Abstract
Muscle differentiation is a crucial process controlling muscle development and homeostasis. Mitochondrial reactive oxygen species (mtROS) rapidly increase and function as critical cell signaling intermediates during the muscle differentiation. However, it has not yet been elucidated how they control myogenic signaling. Autophagy, a lysosome-mediated degradation pathway, is importantly recognized as intracellular remodeling mechanism of cellular organelles during muscle differentiation. Here, we demonstrated that the mtROS stimulated phosphatidylinositol 3 kinase/AKT/mammalian target of rapamycin (mTOR) cascade, and the activated mTORC1 subsequently induced autophagic signaling via phosphorylation of uncoordinated-51-like kinase 1 (ULK1) at serine 317 and upregulation of Atg proteins to prompt muscle differentiation. Treatment with MitoQ or rapamycin impaired both phosphorylation of ULK1 and expression of Atg proteins. Therefore, we propose a novel regulatory paradigm in which mtROS are required to initiate autophagic reconstruction of cellular organization through mTOR activation in muscle differentiation.
Similar content being viewed by others
Introduction
Muscle differentiation is a highly coordinated sequential program to generate mature muscle fibers. The muscle differentiation proceeds through cell cycle withdrawal of myoblasts and increase in the expression of muscle-specific genes, leading to fusion of myoblasts into multinucleate myofibers [1].
In muscle differentiation, phosphatidylinositol 3 kinase (PI3K)/AKT pathway is a crucial signaling cascade that is mainly stimulated by insulin-like growth factors (IGFs) [2]. A key repressor of this signaling, phosphatase and tensin homolog deleted on chromosome 10 (PTEN), is a lipid phosphatase that dephosphorylates phosphatidylinositol 3,4,5-trisphosphate (PIP3) to phosphatidylinositol-4,5-bisphosphate (PIP2), thereby controlling the activity of PI3K/AKT signaling [3]. Enzymatic PTEN activity can be altered by post-translational regulation, including phosphorylation and oxidation [4]. PTEN phosphorylation at serine 308 (S308) prevents its recruitment to the plasma membrane and subsequently decreases its catalytic activity [5]. PTEN activity is also suppressed via oxidation of the essential cysteine 124 (C124) and the formation of a disulfide with C71 by hydrogen peroxide (H2O2) [6]. The elevation of reactive oxygen species (ROS) results in PI3K/AKT signaling activation [7].
Mammalian target of rapamycin (mTOR) is a well-characterized serine/threonine kinase that participates in a wide spectrum of cellular processes [8]. mTOR has emerged as a principal regulator of muscle differentiation [9]. It forms two different complexes, mTORC1 and mTORC2, which are dynamically assembled and cross-regulated with their modulators such as Raptor, Rictor, and LST8 via inter-complex and intra-complex loops [10]. mTORC1 activates ribosomal protein kinase S6 kinase (S6K) and inhibits eukaryotic translation initiation factor 4E-binding proteins (4E-BPs) to regulate protein translation [11]. mTORC2 phosphorylates AKT, protein kinase C and serum-induced and glucocorticoid-induced protein kinases, and regulates cell survival and actin cytoskeleton reorganization [12].
Autophagy is a conserved catabolic process that controls protein and organelle degradation [11]. Generally, active mTORC1 interacts with the uncoordinated-51-like kinase 1 (ULK1), and then directly phosphorylates autophagy-related protein 13 (Atg13) and ULK1 subunits at serine 258 and serine 757 (S757), respectively, to repress ULK1 activity under nutrient-enriched condition. Subsequently, mTORC1 seizes the ULK1/Atg13/focal adhesion kinase family-interacting protein of 200 kDa (FIP200) complex to prevent autophagy induction [13]. Conversely, under nutrient-deprived conditions, mTORC1 is dissociated from the ULK1 complex, and ULK1 is then phosphorylated at serine 317 (S317) and serine 777 (S777) by AMP-activated protein kinase (AMPK) for autophagy initiation [14]. Sequentially, activation of two ubiquitin-like conjugation systems contributes to autophagosomal elongation and completion to recognize and sequester materials destined for degradation, which is controlled by Atg proteins to fully encapsulate the cytosolic cargo [15]. The conjugation of Atg12–Atg5 results in the formation of a multimeric complex. LC3-I is conjugated to phosphatidylethanolamine in a reaction requiring Atg7 and Atg3. LC3-II, which is the lipidated form of LC3-I, is bound to both sides of the phagophore membrane. The autophagosome fuses with the lysosome into the autophagolysosome, and the cargo-containing membrane compartment is then lysed and degraded [16]. However, it has not been explored whether mTOR is involved in the autophagic process to reconstruct intracellular organization during muscle differentiation process, which occurs under nutrient-enriched conditions.
Mitochondria are primary energy producers to generate ATPs via oxidative phosphorylation [17]. Mitochondrial ROS (mtROS) are formed as byproducts of the mitochondrial electron transport chain during the generation of ATP or by an imbalance in cellular oxidant/antioxidant systems [18]. Previous studies reported that the mitochondria-derived H2O2 is crucial as a signaling molecule to induce muscle, adipocyte, keratinocyte, dendritic cell, and stem cell differentiation [19,20,21,22,23]. However, it is still unclear how mtROS practically control cellular differentiation in myogenic signal transduction.
In this study, we provide evidences that mtROS induce PTEN oxidation to stimulate PI3K/AKT/mTOR signaling pathway and thus promote autophagy, which is required for muscle differentiation and regeneration.
Results
Mitochondrial ROS increase during muscle differentiation
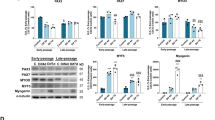
To examine the roles of mtROS in the process of muscle differentiation, we first observed that myotube formation and myosin heavy chain (MHC) expression levels were gradually increased up to 5 days after induction of differentiation in C2C12 cells. Superoxide dismutase 2 (SOD2) expression levels were also elevated without changes in SOD1 expression (Fig. 1a). MitoQ, a mitochondria-targeted antioxidant, significantly suppressed muscle differentiation as well as SOD2 expression in a concentration-dependent manner (Fig. 1b). Mitochondria-specific superoxide fluorescent probe (MitoSOX) fluorescence intensity, indicating mitochondrial superoxide (mtO2•−) content, was also dramatically increased during differentiation, compared to that of proliferation, whereas MitoQ reduced the MitoSOX intensity (Fig. 1c).
