Abstract
Multiple Sclerosis (MS) is a chronic autoimmune demyelinating disease of the central nervous system (CNS), with a largely unknown etiology, where mitochondrial dysfunction likely contributes to neuroaxonal loss and brain atrophy. Mirroring the CNS, peripheral immune cells from patients with MS, particularly CD4+ T cells, show inappropriate mitochondrial phenotypes and/or oxidative phosphorylation (OxPhos) insufficiency, with a still unknown contribution of mitochondrial DNA (mtDNA). We hypothesized that mitochondrial genotype in CD4+ T cells might influence MS disease activity and progression. Thus, we performed a retrospective cross-sectional and longitudinal study on patients with a recent diagnosis of either Clinically Isolated Syndrome (CIS) or Relapsing–Remitting MS (RRMS) at two timepoints: 6 months (VIS1) and 36 months (VIS2) after disease onset. Our primary outcomes were the differences in mtDNA extracted from CD4+ T cells between: (I) patients with CIS/RRMS (PwMS) at VIS1 and age- and sex-matched healthy controls (HC), in the cross-sectional analysis, and (II) different diagnostic evolutions in PwMS from VIS1 to VIS2, in the longitudinal analysis. We successfully performed mtDNA whole genome sequencing (mean coverage: 2055.77 reads/base pair) in 183 samples (61 triplets). Nonetheless, mitochondrial genotype was not associated with a diagnosis of CIS/RRMS, nor with longitudinal diagnostic evolution.
Similar content being viewed by others
Introduction
Multiple Sclerosis (MS) is a chronic neuroinflammatory and neurodegenerative disease with a largely unknown etiology, secondary to an autoimmune demyelination in the central nervous system (CNS). Histopathologically, it is characterized by gliosis, oligodendrocyte death, and neuroaxonal loss1. Worldwide, approximately 2.8 million patients with MS deal with significant levels of disability2.
Several studies have shown that the autoimmune response seen in MS mostly derives from a dysfunctional autoreactive CD4+ T cell compartment3,4,5, although other pro-inflammatory cells, such as B cells and myeloid cells, seem to be implicated as well6. Recently, mitochondria have also been shown to play a role in driving MS disease activity and progression. In the CNS, mitochondrial dysfunction has been found to be a critical trigger for neuroaxonal loss and brain atrophy7,8. In the peripheral immune compartment, CD4+ T cells from patients with MS show oxidative phosphorylation (OxPhos) insufficiency9,10,11, which, in animal models, has been linked to an exacerbation of CNS autoimmune-mediated inflammation12,13.
Mitochondrial DNA (mtDNA) is responsible for encoding 22 transfer RNAs (tRNAs), two ribosomal RNAs (rRNAs), and 13 proteins of the OxPhos chain, where multiple mtDNA molecules coexist, allowing for the existence of multiple genotypes at various variant levels (VLs) within the same cell—heteroplasmy14.
Interestingly, mtDNA mutations, as well as particular haplogroups, which are inherited mutational patterns that may be classified into phylogenetic clusters15, have been associated with an increased risk of MS, albeit not consistently16,17,18,19,20. Additionally, mtDNA polymorphisms have been shown to modulate both metabolism and immunity21,22, and mtDNA variants have significant tissue-specificity, including in T cells23. Nonetheless, whether the aforementioned CD4+ T cell OxPhos dysfunction in patients with MS is an inherent consequence of the particular mitochondrial genotype of this immune subset remains unknown.
We hypothesized that mitochondrial genotype in CD4+ T cells might influence MS disease activity and progression. Thus, we aimed to explore the differences in mtDNA extracted from CD4+ T cells between patients with a recent diagnosis of either Clinically Isolated Syndrome (CIS) or Relapsing–Remitting MS (RRMS) and healthy controls (HC). We also analyzed longitudinal mtDNA changes in patients with CIS/RRMS (PwMS).
Methods
Cohort description
We performed an observational retrospective evaluation of prospectively collected data within the Berlin CIS-Cohort (reference: NCT01371071)24 at the Charité—Universitätsmedizin Berlin, Berlin, Germany. Berlin CIS-Cohort’s inclusion criteria are: (I) age ≥ 18 years and (II) diagnosis of either Clinically Isolated Syndrome (CIS) within 6 months from symptom onset or of Relapsing–Remitting Multiple Sclerosis (RRMS) within two years from symptom onset, according to the 2017 revisions of the McDonald criteria25.
To address the disease activity and progression of patients with CIS/RRMS (PwMS), the following variables were assessed on each clinical visit: number of relapses and time to last relapse; expanded disability status scale (EDSS) and Multiple Sclerosis functional composite (MSFC) scores; brain MRI: T2 hyperintense lesions and T1 gadolinium-enhancing lesions; and optical coherence tomography (OCT): peripapillary retinal nerve fiber layer (RNFL) thickness and ganglion cell-inner plexiform layer (GCIPL) volume. Additionally, we assessed if patients fulfilled the criteria for no evidence of disease activity (NEDA)-3, namely, absence of new relapses, on MRI: no T1 gadolinium-enhancing lesions and no new or enlarging T2 hyperintense lesions and the absence of EDSS worsening. Details of MRI, OCT, and NEDA-3 were described earlier26. Finally, blood samples were collected and, subsequently, peripheral blood mononuclear cells (PBMCs) were isolated. Further details on the methodology used for collection and analysis of clinical data may be found in Sect. 1 of the Supplementary Information.
PwMS were included in this study if biological samples and a clinical assessment were available for two clinical visits: 6 months (VIS1) and 36 months (VIS2) after disease onset. Recruitment of HC without a family history of MS was finalized in May 2019; HC were matched to PwMS at VIS1 in a 1:1 ratio according to sex and a maximum age difference of five years.
