Abstract
Hepatocellular carcinoma (HCC) is one of the most aggressive malignancies and lacks targeted therapies. Here, we reported a novel potential therapeutic target hematological and neurological expressed 1 like (HN1L) in HCC. First, HCC tissue microarray analysis showed that HN1L was frequently up-regulated in cancer tissues than that in normal liver tissues, which significantly associated with tumor size, local invasion, distant metastases, and poor prognosis for HCC patients. Functional studies demonstrated that ectopic expression of HN1L could increase cell growth, foci formation in monolayer culture, colony formation in soft agar and tumorigenesis in nude mice. In addition, HN1L could also promote HCC metastasis by inducing epithelial-mesenchymal transition. Inversely, silencing HN1L expression with shRNA could effectively attenuate its oncogenic function. We further showed that HN1L transcriptionally up-regulated methyltransferase like 13 (METTL13) gene in an AP-2γ dependent manner, which promoted cell proliferation and metastasis by up-regulating TCF3 and ZEB1. Importantly, administration of lentivirus-mediated shRNA interfering HN1L expression could inhibit tumorigenesis and metastasis in mice. Collectively, HN1L-mediated transcriptional axis AP-2γ/METTL13/TCF3-ZEB1 promotes HCC growth and metastasis representing a promising therapeutic target in HCC treatment.
Similar content being viewed by others
Introduction
The human hematopoietic-expressed and neurologic-expressed sequence 1-like (HN1L), also known as jupiter microtubule associated homolog 2, C16orf34 or L11, was originally identified from a mouse fertilized egg cDNA library in 2000 [1] . This gene is located on chromosome 16p13.3, encodes a 190-aa protein, and specifically expressed in certain human tissues, such as the liver, kidney, prostate, testis, and uterus. However, the physiological functions of HN1L in human remain unclear. Overexpression of HN1L in non-small cell lung cancer was firstly identified by suppression subtractive hybridization, but the gene has not been deeply explored in their study [2]. Previous study has found that overexpression of HN1L could promote the malignant proliferation of lung cancer cells by activating MAPK pathway [3]. However, its precise roles in HCC has not been determined.
Hematopoietic-expressed and neurologic-expressed sequence 1 (HN1), a homologous gene of HN1L, is located on chromosome 17q25.2 and encodes a 16.5 kDa protein. It shares 30% identity amino acids with HN1L, and they are both highly conserved among mammal species. In rodents, HN1 is widely expressed in numerous tissues, such as nervous tissues and immature retina, during embryonic development, as well as HN1L [4, 5]. It has been also demonstrated to up-regulate in many human cancers, such as breast cancer [6] and pancreatic carcinoma [7], which is significantly associated with poorer overall survival of these cancer patients. Knockdown of HN1 by siRNA in a murine melanoma cell line B16-F10 promotes cell differentiation and induces cell cycle arrest [4]. Additionally, silencing HN1L murine GL261 glioma cells suppresses the growth of xenografts after intracranial implantation into mice [8], suggesting that HN1 significantly contributes to the cell cycle regulation. In addition, HN1 contributes to prostate cell migration through controlling the stability of β-catenin interaction with E-cadherin in adherent junctions [9]. These evidences suggest that HN1 is critical for regulating cancer cell growth and metastasis.
HN1L belongs to HN1 family, but its roles and regulatory mechanisms in HCC progression have not been investigated. In this study, we showed that overexpression of HN1L led to patients' poor survival and prognosis by increasing tumor growth and metastasis. Mechanism studies revealed that HN1L transcriptionally increase methyltransferase like 13 (METTL13) expression by promoting the transcriptional activity of AP-2γ via a direct protein-protein interaction. Furthermore, up-regulated METTL13 promotes tumor growth and metastasis by increasing the expression of TCF3 and ZEB1. These data suggest that the transcriptional axis HN1L/AP-2γ/METTL13/TCF3-ZEB1 is a novel pathway contributing to the aggressiveness and poor prognosis of HCC.
Results
Aberrant expression of HN1L correlates with poor outcome of HCCs
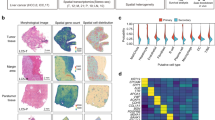
HN1L is evolutionarily conserved among mammal species, but it’s biological function has not been deeply explored (Supplementary Fig. 1A and 1B). Disease Ontology analysis of HN1L in human using Coexpedia indicated that HN1L was closely associated with malignant cancer development (Supplementary Fig. 1C). Moreover, gene expression data from TCGA database and Lim HY cohort (GSE36376) demonstrated that HN1L expression was substantially higher in HCC tissues than that in normal liver tissues (Fig. 1a). Consistent with these biostatistics, we also observed the increased protein expression levels of HN1L in clinical HCC samples than that in the paired non-tumor tissues by western blotting (Fig. 1b) and IHC staining (Fig. 1c).
Overexpression of HN1L predicts the poor survival of HCCs. a The expressions of HN1L in HCC tumor and normal liver tissues were analyzed based on TCGA database and Lim HY’ cohort (GSE36376) dataset. Data are presented as the mean ± SD. b The protein levels of HN1L in twelve pairs of HCC tumors (T) and corresponding normal liver (N) tissues were tested by western blotting. β-Tubulin was served as a loading control. c The expressions of HN1L in HCC and paired normal liver tissues were tested by IHC staining. Scale bars, 50 μm. d IHC staining of HN1L in HCC tissue microarray (n = 139). Scale bars, up: 200 μm; down: 50 μm. Negative and weak levels of HN1L in HCCs were consider as low expression (n = 43), and moderate or strong staining intensities were determined as high expression (n = 96). e HN1L staining scores in HCC tumors and the corresponding non-tumor tissues (n = 139). Data are presented as the mean ± SD. f Correlation analyses between HN1L expressions and clinical characteristics in HCCs. g and h Kaplan–Meier survival curves showed that HN1L expression level was negatively correlated with prognosis prediction of HCCs analyzed by HCC tissue microarray and TCGA database
Next, a tissue microarray containing 139 pairs of primary HCCs was applied to analyze the association of HN1L overexpression with clinicopathological features (Fig. 1d). Firstly, IHC staining analysis indicated the high expression of HN1L in HCC tissues (P < 0.001, Fig. 1e). Furthermore, overexpression of HN1L was significantly associated with tumor size (P < 0.001), adjacent organ invasion (P = 0.003), tumor thrombus (P = 0.001), and metastasis (including intrahepatic and distant metastasis) (P = 0.035, Fig. 1f). Kaplan-Meier analysis revealed that overexpression of HN1L was significantly associated with poorer overall survival (P = 0.012) and progression-free survival rates (P < 0.001) of HCCs (Fig. 1g). In addition, according to the clinical data from TCGA database, HN1L gene expression levels were gradually upregulated from the well differentiated group and the poor differentiated group, as well as from early-stage to stage III of HCC (Supplementary Fig. 2A and 2B). Kaplan-Meier survival curves based on TCGA database suggested that both overall survival (P < 0.001) and progression-free survival (P = 0.003) of HCCs with high HN1L expression were significantly shorter than those with low levels of HN1L expression (Fig. 1h). Taken together, these evidences implicate an aggressive role of HN1L in HCC progression.
