Abstract
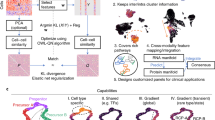
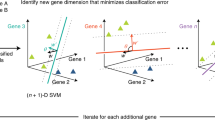
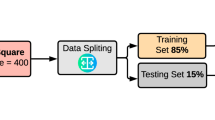
Acute myeloid leukemia (AML) is a type of blood cancer characterized by the rapid growth of immature white blood cells from the bone marrow. Therapy resistance resulting from the persistence of leukemia stem cells (LSCs) are found in numerous patients. Comparative transcriptome studies have been previously conducted to analyze differentially expressed genes between LSC+ and LSC− cells. However, these studies mainly focused on a limited number of genes with the most obvious expression differences between the two cell types. We developed a computational approach incorporating several machine learning algorithms, including Monte Carlo feature selection (MCFS), incremental feature selection (IFS), support vector machine (SVM), Repeated Incremental Pruning to Produce Error Reduction (RIPPER), to identify gene expression features specific to LSCs. One thousand 0ne hudred fifty-nine features (genes) were first identified, which can be used to build the optimal SVM classifier for distinguishing LSC+ and LSC− cells. Among these 1159 genes, the top 17 genes were identified as LSC-specific biomarkers. In addition, six classification rules were produced by RIPPER algorithm. The subsequent literature review on these features/genes and the classification rules and functional enrichment analyses of the 1159 features/genes confirmed the relevance of extracted genes and rules to the characteristics of LSCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ballen KK, Lazarus H. Cord blood transplant for acute myeloid leukaemia. Br J Haematol. 2016;173:25–36.
Chevallier P, Labopin M, Socie G, Rubio MT, Blaise D, Vigouroux S, et al. Comparison of umbilical cord blood allogeneic stem cell transplantation vs. auto-SCT for adult acute myeloid leukemia patients in second complete remission at transplant: a retrospective study on behalf of the SFGM-TC. Eur J Haematol. 2015;94:449–55.
Cheuk D, Chiang A, Ha SY, Chan G. Favorable outcomes of unrelated cord blood transplant for pediatric acute myeloid leukemia In Hong Kong. Pediatr Blood Cancer. 2013;60:78–78.
Huang XX, Xiong M, Jin YJ, Deng CH, Xu H, An CQ, et al. Evidence that high-migration drug-surviving MOLT4 leukemia cells exhibit cancer stem cell-like properties. Int J Oncol. 2016;49:343–51.
Fong CY, Gilan O, Lam EYN, Rubin AF, Ftouni S, Tyler D, et al. BET inhibitor resistance emerges from leukaemia stem cells. Nature. 2015;525:538–42.
Sinclair A, Latif AL, Holyoake TL. Targeting survival pathways in chronic myeloid leukaemia stem cells. Br J Pharmacol. 2013;169:1693–707.
Brenner AK, Tvedt THA, Bruserud O. The complexity of targeting PI3K-Akt-mTOR signalling in human acute myeloid leukaemia: the importance of leukemic cell heterogeneity, neighbouring mesenchymal stem cells and immunocompetent cells. Molecules. 2016;21:1512.
Zoller M. CD44, hyaluronan, the hematopoietic stem cell, and leukemia-initiating cells. Front Immunol. 2015;6:235.
Fong CY, Gilan O, Lam EY, Rubin AF, Ftouni S, Tyler D, et al. BET inhibitor resistance emerges from leukaemia stem cells. Nature. 2015;525:538–42.
Prost S, Relouzat F, Spentchian M, Ouzegdouh Y, Saliba J, Massonnet G, et al. Erosion of the chronic myeloid leukaemia stem cell pool by PPAR gamma agonists. Nature. 2015;525:380–3.
Crews LA, Jamieson CHM. Selective elimination of leukemia stem cells: Hitting a moving target. Cancer Lett. 2013;338:15–22.