Our primary outcomes were a mitochondrial genotype comparison between: (I) PwMS and HC in the cross-sectional analysis and (II) different diagnostic evolutions from VIS1 to VIS2 in the longitudinal analysis, namely, CIS in both clinical visits, a conversion from CIS to RRMS, and RRMS in both clinical visits.
Since mtDNA whole genome sequencing (WGS) in CD4+ T cells from PwMS was unreported in the literature, a pilot study with 20 triplets (20 PwMS at VIS1&VIS2 and 20 age- and sex-matched HC) was performed to determine the appropriateness of the sample size through the Dupont method27.
Ethical approval
The institutional review board (IRB)’s approval was obtained by the Ethics Committee of the Charité—Universitätsmedizin Berlin (application number: EA1/182/10), informed consents were given by every subject, and the study followed the standards of the Declaration of Helsinki28.
Sample processing
CD4 + T cell enrichment and flow cytometry analysis
PBMC sample processing was performed at the same time for each triplet (PwMS at VIS1&VIS2 and HC), to minimize differences within processing. The MojoSort™ Human CD4 T Cell Isolation Kit (#480010, BioLegend, San Diego, CA, USA) was used for CD4+ T cell enrichment, in accordance with the manufacturer’s instructions.
To assess whether CD4+ T cell enrichment was achieved, flow cytometry was performed in a BD LSRFortessa™ X-20 Cell Analyzer (BD Biosciences, Franklin Lakes, NJ, USA) on a subset of samples (before and after magnetic enrichment). The following antibodies were used: CD14-eFluor450 (#48014941, clone 61D3, eBioscience™, Thermo Fisher Scientific, Waltham, MA, USA); CD19-Alexa Fluor 647 (#302222, clone HIB19, BioLegend, San Diego, CA, USA); CD3-PerCP-Cy5.5 (#45003741, clone OKT3, eBioscience™, Thermo Fisher Scientific, Waltham, MA, USA); CD4-BV711 (#317439, clone OKT4, BioLegend, San Diego, CA, USA); and CD56-PE-Cy7 (#25056741, clone CMSSB/NCAM, eBioscience™, Thermo Fisher Scientific, Waltham, MA, USA).
Regarding the gating strategy, CD4+ T cells were defined as CD3+CD4+CD19-CD56- single cells (see Supplementary Figs. S6, 7 online). A detailed protocol and additional details regarding the methodology used for CD4+ T cell enrichment and flow cytometry may be found in Sect. 2 of the Supplementary Information.
DNA extraction and mtDNA sequencing, including bioinformatic processing and data analysis
Similarly, DNA extraction was performed at the same time for each triplet with the QIAamp® DNA Blood Midi Kit (QIAGEN GmbH, Hilden, Germany), according to the manufacturer’s instructions.
Samples were sequenced with the Applied Biosystems™ Precision ID mtDNA Whole Genome Panel (Thermo Fisher Scientific, Waltham, MA, USA) at IPATIMUP—Instituto de Patologia e Imunologia Molecular da Universidade do Porto (Porto, Portugal). Libraries were prepared using the Ion Chef™ automated protocol and samples were run on 530™ chips with the Ion Torrent™ Ion S5™ (Thermo Fisher Scientific, Waltham, MA, USA). Samples with a coverage uniformity < 85% and/or a mean coverage < 1500 reads were resequenced.
Regarding bioinformatic processing, the PrecisionCallerPipeline (PCP) pipeline29 was used, with mutserve v230,31 for variant calling, HaploGrep v2.4.015 for haplogroup calling, and Haplocheck v1.3.331 for a contamination check. To account for possible false positives, variants with a variant level (VL) ≥ 10% were only accepted if they were found with both the PCP pipeline and the Ion Torrent Suite™ Software (TSS), while variants below TSS’s limit of detection (10%) and solely present with PCP where filtered in accordance with: (I) sequencing indicators for variant reliability (normalized coverage, coverage ratio, mean value of reported nuclear insertions of mitochondrial DNA [NUMTs], and the distance to the amplicon’s edge); and (II) previous reports of the same variants in curated mtDNA databases. Variants only present in TSS were excluded29.
Data analysis was performed with R version 4.1.132 and Excel 2016 (Microsoft Corporation, Redmond, WA, USA).
The level of statistical significance was set at a two-sided p-value < 0.05 for all tests and multiple comparison testing was adjusted with false discovery rate (FDR). Whenever data was missing, it was censored from the analysis. Reporting followed the standards from STROBE33 and its extension STREGA34.
A detailed description of the methodology regarding DNA extraction, bioinformatic processing, and data analysis may be found in Sect. 3 of the Supplementary Information.
Results
Cohort description
Overall, 61 PwMS were included in this study cohort (Fig. 1, Table 1). Most patients presented with RRMS at both clinical visits and approximately one third of the PwMS (N = 20) suffered a relapse between VIS1 and VIS2, while PwMS under MS immunomodulatory treatment increased from 26.23% to 47.54%.
Clinical cohort flowchart. 61 newly diagnosed PwMS were included in this study cohort and reassessed after 30.50 ± 1.27 (mean ± SD) months. Additionally, HC without a family history of MS were matched to PwMS at cohort entry in a 1:1 ratio according to sex and age, with an average age difference of 2.06 ± 1.31 (mean ± SD) years. Mean time from disease onset is depicted followed by the minimum and maximum values between brackets. Figures and arrows were adapted from OpenMoji. License: CC BY-SA 4.0. HC healthy control(s); MS Multiple Sclerosis; N number; PwMS patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis; V1 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1; V2 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 2.
As mentioned previously, a pilot study with 20 triplets (20 PwMS at VIS1&VIS2 and 20 age- and sex-matched HC) was performed to determine the appropriateness of the sample size through the Dupont method27. Following WGS data analysis of this subset, a 35% discordance in prevalence was detected for deleterious variants in Complex I—55% for PwMS at VIS1 (discordant prevalence: 45%) vs. 20% for HC (discordant prevalence: 10%). Hence, the sample size was adequate according to this endpoint.