Mitochondrial ROS are increased during muscle differentiation. a Morphological changes were observed during the progression of differentiation in DM for 5 days. Expression levels of MHC, SOD1, and SOD2 were analyzed by Western blot using specific antibodies. b Cells were treated with MitoQ at 125, 250, and 500 nM for 5 days. The protein levels of MHC, SOD1, and SOD2 on the indicated days were assessed by Western blot analysis. c Mitochondrial superoxide (mtO2•−) was stained with 1 μM MitoSOX and observed by confocal microscope. d Morphological changes were observed after treatment of 0.25 mM d-galactose with or without 0.015 U/ml galactose oxidase in DM for 5 days. Under these conditions, cells were treated with 250 nM MitoQ from 2 days after induction of differentiation and further incubated for 3 days. The expression levels of MHC and SOD2 were assessed by Western blot analysis. β-Actin was used as a loading control. e Under the same condition as in c, intracellular ROS levels were detected with CM-H2DCFDA (DCF-DA) using a flow cytometer. Cells were incubated with 5 μM DCF-DA at 37 °C for 30 min. The mean DCF-DA fluorescence intensity was measured with excitation at 488 nm and emission at 525 nm. f mtO2•− levels were measured with MitoSOX using a flow cytometer. Cells were incubated with 1 μM MitoSOX at 37 °C for 30 min. The mean MitoSOX fluorescence intensity was measured with excitation at 510 nm and emission at 580 nm. g Cells were stained with 2 μM Amplex Red, and the optical density at 560 nm for intracellular H2O2 was evaluated using a microplate reader. The data shown represent the mean ± SE of three independent experiments. * p < 0.05, compared to cells in PM. # p < 0.05, compared to cells in DM. Densitometry analyses were assessed using the ImageLab software. MQ MitoQ, GA d-galactose, GO galactose oxidase
H2O2 is the major form of ROS that activates redox-dependent signaling in the cytosol [24]. Thus, we hypothesized that if H2O2 was the major form of ROS required for muscle differentiation, administering exogenous peroxide could rescue the muscle differentiation suppressed by MitoQ. It is well established that galactose oxidase (GO) with d-galactose (GA) produces H2O2 in culture medium, which rapidly enters the cytosol [25]. Thus, to further examine whether mtROS-derived cytosolic H2O2 stimulated muscle differentiation, we employed this exogenous pro-oxidant generation system to muscle differentiation [20]. Treatment with GA alone did not affect muscle differentiation. However, exogenous H2O2 (exH2O2) by co-treatment of GA and GO in differentiation medium (DM) enhanced muscle differentiation, compared to non-treatment in DM. Importantly, the exH2O2 also restored the cellular differentiation even in the presence of MitoQ (Fig. 1d).
Under the same experimental conditions, we measured intracellular ROS (cROS) and mtO2•− using diacetate form of DCFH (DCF-DA) and MitoSOX, respectively, via flow cytometry [26]. The cROS and mtO2•− levels were significantly increased in DM, compared to proliferation medium (PM). Moreover, MitoQ suppressed these ROS productions. The exH2O2 in DM further stimulated cROS, not mtROS, compared to non-treatment in DM. The exH2O2 generation also restored cROS level, but not mtROS generation in the presence of MitoQ (Fig. 1e, f). However, C2C12 myoblasts started to be fused after 1 day of differentiation induction. Since the flow cytometry method could not be used to detect intracellular H2O2 (cH2O2) in fused myotubes due to their cellular sizes, we used Amplex Red reagent to redeem the limitation of flow cytometry analysis in measuring cH2O2 during muscle differentiation. cH2O2 level was significantly increased for 5 days of differentiation (Fig. S1). MitoQ also suppressed the cH2O2 level. The exH2O2 in DM further stimulated cH2O2, compared to non-treatment in DM. The exH2O2 also restored the cH2O2 level in the presence of MitoQ (Fig. 1f).
Mitochondria-derived H2O2 regulates PTEN oxidation and activity during muscle differentiation
Previous studies have demonstrated that PTEN is inactivated via oxidation by H2O2, and its inactivation enhances PI3K/AKT signaling [27, 28]. Thus, we next investigated whether PTEN was oxidized by H2O2 dismutated from mtO2•− by SOD2 during muscle differentiation. Indeed, oxidized PTEN (lower band) was gradually increased without any changes in phosphorylated (at S308) or total PTEN levels during muscle differentiation (Fig. 2a). Consistent with these findings, PTEN activity was attenuated (Fig. 2b). MitoQ decreased the PTEN oxidization with concomitant restoration of PTEN activity in a concentration-dependent manner (Fig. 2c, d). As expected, exH2O2 elevated PTEN oxidation levels and reduced its activity even in the presence of MitoQ. Treatment with GA alone did not affect PTEN oxidation and activity (Fig. 2e, f).
Mitochondrial H2O2 regulates PTEN oxidation and activity during muscle differentiation. a C2C12 cells were incubated in DM for 5 days and the medium was changed every day. Proteins were resolved by non-reducing SDS-PAGE gel. The reduced (upper band) and oxidized (lower band) forms of PTEN protein, phospho-PTEN (p-PTEN) and PTEN, were assessed by Western blot analysis. b PTEN activity was evaluated via detecting PIP3-hydrolyzing level by PTEN, and expressed by percentage, compared to PM. *p < 0.05, compared to cells in PM. c Cells were treated with 125, 250, and 500 nM MitoQ for 5 days, and the reduced and oxidized PTEN proteins were assessed by Western blot analysis. d Under the same condition as in c, the PTEN activity was determined at 450 nm and expressed by percentage. *p < 0.05, compared to cells in PM. #p < 0.05, compared to cells in DM. e Cells were treated with 0.25 mM d-galactose with or without 0.015 U/ml galactose oxidase in DM for 5 days. Under these conditions, cells were treated with 250 nM MitoQ from 2 days after induction of differentiation and further incubated for 3 days. f Under the same conditions as in e, the PTEN activity was determined at 450 nm and expressed by percentage. *p < 0.05, compared to cells in PM. #p < 0.05, compared to cells treated with d-galactose in DM. ¶p < 0.05, compared to cells treated with d-galactose and galactose oxidase in DM. β-Actin was used as a loading control. In graphs, the data shown represent the mean ± SE of three independent experiments. MQ MitoQ, GA d-galactose, GO galactose oxidase
PI3K/AKT/mTOR pathway is activated by mitochondrial ROS during muscle differentiation
The PI3K/AKT/mTOR signaling pathway has an important role in muscle differentiation [29]. Thus, we first tested whether this signaling cascade was activated during muscle differentiation. The phosphorylation of PI3K downstream signaling molecules, such as phosphoinositide-dependent kinase 1 (PDK1), AKT, p70S6K, and 4E-BP1, was gradually increased in differentiating myoblasts up to 5 days. mTOR phosphorylations were also elevated at the sites of S2448 and S2481 for mTORC1 and mTORC2, respectively. However, the mitochondria-targeted antioxidants, MitoQ or MitoTempol, markedly diminished the phosphorylation levels (Fig. 3a). The exH2O2 further stimulated the phosphorylation of PI3K downstream molecules in DM, and recovered the phosphorylation levels even in the presence of MitoQ (Fig. 3b).
