Abstract
Nuclear degradation is a major event during programmed cell death (PCD). The breakdown of nuclear components has been well characterized during apoptosis, one form of PCD. Many nonapoptotic forms of PCD have been identified, but our understanding of nuclear degradation during those events is limited. Here, we take advantage of Drosophila oogenesis to investigate nuclear degeneration during stress-induced apoptotic and developmental nonapoptotic cell death in the same cell type in vivo. We find that nuclear Lamin, a caspase substrate, dissociates from the nucleus as an early event during apoptosis, but remains associated with nuclei during nonapoptotic cell death. Lamin reveals a series of changes in nuclear architecture during nonapoptotic death, including nuclear crenellations and involutions. Stretch follicle cells contribute to these architecture changes, and phagocytic and lysosome-associated machinery in stretch follicle cells promote Lamin degradation. More specifically, we find that the lysosomal cathepsin CP1 facilitates Lamin degradation.
Similar content being viewed by others
Introduction
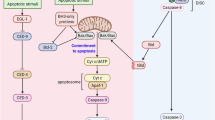
Programmed cell death (PCD) modalities utilize genetically encoded components to destroy the cell; a process essential for organismal development and homeostasis [1]. The importance of apoptosis, one form of PCD, in development and disease has led to its thorough characterization. Apoptosis culminates in the activation of a cascade of caspases, cysteine-directed aspartate-specific proteases that discriminately cleave >1000 substrates [2,3,4]. At least a dozen other forms of PCD have been identified that are nonapoptotic and noncaspase dependent, but our understanding of them is limited [5,6,7,8].
Nuclear breakdown is a major event of PCD. Lingering nuclear material can be toxic to the organism and, therefore, must be effectively eliminated [9, 10]. Nuclear lamins form a meshwork subjacent to the inner nuclear membrane, and consist of A- and B-type lamin proteins which differ biochemically, structurally, and in their expression patterns [11, 12]. In apoptosis, caspase cleavage of lamins facilitates nuclear degradation events in vitro [13]. In addition, lamin cleavage is used to track caspase activity in vivo [14,15,16]. Yet, it remains to be determined how lamins, and the nucleus in general, are dismantled in caspase-independent PCD modalities.
Several examples of nonapoptotic PCD have emerged, including the elimination of alveolar epithelial cells during mammary gland regression post lactation, and removal of uterine epithelial cells during implantation in mammals [17,18,19]. Another well-defined example of nonapoptotic PCD is linker cell death (LCD) in Caenorhabditis elegans [20]. The male-specific linker cell is essential for male fertility, as it promotes gonad elongation and its death enables fusion with the cloaca, allowing sperm to be released. LCD shows distinct nuclear morphology compared with apoptosis: chromatin does not condense and the nucleus becomes crenellated [20]. In their recent review, Kutscher and Shaham [6] suggest that LCD morphology is not unique, as dying cells in various tissues and organisms display similar nonapoptotic morphology. Despite the conservation of morphological features, it is unclear whether these changes in nuclear structure impact nuclear breakdown.
The Drosophila melanogaster ovary provides a tractable model for investigating nuclear degradation dynamics in both apoptotic and nonapoptotic cell deaths of the same cell type, nurse cells (NCs) [21, 22]. Roughly 15–20 ovarioles comprise each ovary, and each ovariole contains egg chambers progressing through 14 stages of oogenesis (Fig. 1a). Each egg chamber contains 16 germ cells: 15 polyploid NCs connected to an oocyte through cytoplasmic bridges [23,24,25]. Throughout oogenesis, NCs deposit proteins, mRNA, and cytoplasm into the oocyte, a process conserved in mammals [26]. Surrounding the germline cyst is an epithelial layer of somatically derived follicle cells (FCs), which differentiate into specialized subgroups during oogenesis. Notably, during stage 9 (s9) and s10 of oogenesis, a subset of FCs called the stretch FCs (SFCs) is established at the anterior end of the egg chamber [27, 28]. This group of ~50 SFCs are morphologically and functionally distinct from main body FCs as their membranes thin and elongate to surround the NCs (Fig. 1a).
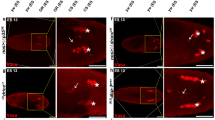
Lamin is degraded as morphological changes in chromatin are observed during caspase-dependent apoptosis. a Schematic of a D. melanogaster ovariole composed of progressively developing egg chambers. Environmental stress, such as nutrient starvation, can induce a caspase-dependent cell death program of nurse cells (NCs) in mid-stage egg chambers. In contrast, NCs are eliminated through a caspase-independent and nonapoptotic developmental death during the late stages of oogenesis. b–g” Representative mid-stage egg chambers of starved wild-type (w1118) flies stained with DAPI to mark DNA (cyan), anticleaved-Dcp-1 (cDcp-1, yellow) to mark egg chambers undergoing apoptosis, and ADL84.12 to mark B-type Lamin (green). Mid-stage phasing previously described by Etchegaray et al. [37]. b–b” Phase 0 (not dying) mid-stage egg chamber: NC chromatin is dispersed (b arrowhead), a layer of follicle cells (FCs) line the perimeter of the egg chamber (b arrow indicating FC nuclei), and cDcp-1 is absent (b’). Both NC (b” inset) and FC nuclei (b” arrow) have continuous peripheral Lamin. c–c” Phase 1 dying egg chamber: NC chromatin is disorganized and cDcp-1 is present in the germline. Lamin begins to appear cytoplasmic and continues to at least partially associate with NC nuclei (c” inset). d–d” Phase 2 egg chamber: chromatin is condensed. Lamin generally no longer associates with condensed NC nuclei and continues to appear cytoplasmic (d” inset). Asterisks indicate a healthy egg chamber anterior to the dying egg chamber. e, f” Phase 3–4 egg chambers: NC chromatin progressively condenses and fragments. Lamin is not associated with degrading chromatin. Asterisks indicate a healthy egg chamber anterior to the dying egg chamber (e–e”). g–g” Phase 5 egg chamber: few remnants of NC chromatin remain (g arrowhead), and FC nuclei are disorganized surrounding the germline material (g arrow). Most of the germline material has been engulfed (g’). Lamin is observed surrounding only FC nuclei (g” arrow compared with arrowhead pointing to NC chromatin). Scale bar = 50 µm
The different forms of PCD in Drosophila oogenesis facilitate a comparative analysis between apoptotic and nonapoptotic nuclear degradation dynamics. In mid-oogenesis (s7–9), environmental stressors like protein starvation and predator threat can induce NC apoptosis (Fig. 1a) [29, 30]. In late oogenesis (s10B–14), the NCs are eliminated to produce a mature oocyte (Fig. 1a). Developmental death of the NCs is nonapoptotic, as chromatin morphology is distinct from apoptosis and blocking caspases does not impair NC death [22, 31,32,33]. Instead, we previously demonstrated that developmental NC death occurs through phagoptosis, a form of PCD dependent on phagocytic machinery [34, 35].
