanonical LIR Motif in NDP52, is required for antibacterial autophagy. Mol Cell. 2012;48:329–42.
Skytte Rasmussen M, Mouilleron S, Kumar Shrestha B, Wirth M, Lee R, et al. ATG4B contains a C-terminal LIR motif important for binding and efficient cleavage of mammalian orthologs of yeast Atg8. Autophagy. 2017;13:834–53.
Wirth M, Zhang W, Razi M, Nyoni L, Joshi D, O’Reilly N, et al. Molecular determinants regulating selective binding of autophagy adapters and receptors to ATG8 proteins. Nat Commun. 2019;10:2055.
Cheng X, Wang Y, Gong Y, Li F, Guo Y, Hu S, Liu J, Pan L. Structural basis of FYCO1 and MAP1LC3A interaction reveals a novel binding mode for Atg8-family proteins. Autophagy. 2016;12:1330–9.
Kirkin V, Rogov VV. A diversity of selective autophagy receptors determines the specificity of the autophagy pathway. Mol Cell. 2019;76:268–85.
Johansen T, Lamark T. Selective autophagy: ATG8 family proteins, LIR motifs and cargo receptors. J Mol Biol. 2020;432:80–103.
Sakurai S, Tomita T, Shimizu T, Ohto U. The crystal structure of mouse LC3B in complex with the FYCO1 LIR reveals the importance of the flanking region of the LIR motif. Acta Crystallogr F Struct Biol Commun. 2017;73:130–7.
Laio A, Parrinello M. Escaping free-energy minima. Proc Natl Acad Sci USA. 2002;99:12562–6.
Bonomi M, Barducci A, Parrinello M. Reconstructing the equilibrium boltzmann distribution from well-tempered metadynamics. J Comput Chem. 2009;30:1615–21.
Bussi G, Laio A. Using metadynamics to explore complex free-energy landscapes. Nat Rev Phys. 2020;2:200–12.
Richter B, Sliter DA, Herhaus L, Stolz A, Wang C, Beli P, et al. Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria. Proc Natl Acad Sci USA. 2016;113:4039–44.
Moore AS, Holzbaur ELF. Dynamic recruitment and activation of ALS-associated TBK1 with its target optineurin are required for efficient mitophagy. Proc Natl Acad Sci USA. 2016;113:3349–58.
Turco E, Fracchiolla D, Martens S. Recruitment and activation of the ULK1/Atg1 kinase complex in selective autophagy. J Mol Biol. 2020;432:123–34.
Karanasios E, Stapleton E, Manifava M, Kaizuka T, Mizushima N, Walker SA, et al. Dynamic association of the ULK1 complex with omegasomes during autophagy induction. J Cell Sci. 2013;126:5224–38.
Sabatino JJ, Pröbstel AK, Zamvil SS. B cells in autoimmune and neurodegenerative central nervous system diseases. Nat Rev Neurosci. 2019;20:728–45.
Wilhelmus MM, van der Pol SM, Jansen Q, Witte ME, van der Valk P, Rozemuller A, et al. Association of Parkinson disease-related protein PINK1 with Alzheimer disease and multiple sclerosis brain lesions. Free Radic Biol Med. 2011;50:469–76.
Ellinghaus D, Zhang H, Zeissig S, Lipinski S, Till A, Jiang T, et al. Association between variants of PRDM1 and NDP52 and crohn’s disease, based on exome sequencing and functional studies. Gastroenterology. 2013;145:339–47.
Verlhac P, Grégoire IP, Azocar O, Petkova DS, Baguet J, Viret C, Faure M. Autophagy receptor NDP52 regulates pathogen-containing autophagosome maturation. Cell Host Microbe. 2015;17:515–25.
van der Burgh R, Nijhuis L, Pervolaraki K, Compeer EB, Jongeneel LH, van Gijn M, et al. Defects in mitochondrial clearance predispose human monocytes to interleukin-1β hypersecretion. J Biol Chem. 2014;289:5000–12.
McNicholas N, Hutchinson M, McGuigan C, Chataway J. 2017 McDonald diagnostic criteria: a review of the evidence. Mult Scler Relat Disord. 2018;24:48–54.
Cascella R, Strafella C, Ragazzo M, Manzo L, Costanza G, Bowes J, et al. KIF3A and IL-4 are disease-specific biomarkers for psoriatic arthritis susceptibility. Oncotarget. 2017;8:95401–11.
Cascella R, Strafella C, Longo G, Ragazzo M, Manzo L, De Felici C, et al. Uncovering genetic and non-genetic biomarkers specific for exudative age-related macular degeneration: significant association of twelve variants. Oncotarget. 2018;9:7812–21.
Rogov VV, Rozenknop A, Rogova NY, Löhr F, Tikole S, Jaravine V, et al. A universal expression tag for structural and functional studies of proteins. ChemBioChem. 2012;13:959–63.
Williamson MP. Using chemical shift perturbation to characterise ligand binding. Prog Nucl Magn Reson Spectrosc. 2013;73:1–16.
Suzuki H, Tabata K, Morita E, Kawasaki M, Kato R, Dobson RC, et al. Structural basis of the autophagy-related LC3/Atg13 LIR complex: recognition and interaction mechanism. Structure. 2014;22:47–58.
Webb B, Sali A. Comparative protein structure modeling using MODELLER. Curr Protoc Bioinformatics. 2016;54:5.6.1–5.6.37.