PI3K/AKT/mTOR pathway activation by mitochondrial ROS during muscle differentiation. a C2C12 cells were cultured in DM for 5 days (left panel). Cells were treated with 250 nM MitoQ or 100 μM MitoTempol for 5 days (right panel). b Cells were treated with 0.25 mM d-galactose with or without 0.015 U/ml galactose oxidase in DM for 5 days. Under these conditions, cells were treated with 250 nM MitoQ from 2 days after induction of differentiation and further incubated for 3 days. The levels of phospho-PDK1 at serine 241 (p-PDK1), PDK1, phospho-AKT at threonine 308 (T308) and serine 473 (S473), AKT, phospho-mTOR at serine 2448 (S2448) and serine 2481 (S2481), phospho-p70S6K at serine 371 (S371), threonine 389 (T389) and threonine 421/serine424 (T421/S424), p70S6K, phospho-4E-BP1 at threonine 37 and 46 (p-4E-BP1), 4E-BP1, and MHC were assessed by Western blot analysis. β-Actin was used as a loading control. MQ MitoQ, MT MitoTempol, GA d-galactose, GO galactose oxidase
Inhibition of PTEN enhances muscle differentiation
To further investigate whether inactivation of PTEN stimulated muscle differentiation, we employed specific small interference RNA (siRNA). PTEN knockdown led to enhanced AKT phosphorylation in PM (Fig. 4a). Moreover, PTEN-knockdown cells exhibited further phosphorylation of AKT and induction of muscle differentiation, compared to scramble siRNA-transfected cells (Fig. 4b). Next, we performed the PTEN recovery experiment in which PTEN was knocked down using its 3′-untranslated region (UTR) targeting siRNA, and then re-expressed by wild-type (WT) or mutant (C124S, phosphatase-inactive form) PTEN (Fig S2a). The PTEN WT overexpression impaired muscle differentiation in scramble-transfected myoblasts of DM, whereas the muscle differentiation was rescued via the countervailing of PTEN WT expression in PTEN-knockdown cells. However, mutant PTEN (PTEN C124S) increased induction of muscle differentiation in scramble-transfected myoblasts of DM, and further elevated the muscle differentiation in PTEN-knockdown cells (Fig. S2b). Consistent with PTEN knockdown, a selective PTEN inhibitor, SF1670, further stimulated AKT phosphorylation and MHC expression in a concentration-dependent manner (Fig. 4c).
Silencing of PTEN enhances muscle differentiation and AKT phosphorylation. a C2C12 cells were transfected with 100, 200, or 400 pM PTEN siRNA (si-PTEN) for 48P, and then PTEN expression and AKT phosphorylations and expression were evaluated by Western blot analysis. b After transfection with 100, 200, and or 400 nM PTEN siRNA, the cells were further incubated in DM for 5 days, and morphological changes were then observed. MHC expression, AKT activities, and expression were assessed by Western blot analysis. c Cells were treated with 30 or 60 nM SF1670 in DM for 5 days. β-Actin was used as a loading control. SC scramble siRNA, si-PTEN PTEN siRNA
Mitochondrial ROS are required for mTORC1 activation to induce myogenic autophagy
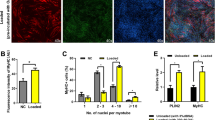
It is well established that mTORC1 stimulates muscle differentiation [30], and the differentiation process requires autophagy to remodel cellular organization for muscle fiber formation [31]. Autophagy also protects against apoptosis during myoblast differentiation [32]. Consistent with this, we found that MitoQ and rapamycin suppressed p70S6K phosphorylation and MHC expression (Fig. 5a). The exH2O2 did not rescue the p70S6K phosphorylations and MHC expression in the presence of rapamycin (Fig. 5b). Under the same experimental conditions, cROS, mtO2•−, and cH2O2 levels were significantly increased in DM, compared to PM. Intriguingly, rapamycin treatment did not influence all of the ROS levels under condition of non-treatment or exH2O2 generation in DM (Fig. 5c–e). Thus, we further investigated the role of mTOR in myogenic autophagy. Autophagosomal or autophagolysosomal LC3 puncta (green and yellow dots) formation was remarkably increased (Fig. S3a), and pro-LC3 messenger RNA (mRNA) level was also continuously upregulated during muscle differentiation (Fig. S3b). But, the autophagy flux was completely attenuated by MitoQ or rapamycin treatment. 3-Methyladenine (3-MA) or wortmannin also inhibits myogenic autophagy by blocking autophagosome formation via inhibition of class III PI3K or pan-PI3K. Furthermore, chloroquine and bafilomycin A1, which suppress vacuolar-type H+-ATPase and autophagolysosome formation, respectively [33], substantially increased the formation of LC3 puncta (Fig. 5f, g). Additionally, chloroquine, bafilomycin A1, or leupeptin (a lysosomal degradation inhibitor) in DM remarkably increased accumulation of LC3 lipidation, diminishing muscle differentiation (Fig. S4a and b).
Mitochondrial ROS-induced mTOR activation is essential for myogenic autophagy. a C2C12 cells were treated with 250 nM MitoQ or 500 pM rapamycin in DM for 5 days. The cells were harvested at 1, 3, and 5 days of differentiation. The indicated proteins were assessed by Western blot analysis. b Cells were treated with 0.25 mM d-galactose with or without 0.015 U/ml galactose oxidase in DM for 5 days. Under these conditions, cells were treated with 500 pM rapamycin in DM for 5 days. c Under the same condition as in b, intracellular ROS levels were detected with CM-H2DCFDA (DCF-DA) using a flow cytometer. Cells were incubated with 5 μM DCF-DA at 37 °C for 30 min. The mean DCF-DA fluorescence intensity was measured with excitation at 488 nm and emission at 525 nm. d mtO2•− levels were measured with MitoSOX using a flow cytometer. Cells were incubated with 1 μM MitoSOX at 37 °C for 30 min. The mean MitoSOX fluorescence intensity was measured with excitation 510 nm and emission 580 nm. e Cells were stained with 2 μM Amplex Red, and the optical density at 560 nm for intracellular H2O2 was evaluated using a microplate reader. f Cells were treated with 250 nM MitoQ, 500 pM rapamycin, 2 mM 3-MA, 20 μM wortmannin, 0.5 nM bafilomycin A1, and 2.5 μM chloroquine were treated in DM for 5 days. The fluorescence was visualized by confocal microscopy. LysoTrackerTM DND-99 was used for lysosome staining. DAPI was used for nuclear staining. g Quantification of autophagosome flux was represented. Autophagic flux were calculated by counting autophagosome (green) and autopahgolysosome (merged form with LysoTracker, yellow) dots in the cells, and expressed as a percentage of cell with green or yellow dots (minimum five dots per cell). *p < 0.05, compared to cells in PM. #p < 0.05, compared to cells in DM. h Cells were transfected with 400 nM mTOR, Raptor, or Rictor siRNA for 48f, and then protein levels were assessed by Western blot analysis. i Fusion index was measured at 5 days after induction of differentiation. *p < 0.05, compared to cells in PM. j After 48-h transfection with si-mTOR, si-Raptor, or si-Rictor, the cells were treated with 0.25 mM d-galactose with or without 0.015 U/ml galactose oxidase in DM for 5 days. Under these conditions, cells were treated with 250 nM MitoQ from 2 days after induction of differentiation and further incubated for 3 days. MHC expression, p70S6K activity and expression, and LC3-II conversion were assessed by Western blot analysis. β-Actin was used as a loading control. The data shown represent the mean ± SE of three independent experiments. MQ MitoQ, Rapa rapamycin, Wort wortmannin, CQ chloroquine, Baf bafilomycin A1, GA d-galactose, GO galactose oxidase, SC scramble siRNA, siT mTOR siRNA, siRa Raptor siRNA, siRi Rictor siRNA
In Fig. 3a, the mTOR phosphorylations at both sites of S2448 and S2481 were increased during muscle differentiation. Thus, to understand which mTOR complex (mTORC1 or mTORC2) is predominantly involved in muscle differentiation, we employed siRNA-mediated mTOR, Raptor, or Rictor knockdown (Fig. 5h), or Raptor knockdown impaired myotube formation, whereas Rictor knockdown relatively less inhibited the muscle differentiation (Fig. 5i). Moreover, the mTOR or Raptor inhibition did not restore MHC expression, p70S6K phosphorylation, or LC3-II conversion even after exH2O2 generation, although the exH2O2 rescued all of these protein regulations under MitoQ-treated condition in scramble siRNA-transfected cells. In case of Rictor knockdown, it less attenuated the MHC expression, p70S6K phosphorylations, or LC3-II conversion even after exH2O2 generation than mTOR or Raptor knockdown (Fig. 5j). The MHC expressions were quantified by densitometry assay (Fig. S5). Additionally, to further elucidate whether mTORC1 activation is essential in the muscle differentiation, we performed the recovery experiment of Rictor in which Rictor was knocked down using its 3′-UTR targeting siRNA, and then mTOR or Raptor was overexpressed, or Rictor was re-expressed (Fig. S6a). The mTOR, Raptor overexpression slightly enhanced muscle differentiation in scramble-transfected myoblasts of DM, or considerably rescued the muscle differentiation in Rictor-knockdown myoblasts of DM. Similarly to Fig. 5i, j, Rictor knockdown itself did not show the severe impairment in muscle differentiation. Reasonably, the muscle differentiation was completely rescued via Rictor re-expression in Rictor-knockdown cells (Fig. S6b). Collectively, these data indicate that mTORC1 activation is more responsible for mtROS-induced myogenic autophagy than mTORC2.