We leveraged this powerful in vivo system to determine how nuclear degradation proceeds in a nonapoptotic, caspase-independent cell death. Specifically, we visualized a nuclear caspase substrate, Lamin (the Drosophila B-type lamin), to analyze comparative changes in nuclear architecture and Lamin breakdown during NC apoptosis and developmental phagoptosis. We identified a series of nuclear architecture changes, wherein NC nuclei become crenellated and form involutions—in contrast to apoptosis, in which Lamin is degraded soon after changes in chromatin morphology are observed. Furthermore, we found that CP1, a lysosomal protease, nonautonomously facilitates Lamin breakdown during caspase-independent NC death. Our work establishes a model for nuclear degradation mechanisms in a nonapoptotic PCD.
Materials and methods
Fly strains and maintenance
The following fly lines were obtained from the Bloomington Drosophila Stock Center: w1118, tub-Gal80ts, luciferase RNAi (JF01355), spinsterRNAi (JF02782), Vha16-1 RNAi (HMS02171), CP1RNAi (HMS00725), Diap1 RNAi (HMS00752). PG150-GAL4/FM7 was received from Ellen LeMosy and GR1-GAL4 was received from Trudi Schupbach. dor4 [36], drpr∆5 [37] mutants have been previously described.
Flies were maintained on standard cornmeal and molasses at 25 °C. If Gal80ts was used, flies were reared at 18 °C, then moved to 29 °C for 2 days immediately preceding dissection; ~18 h at 29 °C when expression of Diap1RNAi was induced in the SFCs. Other RNAi lines were also incubated for 2 days immediately preceding dissection for consistency. For analyses of nonapoptotic developmental death, flies were conditioned with yeast paste to promote oogenesis 2–3 days immediately prior to dissection. To study apoptotic stress-induced death, flies were conditioned for 2 days, then deprived of protein for ~18 h immediately preceding dissection in apple juice agar vials.
Immunohistochemistry and phalloidin staining
Generally, flies <10 days old were conditioned on yeast paste, or conditioned and then starved for ~18 h immediately preceding dissection at 25 °C. Flies expressing RNAi were conditioned for 2 days at 29 °C. Flies were dissected in 1× phosphate-buffered saline (PBS) and ovaries were gently teased apart to separate ovarioles. Ovary tissue was fixed in 2% paraformaldehyde in 1× PBS for 20 min, then rinsed two times in 1% TritonX-100 in 1× PBS (1% PBT). Ovary tissue was then washed three times in 1% PBT for 1 h at room temperature (RT). Next, tissue was blocked in PBANG (0.1% PBT, 0.5% bovine serum albumin (BSA), and 5% normal goat serum) for 1 h at RT, then incubated with primary antibody diluted in PBANG for at least 3 days to aid Lamin penetrance, at 4 °C. Primary antibodies: ADL84.12 (1:10; Developmental Studies Hybridoma Bank) [38] and cleaved Death caspase-1 (cDcp-1) (1:100; Cell Signaling Technology).
After incubation in primary antibody, ovary tissue was briefly rinsed twice with 1% PBT, then washed four times for 2 h in 0.1% PBT + 0.5% BSA. Samples were then incubated in secondary antibody diluted in PBANG for at least 1 h at RT, and protected from light. If actin staining was desired, 2.5% phalloidin conjugated to Alexa Fluor 546 diluted in methanol (Invitrogen; Cat #A22283) was added during this step with the secondary antibody. Secondary antibodies include: goat anti-mouse Cy3 (1:200; Jackson ImmunoResearch, Cat #115-165-003); goat anti-mouse Alexa Fluor 488 (1:400; Jackson ImmunoResearch, Cat #115-545-003); goat anti-rabbit Alexa Fluor 488 (1:500; Jackson ImmunoResearch, Cat #111-545-003); goat anti-mouse Alexa Fluor 647 (1:75; Jackson ImmunoResearch, Cat #115-605-003). Tissue was briefly rinsed two times in 1% PBT, then again washed four times in 0.1% PBT + 0.5% BSA for 2 h at RT. Finally, tissue was rinsed in 1× PBS and stored in Vectashield with DAPI (Vector Labs, Cat #H-1200) overnight at 4 °C before mounting.
LysoTracker staining
After dissection in 1× PBS, flies were incubated in 150 μL of LysoTracker Red DND-99 (Invitrogen, Cat #L7528) freshly diluted 1:50 in 1× PBS, for 3 min at RT. The tube was periodically flicked and kept on a shaker during this time. The LysoTracker solution was then removed and tissues were washed three times in 1× PBS for 30 min. The first wash was quick, and the other two were longer. Samples were protected from light hereafter. After washing, tissues were fixed in 2% paraformaldehyde in 1× PBS and primary and secondary staining proceeded as described above.
Microscopy and image processing
All images were taken on an Olympus FV10i confocal microscope. Images were processed using ImageJ and Photoshop. Figures were made using Adobe Illustrator. For models, NC nuclei were isolated and processed in Microsoft Office PowerPoint, then imported into Adobe Illustrator.
Quantifications and statistics
For all quantifications, s13 egg chambers were identified by the presence of dorsal appendages that had not fully matured as in s14 egg chambers. S13 egg chambers were haphazardly selected, and a single Z-plane was imaged for quantification of NC nuclei. NC nuclei with clearly distinguishable traits were quantified. For example, sometimes Lamin antibody does not fully penetrate egg chambers so if penetrance was poor, nuclei were not quantified.
To quantify the formation of involutions when SFCs were ablated, NC nuclei of s13 egg chambers in experimental and control samples were analyzed for the presence or absence of Lamin involution into the nucleus. Of the nuclei that had identifiable involutions, we quantified whether actin was associated with the involutions. We quantified three independent replicates for control samples, and four independent replicates for experimental samples. Using Excel software, we calculated the average percentage and standard deviation of NC nuclei with involutions, and the average percentage and standard deviation of involutions associated with actin. Using Prism software, we performed unpaired t-tests to determine p-values.
