Abstract
Background
The E3 ubiquitin ligase HECTD3 is a homologue of the E6-related protein carboxyl terminus, which plays a crucial role in biological processes and tumourigenesis. However, the functional characterisation of HECTD3 in glioblastoma is still elusive.
Methods
Determination of the functional role of HECTD3 in glioblastoma was made by a combination of HECTD3 molecular pattern analysis from human glioblastoma databases and subcutaneous and in situ injections of tumours in mice models.
Results
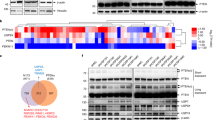
This study reports that the DOC domain of HECTD3 interacts with the DNA binding domain of PARP1, and HECTD3 mediated the K63-linked polyubiquitination of PARP1 and stabilised the latter expression. Moreover, the Cysteine (Cys) 823 (ubiquitin-binding site) mutation of HECTD3 significantly reduced PARP1 polyubiquitination and HECTD3 was involved in the recruitment of ubiquitin-related molecules to PARP1 ubiquitin-binding sites (Lysines 209 and 221, respectively). Lastly, activation of EGFR-mediated signalling pathways by HECTD3 regulates PARP1 polyubiquitination.
Conclusion
Our results unveil the potential role of HECTD3 in glioblastoma and strongly preconise further investigation and consider HECTD3 as a promising therapeutic marker for glioblastoma treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data analysed or generated in this study are included in this article as well as in the Supplementary Information file.
References
Budke M, Isla-Guerrero A, Perez-Lopez C, Perez-Alvarez M, Garcia-Grande A, Bello MJ, et al. A comparative study of the treatment of high grade gliomas. Rev Neurol. 2003;37:912–6.
Riddick G, Fine HA. Integration and analysis of genome-scale data from gliomas. Nat Rev Neurol. 2011;7:439–50.
Zhao Y, He J, Li Y, Lv S, Cui H. NUSAP1 potentiates chemoresistance in glioblastoma through its SAP domain to stabilize ATR. Signal Transduct Target Ther. 2020;5:44.
Jiang Q, Li F, Cheng Z, Kong Y, Chen C. The role of E3 ubiquitin ligase HECTD3 in cancer and beyond. Cell Mol life Sci. 2020;77:1483–95.
Pickart CM. Mechanisms underlying ubiquitination. Annu Rev Biochem. 2001;70:503–33.
Komander D, Rape M. The ubiquitin code. Annu Rev Biochem. 2012;81:203–29.
Tokunaga F, Sakata S, Saeki Y, Satomi Y, Kirisako T, Kamei K, et al. Involvement of linear polyubiquitylation of NEMO in NF-kappaB activation. Nat Cell Biol. 2009;11:123–32.
Trempe JF. Reading the ubiquitin postal code. Curr Opin Struct Biol. 2011;21:792–801.
Li Y, Chen X, Wang Z, Zhao D, Chen H, Chen W, et al. The HECTD3 E3 ubiquitin ligase suppresses cisplatin-induced apoptosis via stabilizing MALT1. Neoplasia. 2013;15:39–48.
Zhang L, Kang L, Bond W, Zhang N. Interaction between syntaxin 8 and HECTd3, a HECT domain ligase. Cell Mol Neurobiol. 2009;29:115–21.
Yu J, Lan J, Zhu Y, Li X, Lai X, Xue Y, et al. The E3 ubiquitin ligase HECTD3 regulates ubiquitination and degradation of Tara. Biochem Biophys Res Commun. 2008;367:805–12.
Shu T, Li Y, Wu X, Li B, Liu Z. Down-regulation of HECTD3 by HER2 inhibition makes serous ovarian cancer cells sensitive to platinum treatment. Cancer Lett. 2017;411:65–73.
Li Y, Wu X, Li L, Liu Y, Xu C, Su D, et al. The E3 ligase HECTD3 promotes esophageal squamous cell carcinoma (ESCC) growth and cell survival through targeting and inhibiting caspase-9 activation. Cancer Lett. 2017;404:44–52.
Li Z, Zhou L, Prodromou C, Savic V, Pearl LH. HECTD3 mediates an HSP90-dependent degradation pathway for protein kinase clients. Cell Rep. 2017;19:2515–28.
Li F, Li Y, Liang H, Xu T, Kong Y, Huang M, et al. HECTD3 mediates TRAF3 polyubiquitination and type I interferon induction during bacterial infection. J Clin Investig. 2018;128:4148–62.
Li Y, Kong Y, Zhou Z, Chen H, Wang Z, Hsieh YC, et al. The HECTD3 E3 ubiquitin ligase facilitates cancer cell survival by promoting K63-linked polyubiquitination of caspase-8. Cell Death Dis. 2013;4:e935.
Wu X, Li L, Li Y, Liu Z. MiR-153 promotes breast cancer cell apoptosis by targeting HECTD3. Am J Cancer Res. 2016;6:1563–71.
Engbrecht M, Mangerich A. The nucleolus and PARP1 in cancer biology. Cancers. 2020;12:1813.
Ghorai A, Mahaddalkar T, Thorat R, Dutt S. Sustained inhibition of PARP-1 activity delays glioblastoma recurrence by enhancing radiation-induced senescence. Cancer Lett. 2020;490:44–53.
Libermann TA, Nusbaum HR, Razon N, Kris R, Lax I, Soreq H, et al. Amplification, enhanced expression and possible rearrangement of EGF receptor gene in primary human brain tumours of glial origin. Nature. 1985;313:144–7.
Furnari FB, Fenton T, Bachoo RM, Mukasa A, Stommel JM, Stegh A, et al. Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes Dev. 2007;21:2683–710.
Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature. 2008;455:1061–8.