CD4 + T cell enrichment, DNA extraction, and WGS quality
Following the magnetic enrichment in CD4+ T cells, we obtained a wide range in the number of cells and cell mortality, for all subject types (Table 2 and online Supplementary Table S6). In comparison with HC, PwMS at VIS1 showed a lower number of cells (mean difference of 0.65 million, 95% confidence interval [CI] 0.16–1.14; paired t-test adjusted with FDR). In comparison with VIS2, cells from PwMS at VIS1 had lower mortality (mean difference of 2.77%, 95% CI 0.56–4.97%; paired t-test adjusted with FDR).
Nonetheless, a significant increase in CD4+ T cells (CD3+ CD4+ CD56- CD19-) was achieved (Fig. 2a and online Supplementary Table S4: mean fold change in paired samples of 2.59, 95% CI 2.31–2.87; paired t-test, N = 49), regardless of subject type (see Supplementary Table S5 online).
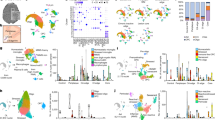
CD4+ T cell enrichment and mitochondrial DNA whole genome sequencing. (a) Percentage of CD4+ T cells (CD3+ CD4+ CD56- CD19-) before and after magnetic enrichment, as assessed with flow cytometry—lines connect the same sample; (b) Mean mappability, per subject type and per definition—error bars denote minimum and maximum values; (c) Mean mtDNA coverage, per subject type; (d) Median mtDNA coverage, per subject type. HC healthy control(s); mtDNA mitochondrial DNA; V1 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1; V2 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 2.
Subsequently, after the extraction of DNA, variable amounts of DNA were achieved (Table 2 and online Supplementary Table S6). In comparison with PwMS at VIS1, HC had higher yields of extracted DNA (mean difference of 4.45 ng/μL for the measurement with Synergy HTX, 95% CI 1.06–7.84, and mean difference of 3.35 ng/μL for the measurement with Qubit®, 95% CI 0.80–5.90; paired t-tests adjusted with FDR).
However, while, as expected, the two DNA measurements showed a correlation between each other (adjusted R2 of 0.88 and a p-value < 2.2 × 10–16, linear regression model) and the number of cells used for DNA extraction correlated with DNA concentration (adjusted R2 of 0.77 and a p-value < 2.2 × 10–16, linear regression model), DNA concentration had no effect on mtDNA coverage, with most variability appearing to arise from each sequencing run (see Supplementary Fig. S10 online). Correspondingly, no differences in mtDNA coverage and mappability were found between different subject types (Fig. 2b–d and online Supplementary Fig. S10).
Taking into account all samples analyzed through WGS (see Supplementary Table S8 online), no contamination was detected and all haplogroups corresponded to European lineages. PwMS had the same haplogroup with two minor exceptions (Triplets #5 and #48) at both visits, albeit without changing their simplified haplogroup (see Supplementary Table S8 online).
Cross-sectional comparison
The total number of variants was similar between different subject types, i.e., between HC and PwMS at VIS1 and between VIS1 and VIS2 for PwMS (Fig. 3 and online Supplementary Table S8: paired t-tests adjusted with FDR). Variant distribution was also bimodal in all subject types (Fig. 3a), likely arising from each sample’s haplogroup (Fig. 3b–c), since the mean number of variants in HC and PwMS at VIS1 was significantly different in haplogroups with at least three samples (p-value 3.34 × 10–16; Kruskal–Wallis test). Haplogroups H and HV had significantly fewer variants (see Supplementary Table S9 online), which is consistent with the revised Cambridge Reference Sequence (rCRS)’s own H haplogroup35. Nonetheless, haplogroup distribution was independent from subject type (HC and PwMS at VIS1; Fisher’s exact test with Monte Carlo simulations).
Total number of variants: Cross-sectional comparison (subject types and haplogroups). (a) Number of variants, per subject type; (b) Number of variants, per subject type and per haplogroup; (c) Expansion of (b) for better visualization. HC healthy control(s); V1 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1; V2 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 2.
When comparing HC and PwMS at VIS1, we observed that the total number of variants did not vary according to age (linear regression model). The difference in the number of mutations between HC and PwMS at VIS1 was also not influenced by sex (two sample t-test, χ2 test of independence, and two-proportions z-test) (see Supplementary Fig. S11 online).
When looking at the differences between PwMS at VIS1 and HC in various mtDNA regions, the highest discordancy in prevalence was in MT-ND3 for PwMS at VIS1 and in MT-RNR2 for HC, while the highest discordant adjusted mutational rate, which is the sum of all VLs in a specific region divided by its region size, was in MT-TA for PwMS at VIS1 and in MT-TT for HC. However, there were no significant differences for discordant prevalence (McNemar’s tests adjusted with FDR); nor for discordant mutational rates (paired t-tests adjusted with FDR) (see Supplementary Fig. S12 online).
Regarding macro regions in mtDNA, the most commonly discordant affected region for PwMS at VIS1 was Other, which refers to positions in rCRS with an overlap between the two strands or left unannotated, and, for HC, transfer RNA (tRNA), while the region with the highest discordant adjusted mutational rate was Complex III for PwMS at VIS1 and Other for HC. Nonetheless, there were no significant differences for discordant prevalence (McNemar’s tests adjusted with FDR); nor for discordant mutational rates (paired t-tests adjusted with FDR) (see Supplementary Fig. S12 online).
We further took all variants into account (see Supplementary Table S10 online), to see if individual variants differed between HC and PwMS at VIS1. However, no significant differences were found (McNemar’s tests adjusted with FDR). Interestingly, a single PwMS had a pathogenic variant, namely, m.11778G > A in the MT-ND4 gene, which is associated with Leber’s hereditary optic neuropathy (LHON) and neuropathy, ataxia, and retinitis pigmentosa (NARP), albeit at low VLs: 5.8% and 7.3% for VIS1 and VIS2, respectively; well below the VLs usually required for pathogenesis36. The patient in question initially presented with pyramidal and sensory complaints, maintaining the latter throughout their disease course, without any visual changes.