To quantify Lamin degradation, NC nuclei in cross section of s13 egg chambers were analyzed. To test whether phagocytosis and lysosome-associated genes affected Lamin degradation, we calculated the proportion of NC nuclei with degrading Lamin versus total nuclei associated with Lamin, and shifted the data by adding 0.1 to test whether the data best fit a normal, lognormal, or gamma distribution using the fitdistrplus package [39] and employing R version 3.5.1 in RStudio [40]. The data best fit a gamma distribution. Using the lme4 package [41], we made a generalized linear mixed model using genotype as a fixed effect and experiment replicate number as a random effect. To compare significance between genotypes, we used the pairwise least-square means tests using the emmeans package [42].
To quantify changes in chromatin, Lamin, and acidification in CP1 knockdowns and controls, images were assigned unique codes and quantified blindly. Multiple correspondence analyses were computed using R, and the script is available at: https://github.com/spholmes/Nuclei. Prism software was used to determine standard deviation and perform pairwise Student’s t-tests comparing the mean percentage between experimental and control nuclear states across three replicate experiments of >7 flies per experiment.
Results
During apoptosis, changes in chromatin morphology are quickly followed by Lamin degradation
To characterize nuclear lamin degradation during apoptosis, flies were starved and egg chambers were labeled with antibodies against cDcp-1 and B-type Lamin (Lamin). We determined the phase of death of mid-stage egg chambers according to Etchegaray et al. [37]. Phase 0 (healthy, nondying) mid-stage egg chambers had dispersed NC chromatin and lacked cDcp-1 (Fig. 1b–b’). NC and FC nuclei of healthy mid-stage egg chambers were encircled by Lamin (Fig. 1b”). In phase 1 (early dying) egg chambers, NC chromatin was disorganized and cDcp-1 was present in the germline (Fig. 1c–c’). Lamin began to appear cytoplasmic, while continuing to associate with NC nuclei (Fig. 1c”). As mid-stage death proceeded, NC chromatin progressively condensed and fragmented (Fig. 1d–g’). Lamin continued to appear cytoplasmic in phase 2 (Fig. 1d”), and rarely associated with NC chromatin (Fig. 1d”, inset). In phase 3, Lamin was still detected in the germline cytoplasm, but was not associated with NC nuclear fragments for the remainder of mid-stage death (Fig. 1e”–g”). In contrast, Lamin in FC nuclei stayed intact through phase 5 (Fig. 1g–g”). Thus, the loss of Lamin from NC nuclei was an early event in NC apoptosis, occurring by phase 2.
During developmental nonapoptotic cell death, Lamin remains associated with NC nuclei despite chromatin changes
To determine the progression of nuclear events during nonapoptotic developmental death [31, 32], we examined egg chambers of well-fed wild-type flies labeled with a Lamin antibody. In s10 egg chambers, NC chromatin was dispersed and peripherally enclosed by Lamin (Fig. 2a–a’). Consistent with previous work showing chromatin changes in s11 egg chambers [43], we found that as NCs rapidly transferred their cytoplasmic contents into the oocyte, NC nuclei began to compress (Fig. 2b). By s12, NC chromatin had compacted (Fig. 2c), but was not as condensed as in mid-stage apoptotic death (for example, Fig. 1d). As these chromatin changes occurred, Lamin remained peripherally associated with NC nuclei and showed striking changes in architecture: as the NC nuclei compressed, the nuclear lamina formed crenellations (Fig. 2b’) and then deep involutions (Fig. 2c’).
Lamin persists after chromatin changes have occurred during caspase-independent developmental PCD in NCs. a–f’ Representative late-stage egg chambers of well-fed wild-type (w1118) flies stained with DAPI to mark chromatin (cyan), and ADL84.12 to mark B-type Lamin (green). a–a’ S10 egg chamber: NC nuclei are spheroid with dispersed chromatin and continuous peripheral Lamin. b–b’ S11 egg chamber: compressed NC nuclei, crenellated Lamin (b’ inset). c–c’ S12 egg chamber: NC chromatin has compacted, but remains associated with Lamin. Lamin involutions extend into NC nuclei (c’ inset, arrowhead). d, e’ S13 egg chambers: NC nuclei asynchronously degrade. Degrading Lamin appears faint (d’ inset), or shows focal disruptions (e’ inset, arrowhead). Some NC nuclei lack Lamin (e, e’ yellow arrowheads). f–f’ S14 egg chamber: characterized by fully extended dorsal appendages (DA, f’ inset). NCs have been eliminated, while FCs remain (arrowhead). g Model of NC Lamin changes (white) in developmental NC PCD. S8 nucleus from Fig. 1b inset. Scale bar = 50 µm
During mid-stage apoptosis, NCs degenerate synchronously. In contrast, developmental NC death had some synchronous and asynchronous processes. NC nuclei compressed, became crenellated, and formed involutions synchronously. However, Lamin and chromatin degradation occurred asynchronously. S13 egg chambers contained NC nuclei with different states of chromatin degradation: in some nuclei, there were regions of condensed chromatin, while in others chromatin had degraded based on weak DAPI staining (Fig. 2d). In addition, different states of Lamin association were observed: in some nuclei Lamin appeared unperturbed, while in others Lamin was degrading. We observed two classes of Lamin degradation which were not mutually exclusive—appearing either less concentrated along the nuclear periphery (Fig. 2d’), or as “focal disruptions” where breaks were observed in peripheral Lamin (Fig. 2e’). In some nuclei with weak DAPI labeling, Lamin had been entirely degraded and was absent (Fig. 2e–e’). By s14, all NC nuclei had been eliminated and only FC nuclei remained (Fig. 2f–f’).
These data demonstrate compelling differences in nuclear degradation dynamics between apoptosis in mid-stage NC death and nonapoptotic developmental NC death. In apoptosis, Lamin disappeared from nuclei soon after chromatin changes were observed (Fig. 1), whereas in the nonapoptotic developmental death, Lamin remained associated with nuclei and revealed a progression of architecture changes (Fig. 2g). Early egg chambers, such as s8, had round NC nuclei (Figs. 1b” and 2g). By s10, NC Lamin architecture began to deviate from a smooth, round perimeter (Fig. 2a’, g). As NC nuclei started to compress, Lamin became crenellated and subsequently involuted, remaining associated with nuclei despite chromatin changes (Fig. 2g). Finally, Lamin asynchronously degraded, yet continued to at least partially associate with chromatin (Fig. 2g).