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol. 2011;7:539.
Abraham MJ, Murtola T, Schultz R, Pall S, Smith JC, Hess B, et al. Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 2015;1:19–25.
Piana S, Lindorff-Larsen K, Shaw DE. How robust are protein folding simulations with respect to force field parameterization? Biophys J. 2011;100:47–9.
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML. Comparison of simple potential functions for simulating liquid water. J Chem Phys. 1983;79:926.
Bussi G, Donadio D, Parrinello M. Canonical sampling through velocity rescaling. J Chem Phys. 2007;126:014101.
Harvey MJ, De Fabritiis G. An implementation of the smooth particle mesh Ewald method on GPU hardware. J Chem Theory Comput. 2009;5:2371–7.
Tribello GA, Bonomi M, Branduardi D, Camilloni C, Bussi G. PLUMED 2: new feathers for an old bird. Comput Phys Commun. 2014;185:604–13.
Tiberti M, et al. PyInteraph: a framework for the analysis of interaction networks in structural ensembles of proteins. J Chem Inf Model. 2014;54:1537–51.
Acknowledgements
We wish to thank Dr Christian Behrends for kindly providing us the construct encoding HA-LC3C; Prof. Honglin Luo for the gift of 3x-Flag-NDP52 plasmid; Dr Anna Kabanova for granting the execution of some revision experiments in her laboratory; Krenare Bruqi for her technical assistance and D.Hodder for his proofreading. We are grateful to Meike Crecelius, Natalia Rogova, and Viktoria Morasch for their help with cloning of NDP52 fragments for ITC and NMR experiments, protein/peptide sample preparation, and ITC data collection.
Funding
This work was supported by grants: ROCHE (Roche per la ricerca 2017) and 5XMILLE Italian Ministry of Health (2017) to FS. The study was partially supported by the Italian Ministry of Health (Progetto di ricerca Finalizzata RF-2018-12366111) and by the Italian Foundation for Multiple Sclerosis (Fism Progetto Speciale 2018/S/5) to LB. The work of NW, VD, and VVR was supported by the DFG-funded Collaborative Research Centre on Selective Autophagy (SFB 1177 “Molecular and Functional Characterization of Selective Autophagy”), Germany. VD and VVR also received funding from Structural Genomic Consortium (SGC). The SGC is a registered charity (no: 1097737) that receives funds from AbbVie, Bayer AG, Boehringer Ingelheim, Canada Foundation for Innovation, Eshelman Institute for Innovation, Genentech, Genome Canada through Ontario Genomics Institute [OGI-196], EU/EFPIA/OICR/McGill/KTH/Diamond, Innovative Medicines Initiative 2 Joint Undertaking [EUbOPEN grant 875510], Janssen, Merck KGaA (aka EMD in Canada and USA), Merck & Co (aka MSD outside Canada and USA), Pfizer, São Paulo Research Foundation-FAPESP, Takeda and Wellcome [106169/ZZ14/Z]. The work of VC was supported by Italian Foundation of Multiple Sclerosis (FISM grant 2017/R/08) and by Ministry of Health, Progetto Giovani Ricercatori (GR-2016-02362380). The work of MK, MT, ML, FL, VD, and EP is supported by Danmarks Grundforskningsfond (DNRF125) and a Carlsberg Foundation Distinguished Fellowship (CF18-0314) to EP’s group. Moreover, the project has been supported by a Netaji Subhash ICAR international fellowship, Government of India to MK to work in EP group. The calculations described in this paper were performed using the DeiC National Life Science Supercomputer Computerome at DTU (Denmark), and a DeiC-Pilot Grant on Abacus (Denmark).
Author information
Authors and Affiliations
Contributions
FS designed the project. ADR and FS wrote the paper. ADR performed analysis in MS patients (Fig. 1), on B cells (Fig. 5a–d) and performed the cloning of NDP52GE. TM investigated the role of NDP52 variants in the TLR signaling (Supplementary Fig. 1) and helped FS in performing Immunofluorescences of Figs. 2, 3, and 5e–g. FS characterized the binding between LC3C and NDP52 variants, performed the mitophagy analysis and Fig. 5e–g. MC helped FS in the characterization of LC3C-NDP52 interaction. PDA participated in the amplification of NDP52-LIR motif from MS patients and HD donors. DFA and LB provided PBMCs of MS patients and the LCL cell line. VC provided PBMCs of HD and MS patients and helped in preparing some cDNAs (Fig. 1). DFA performed FACS analysis (TMRM) and sorted monocytes, B and T cells from healthy patients (Fig. 5a). GC, RC, and VC performed the genetical study. MK, MT, ML, FL, VD, and EP performed Fig. 3e, f. The ITC and NMR experiments were designed by VD and VVR, and performed by NW and FL (Fig. 3 and Supplementary Fig. 2). ADR, EP, VVR, EG, and FS analyzed data. All authors commented on the final draft of the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
Refer to Human samples paragraph in the Materials and methods section.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by H. Zhang
Rights and permissions
About this article
Cite this article
Di Rita, A., Angelini, D.F., Maiorino, T. et al. Characterization of a natural variant of human NDP52 and its functional consequences on mitophagy. Cell Death Differ 28, 2499–2516 (2021). https://doi.org/10.1038/s41418-021-00766-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-021-00766-3
This article is cited by
-
The mitophagy pathway and its implications in human diseases
Signal Transduction and Targeted Therapy (2023)