Subsequently, we took into account variants predicted in silico to be likely deleterious for proteins, defined by a mean value > 0.5 from two independent scores: (I) MutPred37,38; and (II) APOGEE39. Nevertheless, the number of deleterious variants (see Supplementary Fig. S13 online) and the cumulative deleterious burden (Fig. 4a), which corresponds to the total sum of a variant’s VL multiplied by its deleterious score per sample, did not differ significantly between the different subject types (paired t-tests adjusted with FDR).
Cumulative deleterious burden: Cross-sectional comparison (subject types and haplogroups). (a) Cumulative deleterious burden, per subject type; (b) Cumulative deleterious burden, per subject type and haplogroup; (c) Expansion of (b) for better visualization. HC healthy control(s); V1 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1; V2 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 2.
We observed a similar haplogroup-specific effect for both the number of deleterious mutations and cumulative deleterious burden (online Supplementary Fig. S13 and Fig. 4b,c: p-values 4.61 × 10–13 and 1.38 × 10–15, respectively; Kruskal–Wallis tests). Haplogroups J and T had significantly more deleterious variants (see Supplementary Table S11 online) and cumulative deleterious burden (see Supplementary Table S12 online).
When comparing HC and PwMS at VIS1, the number of deleterious variants did not vary according to age (Kendall rank correlation test). The difference in the number of deleterious mutations between HC and PwMS at VIS1 was also not influenced by sex (two sample t-test, χ2 test of independence, and two-proportions z-test). The same was true for cumulative deleterious burden (linear regression model for age and two sample t-test for sex) (see Supplementary Fig. S14 online).
Regarding differences between PwMS at VIS1 and HC in various mtDNA regions, the highest discordancy in prevalence was in MT-ND5 for PwMS at VIS1 and in MT-ATP6 for HC (Fig. 5a). In parallel, the highest discordant adjusted cumulative deleterious rate, which is the cumulative deleterious rate for a specific region relative to its region size, was in MT-ND3 for PwMS at VIS1 and in MT-ATP6 for HC (Fig. 5b). Regardless, no significant differences were found for discordant prevalence (Fig. 5a: McNemar’s tests adjusted with FDR); nor for discordant cumulative deleterious rates (Fig. 5b: paired t-tests adjusted with FDR).
Deleterious variants: Cross-sectional comparison (regions). (a,c) Discordant prevalence of deleterious variants for each mtDNA coding region/locus and for each macro mtDNA coding region, per subject type, respectively; (b,d) Relative cumulative deleterious burden for each mtDNA coding region/locus and for each macro mtDNA coding region, per subject type, respectively. bp base pair; HC healthy control(s); V1 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1.
Regarding macro regions in mtDNA, the most commonly discordant affected region was Complex I for PwMS at VIS1 and Complex V for HC (Fig. 5c), while the region with the highest adjusted cumulative deleterious rate was Complex III for PwMS at VIS1 and Complex V for HC (Fig. 5d). However, there were no significant differences for discordant prevalence (Fig. 5c: McNemar’s tests adjusted with FDR); nor for cumulative deleterious rate (Fig. 5d: paired t-tests adjusted with FDR).
Finally, we looked into tRNA mutations predicted to be pathogenic according to the MitoTIP score40: no variants were found (see Supplementary Fig. S15 online). The cumulative MitoTIP score, which corresponds to the total sum of a variant’s VL multiplied by its MitoTIP score per sample, was similar in all subject types (paired t-tests adjusted with FDR).
Nonetheless, as observed previously, haplogroup had a significant influence in the cumulative MitoTIP score (p-value 3.59 × 10–13; Kruskal–Wallis test), with haplogroups K and U having higher cumulative MitoTIP scores, whereas haplogroups H and HV had lower scores (see Supplementary Table S13 online). No influence was found regarding age (linear regression model) or sex (two sample t-test). The region with the highest discordant adjusted cumulative MitoTIP rate, which is the cumulative MitoTIP rate for a specific region relative to its region size, was MT-TA for PwMS at VIS1 and MT-TT for HC. However, no significant differences were found (paired t-tests adjusted with FDR) (see Supplementary Fig. S15 online).
Longitudinal comparison
As mentioned previously, there were no significant differences between PwMS at VIS1 and at VIS2 (paired t-tests adjusted with FDR) concerning total number of variants (Fig. 3a), number of deleterious variants (see Supplementary Fig. S13 online), cumulative deleterious burden (Fig. 4a), and cumulative MitoTIP score (see Supplementary Fig. S15 online).
The mean proportion of mutations present in a single visit—transient variants—were 4.01% (range: 0.00–53.33%) for VIS1 and 3.38% (range: 0.00–37.14%) for VIS2 (Fig. 6a), with mean VLs of 8.39% (range: 2.50–100.00%) for VIS1 and 4.66% (range: 2.50–19.60%) for VIS2 (Fig. 6b); no significant differences were found between different visits (paired t-tests).
Longitudinal changes in PwMS: Intrapersonal changes. (a) Proportion of transient variants, per subject and clinical visit; (b) Mean VL of transient variants, per subject and clinical visit; (c,d) Density plots of VL change in persistent variants, for all mutations and per subject, respectively. PwMS patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis; V1 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1; V2 patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 2; VL variant level.
Subsequently, we considered the difference in VLs for the previously found variants between the second and the first visits—persistent variants. The mean VL change was 0.06% in all variants (Fig. 6c) and 0.09% per subject (Fig. 6d) (range: − 10.90–50.30%). Both VL changes were statistically similar to zero (one-sample t-test).