SFCs promote Lamin architecture changes and Lamin degradation
Our data revealed that an early step of nuclear degradation during nonapoptotic NC death is a change in Lamin architecture. In wild type, the formation of cytoplasmic actin bundles in s10B egg chambers [43,44,45] precedes the dramatic changes in Lamin architecture. Actin bundles actively position NC nuclei during dumping: as NCs contract, the actin bundles push NC nuclei and laterally associate with them as nuclei rotate [44]. Previously, we demonstrated that SFCs closely associate with NC nuclei in late oogenesis: in s11 egg chambers SFCs begin to invade between NCs, and by s12, SFC membranes surround NC remnants [34, 46]. Moreover, SFCs are required for several events that take place during NC death, including cytoplasmic dumping, acidification, DNA fragmentation, and cytoplasmic actin bundles [34]. Thus, to investigate the role of SFCs in NC nuclear degradation, we genetically ablated SFCs in late-stage egg chambers and examined Lamin and cytoplasmic actin dynamics. We used GAL4-UAS [47] to express Diap1RNAi in SFCs to induce their death [34].
Consistent with previous studies [43, 44, 48], control egg chambers displayed subcortical actin in NCs and FCs (Fig. 3a–a”’), actin filaments in the ring canals (Fig. 3a–a”’), and cytoplasmic actin bundles extending towards NC nuclei in s10–11 egg chambers (Fig. 3a–b”’) [43, 44, 48]. We found actin filaments poking into NC nuclei starting in s11 (Fig. 3b–b”’), and extending into involutions during s12–13 (Fig. 3c, d”’, insets). By s14, NCs were eliminated and only cortical actin in FCs was observed (Fig. 3e–e”’).
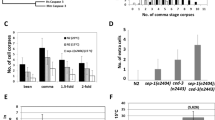
Actin bundles extend into nuclear involutions and nonautonomous components from stretch follicle cells promote nuclear Lamin degradation. a–h”’ Representative late-stage egg chambers of well-fed control (a–e”’, FM7/+; tub-Gal80ts/+; UAS-Diap1RNAi/+) and SFC ablated (f–j”’, PG150-Gal4/+; tub-Gal80ts/+; UAS-Diap1RNAi/+) flies stained with DAPI to mark chromatin (cyan), ADL84.12 to mark B-type Lamin (green), and phalloidin to mark actin (magenta). a–a”’ Control s10 egg chamber: actin staining reveals cortical actin surrounding nurse cells (NCs; arrow), cytoplasmic actin bundles beginning to form (arrowhead), and ring canals between germline cells (dashed arrow). b–d”’ S11–13 egg chambers: cytoplasmic actin bundles extend towards NC nuclei and into nuclear involutions (b–d”’, insets, arrowhead). The muscle sheath surrounding each ovariole is also sometimes seen (d”–d”’, M). e–e”’ S14: NC nuclei have been eliminated and dorsal appendages have fully extended (DA). f, g”’ SFC ablated s10–11 egg chambers: occasionally cytoplasmic actin bundles are present, extending towards NC nuclei and into involuting NC nuclei (arrowheads, insets). h, i”’ S12–13 egg chambers: cytoplasmic actin bundles are largely absent, corresponding with a lack of involutions in NC nuclei. However, cortical actin is apparent (h”, arrow). j–j”’ S14 egg chamber: cortical actin between NCs remains visible. In addition, NC nuclear Lamin remains largely intact. Chromatin in some nuclei appears highly condensed (arrowhead) and possibly degrading, though the appearance deviates from the progression seen in wild type and control egg chambers. k, l Quantification of the mean percentage ± standard deviation of NC nuclei with involutions from three replicate experiments in control and four replicate experiments in experimental flies. k In control egg chambers, 84.6% ± 5.3 of NC nuclei had involutions (120 NC nuclei total). When stretch FCs were ablated, the percentage of NC nuclei with involutions was significantly reduced to 20.4% ± 4.3 (p < 0.0001) (160 NC nuclei total). l Quantification of the mean percentage of involutions associated with actin. In control egg chambers, 87.4% ± 5.9 of involutions were associated with actin. When stretch FCs were ablated, the percentage of involutions associated with actin was significantly reduced to 50.1% ± 13.7 (p = 0.0074). m Quantification of NC nuclei with degrading Lamin. The proportion of NC nuclei with degrading Lamin per egg chamber was determined and plotted as a boxplot. In control egg chambers, the median percentage of NC nuclei with degrading Lamin was 60% (103 NC nuclei total). When stretch FCs were ablated, the median number of NC nuclei with degrading Lamin significantly decreased to 0 (p < 0.001) (112 NC nuclei total). Scale bar = 50 µm
Genetic ablation of SFCs did not entirely disrupt cytoplasmic actin bundles in s10–11 egg chambers (Fig. 3f, g”’). Similar to controls, these actin bundles seemed to poke into some NC nuclei (Fig. 3f”–g”’). However, in s12–13 egg chambers, the actin bundles were less readily observed (Fig. 3h–i”’), although cortical actin remained (Fig. 3h”, arrow). Moreover, the architecture of NC nuclei in these egg chambers deviated from controls: nuclei did not appear synchronously compressed, and failed to form involutions (Fig. 3h–i”’). NCs lingered in s14 egg chambers, and persisting nuclei generally remained associated with Lamin. In some instances, NC nuclei appeared to be degrading as judged by regions of highly condensed chromatin (Fig. 3j); Lamin also appeared to be breaking down in these nuclei (Fig. 3j’).
To determine the effect of SFCs on nuclear events, we quantified involutions and Lamin degradation in NC nuclei of s13 egg chambers. We found a significant decrease in involutions (Fig. 3k), and in the association of involutions with actin (Fig. 3l) when SFCs were ablated. Since actin bundles were seen in s10–11 egg chambers lacking SFCs, it is possible that they contributed to some of the involutions. In addition, Lamin degradation was significantly decreased when SFCs were ablated (Fig. 3m). Altogether, these data suggest that actin dynamics during late oogenesis are associated with nuclear involutions. Furthermore, Lamin degradation relies on nonautonomous components from SFCs.
Phagocytosis and lysosome-associated genes promote Lamin degradation, but not Lamin architecture changes
To better understand how nonautonomous factors promote nuclear degradation events, we investigated phagocytosis and lysosome-associated genes acting in SFCs. The engulfment receptor Draper is nonautonomously required for NC clearance, as FC-specific knockdowns of draper result in a strong persisting nuclei phenotype [34]. Knocking down draper specifically in SFCs blocked NC acidification and DNA fragmentation in s13 [34]. Similarly, the lysosome-associated genes deep orange (dor), spinster (spin), and Vha16-1 are required for NC clearance [34, 46, 49]. dor and Vha16-1 are also required for acidification in s13 egg chambers.