To explore the mechanism of HN1L transcriptionally upregulated in HCC, we used TargetScan to predict the miRNAs that potentially regulated the expression of HN1L. Results indicated that miR-212-5p potentially bind the 3' UTR of HN1L (Supplementary Fig. 2C). To determine if HN1L is regulated by miR-212-5p in HCC, the expression levels of HN1L and miR-212-5p were analyzed in 23 HCC samples with qRT-PCR. Results showed that the expression of HN1L is negatively correlated with miR-212-5p in HCCs (r = −0.4441, P = 0.0005, Supplementary Fig. 2D). Most importantly, it has been reported that miR-212-5p was down-regulated in HCC and overexpression of miR-212-5p inhibited the growth and migration of HCC cells [10]. Therefore, HN1L expression is transcriptionally regulated by miR-212-5p in HCC. In addition, gene expression and cancer patients survival data from TCGA also showed the higher expression of HN1L in some other aggressive cancers, including bladder urothelial carcinoma, breast invasive carcinoma, brain lower grade glioma, lung cancer (squamous cell carcinoma and adenocarcinoma), pancreatic adenocarcinoma and skin cutaneous melanoma, compared with their corresponding normal tissues (Supplementary Fig. 3A). Most importantly, up-regulated HN1L in these cancer tissues was significantly associated with poorer overall survival (Supplementary Fig. 3B).
HN1L has strong oncogenicity function
To investigate the role of HN1L in HCC progression, Gene Ontology Enrichment analysis (biological process) was performed using Coexpedia [11]. Analysis results show that HN1L is significantly related to DNA replication and cell cycle regulation, which hints that HN1L may be associated with cell malignant proliferation in HCC. (Fig. 2a). Hence, two HCC cell lines BEL-7402 and QGY-7703 that relatively low-expressed HN1L were stably transfected with the lentiviral HN1L construct and empty lentivector, respectively (Fig. 2b). Ectopic expression of HN1L was evaluated by western blotting (Fig. 2c). Both in vitro and in vivo functional assays were used to characterize the tumorigenicity of HN1L. Cell growth assay showed that the cell growth rates in HN1L-transfected cells were significantly higher than that in control cells (P < 0.001, Fig. 2d). The foci formation frequency was significantly higher in HN1L-expressing cells compared to the control cells (P < 0.001, Fig. 2e). Non-adherent colony formation assays showed that the formation frequency and volume of microspheres in soft agar were significantly increased in HN1L-transfected cells than that in the control cells (P < 0.001, Fig. 2f). Subcutaneous tumor xenografting assay in nude mice showed that the volume of xenograft tumors developed from HN1L-transfected cells was significantly larger than tumors from control cells (Fig. 2g). Most importantly, results from the IHC staining confirmed the higher expressions of HN1L and the proliferation marker Ki67 in xenograft tumors induced by HN1L-transfected cells, compared with control cells (Fig. 2h). Therefore, upregulation of HN1L facilitates the progression of HCC by promoting malignant proliferation.
Ectopic expression of HN1L increases tumor growth in HCC. a Gene Ontology Enrichment analysis (biological process) of HN1L using Coexpedia internet tool (http://www.coexpedia.org/) that was based on public GEO datasets. b HN1L expressions in two immortalized liver cell lines (L02 and MIHA) and eleven HCC cell lines were tested by western blot. β-Tubulin was served as a loading control. c Western blot analysis showing ectopic expression of HN1L in HN1L-transfected BEL-7402 and QGY-7703 cells. d Cell growth rates between HN1L-transfected and empty vector-transfected BEL-7402 or QGY-7703 cells. Representative images of increased foci formation in monolayer culture (e) and spheres formation in soft agar (f) induced by HN1L overexpression in BEL-7402 or QGY-7703 cells. g Images of the xenograft tumors formed in nude mice injected with HN1L-transfected and empty vector-transfected cells (n = 5). The weights of xenograft tumors were summarized in the right panel. h Representative IHC images of HN1L and Ki67 expressions in xenograft tumors originated from HN1L-transfected and empty vector-transfected BEL-7402 or QGY-7703 cells. Scale bars = 100 μm
Knockdown of HN1L abolishes its tumorigenicity
To further confirm the oncogenic effect of HN1L, one high-efficiency targeted shRNA (shHN1L) was stably transfected into two HCC cell lines Hep3B and HCCLM6 that highly expressed HN1L (Fig. 2b). Western blotting confirmed the silence of HN1L by shRNA at protein levels (Fig. 3a). BrdU incorporation and cell growth assays showed that silencing of HN1L expression significantly inhibited the proliferation of Hep3B and HCCLM6 cells (P < 0.001, Fig. 3b, c). Functional assays revealed that knockdown of HN1L decreased the frequencies of foci and spheres formation (P < 0.001, Fig. 3d, e). In addition, in vivo tumorigenicity assay showed that xenograft tumors induced by shHN1L-transfected cells were significantly smaller than tumors induced by scramble control cells (Fig. 3f). Down-regulation of HN1L and Ki67 were observed in xenograft tumors induced by shHN1L-treated cells than tumors from control cells (Fig. 3g).