Hosen N, Park CY, Tatsumi N, Oji Y, Sugiyama H, Gramatzki M, et al. CD96 is a leukemic stem cell-specific marker in human acute myeloid leukemia. Proc Natl Acad Sci USA. 2007;104:11008–13.
Kinstrie R, Horne GA, Morrison H, Moka HA, Cassels J, Dunn K, et al. CD93 is a novel biomarker of leukemia stem cells in chronic myeloid leukemia. Blood. 2015;126:49.
Horacek JM, Jebavy L, Tichy M, Pudil R, Zak P, Slovacek L, et al. Multiple biomarkers in the assessment of cardiac toxicity during haematopoietic stem cell transplantation in acute leukaemia. Bone Marrow Transplant. 2008;41:S100.
Tohda S. [Biomarker for hematopoietic tumors--aiming for personalized diagnosis of leukemia stem cells]. Rinsho Byori. 2015;63:1110–3.
Ng SWK, Mitchell A, Kennedy JA, Chen WC, McLeod J, Ibrahimova N, et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature. 2016;540:433–7.
Cortes C, Vapnik V. Support-vector networks. Mach Learn. 1995;20:273–97.
Draminski M, Rada-Iglesias A, Enroth S, Wadelius C, Koronacki J, Komorowski J. Monte Carlo feature selection for supervised classification. Bioinformatics. 2008;24:110–7.
Fast effective rule induction. Proc the twelfth international conference on machine learning. Elsevier, Tahoe City, California, USA, 1995.
Dramiński M, Kierczak M, Nowak-Brzezińska A, Koronecki J, Komorowski J. The Monte Carlo feature selection and interdependency discovery is unbiased. Control Cybern. 2011;40:199–211.
Chen L, Li J, Zhang Y-H, Feng K, Wang S, Zhang Y, et al. Identification of gene expression signatures across different types of neural stem cells with the Monte-Carlo feature selection method. J Cell Biochem. 2018;119:3394–403.
Wang D, Li J-R, Zhang Y-H, Chen L, Huang T, Cai Y-D. Identification of differentially expressed genes between original breast cancer and xenograft using machine learning algorithms. Genes. 2018;9:155.
Pan X, Hu X, Zhang Y-h, Feng K, Wang SP, Chen L, et al. Identifying patients with atrioventricular septal defect in down syndrome populations by using self-normalizing neural networks and feature selection. Genes. 2018;9:208.
Wang S, Zhang YH, Lu J, Cui W, Hu J, Cai YD. Analysis and identification of aptamer-compound interactions with a maximum relevance minimum redundancy and nearest neighbor algorithm. Biomed Res Int. 2016:2016:8351204.
A study of cross-validation and bootstrap for accuracy estimation and model selection. International joint Conference on artificial intelligence. Lawrence Erlbaum Associates Ltd, Montreal, QC, Canada, 1995.
Platt J. Sequential minimal optimizaton: a fast algorithm for training support vector machines. Tech. Rep. MSR-TR-98-14. 1998. https://www.microsoft.com/en-us/research/publication/sequential-minimaloptimization-a-fast-algorithm-for-training-support-vector-machines/.
Frank E, Hall M, Trigg L, Holmes G, Witten IH. Data mining in bioinformatics using Weka. Bioinformatics. 2004;20:2479–81.
Chen L, Zhang YH, Lu G, Huang T, Cai YD. Analysis of cancer-related lncRNAs using gene ontology and KEGG pathways. Artif Intell Med. 2017;76:27–36.
Zhang Q, Sun X, Feng K, Wang S, Zhang YH, Wang S, et al. Predicting citrullination sites in protein sequences using mRMR method and random forest algorithm. Comb Chem High Throughput Screen. 2017;20:164–73.
Chen L, Zhang YH, Huang T, Cai YD. Gene expression profiling gut microbiota in different races of humans. Sci Rep. 2016;6:23075.