To characterize the role of phagocytosis and lysosome-associated genes in caspase-independent nuclear degradation, we analyzed chromatin and Lamin changes in draper and dor mutants, and used GAL4-UAS [47] to knock down spin and Vha16-1 in FCs. By s13 in control egg chambers, NC nuclei had compressed, become crenellated, and formed Lamin involutions (Figs. 2d and 4a–a’). NC nuclei degraded asynchronously, with varying degrees of chromatin and Lamin breakdown (Fig. 4a–a’). In s13 egg chambers from draper mutants, nuclear architecture changes proceeded normally: nuclei compressed, and became crenellated and involuted (Fig. 4b–b’). These processes also occurred normally in dor hypomorphs, and spin and Vha16-1 knockdowns (Fig. 4c–e’). Knocking down Vha16-1 additionally resulted in a mild dumpless phenotype, in which NCs did not completely transfer their cytoplasm to the oocyte (Fig. 4e’, j’), suggesting that Vha16-1 may have additional roles in late oogenesis. Both draper and dor significantly impaired Lamin degradation (Fig. 4k). Similarly, knocking down Vha16-1 significantly blocked Lamin degradation (Fig. 4l). In line with these trends, knocking down spinster led to a reduction in Lamin degradation, although this was not statistically significant (Fig. 4l).
Phagocytosis and lysosome-associated genes do not affect Lamin architecture changes, but do affect Lamin degradation during the developmental PCD of NCs. a–j’ Representative egg chambers of well-fed control (GR1-Gal4 /+; UAS-luciferase-RNAi/+) and mutant flies stained with DAPI to mark chromatin (cyan), and ADL84.12 to mark Lamin (green). a–e’ Stage 13 egg chambers from wild type and mutant flies had compressed NC nuclei with peripheral Lamin forming crenellations and involutions. Phase insets show growing dorsal appendages. a–a’ arrow and arrowhead indicate asynchronously degrading NC nuclei. f–f’ Wild-type s14 egg chamber devoid of NC material. g–j’ S14 egg chambers of mutant flies with persisting NC nuclei. Lamin and nuclear architecture, including involutions, are retained (g–j’). Phase insets show fully formed dorsal appendages. k, l Quantification of NC nuclei with degrading Lamin from at least two replicate experiments per genotype. The proportion of NC nuclei with degrading Lamin per egg chamber was determined and plotted as a boxplot. k In control egg chambers (w1118), the median percentage of NC nuclei with degrading Lamin was 50% (118 total NC nuclei). In dor4 and draper null mutants, the median percentage of NC nuclei with degrading Lamin significantly decreased to 14.29% (p = 0.0027) (83 total NC nuclei) and 18.33% (p = 0.0068) (113 NC nuclei total), respectively. l In control egg chambers (GR1-Gal4 /+; UAS-luciferase-RNAi/+), the median percentage of NC nuclei with degrading Lamin was 58.33% (103 NC nuclei total). When spinster was knocked down in FCs, the median percentage of NC nuclei with degrading Lamin decreased to 33.33% (p = 0.06) (72 NC nuclei total). When Vha16-1 was knocked down, the median percentage of NC nuclei with degrading Lamin decreased to 0 (p = 0.0002) (81 NC nuclei total). Scale bar = 50 µm
In control s14 egg chambers, NC DNA and Lamin were completely cleared (Fig. 4f–f’). However, in s14 egg chambers from draper mutants, NC chromatin and Lamin persisted [34]. These persisting nuclei had formed involutions (Fig. 4g–g’), but did not show focal disruptions or degradation of Lamin. Interestingly, asynchronous degradation was less apparent, as variation in chromatin condensation and Lamin degeneration was limited (Fig. 4g–g’). Persisting nuclei in s14 egg chambers from dor hypomorphs, and spin and Vha16-1 knockdowns also proceeded through nuclear architecture changes normally, and generally continued to associate with Lamin (Fig. 4h–j’). However, asynchronous nuclear breakdown events were still observed in contrast to draper mutants. Draper may be required at an earlier step than dor, spin, and Vha16-1; thus, absence of draper disrupted asynchronous degradation, while the other genes did not have as strong of an effect. Alternatively, spin and Vha16-1 could be incompletely knocked down, so perhaps some contribution from these genes remained. Another consideration is that compensatory mechanisms for the targeted lysosome-associated genes may contribute to preserving asynchronous nuclear degradation events. Altogether, these data demonstrate that phagocytic and lysosome-associated genes are important for Lamin breakdown, but not for changes in nuclear architecture (Fig. 2g).
The lysosomal protease CP1 facilitates Lamin degradation
NC Lamin degradation was blocked in draper mutants, and impaired in the absence of lysosome-associated genes. Yet, it remained unclear how Lamin was removed since caspases are not required for developmental NC death [22, 31,32,33]. We recently reported that CP1, the Drosophila ortholog of cathepsin V, is released from SFCs and localizes to degenerating NC nuclei during developmental death [46]. Furthermore, it is required for NC elimination, but not acidification. Thus, we were interested in elucidating whether cathepsins played a role in nuclear degradation during nonapoptotic NC death.
To explore this possibility, we knocked down CP1 in FCs, and colabeled egg chambers with Lamin and LysoTracker to mark acidified compartments. We blindly quantified NC nuclei of s13 egg chambers according to the state of three traits: (a) chromatin morphology, (b) Lamin morphology, and (c) acidification (Table 1; Fig. 5a–b”’). Chromatin state was qualitatively defined as either (i) baseline, in which it appeared evenly dispersed, (ii) degrading, in which chromatin appeared to have regions of condensed chromatin or weak DAPI staining, or (iii) absent, in which DAPI was no longer detected. Lamin state was defined as either (i) continuous, such that a cross section through an NC nucleus had continuous Lamin around the nuclear perimeter, (ii) degrading, such that Lamin appeared faint, or showed focal disruptions, or (iii) absent. Finally, acidification was either (i) present or (ii) absent, based on LysoTracker staining. This produced data sets of different nuclear degradation states in control and FC-specific CP1 knockdowns based on the varying conditions of chromatin, Lamin, and acidification.