When we compared transient and persistent variants, transient variants had a significantly lower VL than persistent variants (adjusted p-value 1.80 × 10–150; two-sample t-test adjusted with FDR) and a lower coverage ratio (adjusted p-value 1.09 × 10–6; two-sample t-test adjusted with FDR). There were no significant differences for other quality control variables29 (two-sample t-tests adjusted with FDR), nor for protein deleterious score and MitoTIP score (two-sample t-tests adjusted with FDR) (see Supplementary Fig. S16 online).
Since each PwMS had a very similar mutational pattern between the two visits, we then moved on from an intrapersonal comparison to an interpersonal comparison, with a focus on exploring the origin of these transient variants and the VL change in persistent variants in the full PwMS cohort. For that purpose, we considered a PwMS as a single data point, where we performed the mean of the results from both sequencing runs.
Afterwards, we considered the various possible causes or cofactors behind differences in the proportion of transient variants, VLs of transient variants or VL change in persistent variants. Haplogroup (Kruskal–Wallis test), age (linear regression model), and sex (two sample t-test) did not influence the aforementioned WGS variables (see Supplementary Figs. S17–19 online). Interestingly, different regions had a significant effect on the proportion of transient variants, excluding patients without transient variants and regions with mutations from fewer than three paired samples (p-value 5.90 × 10–6; Kruskal–Wallis test), where MT-TI had the highest prevalence and D-loop the lowest (see Supplementary Table S14 online). This effect was absent for VLs of transient variants or VL change in persistent variants (Kruskal–Wallis test) (see Supplementary Figs. S18–19 online).
Regarding the clinical variables in the cohort, change in MS medication status, i.e., Yes vs. No (Table 1 and online Supplementary Table S15), was independent from diagnostic evolution (Fisher’s exact test with Monte Carlo simulations) (see Supplementary Fig. S20 online).
To identify associations of clinical disability with diagnostic evolution, we performed batch Kruskal–Wallis tests with diagnostic evolution as a grouping variable for each available clinical variable (see Supplementary Figs. S21, 22 online). Two different approaches were tested: (I) the mean of each clinical variable between VIS1 and VIS2; and (II) the change of each clinical variable from VIS1 to VIS2, by subtracting VIS2 and VIS1. After multiple comparison correction with FDR, only the mean values of the number and volume of T2 hyperintense lesions were associated with diagnostic evolution (see Supplementary Table S16 online).
Similarly, we performed batch Kruskal–Wallis tests with diagnostic evolution as a grouping variable for each WGS variable explained previously (see Supplementary Fig. S23 online); no significant differences were found (see Supplementary Table S17 online). Additionally, haplogroup was independent from diagnostic evolution (Fisher’s exact test with Monte Carlo simulations) (see Supplementary Fig. S24 online). Similarly, as a post hoc analysis, dividing patients according to NEDA-3 status (Table 1 and online Supplementary Fig. S25) or cumulative deleterious burden (see Supplementary Figs. S26, 27 online) yielded null results.
Discussion
The present study investigated whether mitochondrial genotype in CD4+ T cells was associated with a diagnosis of CIS/RRMS or with longitudinal diagnostic evolution. Accordingly, we performed WGS from CD4+ T cells in a matched cohort of PwMS. Overall, however, there were no significant differences regarding number of variants, number of deleterious variants, cumulative protein deleterious burden, or cumulative MitoTIP score between PwMS and HC, nor after a mean of 30.50 months of follow-up between VIS1 and VIS2 for PwMS.
Importantly, the CIS/RRMS cohort herein described is representative of the overall population of PwMS in Germany, regarding mean age at onset, female predominance, and distribution of the diagnoses, with a vast majority of RRMS41.
According to the flow cytometry analysis performed in a subset of magnetically enriched samples, CD4+ T cell enrichment was successful with the MojoSort™ Human CD4 T Cell Isolation Kit, which was expected42. The Precision ID mtDNA Whole Genome Panel, in combination with the PCP pipeline29, proved particularly valuable, as it was able to obtain mtDNA WGS from all samples without signs of contamination, despite a wide range of the number of cells after processing the samples, and, consequently, of DNA yield, which is consistent with previous reports43.
Curiously, amongst the 21 most prevalent mutations in PwMS, five had already been associated with MS, ranging from case reports to large epidemiological studies16,18,44,45,46. Furthermore, remarkably, the high number of deleterious variants in complex I and IV matches the pattern of decreased activity in these complexes in CD4+ T cells of PwMS in previous studies9,47. Moreover, the higher number of deleterious variants and cumulative deleterious burden from haplogroups J and T is consistent with previous reports of increased MS risk for these haplogroups16,17.
Nevertheless, in our cohort, the mitochondrial genotype did not differ significantly between PwMS and HC, nor between VIS1 and VIS2 within PwMS, which is consistent with previous mtDNA WGS studies in PwMS19,20.
In contrast to the 35% discordance in prevalence for deleterious variants in Complex I registered in the pilot study, the final difference was 18%, due to an increase in prevalence of 8% for HC, while in PwMS it decreased by 9%. Thus, our sample size was incompatible with the previously-set endpoint.
A further limitation might have also been the mtDNA WGS of the very heterogeneous CD4+ T cell compartment, as well as its magnetic enrichment, as opposed to other methods aimed at achieving higher cell purity. Nonetheless, previous studies focused on CD4+ T cell OxPhos dysfunction in patients with MS employing magnetic enrichment and fewer subjects than the present cohort were still successful in defining a clear mitochondrial phenotype9,47. Furthermore, several existing disease modifying drugs have been shown to interfere with CD4+ T cell metabolism10,47,48,49,50, which substantiates the relevance of these pathways in MS pathophysiology and points towards new and intriguing possibilities of pharmaceutical development.
Finally, since we have only explored cross-sectional differences between PwMS at VIS1 and HC, we cannot exclude the possibility of having different longitudinal mtDNA mutational rates between PwMS and HC.