Phosphorylations of ULK1 and expressions of Atg proteins are essential for autophagy initiation during muscle differentiation
Recently, it was suggested that autophagy is induced during muscle differentiation, which is accompanied with mTOR activation [34]. Therefore, it is important to understand whether mTOR stimulates the pro-autophagic phosphorylation of ULK1 at S317 and expression of Atg proteins to promote the myogenic autophagy. Interestingly, ULK1 phosphorylations were increased at S317 and anti-autophagic S637 and S757 up to 5 days of differentiation without activations of AMPK (phosphorylation at T172) and acetyl-CoA carboxylase (ACC, phosphorylation at S79). Moreover, the expression levels of Atg proteins and conjugation of Atg12–Atg5 were gradually increased during the differentiation process up to 5 days. Furthermore, conversion of LC3 was enhanced during muscle differentiation (Fig. 6a). MitoQ and rapamycin attenuated the phosphorylations of ULK1, blocked the expression of Atg proteins, and suppressed the conjugation of Atg12–Atg5 and conversion of LC3 in a concentration-dependent manner (Fig. 6b). The exH2O2 generation restored the expression of autophagy and muscle differentiation molecules even in the presence of MitoQ, but it did not affect those in the presence of rapamycin (Fig. 6c). Additionally, ULK1, Atg5, Atg7, or Atg12 knockdown attenuated the muscle differentiation (Fig. S7a and b).
Mitochondrial ROS-induced mTOR activation phosphorylates ULK1 at serine 317 for autophagy initiation and stimulates expression of autophagy-related proteins during muscle differentiation. a C2C12 cells were incubated in differentiation medium up to 5 days and were harvested every day. b Cells were incubated with 125, 250, or 500 nM MitoQ (MQ) or 125, 250, or 500 pM rapamycin for 5 days. MitoQ-containing or rapamycin-containing media were changed every day. c Cells were treated with 0.25 mM d-galactose with or without 0.015 U/ml galactose oxidase in DM for 5 days. Under these conditions, cells were treated with 250 nM MitoQ from 2 days after induction of differentiation and further incubated for 3 days. Cells were treated with 500 pM rapamycin in DM for 5 days. The levels of phospho-ULK1 at serine 317 (S317), serine 637 (S637) and serine 757 (S757), ULK1, phosphor-AMPK (p-AMPK (T172)), AMPK, phospho-ACC (p-ACC (S79)), ACC, Atg3, Atg5, Atg7, Atg12, Atg12–Atg5, LC3 (LC3-I, upper band; LC3-II, lower band), and MHC were assessed by Western blot analysis. β-Actin was used as a loading control. d Co-immunoprecipitation of mTOR (S2448) with phospho-ULK1 (S757 or S317) in cells treated without or with 250 nM MQ or 500 pM rapamycin. Expressions of mTOR and ULK1 were assessed by Western blot analysis. β-Actin was used as a loading control. e ULK1 phosphorylation. L6 or H9c2 cells were incubated in differentiation medium up to 5 days and were harvested every day. f Co-immunoprecipitation of mTOR (S2448) with phospho-ULK1 (S757 or S317) in L6 or H9c2 cells. g mTOR and ULK1 protein expressions in C2C12 and L6 cell lysates or mouse and rat tissue lysates were assessed by Western blot analysis. β-Actin was used as a loading control. C2 C2C12, B brain, H heart, Iʟ large intestine, Is small intestine, K kidney, Lu lung, Li liver, SM skeletal muscle, Sp spleen, and St stomach. h Cells were incubated in proliferation medium with or without 25 mM glucose for 24 g. Western blot analysis was conducted by specific antibodies using cell lysates. i Immunoprecipitation of phospho-ULK1 (S757) in C2C12, L6, or H9c2 cells. Western blot analysis was conducted by specific antibodies using cell lysates. β-Actin was used as a loading control. MQ MitoQ, Rapa or RP rapamycin, GA d-galactose, GO galactose oxidase
Then, we employed the ULK1 recovery experiment in which ULK1 was knocked down using its 3′-UTR targeting siRNA, and then re-expressed by WT ULK1. The ULK1 overexpression did not influence muscle differentiation in scramble-transfected myoblasts of DM, whereas the muscle differentiation was rescued via the ULK1 re-expression in ULK1-knockdown cells (Fig. S8a and b). Also, the ULK1 phosphorylations at S317, S637, and S757 were strictly inhibited by mTOR knockdown (Fig. S9). Next, we performed co-immunoprecipitation to further clarify whether active mTOR phosphorylated ULK1 at S317 to induce autophagy. The result showed that the active mTOR interacted with phosphorylated ULK1 at both sites of S317 and S757 in muscle differentiation. MitoQ or rapamycin disrupted these protein interactions (Fig. 6d). These results were validated via the same experiments using two different myoblast cell lines, L6 and H9c2. Similar to C2C12 cells, ULK1 phosphorylation was increased at both S317 and S757 up to 5 days of differentiation in L6 and H9c2 (Fig. 6e), and the active mTOR interacted with phosphorylated ULK1 at the sites of S317, S637, and S757 during muscle differentiation, whereas MitoQ or rapamycin disrupted these protein interactions (Fig. 6f).
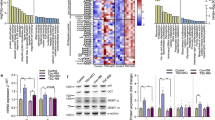
Since our results were strikingly different from previous well-established results [35], we tested protein alterations of mTOR and ULK1 in myoblasts. Western blot assays using lysates from C2C12 or L6 myoblasts and corresponding animal tissues showed the same molecular weights (Fig. 6g). The sequence analysis of mRNAs also showed no differences in both proteins, although there were 14 and 3 wobble mutations in mTOR and ULK1 (yellow highlighted), respectively, without any changes in amino acid sequences (Supplementary Information S1).
To endorse our finding that the increase in ULK1 phosphorylation at the sites of S317, S637, and S757 by active mTOR was specific to the muscle differentiation, we examined the phosphorylation of ULK1 and regulation of Atg proteins under glucose-deprived condition. In contrast to muscle differentiation, ULK1 phosphorylation was rapidly decreased at S757 but gradually elevated at S317 for 24p after glucose deprivation. Autophagy markers were also promptly increased (Fig. 6h). Finally, we explored whether ULK1 was dual-phosphorylated in the muscle differentiation process because ULK1 phosphorylation was increased at the sites of S317, S637, and S757. ULK1 proteins phosphorylated at S757 were pulled down by immunoprecipitation, and then phospho-ULK1 (S317) was assessed by Western blot analysis. ULK1 was indeed dual-phosphorylated during muscle differentiation, but not in proliferation or glucose-deprived condition (Fig. 6i).