Hou J, Deng Q, Zhou J, Zou J, Zhang Y, Tan P, et al. CSN6 controls the proliferation and metastasis of glioblastoma by CHIP-mediated degradation of EGFR. Oncogene. 2017;36:1134–44.
Hatanpaa KJ, Burma S, Zhao D, Habib AA. Epidermal growth factor receptor in glioma: signal transduction, neuropathology, imaging, and radioresistance. Neoplasia. 2010;12:675–84.
Felsberg J, Hentschel B, Kaulich K, Gramatzki D, Zacher A, Malzkorn B, et al. Epidermal growth factor receptor variant III (EGFRvIII) positivity in EGFR-amplified glioblastomas: prognostic role and comparison between primary and recurrent tumors. Clin Cancer Res. 2017;23:6846–55.
Lai Y, Kong Z, Zeng T, Xu S, Duan X, Li S, et al. PARP1-siRNA suppresses human prostate cancer cell growth and progression. Oncol Rep. 2018;39:1901–9.
Peng W, Shi S, Zhong J, Liang H, Hou J, Hu X, et al. CBX3 accelerates the malignant progression of glioblastoma multiforme by stabilizing EGFR expression. Oncogene. 2022;41:3051–63.
Hou J, Xu M, Gu H, Pei D, Liu Y, Huang P, et al. ZC3H15 promotes glioblastoma progression through regulating EGFR stability. Cell Death Dis. 2022;13:55.
Dong Z, Lei Q, Yang R, Zhu S, Ke XX, Yang L, et al. Inhibition of neurotensin receptor 1 induces intrinsic apoptosis via let-7a-3p/Bcl-w axis in glioblastoma. Br J Cancer. 2017;116:1572–84.
Xuan F, Huang M, Liu W, Ding H, Yang L, Cui H. Homeobox C9 suppresses Beclin1-mediated autophagy in glioblastoma by directly inhibiting the transcription of death-associated protein kinase 1. Neuro Oncol. 2016;18:819–29.
McPhee TR, McDonald PC, Oloumi A, Dedhar S. Integrin-linked kinase regulates E-cadherin expression through PARP-1. Dev Dyn. 2008;237:2737–47.
Li F, Liang H, You H, Xiao J, Xia H, Chen X, et al. Targeting HECTD3-IKKalpha axis inhibits inflammation-related metastasis. Signal Transduct Target Ther. 2022;7:264.
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, et al. The somatic genomic landscape of glioblastoma. Cell. 2013;155:462–77.
Furnari FB, Cloughesy TF, Cavenee WK, Mischel PS. Heterogeneity of epidermal growth factor receptor signalling networks in glioblastoma. Nat Rev Cancer. 2015;15:302–10.
Chaffanet M, Chauvin C, Laine M, Berger F, Chedin M, Rost N, et al. EGF receptor amplification and expression in human brain tumours. Eur J Cancer. 1992;28:11–7.
Libermann TA, Razon N, Bartal AD, Yarden Y, Schlessinger J, Soreq H. Expression of epidermal growth factor receptors in human brain tumors. Cancer Res. 1984;44:753–60.
Eskilsson E, Rosland GV, Solecki G, Wang Q, Harter PN, Graziani G, et al. EGFR heterogeneity and implications for therapeutic intervention in glioblastoma. Neuro Oncol. 2018;20:743–52.
Gilbert MR, Dignam JJ, Armstrong TS, Wefel JS, Blumenthal DT, Vogelbaum MA, et al. A randomized trial of bevacizumab for newly diagnosed glioblastoma. N. Engl J Med. 2014;370:699–708.
Preusser M, Lim M, Hafler DA, Reardon DA, Sampson JH. Prospects of immune checkpoint modulators in the treatment of glioblastoma. Nat Rev Neurol. 2015;11:504–14.
Alexander BM, Cloughesy TF. Adult glioblastoma. J Clin Oncol. 2017;35:2402–9.
Acknowledgements
We thank Daping Hospital (Chongqing, China) for providing primary glioblastoma cells (GBM-3). We thank Dr. Ulrich Aymard Ekomi Moure for correcting spelling and grammar errors. We also thank Jifu Li for providing technical support during the use of the Confocal Laser Scanning Microscope (Olympus Fv1000, Japan).
Funding
This research was supported by the Natural Science Foundation of Chongqing (cstc2022ycjh-bgzxm0145) and (cstc2019jcyjzdxmX0033).
Author information
Authors and Affiliations
Contributions
GZ and HC were responsible for the design of the study. GZ, PL, RT, SW, BL, and XH performed the experiments. HC, GZ, XD, RY, and EZ analysed the data and designed the figures. GZ, PL, and JZ wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Animal handling was approved by the Committee for Animal Protection and ethics of Southwest University. All experiments were conducted following the Guidelines for Animal Health and Use of the Ministry of Science and Technology of China (2006). Clinical glioma tissue samples were purchased from Chaoying Biotechnology Co., Ltd, and the study was approved by the Medical Ethics Committee of Tongxu County People’s Hospital of Henan Province and all participants provided informed consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, G., Tan, R., Wan, S. et al. HECTD3 regulates the tumourigenesis of glioblastoma by polyubiquitinating PARP1 and activating EGFR signalling pathway. Br J Cancer 127, 1925–1938 (2022). https://doi.org/10.1038/s41416-022-01970-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-01970-9
This article is cited by
-
OTUD4 promotes the progression of glioblastoma by deubiquitinating CDK1 and activating MAPK signaling pathway
Cell Death & Disease (2024)
-
Predictive value of CCL2 in the prognosis and immunotherapy response of glioblastoma multiforme
BMC Genomics (2023)
-
AKIP1 accelerates glioblastoma progression through stabilizing EGFR expression
Oncogene (2023)