Zhang YH, Xing ZH, Liu CL, Wang SP, Huang T, Cai YD, et al. Identification of the core regulators of the HLA I-peptide binding process. Sci Rep. 2017;7:42768.
Li J, Lu L, Zhang Y, Liu M, Chen L, Huang T, et al. Identification of synthetic lethality based on a functional network by using machine learning algorithms. J Cell Biochem. 2019;120:405–16.
Chen L, Pan X, Hu X, Zhang Y-H, Wang S, Huang T, et al. Gene expression differences among different MSI statuses in colorectal cancer. Int J Cancer. 2018;143:1731–40.
Zhao X, Chen L, Lu J. A similarity-based method for prediction of drug side effects with heterogeneous information. Math Biosci. 2018;306:136–44.
Zhao X, Chen L, Guo Z-H, Liu T. Predicting drug side effects with compact integration of heterogeneous networks. Curr. Bioinforma. 2019. https://www.benthamscience.com/journals/current-bioinformatics/article/170114/.
Wang S, Zhang Q, Lu J, Cai YD. Analysis and prediction of nitrated tyrosine sites with mRMR method and support vector machine algorithm. Curr Bioinforma. 2018;13:3–13.
Li BQ, Cai YD, Feng KY, Zhao GJ. Prediction of protein cleavage site with feature selection by random forest. PloS ONE. 2012;7:e45854.
Chen L, Wang S, Zhang Y-H, Li J, Xing Z-H, Yang J, et al. Identify key sequence features to improve CRISPR sgRNA efficacy. IEEE Access. 2017;5:26582–90.
Liu L, Chen L, Zhang YH, Wei L, Cheng S, Kong X, et al. Analysis and prediction of drug-drug interaction by minimum redundancy maximum relevance and incremental feature selection. J Biomol Struct Dyn. 2017;35:312–29.
Chen L, Chu C, Zhang Y-H, Zheng M-Y, Zhu L, Kong X, et al. Identification of drug-drug interactions using chemical interactions. Curr Bioinforma. 2017;12:526–34.
Cui H, Chen L.. A binary classifier for the prediction of EC numbers of enzymes. Curr. Proteom. 2019;16:381–9.
Matthews BW. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim Biophys Acta. 1975;405:442–51.
Minagawa K, Jamil MO, Al-Obaidi M, Pereboeva L, Salzman D, Erba HP, et al. In vitro pre-clinical validation of suicide gene modified anti-CD33 redirected chimeric antigen receptor T-Cells for acute myeloid leukemia. PloS ONE. 2016;11:e0166891.
Lakshmikanth T, Olin A, Chen Y, Mikes J, Fredlund E, Remberger M, et al. Mass cytometry and topological data analysis reveal immune parameters associated with complications after allogeneic stem cell transplantation. Cell Rep. 2017;20:2238–50.
Hashiguchi K, Ozaki M, Kuraoka I, Saitoh H. Establishment of a human cell line stably overexpressing mouse Nip45 and characterization of Nip45 subcellular localization. Biochem Biophys Res Commun. 2013;430:72–7.
Kozlowska M, Smolenski RT, Makarewicz W, Hoffmann C, Jastorff B, Swierczynski J. ATP depletion, purine riboside triphosphate accumulation and rat thymocyte death induced by purine riboside. Toxicol Lett. 1999;104:171–81.
Sekelova Z, Polansky O, Stepanova H, Fedr R, Faldynova M, Rychlik I. et al. Different roles of CD4, CD8 and gammadelta T-lymphocytes in naive and vaccinated chickens during Salmonella Enteritidis infection. Proteomics. 2017;17:13–4.
Huang W, Li H, Luo R. The microRNA-1246 promotes metastasis in non-small cell lung cancer by targeting cytoplasmic polyadenylation element-binding protein 4. Diagn Pathol. 2015;10:127.
Charlesworth A, Meijer HA, de Moor CH. Specificity factors in cytoplasmic polyadenylation. Wiley Inter Rev RNA. 2013;4:437–61.