CP1 facilitates Lamin degradation. a, b”’ Representative s13 egg chambers (phase insets) of well-fed control (tub-Gal80ts/+; UAS-luciferase-RNAi/GR1-Gal4) and CP1 knockdown (tub-Gal80ts/+; UAS-CP1-RNAi/GR1-Gal4) flies stained with DAPI to mark DNA (cyan), ADL84.12 to mark Lamin (green), and LysoTracker (magenta) to mark acidified compartments. NC nuclei display different combinations of chromatin, Lamin, and acidification states (arrows and arrowheads). c Rationale for performing multiple correspondence analyses: between s10 and s14, DNA and Lamin are degraded and NC nuclei become acidified. We investigated whether these events correlate in degrading nuclei. d Multiple correspondence analyses of nuclear states in control and CP1 knockdown flies. Note the correlation of baseline chromatin (Chrom_Base) with continuous Lamin (Lam_Cont), degrading chromatin (Chrom_Deg) with degrading Lamin (Lam_Deg), and absent chromatin (Chrom_Abs) with absent Lamin (Lam_Abs), which in turn correlate with acidification (LT1). N = 3 replicates per genotype, >7 flies per replicate. Scale bar = 50 µm
We wanted to use an unbiased approach to explore these data sets in their entirety and determine whether there was a relationship between the different states of chromatin, Lamin, and acidification. We performed multiple correspondence analyses which confirmed our hypothesis that in control NC nuclei, baseline chromatin correlated with continuous Lamin, degraded chromatin correlated with degraded Lamin, and the absence of chromatin correlated with the absence of Lamin (Fig. 5c). Nuclei in which chromatin and Lamin were absent also correlated more closely with being acidified (Fig. 5c). When CP1 was knocked down, these states were less closely correlated, suggesting nuclear changes occurred through more intermediate states (Fig. 5c). As expected, acidification state did not change in CP1 knockdowns versus controls. However, in addition to being less closely correlated, the relationship between degraded chromatin and Lamin was inverted in CP1 knockdowns relative to controls. A similar inversion was noted in chromatin and Lamin absence. This suggests that when CP1 is knocked down in FCs, Lamin degradation, and elimination lags.
Next, we wanted to examine how individual nuclear degradation states differed upon CP1 knockdown by doing pairwise comparisons. Statistically significant differences in nuclear states were only observed in acidified NC nuclei (Table 2). Notably, CP1 knockdown led to a significant increase in acidified NC nuclei with degrading chromatin and degrading Lamin (Table 2). There was also a significant decrease in acidified NC nuclei with degrading chromatin and absent Lamin (Table 2). Finally, knocking down CP1 resulted in a significant increase in acidified nuclei in which chromatin was absent, but Lamin was still degrading, and thus still present and partially associated with nuclei (Table 2; Fig. 5b–b”’). These findings suggest that NC nuclei with degrading chromatin and degrading Lamin generally progress to a state of degrading chromatin and absent Lamin (Table 2). When CP1 is knocked down, there is an increase of NC nuclei with degrading chromatin and degrading Lamin, possibly because they have not progressed to the degrading chromatin and absent Lamin state (Table 2). There is also an increase of NC nuclei with absent chromatin and degrading Lamin (Table 2). Altogether, these data suggest that CP1 facilitates NC Lamin degradation.
Discussion
In this study, we characterized nuclear degradation in a novel nonapoptotic cell death that occurs during Drosophila oogenesis. Our analysis revealed differences in nuclear degradation morphology between apoptosis and nonapoptotic developmental death in vivo. We established a series of nuclear breakdown events, and identified factors that contribute to each stage (Fig. 6). We found that SFCs promote NC nuclear architectural changes, and are required for Lamin degradation. Moreover, phagocytosis and lysosome-associated genes promoted Lamin degradation, but were not required for mediating changes in nuclear architecture. Finally, we showed that the cathepsin CP1 facilitates Lamin degradation. Altogether, our work provides a molecular and morphological foundation for further interrogating nuclear degradation processes during nonapoptotic NC death.
Model of nuclear degradation during nonapoptotic nurse cell death. Relative to earlier stages in which NC nuclei are spheroid, nuclei of stage 10B NCs are irregular. As NCs dump their cytoplasmic contents into the oocyte, NC nuclei become deformed by cytoplasmic actin bundles extending towards them. By s12, NC nuclei form involutions associated with actin, a process that relies on stretch follicle cells. Lysosomal machinery and Draper promote nuclear acidification and Lamin breakdown, but are not required in earlier steps for nuclear architecture changes. Finally, CP1, which does not impair acidification, facilitates Lamin degradation
The nuclear architecture changes we identified complement similar findings observed by electron microscopy [43]. Several other changes in late-stage NC nuclei have been reported: gaps form in the nuclear membrane, and the nuclear surface area increases without a corresponding increase in volume [43, 50]. Conceivably, the gaps and increased surface area confer flexibility to the nuclear membrane, facilitating ensuing nuclear architecture changes. Actin filaments continue to elongate once they reach NC nuclei, actively pushing and rotating the nuclei resulting in the association of the actin filaments with deformed NC nuclei [44]. We propose these actin dynamics contribute to the Lamin involutions we identified, which associate with actin (for example, Fig. 3c”, d”). Given that late-stage NC nuclei become acidified, an increased surface area could render more nuclear material exposed to an acidified environment and proteases.
Our data suggest that the final Lamin degradation is facilitated by cathepsins. Cathepsins are lysosomal enzymes with different catalytic mechanisms, as they can be cysteine, serine, or aspartate directed [51]. Their substrate specificity is less stringent than caspases, and they have also been shown to play a role in several forms of cell death, including apoptosis [51]. Some cathepsins translocate to the nucleus during cell death [52], and are specifically required for protein cleavage in vitro [53]. Yet, the activities of cathepsins in the nucleus during PCD remain elusive. Our work presents an in vivo model to further investigate nuclear cathepsin function during PCD.
Another important consideration is the role of phosphorylation in regulating intermediate filaments, including lamins. Phosphorylation state affects myriad activities of intermediate filaments [54]. One example is lamin phosphorylation during mitosis, enabling their dissociation from the nucleus [55,56,57,58]. Lamin phosphorylation has been observed during apoptosis, wherein the kinase PKC-δ was found to facilitate lamin disassembly [59], and in nuclear breakdown during terminal differentiation of lens fiber cells [60]. However, the role of phosphorylation in PCD remains largely unexplored. Interestingly, Smith and Fisher [61] discovered the presence of a soluble Lamin isoform in late stages of Drosophila oogenesis, coinciding with NC degradation. Thus, future research could determine whether a kinase works in tandem with proteases to efficiently remove Lamin during nonapoptotic NC death.
While the changes we have identified in nuclear architecture are just beginning to be explored in cell death, altered nuclear morphology has been appreciated as a feature of cancer cells and laminopathies, such as progeria [62, 63]. Nuclear morphology is integral to determining tumor grade and prognosis [64], and involutions revealed by lamin staining are a feature of high-grade ductal carcinomas [65]. Thus, future endeavors elucidating changes in nuclear morphology and the mechanisms through which they occur may further our understanding of two opposing cell fates—death and immortality.