PTEN oxidation and mTOR activation are required for muscle regeneration
Previous studies have shown that PTEN inhibition and ROS are required for muscle regeneration and the wound healing response [36, 37]. Thus, we injected cardiotoxin (CTX), which induced skeletal muscle necrosis [38], into the left tibialis anterior (TA) muscle. As shown in Fig. 7a, muscle damage (white wounds) was increased in the left TA muscle up to 4 days after CTX injection, and then the recovery was gradually observed for the next 4 days. The phosphate-buffered saline (PBS)-injected right TA muscle did not show any changes. PTEN oxidation levels were gradually increased without any changes in total and phosphorylated PTEN level. MHC and SOD2 expression levels were also increased during muscle regeneration. (Fig. 7b). AKT/mTOR signaling molecules and their downstream molecules, including p70S6K and 4E-BP1, were significantly modulated after CTX injection (Fig. 7c). ULK1 phosphorylations at S317, S637, and S757, as well as other autophagy markers were upregulated during TA muscle regeneration (Fig. 7d).
PTEN oxidation and mTOR activation are induced during muscle regeneration in mouse model. a Mouse left tibialis anterior (TA) muscles were injected with 50 μl (10 mM) cardiotoxin (CTX). Mice were sacrificed at indicated days after the injection (N = 5). b Mouse TA muscle tissues were homogenized and assessed by Western blot analysis. Oxidized and reduced PTEN proteins were resolved by non-reducing SDS-PAGE. The levels of phospho-PTEN (p-PTEN), PTEN, SOD2, and MHC proteins were assessed by Western blot analysis. c The levels of phospho-AKT at threonine 308 (T308) and serine 473 (S473), AKT, phospho-mTOR at serine 2448 (S2448) and serine 2481 (S2481), phospho-p70S6K at serine 371 (S371), threonine 389 (T389) and threonine 421/serine424 (T421/S424), p70S6K, phospho-4E-BP1 at threonine 37 and 46 (p-4E-BP1), and 4E-BP1 were determined by Western blot analysis. d The levels of phospho-ULK1 at serine 317 (S317), serine 637 (S637) and serine 757 (S757), ULK1, Atg3, Atg5, Atg7, Atg12, Atg12–Atg5, LC3 (LC3-I, upper band; LC3-II, lower band) were assessed by Western blot analysis. β-Actin was used as a loading control
Discussion
Here, we reported for the first time that the dramatic increase in mtROS during muscle differentiation stimulated the PI3K/AKT/mTOR pathway through PTEN oxidative inactivation, and that mTOR activation led to myogenic autophagy via phosphorylation of ULK1 at S317 and expressions of Atg proteins.
Intracellular ROS have been recognized as essential molecules for myogenesis [39], and the preferential mitochondrial origin of ROS has been further substantiated [40]. We previously reported that a gradual increase in mitochondria-derived H2O2 is required to induce muscle differentiation [19]. It was authenticated that mitochondria are the most important ROS source in muscle differentiation process [41]. In the current study, we firmly validated that mitochondrial superoxide stimulated muscle differentiation after dismutation to H2O2. Moreover, MitoQ impaired muscle differentiation. We further showed that exogenously generated intracellular H2O2 rescued muscle differentiation in the presence of MitoQ. These data demonstrated that intracellular H2O2, which was dismutated from mtO2•− by MnSOD, was a key signaling molecule in muscle differentiation.
PTEN activity can be post-transcriptionally inactivated via its reversible oxidative inactivation, resulting in activation of the PI3K/AKT cascade [42]. We found that the gradual elevation of mtROS provoked PTEN oxidative inactivation, by which the PI3K/AKT signaling pathway was gradually activated for several days until muscle differentiation was terminated. In addition, MitoQ completely prevented PTEN oxidative inactivation. In agreement with previous studies [43, 44], the chemical inhibition or knockdown of PTEN facilitated myoblast differentiation. Collectively, our findings supported that mtROS-imposed PTEN oxidation was essential for stimulating PI3K/AKT/mTOR signaling and thus promoting muscle differentiation.
mTOR is a key myogenic regulator involved in the development and physiology. It was clearly reported that rapamycin treatment or mTOR knockdown blocks C2C12 myoblast differentiation in vitro and in vivo [45,46,47]. Consistently, we found that both mTORC1 and mTORC2 complex were activated, and inhibition of mTORC1 by rapamycin or RNA interference diminished muscle differentiation. However, Rictor knockdown did not exhibit severe impairment of muscle differentiation in our experiments, which is contrary to the previous studies that Rictor is essential for muscle differentiation [48, 49]. Thus, we argue that the mTORC1 is predominant in the process of muscle differentiation, whereas mTORC2 plays a minor role in myogenesis.
Autophagy is known to facilitate cellular development [50]. The autophagic complex process begins at the phagophore assembly site (PAS) where proteins of the ULK1 complex assemble to initiate autophagosome formation [13]. Under nutrient-enriched condition, mTOR directly phosphorylates ULK1 at S757, and thereby prevents ULK1 interaction with AMPK. In contrast, activated AMPK under nutrient-deprived condition phosphorylates ULK1 at S317 and S777 for initiating autophagy [14]. Consequently, much of our current understanding of mTOR is limited in the context of inhibitory effects on autophagy in nutrient-enriched condition. However, myoblast differentiation requires autophagy to facilitate the elimination of pre-existing structures and promote cellular remodeling [50], indicating that the basic autophagy mechanism underlying muscle differentiation is completely different. In our results, mTOR activity was elevated to amplify autophagic processes during muscle differentiation. Consistently, mTOR inhibition attenuated autophagy induction via suppressing pro-autophagic or anti-autophagic phosphorylations of ULK1 and expressions of Atg proteins. Furthermore, MitoQ-induced mTOR inactivation impaired the phosphorylations of ULK1 and subsequent induction of Atg proteins in muscle differentiation, indicating that mTOR functioned in not only initiation but also elongation and completion of autophagic processes. Intriguingly, we also found that AMPK, a well-known autophagy inducer, was not activated during muscle differentiation. Thus, we could exclude the possibility that AMPK phosphorylated ULK1 at S317 during muscle differentiation. Additionally, we showed that active mTOR (S2448) interacted with phosphorylated ULK1 at both sites of S317 and S757 during muscle differentiation. The same results were also observed in other myoblasts. From these findings, we concluded that active mTOR could simultaneously phosphorylate ULK1 at pro-autophagic and anti-autophagic sites during muscle differentiation, despite the fact that these phosphorylations lie on the antagonistic relation. This simultaneous phosphorylations of ULK1 at pro-autophagic and anti-autophagic sites have never been reported before and was not due to protein alterations in mTOR and ULK1 in C2C12 myoblasts, since both proteins were exactly same as the corresponding proteins in other tissues. More interestingly, phosphorylations of ULK1 at the both sites of S637 and S757 could not block autophagy when ULK1 was phosphorylated at S317, indicating that its pro-autophagic phosphorylation was predominant during muscle differentiation. Accordingly, it is possible that activated mTOR induces autophagic reorganization of cellular components, leading to subsequent achievement of differentiated muscle fibers. In this context, our current study provides insights into new understanding that mTOR is inseparable from the phosphorylation of ULK1 and expression of Atg proteins, irrespective of AMPK activation in muscle differentiation. At present, another co-regulator in myogenic autophagy remains to be sought out. It is therefore valuable to further study how mTOR induces ULK1 phosphorylations to initiate autophagy during myogenesis.