Maillo C, Martin J, Sebastian D, Hernandez-Alvarez M, Garcia-Rocha M, Reina O, et al. Circadian- and UPR-dependent control of CPEB4 mediates a translational response to counteract hepatic steatosis under ER stress. Nat Cell Biol. 2017;19:94–105.
Crowther-Swanepoel D, Broderick P, Di Bernardo MC, Dobbins SE, Torres M, Mansouri M, et al. Common variants at 2q37.3, 8q24.21, 15q21.3 and 16q24.1 influence chronic lymphocytic leukemia risk. Nat Genet. 2010;42:132–6.
Yin J, Park G, Lee JE, Park JY, Kim TH, Kim YJ, et al. CPEB1 modulates differentiation of glioma stem cells via downregulation of HES1 and SIRT1 expression. Oncotarget. 2014;5:6756–69.
Coleman EA, Lee JY, Erickson SW, Goodwin JA, Sanathkumar N, Raj VR, et al. GWAS of 972 autologous stem cell recipients with multiple myeloma identifies 11 genetic variants associated with chemotherapy-induced oral mucositis. Support Care Cancer. 2015;23:841–9.
Sanders MA, Madoux F, Mladenovic L, Zhang H, Ye X, Angrish M, et al. Endogenous and synthetic ABHD5 ligands regulate ABHD5-perilipin interactions and lipolysis in fat and muscle. Cell Metab. 2015;22:851–60.
Vieyres G, Welsch K, Gerold G, Gentzsch J, Kahl S, Vondran FW, et al. ABHD5/CGI-58, the Chanarin-Dorfman syndrome protein, mobilises lipid stores for hepatitis C virus production. PLoS Pathog. 2016;12:e1005568.
Demignot S, Beilstein F, Morel E. Triglyceride-rich lipoproteins and cytosolic lipid droplets in enterocytes: key players in intestinal physiology and metabolic disorders. Biochimie. 2014;96:48–55.
Gal H, Amariglio N, Trakhtenbrot L, Jacob-Hirsh J, Margalit O, Avigdor A, et al. Gene expression profiles of AML derived stem cells; similarity to hematopoietic stem cells. Leukemia. 2006;20:2147–54.
Xiao S, Li R, Diao H, Zhao F, Ye X. Progesterone receptor-mediated regulation of N-acetylneuraminate pyruvate lyase (NPL) in mouse uterine luminal epithelium and nonessential role of NPL in uterine function. PloS ONE. 2013;8:e65607.
Byers HL, Homer KA, Beighton D. Utilization of sialic acid by viridans streptococci. J Dent Res. 1996;75:1564–71.
Minelli A, Maserati E, Rossi G, Bernardo ME, De Stefano P, Cecchini MP, et al. Familial platelet disorder with propensity to acute myelogenous leukemia: genetic heterogeneity and progression to leukemia via acquisition of clonal chromosome anomalies. Genes Chromosomes Cancer. 2004;40:165–71.
Horwitz M, Benson KF, Li FQ, Wolff J, Leppert MF, Hobson L, et al. Genetic heterogeneity in familial acute myelogenous leukemia: evidence for a second locus at chromosome 16q21-23.2. Am J Hum Genet. 1997;61:873–81.
Sharma HS, Muresanu DF, Lafuente JV, Patnaik R, Tian ZR, Ozkizilcik A, et al. Co-Administration of TiO2 Nanowired Mesenchymal Stem Cells with Cerebrolysin Potentiates Neprilysin Level and Reduces Brain Pathology in Alzheimer’s Disease. Mol Neurobiol. 2018;55:300–11.
Wong SH, Goode DL, Iwasaki M, Wei MC, Kuo HP, Zhu L, et al. The H3K4-methyl epigenome regulates leukemia stem cell oncogenic potential. Cancer Cell. 2015;28:198–209.