References
Fuchs Y, Steller H. Programmed cell death in animal development and disease. Cell. 2011;147:742–58.
Kumar S, van Raam BJ, Salvesen GS, Cieplak P. Caspase cleavage sites in the human proteome: CaspDB, a database of predicted substrates. PLoS ONE. 2014;9:e110539.
Crawford ED, Wells JA. Caspase substrates and cellular remodeling. Annu Rev Biochem. 2011;80:1055–87.
Nagata S. Apoptosis and clearance of apoptotic cells. Annu Rev Immunol. 2018;36:489–517.
Fuchs Y, Steller H. Live to die another way: modes of programmed cell death and the signals emanating from dying cells. Nat Rev Mol Cell Biol. 2015;16:329–44.
Kutscher LM, Shaham S. Non-apoptotic cell death in animal development. Cell Death Differ. 2017;24:1326–36.
Yalonetskaya A, Mondragon AA, Elguero J, McCall K. I spy in the developing fly a multitude of ways to die. J Dev Biol. 2018; 6. https://doi.org/10.3390/jdb6040026.
Galluzzi L, Vitale I, Aaronson SA, Abrams JM, Adam D, Agostinis P, et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on cell death 2018. Cell Death Differ 2018;25:486–541.
Kawane K, Motani K, Nagata S. DNA degradation and its defects. Cold Spring Harb Perspect Biol. 2014; 6. https://doi.org/10.1101/cshperspect.a016394.
Nishimoto S, Kawane K, Watanabe-Fukunaga R, Fukuyama H, Ohsawa Y, Uchiyama Y, et al. Nuclear cataract caused by a lack of DNA degradation in the mouse eye lens. Nature. 2003;424:1071–4.
De Leeuw R, Gruenbaum Y, Medalia O. Nuclear lamins: thin filaments with major functions. Trends Cell Biol. 2018;28:34–45.
Dechat T, Adam SA, Taimen P, Shimi T, Goldman RD. Nuclear lamins. Cold Spring Harb Perspect Biol. 2010;2:a000547.
Rao L, Perez D, White E. Lamin proteolysis facilitates nuclear events during apoptosis. J Cell Biol. 1996;135:1441–55.
Martin DN, Baehrecke EH. Caspases function in autophagic programmed cell death in Drosophila. Development. 2004;131:275–84.
Berry DL, Baehrecke EH. Growth arrest and autophagy are required for salivary gland cell degradation in Drosophila. Cell. 2007;131:1137–48.
Lin L, Rodrigues FSLM, Kary C, Contet A, Logan M, Baxter RHG, et al. Complement-related regulates autophagy in neighboring cells. Cell. 2017;170:158–71.
Kreuzaler PA, Staniszewska AD, Li W, Omidvar N, Kedjouar B, Turkson J, et al. Stat3 controls lysosomal-mediated cell death in vivo. Nat Cell Biol. 2011;13:303–9.
Sargeant TJ, Lloyd-Lewis B, Resemann HK, Ramos-Montoya A, Skepper J, Watson CJ. Stat3 controls cell death during mammary gland involution by regulating uptake of milk fat globules and lysosomal membrane permeabilization. Nat Cell Biol. 2014;16:1057–68.
Li Y, Sun X, Dey SK. Entosis allows timely elimination of the luminal epithelial barrier for embryo implantation. Cell Rep. 2015;11:358–65.
Abraham MC, Lu Y, Shaham S. A morphologically conserved nonapoptotic program promotes linker cell death in Caenorhabditis elegans. Cell. 2007;12:73–86.
Jenkins VK, Timmons AK, McCall K. Diversity of cell death pathways: insight from the fly ovary. Trends Cell Biol. 2013;23:567–74.
Peterson JS, Timmons AK, Mondragon AA, McCall K. The end of the beginning: cell death in the germline. In: Steller H editors. Current Topics in Developmental Biology. Burlington: Academic Press; 2015. p. 93–119.
King RC, Rubinson AC, Smith RF. Oogenesis in adult Drosophila melanogaster. Growth. 1956;20:121–57.
King RC. Ovarian development in Drosophila melanogaster. New York: Academic Press; 1970.
Spradling AC. Developmental genetics of oogenesis. In: Bate M, Martinez Arias A (eds). The Development of Drosophila melanogaster. New York, USA: Cold Spring Harbor Laboratory Press; 1993. p. 1–70.
Lei L, Spradling AC. Mouse oocytes differentiate through organelle enrichment from sister cyst germ cells. Science. 2016;352:95–99.
González-Reyes A, St Johnston D. Patterning of the follicle cell epithelium along the anterior-posterior axis during Drosophila oogenesis. Development. 1998;125:2837–46.
Boyle MJ, Berg CA. Control in time and space: Tramtrack69 cooperates with Notch and Ecdysone to repress ectopic fate and shape changes during Drosophila egg chamber maturation. Development. 2009;136:4187–97.
Drummond-Barbosa D, Spradling AC. Stem cells and their progeny respond to nutritional changes during Drosophila oogenesis. Dev Biol. 2001;231:265–78.
Kacsoh BZ, Bozler J, Ramaswami M, Bosco G. Social communication of predator-induced changes in Drosophila behavior and germ line physiology. elife. 2015. https://doi.org/10.7554/eLife.07423.001.
Mazzalupo S, Cooley L. Illuminating the role of caspases during Drosophila oogenesis. Cell Death Differ. 2006;13:1950–9.
Peterson JS, McCall K. Combined inhibition of autophagy and caspases fails to prevent developmental nurse cell death in the Drosophila melanogaster ovary. PLoS ONE. 2013;8:e76046.
Peterson JS, Barkett M, McCall K. Stage-specific regulation of caspase activity in Drosophila oogenesis. Dev Biol. 2003;260:113–23.
Timmons AK, Mondragon AA, Schenkel CE, Yalonetskaya A, Taylor JD, Moynihan KE, et al. Phagocytosis genes nonautonomously promote developmental cell death in the Drosophila ovary. Proc Natl Acad Sci USA. 2016; 113. https://doi.org/10.1073/pnas.1522830113.
Brown GC, Neher JJ. Eaten alive! cell death by primary phagocytosis: ‘Phagoptosis’. Trends Biochem Sci. 2012;37:325–32.