In summary, we demonstrated for the first time that increased mitochondrial ROS were strongly involved in PTEN oxidative inactivation to enhance PI3K/AKT/mTOR signaling activity. We also showed that mTOR activity induced myogenesis-specific autophagy by phosphorylating ULK1 at S317 without involvement of AMPK during muscle differentiation.
Materials and methods
Cell culture
C2C12 and L6 cells were maintained in Dulbeco modified Eagle’s medium (DMEM) (Gibco) supplemented with 10% (v/v) fetal bovine serum, 100 U/ml penicillin, and 100 μg/ml streptomycin (PM). H9c2 cells were cultured in Dulbecco's modified Eagle's medium and Ham’s F12 medium (DMEM/F12) (Gibco), supplemented with 10% bovine serum, 100 U/ml penicillin, and 100 mg/ml streptomycin (PM). C2C12 and L6 cells were induced to differentiate by replacing them in DMEM containing 2% (v/v) horse serum, 100 U/ml penicillin, and 100 μg/ml streptomycin (DM). H9c2 cells were induced to differentiate by replacing them in DMEM/F12 containing 2 % (v/v) horse serum, 100 U/ml penicillin, and 100 μg/ml streptomycin (DM). Full differentiation was achieved after 5 days in DM.
Reagents and antibodies
MitoQ was acquired from Dr. Micheal P Murpy (University of Cambridge). MitoTempol, chloroquine, bafilomycin A1, 3-MA, wortmannin, rapamycin, d-galactose, and galactose oxidase were purchased from Sigma-Aldrich. Specific antibodies for MHC and β-Actin were purchased from Santa Cruz Biotechnology. Anti-SOD1 and SOD2 antibodies were purchased from Abfrontier. Specific antibodies for phospho-PTEN (S308/T381/T383), PTEN, phospho-PDK1 (S241), PDK1, phospho-AKT (T308 and S473), AKT, phospho-mTOR (S2448 and S2481), mTOR, Raptor, Rictor, phospho-p70S6K (S371, T389 and T421/S424), p70S6K, phospho-4E-BP1 (T37/46), 4E-BP1, phospho-ULK1 (S317, S637 and S757), ULK1, phosphor-AMPK (T172), AMPK, phosphop-ACC (S79), ACC, Atg3, Atg5, Atg7, Atg12, Atg12-5, and LC3 were purchased from Cell Signaling.
Transient transfection
pcDNA3.0 containing PTEN WT or mutant PTEN (C124S) Myc-tagged constructs, pcDNA3.0 containing mTOR HA-tagged constructs, and pRK5 containing Raptor or Rictor Myc-tagged constructs, pcDNA3.0 containing ULK1 HA-tagged constructs, were transfected into C2C12 cells by using TransIT®-2020 Transfection Reagent (Mirus Corp.), according to the manufacturer’s instructions. The efficiency of transfection was monitored by change of each protein expression level using Western blot analysis. Empty plasmid vectors were used as a control.
Small interference RNA
siRNA for PTEN was acquired from Cell Signaling. siRNAs for mTOR and Raptor were obtained from Sigma-Aldrich. siRNAs for Rictor, ULK1, Atg5, Atg7, and Atg12 were purchased from Bioneer. The siRNAs (400 nM/60 mm dish) were transfected into C2C12 cells using TransIT®-2020 Transfection Reagent (Mirus Corp.), according to the manufacturer’s instructions. The efficiency of knockdown were monitored by change of each protein expression level using Western blot analysis. A non-silencing control siRNA from Cell Signaling was used as a control.
Flow cytometry analysis
Total intracellular ROS levels were measured with CM-H2DCFDA (DCF-DA) using a flow cytometer (Beckman Coulter, Inc.). C2C12 cells were incubated with 5 μM DCF-DA at 37 °C for 30 min. The mean DCF-DA fluorescence intensity was measured with excitation at 488 nm and emission at 525 nm. Mitochondrial superoxide (mtO2•−) levels were measured with MitoSOX using a flow cytometer (Beckman Coulter, Inc.). C2C12 cells were incubated with 1 μM MitoSOX at 37 °C for 30 min. The mean MitoSOX fluorescence intensity was measured with excitation at 510 nm and emission at 580 nm.
Intracellular H2O2 measurement
To quantify intracellular H2O2 level, cells were lysed with lysis buffer (120 mM KCl, 3 mM HEPES free acid, pH 7.2, 1 mM EGTA, and 0.3 % bovine serum albumin (BSA)). The lysates were added to a microplate well and pre-warmed at 37 °C for 10 min. Then, the reaction was started by adding 30 μg (50 μl) of cell lysate resuspended in 50 μl of reaction buffer containing 100 μM Amplex Red and 0.01 U/ml horseradish peroxidase. After 30 min, the absorbance of the reaction mixtures was measured at 560 nm using a microplate reader (Bio-Rad). A standard curve was prepared by the addition of known amounts of H2O2.
Measurement of PTEN redox state and activity
Cells were harvested, washed twice with PBS, and resuspended in cell lysis buffer (1% sodium dodecyl sulfate (SDS), 10% glycerol, 32 mM Tris-Cl, pH 6.8). Protein samples were subjected to SDS-polyacrylamide gel electrophoresis (PAGE) under non-reducing conditions. The separated proteins were then transferred onto polyvinylidene difluoride membranes and subjected to Western blotting with an anti-PTEN antibody. The samples for PTEN phosphatase activity assay were prepared according to the manufacturer’s protocol (Echelon Bioscience).
Fusion assay (fusion index)
Cells were seeded in 60 mm dishes, cultivated in DM for 5 days, and stained with Giemsa staining. The fusion index (percentage of nuclei within myotubes and mononucleated cells) was determined using a microscope (Nikon). The total number of nuclei and myotubes was counted in five random fields (100 total nuclei) per each condition. A myotube was defined by the presence of at least three nuclei within a continuous cell membrane. Proliferating cells were used as a control.
Immunoprecipitation
Cells were lysed with lysis buffer (50 mM Tris-Cl, pH 7.4, 0.1 % NP40, and 150 mM NaCl) containing protease inhibitors and phosphatase inhibitors. For immunoprecipitation, the indicated antibody was coupled with protein A/G plus agarose bead (Santa Cruz) in 5% BSA in TBST (20 mM Tris-Cl, pH 8.0, 170 mM NaCl, and 0.05% Tween-20) for 2 h. This immune complex was added to the cell lysates (500 μg) and incubated at 4 °C overnight. The resulting beads were washed with TBST five times before analysis.