Roos J, Oancea C, Heinssmann M, Khan D, Held H, Kahnt AS, et al. 5-Lipoxygenase is a candidate target for therapeutic management of stem cell-like cells in acute myeloid leukemia. Cancer Res. 2014;74:5244–55.
Shlush LI, Zandi S, Mitchell A, Chen WC, Brandwein JM, Gupta V, et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature. 2014;506:328–33.
Maekawa S, Imamachi N, Irie T, Tani H, Matsumoto K, Mizutani R, et al. Analysis of RNA decay factor mediated RNA stability contributions on RNA abundance. BMC Genom. 2015;16:154.
Yeung ML, Houzet L, Yedavalli VS, Jeang KT. A genome-wide short hairpin RNA screening of jurkat T-cells for human proteins contributing to productive HIV-1 replication. J Biol Chem. 2009;284:19463–73.
Schwartz EL, Nilson LA. Activation of 2’,5’-oligoadenylate synthetase activity on induction of HL-60 leukemia cell differentiation. Mol Cell Biol. 1989;9:3897–903.
Ksionda O, Melton AA, Bache J, Tenhagen M, Bakker J, Harvey R, et al. RasGRP1 overexpression in T-ALL increases basal nucleotide exchange on Ras rendering the Ras/PI3K/Akt pathway responsive to protumorigenic cytokines. Oncogene. 2016;35:3658–68.
Oki T, Kitaura J, Watanabe-Okochi N, Nishimura K, Maehara A, Uchida T, et al. Aberrant expression of RasGRP1 cooperates with gain-of-function NOTCH1 mutations in T-cell leukemogenesis. Leukemia. 2012;26:1038–45.
Wang Y, Wu P, Lin R, Rong L, Xue Y, Fang Y. LncRNA NALT interaction with NOTCH1 promoted cell proliferation in pediatric T cell acute lymphoblastic leukemia. Sci Rep. 2015;5:13749.
Chen T, Meng Z, Gan Y, Wang X, Xu F, Gu Y, et al. The viral oncogene Np9 acts as a critical molecular switch for co-activating beta-catenin, ERK, Akt and Notch1 and promoting the growth of human leukemia stem/progenitor cells. Leukemia. 2013;27:1469–78.
Choi DB, Park MR, Kim HR, Jun CD, Kim HJ, Shim H, et al. Aberrant proteomic expression of NSRP70 and its clinical implications and connection to the transcriptional level in adult acute leukemia. Leuk Res. 2014;38:1252–9.
Liu J, Huang B, Xiao Y, Xiong HM, Li J, Feng DQ, et al. Aberrant expression of splicing factors in newly diagnosed acute myeloid leukemia. Onkologie. 2012;35:335–40.
Niemsiri V, Wang X, Pirim D, Radwan ZH, Bunker CH, Barmada MM, et al. Genetic contribution of SCARB1 variants to lipid traits in African Blacks: a candidate gene association study. BMC Med Genet. 2015;16:106.
Kobayashi S, Suzuki T, Kawaguchi A, Phongphaew W, Yoshii K, Iwano T, et al. Rab8b regulates transport of West Nile virus particles from recycling endosomes. J Biol Chem. 2016;291:6559–68.
Hein MY, Hubner NC, Poser I, Cox J, Nagaraj N, Toyoda Y, et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell. 2015;163:712–23.
Usman H, Ameer F, Munir R, Iqbal A, Zaid M, Hasnain S, et al. Leukemia cells display lower levels of intracellular cholesterol irrespective of the exogenous cholesterol availability. Clin Chim Acta. 2016;457:12–7.
Capalbo G, Mueller-Kuller T, Koschmieder S, Klein HU, Ottmann OG, Hoelzer D, et al. Endoplasmic reticulum protein GliPR1 regulates G protein signaling and the cell cycle and is overexpressed in AML. Oncol Rep. 2013;30:2254–62.