Knockdown of HN1L inhibits HCC cell growth in vitro and in vivo. a One shRNA against HN1L effectively down-regulated the expression of HN1L in Hep3B and HCCLM6 cells detected with western blotting. Scrambled shRNA was used as negative control. BrdU incorporation (b) and Cell growth (c) assays showed that knockdown of HN1L deceased cell proliferation rate in Hep3B and HCCLM6 cells. Scale bar = 50 μm in panel b. Silencing HN1L expression could effectively inhibit the foci formation in monolayer culture (d) spheres formation in soft agar (e) and tumor formation in nude mice (f, n = 4). g IHC staining confirmed the down-regulation of HN1L and Ki67 in xenograft tumors from shHN1L-transfected Hep3B or HCCLM6 cells. Scale bars, 200 μm
In addition, flow cytometry assay showed that overexpression of HN1L could accelerate the cell cycle in HN1L-transfected BEL-7402 and QGY-7703 cells (Supplementary Fig. 4A). Moreover, western blotting showed the up-regulation of cyclin D1, cyclin E1, CDK2, CDK4, and CDK6 in HN1L-overexpressed cells (Supplementary Fig. 4B). Inversely, knockdown of HN1L could induce cell cycle arrest and down-regulate this cell cycle-related proteins in HN1L-silenced HCCLM6 and Hep3B cells (Supplementary Fig. 4A and 4B). However, overexpression or knockdown of HN1L in HCC cells did not affect cell apoptosis (Supplementary Fig. 4C and 4D). Taken together, our data suggest the important role of HN1L in HCC progression.
HN1L drives cell migration and metastasis by inducing epithelial-mesenchymal transition (EMT)
Since overexpression of HN1L was closely associated with unfavorable progression-free survival of HCCs, the effect of HN1L on HCC metastasis was also investigated in vitro and in vivo. 3D tumor spheroid invasion assay in Matrigel showed the invasive morphological characteristics of tumor spheroid in HN1L-tansfected cells (Fig. 4a). Transwell migration and invasion assays also revealed that overexpression of HN1L could significantly increase cell motility and invasion (Fig. 4b). Conversely, the wound-healing assay showed that HN1L-silenced cells had slower closure of the scratched “wound”, compared with the control cells (Fig. 4c). Moreover, silencing HN1L could significantly decreased the migratory and invasive abilities of Hep3B and HCCLM6 cells (Fig. 4d). Remarkably, silencing of HN1L could convert the high-invasive Hep3B and HCCLM6 cells into lowly metastatic entities as assessed using spleen to liver metastasis model (Fig. 4e).
HN1L drives HCC cell migration and metastasis by inducing epithelial-mesenchymal transition (EMT). a 3D tumor spheroid invasion assay in Matrigel showed the invasive morphological characteristics of tumor spheroid when BEL-7402 or QGY-7703 cells overexpressed HN1L. Scale bar, 100 μm. b Transwell assays showed that ectopic expression of HN1L promoted cell migration and invasion in BEL-7402 and QGY-7703 cells. c Wound-healing assay showed that knockdown of HN1L inhibited cell migration. Representative images were taken at 0, 24, and 48 h after scratching. d Silencing HN1L expression could inhibit cell migration and invasion in Hep3B and HCCLM6 cells. e In vivo liver metastasis assay via spleen injection showed the lesser metastatic tumor nodules from Hep3B and HCCLM6 HN1L-silenced cells, compared with scramble control-transfected cells (n = 5). f IF staining with antibodies against HN1L and skeleton protein F-actin showed that silencing HN1L in HCCLM6 cells induced the morphological changes. Scales bar = 50 μm. g Western blotting showed that several key markers involved in EMT were regulated by HN1L. β-Tubulin was used as a loading control. h Representative IF images showing the decreased expression of E-cadherin and up-regulation of vimentin in HN1L-transfected QGY-7703 cells. Scales bar = 50 μm. In all panels, data are presented as the mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001
EMT was shown to strongly enhance cancer cell motility and metastasis [12]. Knockdown of HN1L in HCCLM6 cells obviously induced the phenotypes changes from leptosomatic to epithelioid shape (Fig. 4f). Hence, we analyzed the expression changes of several EMT-associated proteins in HN1L-transfected and HN1L-knockdown cells with western blotting. Results demonstrated that overexpression of HN1L could down-regulate the levels of the epithelial markers E-cadherin and ZO-1, and up-regulate the mesenchymal markers N-cadherin and vimentin (Fig. 4g). Immunofluorescence (IF) staining also confirmed the decreased expression of E-cadherin and increased expression of vimentin in HN1L-transfected QGY-7703 cells (Fig. 4h). Collectively, these data indicate that HN1L promotes HCC cell migration and tumor metastasis by inducing EMT.
HN1L increases HCC growth and metastasis by up-regulating METTL13
To investigate the mechanisms of HN1L promoting HCC cell growth and metastasis, we surveyed the genes that were positively co-expressed with HN1L in human using the Coexpedia. Analysis results showed that one gene METTL13 was the top differentially co-expressed gene with HN1L (Fig. 5a, b). Importantly, the expression of METTL13 was obviously increased in HN1L-transfected cells, while decreased in HN1L-knockdown cells (Fig. 5c). However, silencing METTL13 with siRNA did not affect the expression of HN1L in Hep3B and HCCLM6 cells (Fig. S5), suggesting that METTL13 was a downstream target gene of HN1L in HCC.
HN1L promotes HCC cell growth and metastasis by up-regulating METTL13. a Co-expressed genes with HN1L were surveyed using the Coexpedia internet tool (http://www.coexpedia.org/). b Correlation analyses the expressions of HN1L and METTL13 in HCC based on the TCGA database. c RT-qPCR analyzed the effects of overexpressing HN1L in BEL-7402 and QGY-7703 cells or silencing HN1L with siRNA in Hep3B and HCCLM6 cells on the expression of METTL13. d Expression of METTL13 in HCC tumor and normal liver tissues were analyzed in TCGA datasheet and Lim HY’ cohort (GSE36376) dataset (independent Student’s t-test). e IHC staining of METTL13 in HCC tumor and paired normal liver tissues. Scale bar, 200 μm. f Kaplan–Meier survival curves showed that METTL13 expression was negatively correlated with prognosis prediction of HCC patients analyzed by TCGA database. g BrdU incorporation assay showed that silencing METTL13 by siRNA could abrogate the proliferation promotion effects of HN1L overexpression in QGY-7703 cells. Scale bar, 100 μm. h Transwell migration assay showed that knockdown of METTL13 by siRNA abolished the HN1L induced migration ability in QGY-7703 cells. In all panels, data are presented as the mean ± SD. **P < 0.01, ***P < 0.001, ns no statistical significance
METTL13 is uniformly overexpressed in human colon, brain, breast, and lung cancers compared with the corresponding normal tissues [13]. METTL13 could induce HCC cells metastasize to the lung in mice, but its mechanism remains unclear [14]. Here, we showed that METTL13 was significantly overexpressed in HCCs according to TCGA database and Lim HY’ cohort (GSE36376) (Fig. 5d). IHC staining also showed the high expression of HN1L in HCC compared to normal liver tissue (Fig. 5e). Moreover, overexpression of HN1L predicted the worse overall survival and progression-free survival in HCCs (Fig. 5f). This evidences indicates the important roles of METTL13 in HCC progression. Importantly, silencing METTL13 by siRNA could abolish the promotion effects in cell proliferation and metastasis caused by HN1L overexpression in QGY-7703 cells (Fig. 5g, h). Taken together, HN1L facilitates HCC growth and metastasis through the transcriptional up-regulation of METTL13.