Confocal microscopy observation
For mtROS determination, C2C12 cells were stained using an mtO2•− indicator 2 μM MitoSOX for 30 min. After washing twice with 1× PBS, MitoSOX intensity was observed at 510 nm. To quantify LC3 puncta, C2C12 cells were plated onto glass coverslips. After 5 days of differentiation in DM, the cells were incubated with 1 μM of LysoTracker™ Red DND-99 (Thermo Fisher Scientific) for 30 min, and then fixed with 4% paraformaldehyde for 30 min and rinsed with PBS twice. The coverslips were then permeablized and blocked with 5% BSA in PBS for 1 h at room temperature, and were incubated with anti-rabbit LC3 (1:200, Cell Signaling) for overnight at 4 °C, followed by secondary antibody (1:300, FITC-conjugated goat anti-rabbit IgG, Thermo Fisher Scientific). Nuclei were stained with 4',6-diamidino-2-phenylindole (DAPI), followed by rinsing and mounting in Vectashield mounting medium (Vecta Laboratories). Images were acquired using Zeiss LSM800 confocal laser scanning microscope (Carl Zeiss).
Muscle injury
The mouse left TA muscle was injured by CTX injection. Before the procedure, the mice were anesthetized. TA muscles were injected with 50 μl of 10 mM CTX solution in 0.9 % NaCl. CTX-injured muscles were dissected on 0 day and 1, 2, 4, 6, and 8 days following injury. The right TA muscles injected with PBS were used as a control. Immediately after isolation, the muscles were frozen in liquid nitrogen and preserved at −80 °C. These tissues were homogenized and lysed with lysis buffer (50 mM Tris-Cl, pH 7.4, 0.1 % NP40, and 150 mM NaCl) containing protease and phosphatase inhibitors and assessed by Western blot analysis. In every experiment, at least three animals were analyzed for each time point after CTX injury. Animal experimentations were conducted according to the protocol approved by the Institutional Animal Care and Use Committee of Kyung Hee University (KHUASP(SE)-17-141).
Sanger-based DNA sequencing of murine mTOR and ULK1 mRNAs
Total mRNA was extracted from C2C12 cells, and the cDNA was synthesized. The translational regions of mTOR and ULK1 were commercially Sanger sequenced by Cosmo Genetech. Then, the sequencing data, of which chromatograms were reviewed by FinchTV software (version 1.4, Perkin Elmer), were aligned with their reference sequences (NM_020009 for mTOR and NM_009469 for ULK1) by Global Alignment (https://blast.ncbi.nlm.nih.gov/).
Real-time quantitative RT-PCR
The mRNA levels of pro-LC3 and GAPDH were measured using a SYBR Green-based real-time RT-PCR using SYBR® Green PCR Master Mix (Applied Biosystems). The thermal cycler conditions were as follows: 10 min at 95 °C followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Finally, the samples were held at 65 °C and melting curves were conducted from 65 to 95.1 °C. All tests were performed in triplicate and all experiments were repeated three times. Expression data were normalized to the geometric mean of housekeeping gene GAPDH to control the variability in expression levels and were analyzed using the 2−ΔΔCT method. The primer sequences, designed by Primer3 (http://primer3.ut.ee/), were as follows: pro-LC3 forward, 5′-AGCAGCATCCAACCAAAATC-3′ and reverse 5′-CTGTGTCCGTTCACCAACAG-3′; GAPDH forward, 5′-TGACCACAGTCCATGCCAT-3′ and reverse, 5′-TTCTAGACGGCAGGTCAGGT-3′.
Western blot analysis
Cell or tissue lysates were collected in cell lysis buffer (1% SDS, 10% glycerol, 32 mM Tris-Cl, pH 6.8) containing phosphatase and protease inhibitors, loaded on an SDS-PAGE gel, and electroblotted onto PVDF membrane. The primary antibodies were applied for overnight in 5% BSA at 4 °C. A Femto Chemiluminescence Kit (Thermo Fisher Scientific) was used for visualization of the signal. Western blots were analyzed using ImageLab (Bio-Rad).
Statistical analysis
The results are expressed as mean ± standard error of the mean (SEM) of at least three independent experiments. The differences between two means were analyzed with the Student's t test and were considered statistically significant when the p values were <0.05.
References
Bassel-Duby R, Olson EN. Signaling pathways in skeletal muscle remodeling. Annu Rev Biochem. 2006;75:19–37.
Florini JR, Ewton DZ, Coolican SA. Growth hormone and the insulin-like growth factor system in myogenesis. Endocr Rev. 1996;17:481–517.
Rodon J, Dienstmann R, Serra V, Tabernero J. Development of PI3K inhibitors: lessons learned from early clinical trials. Nat Rev Clin Oncol. 2013;10:143–53.
Vazquez F, Grossman SR, Takahashi Y, Rokas MV, Nakamura N, Sellers WR. Phosphorylation of the PTEN tail acts as an inhibitory switch by preventing its recruitment into a protein complex. J Biol Chem. 2001;276:48627–30.
Das S, Dixon JE, Cho W. Membrane-binding and activation mechanism of PTEN. Proc Natl Acad Sci USA. 2003;100:7491–6.
Kwon J, Lee SR, Yang KS, Ahn Y, Kim YJ, Stadtman ER, et al. Reversible oxidation and inactivation of the tumor suppressor PTEN in cells stimulated with peptide growth factors. Proc Natl Acad Sci USA. 2004;101:16419–24.
Kang KH, Lemke G, Kim JW. The PI3K-PTEN tug-of-war, oxidative stress and retinal degeneration. Trends Mol Med. 2009;15:191–8.
Laplante M, Sabatini DM. mTOR signaling in growth control and disease. Cell. 2012;149:274–93.
Braun T, Gautel M. Transcriptional mechanisms regulating skeletal muscle differentiation, growth and homeostasis. Nat Rev Mol Cell Biol. 2011;12:349–61.
Ma XM, Blenis J. Molecular mechanisms of mTOR-mediated translational control. Nat Rev Mol Cell Biol. 2009;10:307–18.
Nakatogawa H, Suzuki K, Kamada Y, Ohsumi Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol. 2009;10:458–67.
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science. 2005;307:1098–101.
Kim YC, Guan KL. mTOR: a pharmacologic target for autophagy regulation. J Clin Invest. 2015;125:25–32.
Kim J, Kundu M, Viollet B, Guan KL. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13:132–41.
Ravikumar B, Sarkar S, Davies JE, Futter M, Garcia-Arencibia M, Green-Thompson ZW, et al. Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev. 2010;90:1383–435.
Janku F, McConkey DJ, Hong DS, Kurzrock R. Autophagy as a target for anticancer therapy. Nat Rev Clin Oncol. 2011;8:528–39.
Krauss S, Zhang CY, Lowell BB. The mitochondrial uncoupling-protein homologues. Nat Rev Mol Cell Biol. 2005;6:248–61.
Murphy MP. How mitochondria produce reactive oxygen species. Biochem J. 2009;417:1–13.
Lee S, Tak E, Lee J, Rashid MA, Murphy MP, Ha J, et al. Mitochondrial H2O2 generated from electron transport chain complex I stimulates muscle differentiation. Cell Res. 2011;21:817–34.
Tormos KV, Anso E, Hamanaka RB, Eisenbart J, Joseph J, Kalyanaraman B, et al. Mitochondrial complex III ROS regulate adipocyte differentiation. Cell Metab. 2011;14:537–44.
Hamanaka RB, Glasauer A, Hoover P, Yang S, Blatt H, Mullen AR, et al. Mitochondrial reactive oxygen species promote epidermal differentiation and hair follicle development. Sci Signal. 2013;6:ra8.