Despite these limitations, this study constitutes a thorough cross-sectional and longitudinal analysis of mtDNA WGS in PwMS, in addition to surveying mitochondrial genotype specifically in CD4+ T cells. Moreover, we were able to assess the longitudinal dynamics of mtDNA, which have only very recently started to be unveiled51,52.
Overall, CD4+ T cell mitochondrial genotype was not associated with a diagnosis of CIS/RRMS, nor with longitudinal diagnostic evolution. We further postulate that mitochondrial dysfunction in CD4+ T cells is unlikely to derive from mitochondrial genotype.
Data availability
Participant consent and sample collection mostly preceded the enforcement of the European Union’s General Data Protection Regulation (EU GDPR) and did not include sharing of individual study data, particularly the publication of highly sensitive genetic data. In compliance with both the EU GDPR, German law, and Charité’s internal policies, pseudonymized data may be made available for the purpose of replication of results upon reasonable request from the corresponding authors. Requests for data access will be considered from nonprofit research institutions and subject to approval by the Charité’s Data Protection Officer within 120 days of submission. Co-authorship is expected in subsequent publications utilizing data from this manuscript.
Abbreviations
- bp(s):
-
Base pair(s)
- CI:
-
Confidence interval
- CIS:
-
Clinically Isolated Syndrome
- EDSS:
-
Expanded disability status scale
- FDR:
-
False discovery rate
- GCIPL:
-
Ganglion cell-inner plexiform layer
- gd:
-
Gadolinium
- HC:
-
Healthy control(s)
- IRB:
-
Institutional review board
- les:
-
Lesions
- LHON:
-
Leber’s hereditary optic neuropathy
- MS:
-
Multiple Sclerosis
- MSFC:
-
Multiple Sclerosis functional composite
- mtDNA:
-
Mitochondrial DNA
- N:
-
Number
- NARP:
-
Neuropathy, ataxia, and retinitis pigmentosa
- NEDA:
-
No evidence of disease activity
- NUMT(s):
-
Nuclear insertion(s) of mitochondrial DNA
- OCT:
-
Optical coherence tomography
- OxPhos:
-
Oxidative phosphorylation
- PBMC:
-
Peripheral blood mononuclear cell
- PCP:
-
PrecisionCallerPipeline
- PwMS:
-
Patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis
- rCRS:
-
Revised Cambridge Reference Sequence
- RRMS:
-
Relapsing-Remitting Multiple Sclerosis
- rRNA:
-
Ribosomal RNA
- tRNA:
-
Transfer RNA
- TSS:
-
Ion Torrent Suite™ Software
- V:
-
Volume
- V1:
-
Patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 1, 6 months after disease onset
- V2:
-
Patient(s) with Clinically Isolated Syndrome/Relapsing–Remitting Multiple Sclerosis at visit 2, 36 months after disease onset
- VIS1:
-
Visit 1
- VIS2:
-
Visit 2
- VL(s):
-
Variant level(s)
- WGS:
-
Whole genome sequencing
References
Dendrou, C. A., Fugger, L. & Friese, M. A. Immunopathology of multiple sclerosis. Nat. Rev. Immunol. 15, 545–558 (2015).
Multiple Sclerosis International Federation. Atlas of MS – 3rd Edition. Part 1: Mapping multiple sclerosis around the world. Key epidemiology findings. (2020).
Gerdes, L. A. et al. Immune signatures of prodromal multiple sclerosis in monozygotic twins. Proc. Natl. Acad. Sci. 117, 21546–21556 (2020).
Cruciani, C. et al. T-cell specificity influences disease heterogeneity in Multiple Sclerosis. Neurol. Neuroimmunol. Neuroinflammation 8, e1075 (2021).
Böttcher, C. et al. Multi-parameter immune profiling of peripheral blood mononuclear cells by multiplexed single-cell mass cytometry in patients with early multiple sclerosis. Sci. Rep. 9, 19471 (2019).
Bar-Or, A. & Li, R. Cellular immunology of relapsing multiple sclerosis: Interactions, checks, and balances. Lancet Neurol. 20, 470–483 (2021).
Campbell, G. R. et al. Mitochondrial DNA deletions and neurodegeneration in multiple sclerosis. Ann. Neurol. 69, 481–492 (2011).
Witte, M. E. et al. Reduced expression of PGC-1α partly underlies mitochondrial changes and correlates with neuronal loss in multiple sclerosis cortex. Acta Neuropathol. (Berl.) 125, 231–243 (2013).
De Riccardis, L. et al. Bioenergetics profile of CD4 + T cells in relapsing remitting multiple sclerosis subjects. J. Biotechnol. 202, 31–39 (2015).
La Rocca, C. et al. Immunometabolic profiling of T cells from patients with relapsing-remitting multiple sclerosis reveals an impairment in glycolysis and mitochondrial respiration. Metabolism 77, 39–46 (2017).
De Biasi, S. et al. Mitochondrial functionality and metabolism in T cells from progressive multiple sclerosis patients. Eur. J. Immunol. 49, 2204–2221 (2019).
Zhang, D. et al. High glucose intake exacerbates autoimmunity through reactive-oxygen-species-mediated TGF-β cytokine activation. Immunity 51, 671-681.e5 (2019).
Luu, M. et al. The short-chain fatty acid pentanoate suppresses autoimmunity by modulating the metabolic-epigenetic crosstalk in lymphocytes. Nat. Commun. 10, 760 (2019).
Taylor, R. W. & Turnbull, D. M. Mitochondrial DNA mutations in human disease. Nat. Rev. Genet. 6, 389–402 (2005).
Weissensteiner, H. et al. HaploGrep 2: Mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res. 44, W58-63 (2016).
Yu, X. et al. mtDNA nt13708A Variant Increases the Risk of Multiple Sclerosis. PLOS ONE 3, e1530 (2008).
Tranah, G. J. et al. Mitochondrial DNA sequence variation in multiple sclerosis. Neurology 85, 325–330 (2015).