Sevrioukov EA, He J-P, Moghrabi N, Sunio A, Krämer H. A role for the deep orange and carnation eye color genes in lysosomal delivery in Drosophila. Mol Cell. 1999;4:479–86.
Etchegaray JI, Timmons AK, Klein AP, Pritchett TL, Welch E, Meehan TL, et al. Draper acts through the JNK pathway to control synchronous engulfment of dying germline cells by follicular epithelial cells. Development. 2012;139:4029–39.
Stuurman N, Maus N, Fisher PA. Interphase phosphorylation of the Drosophila nuclear lamin: site-mapping using a monoclonal antibody. J Cell Sci. 1995;108:3137–44.
Delignette-Muller ML, Dutang C. Fitdistrplus: an R package for fitting distributions. J Stat Softw 2015;64:1–34.
RStudio Team. RStudio: integrated development environment for R. Boston, MA. https://www.rstudio.com/
Bates D, Maechler M, Bolker B, Walker S. Fitting linear mixed-effects models using lme4. J Stat Softw 2015;67:1–48.
Lenth RV. Emmeans: estimated marginal means, aka least-squares means. 2019; R package version 1.3.4.
Guild GM, Connelly PS, Shaw MK, Tilney LG. Actin filament cables in Drosophila nurse cells are composed of modules that slide passively past one another during dumping. J Cell Biol. 1997;138:783–97.
Huelsmann S, Ylänne J, Brown NH. Filopodia-like actin cables position nuclei in association with perinuclear actin in Drosophila nurse cells. Dev Cell. 2013;26:604–15.
Gutzeit HO. The role of microfilaments in cytoplasmic streaming in Drosophila follicles. J Cell Sci. 1986;80:159–69.
Mondragon AA, Yalonetskaya A, Ortega AJ, Elguero J, Chung W-S, McCall K. Lysosomal machinery drives extracellular acidification to direct non-apoptotic cell death. Cell Rep. 2019; 27. https://doi.org/10.1016/j.celrep.2019.03.034.
Brand AH, Perrimon N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 1993;118:401–15.
Cooley L, Verheyen E, Ayers K. chickadee encodes a profilin required for intercellular cytoplasm transport during Drosophila oogenesis. Cell. 1992;69:173–84.
Bass BP, Tanner EA, Mateos San Martín D, Blute T, Kinser RD, Dolph PJ, et al. Cell-autonomous requirement for DNaseII in nonapoptotic cell death. Cell Death Differ. 2009;16:1362–71.
Okada E, Waddington CH. The submicroscopic structure of the Drosophila egg. J Embryol Exp Morphol. 1959;7:583–97.
Chwieralski CE, Welte T. Cathepsin-regulated apoptosis. Apoptosis. 2006;11:143–9.
Roberts LR, Kurosawa H, Bronk SF, Fesmier PJ, Agellon LB, Leung Y, et al. Cathepsin B contributes to bile salt-induced apoptosis of rat hepatocytes. Gastroenterology. 1997;4113:1714–26.
Biggs JR, Yang J, Gullberg U, Muchardt C, Yaniv M, Kraft AS. The human brm protein is cleaved during apoptosis: The role of cathepsin G. Proc Natl Acad Sci USA. 2001;98:3814–9.
Omary MB, Ku N-O, Tao G-Z, Toivola DM, Liao J. ‘Heads and tails’ of intermediate filament phosphorylation: multiple sites and functional insights. Trends Biochem Sci. 2006;31:383–94.
Eggert M, Radomski N, Linder D, Tripier D, Traub P, Jost E. Identification of novel phosphorylation sites in murine A‐type lamins. Eur J Biochem. 1993;213:659–71.
Goss VL, Hocevar BA, Thompson LJ, Stratton CA, Burns DJ, Fields AP. Identification of nuclear β(II) protein kinase C as a mitotic lamin kinase. J Biol Chem. 1994;269:19074–80.
Heald R, McKeon F. Mutations of phosphorylation sites in lamin A that prevent nuclear lamina disassembly in mitosis. Cell. 1990;61:579–89.
Gerace L, Blobel G. The nuclear envelope lamina is reversibly depolymerized during mitosis. Cell. 1980;19:277–87.
Cross T, Gri G, Deacon E, Sallis R, Gough M, Watters D, et al. PKC-d is an apoptotic lamin kinase. Oncogene. 2000;19:2331–7.
Rowan S, Chang M-L, Reznikov N, Taylor A. Disassembly of the lens fiber cell nucleus to create a clear lens: the p27 descent. 2017. https://doi.org/10.1016/j.exer.2016.02.011.
Smith DE, Fisher PA. Interconversion of Drosophila nuclear lamin isoforms during oogenesis, early embryogenesis, and upon entry of cultured cells into mitosis. J Cell Biol. 1989;108:255–65.
de las Heras JI, Schirmer EC. The nuclear envelope and cancer: a diagnostic perspective and historical overview. In: Schirmer E, de las Heras J (eds). Cancer Biology and the Nuclear Envelope. Advances in Experimental Medicine and Biology, Vol. 773. New York: Springer; 2014. p. 5–26.
Eriksson M, Brown WT, Gordon LB, Glynn MW, Singer J, Scott L, et al. Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature. 2003;423:293–8.
Bell ES, Lammerding J. Causes and consequences of nuclear envelope alterations in tumour progression. Eur J Cell Biol. 2016;95:449–64.
Bussolati G. Proper detection of the nuclear shape: ways and significance. Rom J Morphol Embryol. 2008;49:435–39.
Acknowledgements
We thank the members of our lab for valuable discussions and comments, especially Anoush Calikyan and Jeanne Peterson for assistance. We thank Todd Blute for microscopy help, Katia Oleinik for helping AY with R, and Nicholas Ray for help with Lamin degradation analyses. The Lamin ADL84.12 antibody developed by Paul A. Fisher was obtained from the Developmental Studies Hybridoma Bank developed under the auspices of the NICHD and maintained by The University of Iowa, Department of Biology, Iowa City, IA 52242. We thank our funding sources: NIH Grants R01 GM060574 and R35 GM127338 to KM, NIH F31 GM115177 to AAM, and the AAUW Dissertation Fellowship to AY.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by E. Baehrecke
Rights and permissions
About this article
Cite this article
Yalonetskaya, A., Mondragon, A.A., Hintze, Z.J. et al. Nuclear degradation dynamics in a nonapoptotic programmed cell death. Cell Death Differ 27, 711–724 (2020). https://doi.org/10.1038/s41418-019-0382-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0382-x
This article is cited by
-
Scaling behaviour and control of nuclear wrinkling
Nature Physics (2023)