HN1L up-regulates METTL13 by activating its transcription factor AP-2γ
To identify the transcriptional factor of METTL13 positively regulated by HN1L, we surveyed one protein-protein interaction database IntAct [15] that displayed 11 unique HN1L interactors including the AP-2γ, the potential transcription factor of METTL13 (Fig. 6a, Supplementary Fig. 6A). IF double staining (Fig. 6b, Supplementary Fig. 6B) and co-immunoprecipitation (Fig. 6c) showed that HN1L directly bound to AP-2γ protein in HCC cells. Most importantly, chromatin immunoprecipitation (ChIP)-qPCR confirmed that AP-2γ indeed bound to the promoter of METTL13, and overexpression of HN1L promoted this binding (Fig. 6d). In addition, luciferase assay further verified that the complex of HN1L and AP-2γ promoted METTL13 expression in BEL-7402 and QGY-7703 cells (Fig. 6e). Moreover, ectopic expression of AP-2γ in BEL-7402 and QGY-7703 cells increased the expression of METTL13 (Fig. 6f). Silencing AP-2γ by siRNA could abolish the up-regulation of METTL13 induced by HN1L ectopic expression in BEL-7402 and QGY-7703 cells (Fig. 6g). Furthermore, the enhanced cell proliferation and migration abilities in HN1L-transfected QGY-7703 cells were abrogated upon AP-2γ knockdown (Fig. 6h, i). However, there are no transcriptional correlation between HN1L and AP-2γ in HCC (Supplementary Fig. 6C). Collectively, these data show that transcription factor AP-2γ is required for HN1L to promote the transcription of METTL13 that leads to the increased proliferation and metastasis.
HN1L up-regulates METTL13 expression via interaction of AP-2γ. a The datasheet showed HN1L interacting proteins obtained from IntAct database (https://www.ebi.ac.uk/intact/). b IF double staining of HN1L and AP-2γ in HN1L-transfected BEL-7402 cells. Scale bar, 20 μm. c Co-immunoprecipitation showed that HN1L directly bound to transcription factor AP-2γ in HN1L-transfected BEL-7402 and QGY-7703 cells. d AP-2γ binding to the promoters of METTL13 detected by ChIP-qPCR in BEL-7402 and QGY-7703 cells. e BEL-7402 and QGY-7703 cells were co-transfected with METTL13 promoter-luciferase reporter and AP-2γ or HN1L plasmids followed by analysis of luciferase activity. f Ectopic expression of AP-2γ in BEL-7402 and QGY-7703 cells could increase the expression of METTL13. g Silencing AP-2γ by siRNA could abolish the up-regulation of METTL13 induced by HN1L ectopic expression in BEL-7402 and QGY-7703 cells. h BrdU incorporation assay showed that the enhanced cell proliferation of QGY-7703 cells induced by HN1L overexpression was abrogated upon AP-2γ knockdown. Scale bar, 100 μm. i Transwell migration assay showed that silencing AP-2γ could decrease the metastatic ability of HN1L-transfected QGY-7703 cells. In all panels, data are presented as the mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, ns no statistical significance
METTL13 facilitates tumorigenicity and metastasis via up-regulation of TCF3 and ZEB1
Next, we investigated the mechanisms of METTL13 as a crucial mediator of HN1L in regulating HCC cell growth and metastasis. Correlation analysis in HCCs using GEPIA web server showed that three key transcription factors, transcription factor 3 (TCF3), transcription factor 4 (TCF4), and zinc finger E-box binding homeobox 1 (ZEB1), were strongly associated with the expressions of HN1L, AP-2γ, and METTL13, respectively (Fig. 7a, Supplementary Fig. 7A). Moreover, TCF3, TCF4 and ZEB1 were up-regulated in HCCs than those in normal liver tissues according to the TCGA datasheet (Fig. 7b). However, only the high expressions of TCF3 and ZEB1 were significantly associated with the unfavorable overall survival and progression-free survival of HCCs (Fig. 7c). Importantly, ectopic expression of HN1L in BEL-7402 and QGY-7703 cells dramatically increased the transcription of TCF3 and ZEB1, but not TCF4 (Fig. 7d). Conversely, knockdown of HN1L or METTL13 could induce the down-regulation of TCF3 and ZEB1 without affecting the expression of TCF4 (Fig. 7e). In summary, METTL13 can transcriptionally up-regulate TCF3 and ZEB1 that are known and vital regulators in tumor growth and metastasis.