Bansal AK, Vishnubhatla S, Bakhshi S. Correlation of serum immunoglobulins with infection-related parameters during induction chemotherapy of pediatric acute myeloid leukemia: a prospective study. Pedia Hematol Oncol. 2015;32:129–37.
Jefferson AL, Woodhead HJ, Fyfe S, Briody J, Bebbington A, Strauss BJ, et al. Bone mineral content and density in Rett syndrome and their contributing factors. Pedia Res. 2011;69:293–8.
Cho WK, Ahn MB, Lee JW, Chung NG, Jung MH, Cho B, et al. Low bone mineral density in adolescents with leukemia after hematopoietic stem cell transplantation: prolonged steroid therapy for GvHD and endocrinopathy after hematopoietic stem cell transplantation might be major concerns? Bone Marrow Transpl. 2017;52:144–6.
Hopkins M, Tyson JJ, Novak B. Cell-cycle transitions: a common role for stoichiometric inhibitors. Mol Biol Cell. 2017;28:3437–46.
Kar S. Unraveling cell-cycle dynamics in cancer. Cell Syst. 2016;2:8–10.
Wang W, Stiehl T, Raffel S, Hoang VT, Hoffmann I, Poisa-Beiro L, et al. Reduced hematopoietic stem cell frequency predicts outcome in acute myeloid leukemia. Haematologica. 2017;102:1567–77.
Thomas D, Majeti R. Biology and relevance of human acute myeloid leukemia stem cells. Blood. 2017;129:1577–85.
Jeong YH, Sekiya M, Hirata M, Ye M, Yamagishi A, Lee SM, et al. The low-density lipoprotein receptor-related protein 10 is a negative regulator of the canonical Wnt/beta-catenin signaling pathway. Biochem Biophys Res Commun. 2010;392:495–9.
Giambra V, Jenkins CE, Lam SH, Hoofd C, Belmonte M, Wang X, et al. Leukemia stem cells in T-ALL require active Hif1alpha and Wnt signaling. Blood. 2015;125:3917–27.
Dong L, Yu WM, Zheng H, Loh ML, Bunting ST, Pauly M, et al. Leukaemogenic effects of Ptpn11 activating mutations in the stem cell microenvironment. Nature. 2016;539:304–8.
Raffel S, Falcone M, Kneisel N, Hansson J, Wang W, Lutz C, et al. BCAT1 restricts alphaKG levels in AML stem cells leading to IDHmut-like DNA hypermethylation. Nature. 2017;551:384–8.
Fukawa T, Ono M, Matsuo T, Uehara H, Miki T, Nakamura Y, et al. DDX31 regulates the p53-HDM2 pathway and rRNA gene transcription through its interaction with NPM1 in renal cell carcinomas. Cancer Res. 2012;72:5867–77.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: archive for functional genomics data sets--update. Nucleic acids Res. 2013;41(Database issue):D991–5.
Zanoni P, Khetarpal SA, Larach DB, Hancock-Cerutti WF, Millar JS, Cuchel M, et al. Rare variant in scavenger receptor BI raises HDL cholesterol and increases risk of coronary heart disease. Science. 2016;351:1166–71.
Helgadottir A, Sulem P, Thorgeirsson G, Gretarsdottir S, Thorleifsson G, Jensson BO, et al. Rare SCARB1 mutations associate with high-density lipoprotein cholesterol but not with coronary artery disease. Eur Heart J. 2018;39:2172–8.
Toomey MB, Lopes RJ, Araujo PM, Johnson JD, Gazda MA, Afonso S, et al. High-density lipoprotein receptor SCARB1 is required for carotenoid coloration in birds. Proc Natl Acad Sci USA. 2017;114:5219–24.
Zhang W, Dong R, Diao S, Du J, Fan Z, Wang F. Differential long noncoding RNA/mRNA expression profiling and functional network analysis during osteogenic differentiation of human bone marrow mesenchymal stem cells. Stem Cell Res Ther. 2017;8:30.