Yonova-Doing, E. et al. An atlas of mitochondrial DNA genotype-phenotype associations in the UK Biobank. Nat. Genet. 53, 982–993 (2021).
Souren, N. Y. P. et al. Mitochondrial DNA variation and heteroplasmy in monozygotic twins clinically discordant for Multiple Sclerosis. Hum. Mutat. 37, 765–775 (2016).
Pienaar, I. S. et al. Investigation of the correlation between mildly deleterious mtDNA Variations and the clinical progression of multiple sclerosis. Mult. Scler. Relat. Disord. 53, 103055 (2021).
Uittenbogaard, M. et al. The m.11778 A > G variant associated with the coexistence of Leber’s hereditary optic neuropathy and multiple sclerosis-like illness dysregulates the metabolic interplay between mitochondrial oxidative phosphorylation and glycolysis. Mitochondrion 46, 187–194 (2019).
Beadnell, T. C. et al. Mitochondrial genetics cooperate with nuclear genetics to selectively alter immune cell development/trafficking. Biochim. Biophys. Acta BBA Mol. Basis Dis. 1866, 165648 (2020).
Walker, M. A. et al. Purifying selection against pathogenic mitochondrial DNA in human T cells. N. Engl. J. Med. 383, 1556–1563 (2020).
Charite University, Berlin, Germany. Clinically Isolated Syndrome and Newly Diagnosed Multiple Sclerosis: Diagnostic, Prognostic and Therapy - Response Markers - a Prospective Observational Study (Berlin CIS-COHORT). https://clinicaltrials.gov/ct2/show/NCT01371071 (2011).
Thompson, A. J. et al. Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. Lancet Neurol. 17, 162–173 (2018).
Lin, T.-Y. et al. Increased serum neurofilament light and thin ganglion cell-inner plexiform layer are additive risk factors for disease activity in early Multiple Sclerosis. Neurol. Neuroimmunol. Neuroinflammation 8, e1051 (2021).
Dupont, W. D. Power calculations for matched case-control studies. Biometrics 44, 1157–1168 (1988).
World Medical Association. World Medical Association Declaration of Helsinki: Ethical principles for medical research involving human subjects. JAMA 310, 2191–2194 (2013).
Cortes-Figueiredo, F. et al. From forensics to clinical research: Expanding the variant calling pipeline for the precision ID mtDNA whole genome panel. Int. J. Mol. Sci. 22, 12031 (2021).
Weissensteiner, H. et al. mtDNA-Server: Next-generation sequencing data analysis of human mitochondrial DNA in the cloud. Nucleic Acids Res. 44, W64-69 (2016).
Weissensteiner, H. et al. Contamination detection in sequencing studies using the mitochondrial phylogeny. Genome Res. https://doi.org/10.1101/gr.256545.119 (2021).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing (2021).
von Elm, E. et al. The strengthening the reporting of observational studies in epidemiology (STROBE) statement: Guidelines for reporting observational studies. Epidemiol. Camb. Mass 18, 800–804 (2007).
Little, J. et al. STrengthening the REporting of Genetic Association Studies (STREGA): An extension of the STROBE statement. PLoS Med. 6, e22 (2009).
Bandelt, H.-J., Kloss-Brandstätter, A., Richards, M. B., Yao, Y.-G. & Logan, I. The case for the continuing use of the revised Cambridge Reference Sequence (rCRS) and the standardization of notation in human mitochondrial DNA studies. J. Hum. Genet. 59, 66–77 (2014).
Chung, C.-Y., Valdebenito, G. E., Chacko, A. R. & Duchen, M. R. Rewiring cell signalling pathways in pathogenic mtDNA mutations. Trends Cell Biol. https://doi.org/10.1016/j.tcb.2021.10.005 (2021).
Li, B. et al. Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinforma. Oxf. Engl. 25, 2744–2750 (2009).
Pereira, L., Soares, P., Radivojac, P., Li, B. & Samuels, D. C. Comparing phylogeny and the predicted pathogenicity of protein variations reveals equal purifying selection across the global human mtDNA diversity. Am. J. Hum. Genet. 88, 433–439 (2011).
Castellana, S. et al. High-confidence assessment of functional impact of human mitochondrial non-synonymous genome variations by APOGEE. PLoS Comput. Biol. 13, e1005628 (2017).
Sonney, S. et al. Predicting the pathogenicity of novel variants in mitochondrial tRNA with MitoTIP. PLOS Comput. Biol. 13, e1005867 (2017).
Ohle, L.-M. et al. Chances and challenges of a long-term data repository in multiple sclerosis: 20th birthday of the German MS registry. Sci. Rep. 11, 13340 (2021).
Godinho-Santos, A. et al. Follicular helper T cells are major human immunodeficiency virus-2 reservoirs and support productive infection. J. Infect. Dis. 221, 122–126 (2020).
Faccinetto, C. et al. Internal validation and improvement of mitochondrial genome sequencing using the Precision ID mtDNA Whole Genome Panel. Int. J. Legal Med. 135, 2295–2306 (2021).
Kalman, B., Lublin, F. D. & Alder, H. Characterization of the mitochondrial DNA in patients with multiple sclerosis. J. Neurol. Sci. 140, 75–84 (1996).
Vyshkina, T. et al. Association of common mitochondrial DNA variants with multiple sclerosis and systemic lupus erythematosus. Clin. Immunol. Orlando Fla 129, 31–35 (2008).
Andalib, S. et al. MtDNA T4216C variation in multiple sclerosis: A systematic review and meta-analysis. Acta Neurol. Belg. 116, 439–443 (2016).
De Riccardis, L. et al. Metabolic response to glatiramer acetate therapy in multiple sclerosis patients. BBA Clin. 6, 131–137 (2016).
Haghikia, A. et al. Interferon-beta affects mitochondrial activity in CD4 + lymphocytes: Implications for mechanism of action in multiple sclerosis. Mult. Scler. J. 21, 1262–1270 (2015).