TCF3 and ZEB1 are key performers in HN1L promoting tumorigenicity and metastasis. a Correlation analysis between the expressions of HN1L, AP-2γ or METTL13 and 15 key transcription factors in regulating tumorigenicity and metastasis in HCC using GEPIA web server that is based on TCGA database (http://gepia.cancer-pku.cn/). b TCF3, TCF4 and ZEB1 were up-regulated in HCC tissues than those in normal liver tissues according to the TCGA datasheet. c Survival curves from TCGA database showed the correlations between TCF3, TCF4 or ZEB1 expression levels and HCCs outcomes. d Ectopic expression of HN1L in BEL-7402 and QGY-7703 cells dramatically increased the transcription of TCF3 and ZEB1, but not TCF4. e Knockdown of HN1L or METTL13 induces the down-regulation of TCF3 and ZEB1 without affecting the expression of TCF4 in Hep3B and HCCLM6 cells. f Schematic diagram summarizes that high expressed HN1L up-regulates the gene METTL13 via the interaction of AP-2γ at transcriptional level, which further induces the high expressions of TCF3 and ZEB1 that are two key performers accelerate proliferation and metastasis in HCC cells. In all panels, data are presented as the mean ± SD. **P < 0.01, ***P < 0.001, ns no statistical significance
Binding sites analysis with Predict Protein, an open resource for online prediction of protein structural features [16], showed that METTL13 had many protein-protein binding sites without polynucleotide binding domains, which suggested that METTL13 might interact with the transcription factor of TCF3 and ZEB1 (Supplementary Fig. 7B). Therefore, using APID proteins interaction database, we found that METTL13 directly interacted with c-Myc, which was confirmed by co-immunoprecipitation in HCCLM6 cells (Supplementary Table S1, Supplementary Fig. 7C). Most importantly, c-Myc as a transcription factor could promote the high constitutive expression of TCF3 and ZEB1 [17, 18]. Therefore, up-regulation of METTL13 by HN1L further induces the high expressions of TCF3 and ZEB1 via interaction of c-Myc, facilitating HCC cell proliferation and metastasis (Fig. 7f).
Administration of lentivirus-shRNA suppresses tumor growth and metastasis
As HN1L is a crucial instigator in regulating HCC cell growth and metastasis, we next investigate the potential of targeting HN1L by the lentivirus containing the shRNA targeting HN1L (LV-shHN1L) for suppressing HCC progression. Using flow cytometry, we firstly confirmed the high infection efficiency (~100%) of lentiviral particle containing GFP in HCCLM6 and Hep3B cells in vitro before intratumor injection and tail intravenous injection (Supplementary Fig. 8A). Firstly, lentivirus containing the scramble control shRNA (LV-scramble) or LV-shHN1L was orthotopically injected into Hep3B-derived and HCCLM6-derived tumors, respectively. Results showed that the tumors treated with LV-shHN1L exhibited a decrease in volume by comparison with LV-scramble control (Fig. 8a). Furthermore, xenograft sections subjected to HN1L or Ki67 staining showed that treatment with LV-shHN1L resulted in a significant decreased expression of HN1L and Ki67 compared with LV-scramble treatment (Fig. 8b). In addition, tail intravenous administration of LV-shHN1L could also decrease the number of metastatic tumor nodules in liver using spleen to liver metastasis model for Hep3B cells (Fig. 8c). H&E staining showed a smaller metastatic tumor in liver treated with LV-shHN1L, compared with LV-scramble control (Fig. 8d). IHC staining and western blotting also confirmed the down-regulation of HN1L in subcutaneous and metastatic tumors treated with LV-shHN1L (Supplementary Fig. 8B-D). Importantly, intratumoral injection of LV-shHN1L could also effectively inhibit the growth of HCC patient-derived tumor xenograft (PDX) that high expressing HN1L (Fig. 8e, Supplementary Fig. 8E). IHC staining also confirmed the down-regulations of HN1L and Ki67 proteins in PDX treated with LV-shHN1L (Fig. 8f). Taken together, our data suggest that HN1L is a promising therapeutic target for suppressing HCC cell growth and metastasis.
Lentivirus-shHN1L diminished the tumorigenesis and metastasis in vivo. a Lentivirus containing the shRNA targeting HN1L (LV-shHN1L) or scramble control shRNA (LV-scramble) was infected intratumorally into Hep3B-derived and HCCLM6-derived tumors, respectively. Treatments were performed every two days for four times. b Xenograft tumor sections were stained with antibodies against HN1L and Ki67. Scale bars, 200 μm. c Tail intravenous administration of LV-shHN1L could suppress liver metastasis using spleen to liver metastasis model for Hep3B cells (n = 6). d H&E staining showed the metastatic tumor nodules in liver of nude mice treated with LV-shHN1L or LV-scramble control. e Subcutaneous transplanted HCC patient-derived xenograft (PDX) in nude mice were treated with LV-scramble or LV-shHN1L via intratumoral injection (n = 4). Xenograft tumors were indicated by arrows. f IHC staining confirmed the down-regulations of HN1L and Ki67 proteins in PDX treated with LV-shHN1L. Scale bars, 200 μm. In all panels, data are presented as the mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001
Discussion
Although HN1L gene has been cloned and characterized in human for over a decade, but there was still a lack of research reports about the functions of HN1L under physiological or pathophysiological conditions [19]. In this study, we investigated the oncogenic effect and underlying mechanism of HN1L in HCC progression. Correlation analysis indicated that overexpression of HN1L was positively associated with tumor size, adjacent organs invasion, tumor thrombus, and distant organs metastasis. Ectopic expression of HN1L could increase cell growth rate, frequencies of foci formation and tumor spheres in soft agar, and tumorigenesis in nude mice. Inversely, knockdown of HN1L by shRNA markedly inhibited cell proliferation in vitro and tumor formation in vivo. In other hand, overexpression of HN1L stimulates the metastatic potential of BEL-7402 and QGY-7703 cells, and silencing HN1L abrogates the cell invasive ability of metastatic Hep3B and HCCLM6 cells using transwell invasion assay in vitro and spleen to liver metastasis model in nude mice. Taken together, our functional analyses demonstrate the pro-proliferation and pro-metastasis roles of HN1L in HCC progression.
Gene co-expression analysis and knockdown experiments confirmed that METTL13 is a key mediator for HN1L in promoting tumor growth and metastasis in HCC. METTL13 was first cloned from rat brain, but its function was rarely reported [20]. Initial study hinted that METTL13 attenuated apoptotic cell death [13]. Moreover, GEO profile databases showed dysregulation of METTL13 gene was associated with tumor malignancy, metastasis, chemosensitivity, and microsatellite instability [21]. However, Zhang et al. showed that the overexpression of METTL13 hinders cellular migration and invasion in bladder cancer cells [21]. Importantly, METTL13 has been linked with cancer stemness [22]. In the present study, we showed that the transcriptional level of METTL13 was up-regulated by HN1L via the activation of its transcription factor AP-2γ in HCC, and overexpression of METTL13 was significantly associated with poorer survival of HCCs. Further study revealed that TCF3 and ZEB1 were the downstream targets of HN1L/AP-2γ/METTL13 pathway.