Del Prete A, Zaccagnino P, Di Paola M, Saltarella M, Oliveros Celis C, Nico B, et al. Role of mitochondria and reactive oxygen species in dendritic cell differentiation and functions. Free Radic Biol Med. 2008;44:1443–51.
Khacho M, Clark A, Svoboda DS, Azzi J, MacLaurin JG, Meghaizel C, et al. Mitochondrial dynamics impacts stem cell identity and fate decisions by regulating a nuclear transcriptional program. Cell Stem Cell. 2016;19:232–47.
Rhee SG. Cell signaling. H2O2, a necessary evil for cell signaling. Science. 2006;312:1882–3.
Wang Y, DuBois JL, Hedman B, Hodgson KO, Stack TD. Catalytic galactose oxidase models: biomimetic Cu(II)-phenoxyl-radical reactivity. Science. 1998;279:537–40.
Kalyanaraman B, Darley-Usmar V, Davies KJ, Dennery PA, Forman HJ, Grisham MB, et al. Measuring reactive oxygen and nitrogen species with fluorescent probes: challenges and limitations. Free Radic Biol Med. 2012;52:1–6.
Connor KM, Subbaram S, Regan KJ, Nelson KK, Mazurkiewicz JE, Bartholomew PJ, et al. Mitochondrial H2O2 regulates the angiogenic phenotype via PTEN oxidation. J Biol Chem. 2005;280:16916–24.
Wu KL, Wu CA, Wu CW, Chan SH, Chang AY, Chan JY. Redox-sensitive oxidation and phosphorylation of PTEN contribute to enhanced activation of PI3K/Akt signaling in rostral ventrolateral medulla and neurogenic hypertension in spontaneously hypertensive rats. Antioxid Redox Signal. 2013;18:36–50.
Crespo FL, Sobrado VR, Gomez L, Cervera AM, McCreath KJ. Mitochondrial reactive oxygen species mediate cardiomyocyte formation from embryonic stem cells in high glucose. Stem Cells. 2010;28:1132–42.
Yoon MS. mTOR as a key regulator in maintaining skeletal muscle mass. Front Physiol. 2017;8:788.
Sandri M. Autophagy in skeletal muscle. FEBS Lett. 2010;584:1411–6.
Martinez-Lopez N, Athonvarangkul D, Sahu S, Coletto L, Zong H, Bastie CC, et al. Autophagy in Myf5+ progenitors regulates energy and glucose homeostasis through control of brown fat and skeletal muscle development. EMBO Rep. 2013;14:795–803.
Forgac M. Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat Rev Mol Cell Biol. 2007;8:917–29.
Fortini P, Ferretti C, Iorio E, Cagnin M, Garribba L, Pietraforte D, et al. The fine tuning of metabolism, autophagy and differentiation during in vitro myogenesis. Cell Death Dis. 2016;7:e2168.
Kim J, Guan KL. Regulation of the autophagy initiating kinase ULK1 by nutrients: roles of mTORC1 and AMPK. Cell Cycle. 2011;10:1337–8.
Hu Z, Wang H, Lee IH, Modi S, Wang X, Du J, et al. PTEN inhibition improves muscle regeneration in mice fed a high-fat diet. Diabetes. 2010;59:1312–20.
Vasilaki A, Jackson MJ. Role of reactive oxygen species in the defective regeneration seen in aging muscle. Free Radic Biol Med. 2013;65:317–23.
Ownby CL, Fletcher JE, Colberg TR. Cardiotoxin 1 from cobra (Naja naja atra) venom causes necrosis of skeletal muscle in vivo. Toxicon. 1993;31:697–709.
Lee J, Choi KJ, Lim MJ, Hong F, Choi TG, Tak E, et al. Proto-oncogenic H-Ras, K-Ras, and N-Ras are involved in muscle differentiation via phosphatidylinositol 3-kinase. Cell Res. 2010;20:919–34.
Remels AH, Langen RC, Schrauwen P, Schaart G, Schols AM, Gosker HR. Regulation of mitochondrial biogenesis during myogenesis. Mol Cell Endocrinol. 2010;315:113–20.
Malinska D, Kudin AP, Bejtka M, Kunz WS. Changes in mitochondrial reactive oxygen species synthesis during differentiation of skeletal muscle cells. Mitochondrion. 2012;12:144–8.
Song MS, Salmena L, Pandolfi PP. The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol. 2012;13:283–96.
White ES, Atrasz RG, Hu B, Phan SH, Stambolic V, Mak TW, et al. Negative regulation of myofibroblast differentiation by PTEN (Phosphatase and Tensin Homolog Deleted on chromosome 10). Am J Respir Crit Care Med. 2006;173:112–21.
Yue F, Bi P, Wang C, Li J, Liu X, Kuang S. Conditional loss of Pten in myogenic progenitors leads to postnatal skeletal muscle hypertrophy but age-dependent exhaustion of satellite cells. Cell Rep. 2016;17:2340–53.
Erbay E, Chen J. The mammalian target of rapamycin regulates C2C12 myogenesis via a kinase-independent mechanism. J Biol Chem. 2001;276:36079–82.
Sun Y, Ge Y, Drnevich J, Zhao Y, Band M, Chen J. Mammalian target of rapamycin regulates miRNA-1 and follistatin in skeletal myogenesis. J Cell Biol. 2010;189:1157–69.
Ohanna M, Sobering AK, Lapointe T, Lorenzo L, Praud C, Petroulakis E, et al. Atrophy of S6K1(−/−) skeletal muscle cells reveals distinct mTOR effectors for cell cycle and size control. Nat Cell Biol. 2005;7:286–94.
Shu L, Houghton PJ. The mTORC2 complex regulates terminal differentiation of C2C12 myoblasts. Mol Cell Biol. 2009;29:4691–4700.
Jaafar R, Zeiller C, Pirola L, Di Grazia A, Naro F, Vidal H, et al. Phospholipase D regulates myogenic differentiation through the activation of both mTORC1 and mTORC2 complexes. J Biol Chem. 2011;286:22609–21.
Mizushima N, Levine B. Autophagy in mammalian development and differentiation. Nat Cell Biol. 2010;12:823–30.
Acknowledgements
This study was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (NRF-2016R1D1A1B03933763 to TGC, and NRF-2011-0030072, NRF-2017R1A2B2007870, and NRF-2018R1A6A1A03025124 to SSK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Edited by E. Baehrecke
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kim, JH., Choi, T.G., Park, S. et al. Mitochondrial ROS-derived PTEN oxidation activates PI3K pathway for mTOR-induced myogenic autophagy. Cell Death Differ 25, 1921–1937 (2018). https://doi.org/10.1038/s41418-018-0165-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-018-0165-9
This article is cited by
-
ROS production by mitochondria: function or dysfunction?
Oncogene (2024)
-
Different levels of autophagy induced by transient serum starvation regulate metabolism and differentiation of porcine skeletal muscle satellite cells
Scientific Reports (2023)
-
ATGL deficiency aggravates pressure overload-triggered myocardial hypertrophic remodeling associated with the proteasome-PTEN-mTOR-autophagy pathway
Cell Biology and Toxicology (2023)
-
Nutri-stress, mitochondrial dysfunction, and insulin resistance—role of heat shock proteins
Cell Stress and Chaperones (2023)
-
Macropinocytosis is an alternative pathway of cysteine acquisition and mitigates sorafenib-induced ferroptosis in hepatocellular carcinoma
Journal of Experimental & Clinical Cancer Research (2022)