Martineau C, Martin-Falstrault L, Brissette L, Moreau R. Gender- and region-specific alterations in bone metabolism in Scarb1-null female mice. J Endocrinol. 2014;222:277–88.
Oehler VG, Yeung KY, Choi YE, Bumgarner RE, Raftery AE, Radich JP. The derivation of diagnostic markers of chronic myeloid leukemia progression from microarray data. Blood. 2009;114:3292–8.
Wang S, Hu B, Ding Z, Dang Y, Wu J, Li D, et al. ATF6 safeguards organelle homeostasis and cellular aging in human mesenchymal stem cells. Cell Disco. 2018;4:2.
Wang HH, Cui Q, Zhang T, Wang ZB, Ouyang YC, Shen W, et al. Rab3A, Rab27A, and Rab35 regulate different events during mouse oocyte meiotic maturation and activation. Histochem Cell Biol. 2016;145:647–57.
Li L, Bhatia R. Role of SIRT1 in the growth and regulation of normal hematopoietic and leukemia stem cells. Curr Opin Hematol. 2015;22:324–9.
Kleppe M, Spitzer MH, Li S, Hill CE, Dong L, Papalexi E, et al. Jak1 integrates cytokine sensing to regulate hematopoietic stem cell function and stress hematopoiesis. Cell Stem Cell. 2018;22:277.
Chen G, Zhou G, Aras S, He Z, Lucas S, Podgorski I, et al. Loss of ABHD5 promotes the aggressiveness of prostate cancer cells. Sci Rep. 2017;7:13021.
Peng Y, Miao H, Wu S, Yang W, Zhang Y, Xie G, et al. ABHD5 interacts with BECN1 to regulate autophagy and tumorigenesis of colon cancer independent of PNPLA2. Autophagy. 2016;12:2167–82.
Ichim CV. Kinase-independent mechanisms of resistance of leukemia stem cells to tyrosine kinase inhibitors. Stem Cells Transl Med. 2014;3:405–15.
Liu J, Zhang X, Zhong JF, Zhang C. CAR-T cells and allogeneic hematopoietic stem cell transplantation for relapsed/refractory B-cell acute lymphoblastic leukemia. Immunotherapy. 2017;9:1115–25.
Danis E, Yamauchi T, Echanique K, Zhang X, Haladyna JN, Riedel SS, et al. Ezh2 controls an early hematopoietic program and growth and survival signaling in early T cell precursor acute lymphoblastic leukemia. Cell Rep. 2016;14:1953–65.
Funding
This study was supported by the National Natural Science Foundation of China (31701151), Natural Science Foundation of Shanghai (17ZR1412500), National Key R&D Program of China (2018YFC0910403), Shanghai Sailing Program (16YF1413800), the Youth Innovation Promotion Association of Chinese Academy of Sciences (CAS) (2016245), the fund of the key Laboratory of Stem Cell Biology of Chinese Academy of Sciences (201703), Science and Technology Commission of Shanghai Municipality (STCSM) (18dz2271000).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, J., Lu, L., Zhang, YH. et al. Identification of leukemia stem cell expression signatures through Monte Carlo feature selection strategy and support vector machine. Cancer Gene Ther 27, 56–69 (2020). https://doi.org/10.1038/s41417-019-0105-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-019-0105-y
This article is cited by
-
A new Monte Carlo Feature Selection (MCFS) algorithm-based weighting scheme for multi-model ensemble of precipitation
Theoretical and Applied Climatology (2024)
-
Machine Learning Approaches for Stem Cells
Current Stem Cell Reports (2023)
-
IGFBP5 is an ROR1 ligand promoting glioblastoma invasion via ROR1/HER2-CREB signaling axis
Nature Communications (2023)
-
Multimetric feature selection for analyzing multicategory outcomes of colorectal cancer: random forest and multinomial logistic regression models
Laboratory Investigation (2022)
-
Cancer gene therapy 2020: highlights from a challenging year
Cancer Gene Therapy (2022)