TCF3 and ZEB1 were common transcription factors in regulating cancer cell proliferation, migration, and invasion [23, 24]. Silencing of TCF3 dramatically decreased the ability of breast cancer cells to initiate tumor formation, and led to decreased tumor growth rates. In culture, TCF3 promotes the sphere formation capacity of breast cancer cells and their self-renewal [25]. Moreover, TCF3 and TCF4 are the key transcription factors that collaborate with β-catenin in its established role in hair follicle stem cell activation [26]. ZEB1 is a central transcription factor that promotes tumor invasion and metastasis by inducing EMT in cancer cells [27, 28]. Therefore, we illuminate the molecular mechanism of HN1L in promoting HCC cell growth and metastasis by up-regulating TCF3 and ZEB1 factors through the transcriptional activation of METTL13.
RNA interference is a specific gene silencing phenomenon induced by double-stranded RNA. Lentiviral vector permits efficient delivery and stable transfection of a sequence of interest, efficiently infects both dividing and non-dividing cells, and shows minimal immunogenicity [29]. Lentivirus-mediated knockdown of oncogenes can inhibit HCC cell growth and invasion, which is probably a promising target for HCC treatment [30]. Here, our study demonstrated that knockdown of HN1L by LV-shHN1L led to decreased tumorigenesis and metastasis of HCC cells in vivo. Moreover, LV-shHN1L could also markedly diminished the growth of HCC PDX in nude mice, suggesting its potential therapeutic value in HCC treatment. However, silencing HN1L could not induce HCC cell apoptosis. The vector containing shRNA can be obtained through bacterial transformation, and lentivirus can be abundantly produced with lentivirus packaging system in vitro. The high transfection efficiency can also meet the need for gene silencing [31]. Therefore, the combination of lentivirus-mediated shRNA and chemotherapy may be a potential therapeutic approach for HCC treatment. In addition, screening the small molecule inhibitors targeting HN1L is also a promising strategy to treat HCC.
Collectively, we reported that HN1L was frequently overexpressed in HCC, and it played an important role in HCC cell proliferation and metastasis by activating AP-2γ/METTL13/TCF3-ZEB1 signaling axis. Importantly, administration of LV-shHN1L could suppress tumor growth and metastasis in vivo, which may lead to develop a novel therapeutic approach in HCC treatment.
Methods
HCC samples and cell lines
Primary HCC specimens and their corresponding non-tumor tissues were obtained with informed consent from patients who underwent hepatectomy for HCC at the Sun Yat-sen University Cancer Center (Guangzhou, China). A tissue microarray containing 139 pairs of matched primary HCC tumor tissues and corresponding non-tumor tissues were retrieved from the archive of paraffin-embedded tissues obtained between 2003 and 2010 at the Sun Yat-sen University Cancer Center (Guangzhou, China) [32]. All samples used in this study were approved by the Committees for Ethical Review at the Sun Yat-sen University Cancer Center. HCC cell line HCCLM6 was obtained from Liver Cancer Institute and Zhongshan Hospital of Fudan University (Shanghai, China). Hep3B was purchased from the American Type Culture Collection (ATCC, Manassas, Virginia, USA). HCC cell lines BEL-7402 and QGY-7703 were obtained from the Institute of Virology, Chinese Academy of Medical Sciences (Beijing, China). Immortalized human hepatocyte line MIHA was provided by Dr. J. R. Chowdhury (Albert Einstein College of Medicine, New York). All cell lines were cultured in high-glucose DMEM (GibcoBRL, Grand Island, NY) supplemented with 10% fetal bovine serum (FBS, GibcoBRL, Grand Island, NY) at 37 °C with 5% CO2.
In vitro oncogenic assays
In vitro tumorigenicity was assessed by cell growth, foci formation, and soft agar assays. For cell growth assay, cells were seeded at a density of 1000 per well onto 96-well plates. The cell growth rate was monitored using a CCK-8 assay kit (Dojindo Corp. Japan) according to the manufacturer’s instruction. For foci formation assay, 1000 cells were seeded onto 6-well plates and then cultured for one week. Surviving colonies were stained counted with 1% crystal violet and colony consisted of >50 cells were counted. Anchorage-independent growth was assessed by colony formation ability in soft agar. Briefly, 5000 cells were suspended in soft agar mixture (DMEM, 10% FBS and 0.4% Sea Plaque agarose) and were subsequently overlaid on the solidified 0.6% agar base. After 2 weeks, colonies (≥10 cells) were counted under the microscope in 10 fields per well.
In vivo xenograft assay
All animal experiments were approved by Animal Ethics Committee at Sun Yat-sen University Cancer Center. Five-week-old female BALB/c nude mice were purchased from the Guangdong Medical Laboratory Animal Center (Guangzhou China). Samples with different numbers of HCC cells (BEL-7402: 5 × 106; QGY-7703: 5 × 106; Hep3B: 3 × 106; HCCLM6: 4 × 106) with HN1L overexpression or knockdown in 100 μl phosphate buffered saline (PBS) were injected subcutaneously into nude mice. The length (L) and width (W) of tumor were measured every week by calipers for 4 weeks, and tumor volumes were calculated as volume (mm3) = L × W2 × 0.5.
Cell migration and invasive assays in vitro
For wound healing assay, cells were grown as a confluent monolayer in 6-well plates. The wounds of cell layer were introduced by scraping the confluent cell with a 200 μl pipette tip. Next, floating cells were carefully removed with DMEM before normal medium was added. The wound healing process was monitored under an inverted light microscope (Olympus, Lake Success, NY). The migration abilities were quantified and normalized by relative gap distance. Cell motility was also assessed by cell migration and invasion arrays using transwell chambers (pore size 8 μm) with or without Matrigel membrane (Corning, NY, USA). Briefly, after serum starvation for 24 h, cells (5 × 104 cells for BEL-7402 and QGY-7703; 1 × 104 cells for Hep3B and HCCLM6) in DMEM medium without FBS were layered in the upper chamber, and medium containing 10% FBS was applied to the lower chamber. The chamber was then incubated for 24 h for cell migration (using transwell without Matrigel) or 42 h for invasion (using transwell with Matrigel) at 37 °C. After removing the cells in the upper surface of filter with cotton swab, the invasive cells attached to the lower surface of the membrane were fixed with 4% paraformaldehyde solution, stained with 0.1% crystal violet and then quantified by counting the cell number at 6 random fields under a microscope.