Kornberg, M. D. et al. Dimethyl fumarate targets GAPDH and aerobic glycolysis to modulate immunity. Science 360, 449–453 (2018).
Klotz, L. et al. Teriflunomide treatment for multiple sclerosis modulates T cell mitochondrial respiration with affinity-dependent effects. Sci. Transl. Med. 11, eaao5563 (2019).
Penter, L. et al. Longitudinal single-cell dynamics of chromatin accessibility and mitochondrial mutations in chronic lymphocytic leukemia mirror disease history. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-21-0276 (2021).
Wang, Y., Guo, X., Ye, K., Orth, M. & Gu, Z. Accelerated expansion of pathogenic mitochondrial DNA heteroplasmies in Huntington’s disease. Proc. Natl. Acad. Sci. 118, e2014610118 (2021).
Acknowledgements
We would like to thank Ana Mafalda Rocha, Bettina Vogelreuter, Bibiane Seeger-Schwinge, Filipa S. Carvalho, Jana Hermann, René Gieß, and Rosalie Schmidt for their support in the conduction of this study. Additionally, we would like to thank Susan Pikol and Cynthia Kraut for their MRI lesion segmentations and collection of MRI data related to this study. We would also like to thank the Flow Cytometry Facility of the Instituto de Medicina Molecular João Lobo Antunes for their technical support, particularly Diana Macedo. Finally, we would like to express our most sincere gratitude to all the subjects who have participated in this research project.
Funding
This research was funded by Merck Germany (restricted research grant), the National Multiple Sclerosis Society (NMSS), NMSS Pilot Research Grant (PP-1712-29466), and Fundação para a Ciência e Tecnologia (FCT) (FCT/PTDC/MED-NEU/7976/2020). F.C.F.’s stipend was supported by FCT (PD/BD/114122/2015) and by Merck Germany (restricted research grant). V.A.M. is an iFCT researcher (IF/01693/2014; IMM/CT/27-2020). The NeuroCure Clinical Research Center (NCRC) is funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany's Excellence Strategy—EXC-2049—390688087 (granted to F.P.). We would also like to acknowledge the Fundos Europeus Estruturais e de Investimento (FEEI) from Programa Operacional Regional de Lisboa 2020 and FCT, grants LISBOA-01-0145-FEDER-016394, LISBOA-01-0145-FEDER-016417, and POCI-01-0145-FEDER-022184, as well as PPBI-POCI-01-0145-FEDER-022122. Funding sources had no influence on the collection, analysis, and interpretation of data, nor in writing the manuscript.
Author information
Authors and Affiliations
Contributions
F.C.F. was responsible for: Conceptualization of the research study, execution of the experimental tasks, data analysis, and manuscript draft preparation. S.A. was responsible for: Clinical data curation and collection, subject recruitment, critical review of the study’s conceptualization, and manuscript revision. C.C. was responsible for: MRI data collection and management, MRI data analysis, and manuscript revision. H.G.Z. was responsible for: OCT data analysis and management, and manuscript revision. K.R. was responsible for: Clinical data collection, subject recruitment, and manuscript revision. T.S.-H. was responsible for: Cohort management, clinical data collection, subject recruitment, project administration, and manuscript revision. J.B.-S. was responsible for: Cohort management, clinical data collection, subject recruitment, project administration, critical review of the study’s conceptualization, and manuscript revision. F.P. and V.A.M. were responsible for: Conceptualization of the research study, funding acquisition, provision of resources, supervision of data interpretation, and manuscript revision (co-senior authors). All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.C.F. reports no disclosures relevant to the manuscript. S.A. received speaker’s honoraria from Alexion, Bayer and Roche. C.C. received speaking honoraria from Bayer and research funding from Novartis, unrelated to the current study. H.G.Z. received speaking honoraria from Bayer and Novartis and research funding from Novartis, unrelated to the current study. K.R. received research support from Novartis Pharma, Merck Serono, German Ministry of Education and Research, European Union (821283-2), Stiftung Charité, Guthy Jackson Charitable Foundation and Arthur Arnstein Foundation; received travel grants from Guthy Jackson Charitable Foundation. K.R. is a participant in the BIH Clinical Fellow Program funded by Stiftung Charité. T.S.-H. reports no disclosures relevant to the manuscript. J.B.-S. received speaking honoraria and travel grants from Bayer Healthcare, and sanofi-aventis/Genzyme, in addition received compensation for serving on a scientific advisory board of Roche, unrelated to the presented work. F.P. serves as an Associate Editor for Neurology: Neuroimmunology & Neuroinflammation and PLoS ONE, reports speaker honoraria and/or travel grants from Bayer, Novartis, Biogen Idec, Teva, Sanofi-Aventis / Genzyme, Merck Serono, Alexion, Chugai, MedImmune, Shire, Roche, Actelion, and Celgene, research support from Bayer, Novartis, Biogen Idec, Teva, Sanofi-Aventis / Genzyme, Alexion, Merck Serono, German Research Council (DFG Exc 257), Werth Stiftung of the City of Cologne, German Ministry of Education and Research (BMBF Competence Network Multiple Sclerosis), Arthur Arnstein Stiftung Berlin, EU FP7 Framework Program (combims.eu), Arthur Arnstein Foundation Berlin, Guthy Jackson Charitable Foundation, and National Multiple Sclerosis Society of the USA, consultancies for Sanofi-Aventis / Genzyme, Biogen Idec, MedImmune, Shire, and Alexion, and is a member of the steering committee of the OCTIMS study (Novartis). V.A.M. reports no disclosures relevant to the manuscript.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cortes-Figueiredo, F., Asseyer, S., Chien, C. et al. CD4+ T cell mitochondrial genotype in Multiple Sclerosis: a cross-sectional and longitudinal analysis. Sci Rep 14, 7507 (2024). https://doi.org/10.1038/s41598-024-57592-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-57592-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.