Injection of lentivirus-mediated shHN1L to inhibit tumor growth and metastasis
The lentivirus was concentrated by ultracentrifuging, and the lentivirus titer was performed as described previously [33]. To assess the inhibition effects of virus particle including shHN1L on established tumors, male BALB/c nude mice at 4 weeks of age were injected subcutaneously into the right dorsal flanks with 100 μl PBS containing 3 × 106 Hep3B or 4 × 106 HCCLM6 cells. The mice were divided randomly into two groups. Once tumors reached a volume of 300 mm3, mice received an intratumoral injection of 50 μl (4 × 108 TU/ml) of virus containing shScramble or virus containing shHN1L. The treatments were performed every two days for four times. At the same time, the volume of tumors were measured by calipers. To explore the inhibition of LV-shHN1L in HCC patient-derived xenograft (PDX), fresh HCC tissue was subcutaneously transplanted into nude mice as soon as possible after hepatectomy. The mice were treated with an intratumoral injection of 50 μl (4 × 108 TU/ml) of virus particle every two days for 10 times. The length (L) and width (W) of tumor were measured every week by calipers, and tumor volumes were calculated as volume (mm3) = L × W2 × 0.5. To test the inhibition of lentivirus-mediated shHN1L on tumor metastasis, we used the liver and spleen metastasis model. After 2 × 106 Hep3B cells were injected into the spleen of the tested nude mouse, 100 μl concentrated lentivirus containing shScramble or shHN1L (4 × 108 TU/ml) was administrated by tail intravenous injection. The treatments were performed every two days for four weeks.
Statistical analysis
SPSS version 17.0 (Chicago, IL) was used for all data analyses. Pearson chi-square test was used for the categorical variables, and an independent Student t test was used for continuous data. The prognostic value was calculated by the Kaplan-Meier analysis with log-rank test. Gene expression levels in non-tumor liver and HCC tissues were directly obtained from UALCAN (http://ualcan.path.uab.edu/) that provided access to publicly available cancer transcriptome data (TCGA) or from GEO datasheet (GSE36376) [34, 35], which have been normalized using the locally weighted scatter plot smoothing (Lowess). Survival curves in TCGA database were directly produced from GEPIA (http://gepia.cancer-pku.cn/) basing on a suitable expression threshold for splitting the high-expression and low-expression cohorts [36]. Co-expressed genes, Gene Ontology and Disease Ontology analyses were analyzed with Coexpedia (http://www.coexpedia.org/) that was based on a series of GEO dataset [11]. Results were considered statistically significant when P < 0.05.
References
Ko MS, Kitchen JR, Wang X, Threat TA, Wang X, Hasegawa A, et al. Large-scale cDNA analysis reveals phased gene expression patterns during preimplantation mouse development. Development. 2000;127:1737–49.
Petroziello J, Yamane A, Westendorf L, Thompson M, McDonagh C, Cerveny C, et al. Suppression subtractive hybridization and expression profiling identifies a unique set of genes overexpressed in non-small-cell lung cancer. Oncogene. 2004;23:7734–45.
Li L, Zeng TT, Zhang BZ, Li Y, Zhu YH, Guan XY. Overexpression of HN1L promotes cell malignant proliferation in non-small cell lung cancer. Cancer Biol Ther. 2017;18:904–15.
Laughlin KM, Luo D, Liu C, Shaw G, Warrington KH Jr., Law BK, et al. Hematopoietic- and neurologic-expressed sequence 1 (Hn1) depletion in B16.F10 melanoma cells promotes a differentiated phenotype that includes increased melanogenesis and cell cycle arrest. Differ Res Biol Divers. 2009;78:35–44.
Goto T, Hisatomi O, Kotoura M, Tokunaga F. Induced expression of hematopoietic- and neurologic-expressed sequence 1 in retinal pigment epithelial cells during newt retina regeneration. Exp Eye Res. 2006;83:972–80.
Zhang ZG, Chen WX, Wu YH, Liang HF, Zhang BX. MiR-132 prohibits proliferation, invasion, migration, and metastasis in breast cancer by targeting HN1. Biochem Biophys Res Commun. 2014;454:109–14.
Zhang C, Xu B, Lu S, Zhao Y, Liu P. HN1 contributes to migration, invasion, and tumorigenesis of breast cancer by enhancing MYC activity. Mol Cancer. 2017;16:90.
Laughlin KM, Luo D, Liu C, Shaw G, Warrington KH Jr., Qiu J, et al. Hematopoietic- and neurologic-expressed sequence 1 expression in the murine GL261 and high-grade human gliomas. Pathol Oncol Res. 2009;15:437–44.
Varisli L, Ozturk BE, Akyuz GK, Korkmaz KS. HN1 negatively influences the beta-catenin/E-cadherin interaction, and contributes to migration in prostate cells. J Cell Biochem. 2015;116:170–8.
Jia P, Wei G, Zhou C, Gao Q, Wu Y, Sun X, et al. Upregulation of MiR-212 inhibits migration and tumorigenicity and inactivates Wnt/beta-Catenin signaling in human hepatocellular carcinoma. Technol Cancer Res Treat. 2018;17:1533034618765221.
Yang S, Kim CY, Hwang S, Kim E, Kim H, Shim H, et al. COEXPEDIA: exploring biomedical hypotheses via co-expressions associated with medical subject headings (MeSH). Nucleic Acids Res. 2017;45(D1):D389–D96.
Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139:871–90.
Takahashi A, Tokita H, Takahashi K, Takeoka T, Murayama K, Tomotsune D, et al. A novel potent tumour promoter aberrantly overexpressed in most human cancers. Sci Rep. 2011;1:15.
Rebouissou S, Amessou M, Couchy G, Poussin K, Imbeaud S, Pilati C, et al. Frequent in-frame somatic deletions activate gp130 in inflammatory hepatocellular tumours. Nature. 2009;457:200–4.
Orchard S, Ammari M, Aranda B, Breuza L, Briganti L, Broackes-Carter F, et al. The MIntAct project–IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 2014;42(Database issue):D358–63.
Yachdav G, Kloppmann E, Kajan L, Hecht M, Goldberg T, Hamp T, et al. Predict Protein–an open resource for online prediction of protein structural and functional features. Nucleic Acids Res. 2014;42(Web Server issue):W337–43.
Braso-Maristany F, Filosto S, Catchpole S, Marlow R, Quist J, Francesch-Domenech E, et al. PIM1 kinase regulates cell death, tumor growth and chemotherapy response in triple-negative breast cancer. Nat Med. 2016;22:1303–13.
Scheller H, Tobollik S, Kutzera A, Eder M, Unterlehberg J, Pfeil I, et al. c-Myc overexpression promotes a germinal center-like program in Burkitt's lymphoma. Oncogene. 2010;29:888–97.
Zhou G, Wang J, Zhang Y, Zhong C, Ni J, Wang L, et al. Cloning, expression and subcellular localization of HN1 and HN1L genes, as well as characterization of their orthologs, defining an evolutionarily conserved gene family. Gene. 2004;331:115–23.
Nagase T, Ishikawa K, Suyama M, Kikuno R, Hirosawa M, Miyajima N, et al. Prediction of the coding sequences of unidentified human genes. XII. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. DNA Res. 1998;5:355–64.
Zhang Z, Zhang G, Kong C, Zhan B, Dong X, Man X. METTL13 is downregulated in bladder carcinoma and suppresses cell proliferation, migration and invasion. Sci Rep. 2016;6:19261.
Kim J, Woo AJ, Chu J, Snow JW, Fujiwara Y, Kim CG, et al. A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell. 2010;143:313–24.
Shah M, Rennoll SA, Raup-Konsavage WM, Yochum GS. A dynamic exchange of TCF3 and TCF4 transcription factors controls MYC expression in colorectal cancer cells. Cell Cycle. 2015;14:323–32.
Song XF, Chang H, Liang Q, Guo ZF, Wu JW. ZEB1 promotes prostate cancer proliferation and invasion through ERK1/2 signaling pathway. Eur Rev Med Pharmacol Sci. 2017;21:4032–8.
Slyper M, Shahar A, Bar-Ziv A, Granit RZ, Hamburger T, Maly B, et al. Control of breast cancer growth and initiation by the stem cell-associated transcription factor TCF3. Cancer Res. 2012;72:5613–24.
Nguyen H, Merrill BJ, Polak L, Nikolova M, Rendl M, Shaver TM, et al. Tcf3 and Tcf4 are essential for long-term homeostasis of skin epithelia. Nat Genet. 2009;41:1068–75.
Zhang P, Sun Y, Ma L. ZEB1: at the crossroads of epithelial-mesenchymal transition, metastasis and therapy resistance. Cell Cycle. 2015;14:481–7.
Ma P, Ni K, Ke J, Zhang W, Feng Y, Mao Q. miR-448 inhibits the epithelial-mesenchymal transition in breast cancer cells by directly targeting the E-cadherin repressor ZEB1/2. Exp. Biol. Med. 2018;1535370218754848.
Sinn PL, Arias AC, Brogden KA, McCray PB Jr. Lentivirus vector can be readministered to nasal epithelia without blocking immune responses. J Virol. 2008;82:10684–92.
Bie CQ, Liu XY, Cao MR, Huang QY, Tang HJ, Wang M, et al. Lentivirus-mediated RNAi knockdown of insulin-like growth factor-1 receptor inhibits the growth and invasion of hepatocellular carcinoma via down-regulating midkine expression. Oncotarget. 2016;7:79305–18.
Xie S, Wang G, Chen G, Zhu M, Lv G. Lentivirus-mediated knockdown of P27RF-Rho inhibits hepatocellular carcinoma cell growth. Contemp Oncol. 2017;21:35–41.
Jiang L, Yan Q, Fang S, Liu M, Li Y, Yuan YF, et al. Calcium-binding protein 39 promotes hepatocellular carcinoma growth and metastasis by activating extracellular signal-regulated kinase signaling pathway. Hepatology. 2017;66:1529–45.
Seo JH, Jeong ES, Choi YK. Therapeutic effects of lentivirus-mediated shRNA targeting of cyclin D1 in human gastric cancer. BMC Cancer. 2014;14:175.
Lim HY, Sohn I, Deng S, Lee J, Jung SH, Mao M, et al. Prediction of disease-free survival in hepatocellular carcinoma by gene expression profiling. Ann Surg Oncol. 2013;20:3747–53.
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19:649–58.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–W102.
Acknowledgements
This work was supported by grants from the National Basic Research Program of China (2012CB967001), the China National Key Sci-Tech Special Project of Infectious Diseases (2018ZX10723204-006-005), the National Natural Science Foundation of China (81772554, 81472250, and 81472255), the China Postdoctoral Science Fund (2018M631030), the Hong Kong Research Grant Council General Research Fund (HKU/7668/11M, 767313), the Hong Kong Theme-based Research Scheme Fund (T12-704/16-R), and the Hong Kong Research Grant Council Collaborative Research Funds (C7027-14G and C7038-14G). Professor X.-Y.G. is Sophie YM Chan Professor in Cancer Research.
Author contributions:
L.L., Y.-L.Z., C.J., and S.F.: acquisition, analysis and interpretation of data; L.L.: drafting of the manuscript; T.-T.Z., Y.-H.Z., Y.L., and D.X.: technical and material support; X.-Y.G.: study design and supervision.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by J.P. Medema
Supplementary information
Rights and permissions
About this article
Cite this article
Li, L., Zheng, YL., Jiang, C. et al. HN1L-mediated transcriptional axis AP-2γ/METTL13/TCF3-ZEB1 drives tumor growth and metastasis in hepatocellular carcinoma. Cell Death Differ 26, 2268–2283 (2019). https://doi.org/10.1038/s41418-019-0301-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0301-1
This article is cited by
-
Methyltransferase-like proteins in cancer biology and potential therapeutic targeting
Journal of Hematology & Oncology (2023)
-
METTL13 facilitates cell growth and metastasis in gastric cancer via an eEF1A/HN1L positive feedback circuit
Journal of Cell Communication and Signaling (2023)
-
Mettl13 protects against cardiac contractile dysfunction by negatively regulating C-Cbl-mediated ubiquitination of SERCA2a in ischemic heart failure
Science China Life Sciences (2023)
-
METTLing in Stem Cell and Cancer Biology
Stem Cell Reviews and Reports (2023)
-
Exportin XPO7 acts as an oncogenic factor in prostate cancer via upregulation of TCF3
Journal of Cancer Research and Clinical Oncology (2023)